Abstract
Severe acute respiratory syndrome coronavirus 2 is an RNA virus currently causing a pandemic. Due to errors during replication, mutations can occur and result in cell adaptation by the virus or in the rise of new variants. This can change the attachment receptors' usage, result in different morphology of plaques, and can affect as well antiviral development. Indeed, a molecule can be active on laboratory strains but not necessarily on circulating strains or be effective only against some viral variants. Experiments with clinical samples with limited cell adaptation should be performed to confirm the efficiency of drugs of interest. In this protocol, we present a method to culture severe acute respiratory syndrome coronavirus 2 from nasopharyngeal swabs, obtain a high viral titer while limiting cell adaptation, and assess antiviral efficiency.
Keywords
Introduction
At the end of 2019, a new coronavirus, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), was identified in Wuhan. 1 Since then, it has caused millions of deaths. 2 Different strategies of treatment are available or under investigation such as protease inhibitor, nucleoside analogs, monoclonal antibodies, or compounds targeting the secondary structure of viral RNA.3–7 Several cell lines can be infected by this virus, which has as primary entry receptor angiotensin-converting enzyme 2 (ACE-2).1,8–10 Vero E6, a monkey epithelial kidney cell line, is the most used for viral culture due to the extensive cytopathic effect after infection and fast viral growth,8,9,11 also if it is not representative of the natural tropism of the virus.
When viruses replicate inside the host cell, mutations can occur and may be selected due to an advantage in replication in the cell line used in the laboratory. The same mutations will not occur in vivo and might cause an evolutionary divergence. Viruses can acquire the ability to bind different receptors and increase their affinity for cells that do not represent the natural tropism. For instance, these changes might affect viral entry. Cell adaptation is well known for several viruses: the use of heparan sulfate as an attachment receptor is increased after cell passaging12–28; human immunodeficiency virus 1 (HIV-1) can adapt to have a higher fitness for infection of MT4 cells 29 ; and SARS-CoV-2 was reported as well to adapt to cell culture: increasing plaques size was observed due to a mutation in the furin-like S1/S2 cleavage site. 11 Adaptations can be an issue during antiviral design since the antiviral activity of the drug of interest can rely on the cause or the consequences of this effect. For example, antivirals acting on the viral entry of yellow fever virus might be effective against the Asibi derived strain (17D), but not on the original strain, because they have a distinct entry route. 21 Experiments with circulating strains with low cell passaging are therefore essential to validate antivirals' activity.
At the time of writing the emergence of new variants is the major concern of vaccine escape and increased transmissibility of the virus.30–32 Moreover, this phenomenon is likely to increase in the future, due to the selective pressure caused by the increase of vaccine coverage and exposure of the population to natural infection. 33 Hence, it is important to assess the antiviral efficacy of a molecule against multiple variants of interest circulating in the population, rapidly isolated from patient samples.
In this context, we established a simple pipeline to culture a clinical sample of SARS-CoV-2 for antiviral assays. With our protocol, it is possible to isolate SARS-CoV-2 with a minimal number of passages from a nasopharyngeal sample. We present as well methods to characterize the stock by titration, reverse transcription-quantitative polymerase chain reaction (RT-qPCR) and sequencing (Figure 1). In the end, we show how to test antiviral activity in Vero E6 cells with isolated clinical strains. Other protocols are available for the isolation of SARS-CoV-2 from different clinical samples, but the goal was to assess the reliability of qPCR and the correlation among RNA copies and infectious viruses.34–37 In opposition, our protocol achieves a better success rate from nasopharyngeal RT-qPCR positive samples and can produce viral stocks with high titer and minimal adaptations.
Pipeline to culture severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) from the clinical swab. Vero E6 cells are infected in a 6 well plate with the transport liquid of a nasopharyngeal sample. After 3 days of infection, cells and supernatants are split into flasks. When the cells display an extensive cytopathic effect, the supernatant is harvested. Virus isolates are characterized by titration, RNA quantification, and sequencing. Created with biorender.com.
This protocol must be performed by personnel who received specific training for a biosafety level (BSL) 3 laboratory authorized to work with SARS-CoV-2.
Materials and reagents
24 well plate (Costar, catalog number: 3524). 6 well plate (Costar, catalog number: 3506). Amphotericin B (Thermo Fisher Scientific, Gibco, catalog number: 15290018). Aspirating pipet (Falcon, catalog number: 357558). Avicel GP3515 (SelectChemie, DuPont, catalog number: 41094201). Cell culture flask 75 cm2, (Falcon, catalog number: 353135). Cell spatula (TPP Techno Plastic Products AG, catalog number: 99010). Cercopithecus aethiops epithelial kidney cells (Vero E6), (ATCC, catalog number: CRL-1586). Crystal violet (Merck, Sigma-Aldrich, catalog number: 61135-100 g). Disinfectants (bleach 13–14%, ethanol 70%). dNTP Mix 10 mM each (Promega, catalog number: U1511). Dulbecco's Modified Eagle Medium (DMEM) + GlutaMAX (Thermo Fisher Scientific, Gibco, catalog number: 31966-021). E.Z.N.A Total RNA Kit I (Omega Bio-Tek, catalog number: R6834-02). Eppendorf tubes 2 mL, (Sarstedt AG & Co, catalog number: 72.694.006). Ethanol absolute (Merck, Sigma-Aldrich, catalog number: 1.00983.2500). extrAXON DNA Clean-up kit (Axonlab, catalog number: 685674). Fetal bovine serum (FBS), (Pan Biotech, catalog number: P40-47500). Formaldehyde 37% (Merck, Sigma-Aldrich, catalog number: F1635-500 mL). GoScript™ Reverse Transcriptase (Promega, catalog number: A5002). GoTaq qPCR Master Mix kit (used as PCR kit) (Promega, catalog number: A6000). Millex-GS Filter 0.22 µm, (Merck, Millipore, catalog number: SLGS033SS). Penicillin-Streptomycin (P/S), (Merck, Sigma-Aldrich, catalog number: P4333-100 ml). Personal equipment for a BSL-2 laboratory (gloves, lab coat). Personal equipment for a BSL-3 laboratory (gloves, Tyvek, FFP3 mask, glasses, shoes, mobcap). Phosphate buffered saline (PBS) pH 7.4 (Thermo Fisher Scientific, Gibco, catalog number: 10010-015). Pipet tips (10 µL, 200 µL) (Thermo Fisher Scientific, ThermoScientific, catalog number: 2149E, 2069). Pipet tips 1′000 µL, (Sorenson BioScience Inc., catalog number: 10380) Primer PCR forward: 5′-TCTCTTCTTAGTAAAGGTAGACTT-3′ (Spike gene). Primer PCR reverse: 5′-CTAACAATAGATTCTGTTGGTTG-3′ (Spike gene). Primer RT-qPCR forward: 5′-ACAGGTACGTTAATAGTTAATAGCGT-3′ (E gene). Primer RT-qPCR reverse: 5′-ATATTGCAGCAGTACGCACACA-3′ (E gene). Probe RT-qPCR: 5′-ACACTAGCCATCCTTACTGCGCTTCG-3′ (E gene). QuantiTect Probe RT-PCR kit (Qiagen, catalog number: 204443). Random primers 3 µg/mL (Thermo Fisher Scientific, Invitrogen, catalog number: 48190011). RNasin Plus RNase Inhibitor (Promega, catalog number: N2611). Serological pipettes (5 mL, 10 mL, 25 mL) (Falcon, catalog number: 357543, 357551, 357525). Stericup Quick Release Filter 0.22 µm (Merck, Millipore, catalog number: S2GPU05RE). Syringe 10 mL, (Braun, catalog number: 4616103V). TRK lysis buffer (Omega Bio-Tek, catalog number: PR021). Trypsin-EDTA PBS 1:250 (Amimed, catalog number: 5-51F00-H). Tube (50 mL, 15 mL) (Falcon, catalog number: 352070, 352096). Water (Bichsel, catalog number: FE1001318).
Equipment
−20°C freezer.
−80°C freezer.
Biosafety hood in a BSL-2 laboratory.
Biosafety hood in a BSL-3 laboratory.
Cell counter.
Cell Locker (Thermo Fisher Scientific, ThermoScientific, catalog number: 50151650).
Centrifuge for 15 – 50 mL tubes refrigerated.
Fridge.
Incubator.
Inverted light microscope.
Pipet set (10 µL, 20 µL, 200 µL, 1000 µL).
Pipetboy.
RT, PCR, RT-qPCR workstation.
Vortex.
Procedures
In the following section, we describe the different steps to culture SARS-CoV-2 from clinical samples, characterize the newly produced virus stock (titration, RNA quantification, sequencing) (Figure 1), and test the antiviral activity of the drug of interest.
Antiviral efficiency by plaque assay. Cells are infected with 100 PFU/well of clinical isolate. After 1 h of incubation at 37°C, the inoculum is removed and serial dilutions of the antiviral are added to the wells in a medium supplemented with Avicel. Cells are incubated for 2 days before fixation and staining. Plaques are counted and IC50 evaluated. Created with biorender.com.
a. Culture of Vero E6
Note: the culture of Vero E6 is done under a biosafety hood.
Vero E6 cells are thawed in a culture flask 75 cm2 in 10% FBS DMEM (see recipes). Vero E6 cells are grown in an incubator at 37°C with 5% CO2. Cells are washed with PBS and detached with trypsin-EDTA when they reach 90–100% confluence.
b. Culture of clinical sample SARS-CoV-2
Note: This work needs to be done under a biosafety hood in a BSL-3 laboratory. Due to safety measures, all the infected cells are incubated and transported into a Cell Locker. To increase the success of viral isolation, clinical specimens with a low Cycle Threshold (CT) (< 25) should be used. The nasopharyngeal sample should be stored at −80°C until use and should not be frozen-thawed several times.
Vero E6 cells are plated in a 6 well plate (3.5*105 cells/well) to reach 75% confluence after 24 h. The next day, cells are infected with 100–300 µL of clinical nasopharyngeal SARS-CoV-2 sample in a total volume of 1 mL with 2.5% FBS DMEM (see recipes). Contamination by bacteria or fungi is possible. To limit the risk, the addition of antifungals or antibiotics and filtration of the sample might be beneficial. Cells are incubated for 3 h at 37°C in a CO2 incubator. The infection mix is then removed and 2 mL of 2.5% FBS DMEM is added to the cells for 3 days in a CO2 incubator. After this period, a beginning of cytopathic effect might be observed, but the protocol can be followed as well in the absence of it. Infected cells look initially rounder and with different light refraction. At later stages, cells detach from the flask. With some isolates, it is possible to observe the formation of syncytia (i.e. fusion of neighbor cells). The supernatant is collected, and the cells are washed with 1 mL of PBS and detached with 1 mL of trypsin-EDTA followed by the addition of 3 mL of 10% FBS DMEM to neutralize the trypsin effect. Two milliliters of uninfected Vero E6 cells and detached infected Vero E6 cells are added in a culture flask 75 cm2 with 1 mL of supernatant collected and fresh media at 2.5% FBS to reach the final volume of 12 mL. Vero E6 cells are incubated at 37°C in a CO2 incubator until they display extensive cytopathic effect (generally after 4–5 days). Supernatant and scraped cells are collected and vortexed. Cells and debris are removed by centrifugation at 2*103 r/min for 5 min. The supernatant is aliquoted and stored at −80°C. The aliquot should be frozen-thawed only once.
c. Characterization of the isolate
c.1 Titration
Vero E6 cells are plated in a 24 well plate (105 cells/well) to reach 100% confluence after 24 h.
Cells are infected in duplicates with serial dilutions of viruses (e.g. 10−3–10−7) in 2.5% FBS DMEM with a total volume of 200 µL/well for 1 h at 37°C. A non-infected control is added for each titration.
The infection mix is removed and 500 µL of 0.4% Avicel GP3515 in 2.5% FBS DMEM is added in each well.
Cells are incubated for 2 days at 37°C with 5% CO2.
The supernatant is removed and 4% formaldehyde (see recipes) is added to the wells (500 µL) for 20 min at room temperature.
Formaldehyde is removed and 500 µL of Crystal violet (see recipes) is added for 20 min to stain the wells.
The wells are washed once with water and dried.
The number of plaques in each well is counted under an inverted light microscope (magnification 4 × ) (Figure 2).
The titer of virus stock in plaque-forming unit (PFU)/mL is the average of individual titration for each dilution:
c.2 Quantification of RNA copies
Fifty microliters of virus stock are lysed with 300 µL of TRK lysis buffer.
Extraction of total RNA is done as described by the fabricant Omega Bio-Tek.
RT-qPCR is performed according to the protocol of Corman et al. 38
c.3 Sequencing
For a small region of interest, RNA is extracted and reverse transcription is performed according to the protocol of the manufacturer. Afterward, a PCR is performed with the desired primers. To identify mutations on the spike gene, the protocol of the University Hospital of Geneva can be followed. 39 After the PCR clean-up column, the amplification product can be subjected to Sanger sequencing.
For complete genome sequencing, we recommend to follow the method of Kubik et al. 40
d. Antiviral activity
Note: We do not describe the assessment of toxicity of the compounds since it is not the main goal of our protocol and we assume that the antiviral evaluation is performed at non-toxic doses. Moreover, any effect of the solvent on cell viability or the virus should be excluded. In the following section, we used a 24 well plate for our antiviral assays since we are able to count plaques under the microscope in 48 h. Other sizes of a plate can be used, that is, 6 well plate and longer incubation times to have larger plaques countable by eye, or smaller formats in which perform immunostaining to detect infection as described by others.41,42
The aspect of plaques after infection with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). At the top, representative wells of non-infected and infected by SARS-CoV-2 Vero E6 after staining with crystal violet. At the bottom, non-infected cells and a plaque were observed under an inverted light microscope (magnification 10 × ).
d.1 Plaque assay
Note: Here we describe a protocol in which the molecule is added after infection until fixation. The treatment can differ accordingly to the mechanism of action of the drug of interest. However, treatment post-infection is more relevant for future clinical use. This method is based on previously published articles.43,44
One hundred thousand Vero E6 cells per well are plated in a 24 well plate. The next day, cells are infected with 100 PFU/well in 2.5% FBS DMEM for 1 h at 37°C. Cells are treated with 500 µL of serial dilutions of the drug, at non-toxic concentrations, in 0.4% Avicel diluted in 2.5% FBS DMEM. Vero E6 cells are incubated for 2 days in a CO2 incubator. Cells are fixed with 4% formaldehyde and stained with crystal violet as previously described. Plaques of SARS-CoV-2 are counted (Figure 3). Percentages of infection are calculated by comparing the number of plaques in the treated wells with the wells containing equal volumes of solvent. The half-maximal inhibitory concentration (IC50) is calculated with Prism 8 (GraphPad, San Diego, US).
d.2 Other methods
The inhibitory activity of the compound of interest can be assessed by alternative methods.
Viral yield reduction can be used to evaluate the release of viral particles by RT-qPCR, or titration of supernatants.45–47
The percentage of infected cells or levels of viral protein can be assessed by immunofluorescence or flow cytometry with a primary antibody targeting viral proteins or with reporter genes.46, 48
Expected results
After isolation of different clinical isolates as described above, examples of titers obtained for antiviral assays and mutation that occurred during the isolation are shown in Table 1.
Lineage, titer and mutation of different strains after isolation using this protocol.

Sixteen clinical samples out of 18 were isolated with this protocol. The two non-isolated samples had high initial CT and low starting material. The percentage of samples without any mutation after the isolation represents 54%. Samples with at least two mutations represent 15%.
An example of dose–response antiviral activity against SARS-CoV-2 is shown in Figure 4. The results were assessed by plaque assay, as described in this protocol, using merafloxacin that was previously reported to inhibit SARS-CoV-2 by interacting with its viral RNA. 7 The data are presented as a percentage of infection calculated in comparison to untreated cells. The dose axis is represented in a logarithmic scale. IC50 can be calculated with GraphPad Prism software (nonlinear regression -> log(inhibitor) vs. response (four parameters) -> bottom and top constraints are 0, 100, respectively). R2 values and confidence interval have to be critically observed. IC50s with R2 values < 0.5 and a large confidence interval represent high variability between the replicates and require additional experiments. IC50s greater than the higher dose tested should be discarded. Drugs with low IC50s and high selectivity index (ratio between the half-maximal cytotoxic concentration (CC50) and the IC50) (e.g. > 10) warrant further investigation.
Example of results obtained following this protocol. Vero E6 cells were infected with a B 1.1.7 (alpha variant) severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and cells were treated with merafloxacin as described in this protocol. The x-axis represents the concentration of the drug on a logarithmic scale and the y-axis, the percentage of the number of plaques normalized to the untreated wells. Metrics were calculated with non-linear regression analysis with GraphPad Prism: IC50 of 23.68 µM with a confidence interval between 16.81 and 33.83 µM, an R2 value of 0.7759 and a Hill slope of −1.385. The result represents the mean and standard error of the mean of two independent experiments.
Conclusion
SARS-CoV-2 caused an important pandemic and treatments are still under investigation.3,4 Since the adaptation of viruses to cell culture can bias the results, and viral variants are emerging with high frequency, drug efficiency should be confirmed with circulating strains. However, material from a nasopharyngeal sample is insufficient for characterization and antiviral testing.
Here, we presented a simple way to culture clinical SARS-CoV-2 without extensive passaging. Our method allowed us to isolate clinical strains from nasopharyngeal samples and limit cell adaptation. Our strategy can be applied as well to other viruses. The limitation of our protocol is that cell adaptation cannot be fully omitted since a balance between fast viral growth and cytopathic effect evaluation obliged us to use Vero E6 cells. These cells are not the natural host cell for SARS-CoV-2. Therefore, confirmation of the antiviral activity should be performed in a representative model such as human lung adenocarcinoma cell line (Calu-3)42,49 or respiratory tissues. 50 This protocol was used to isolate different SARS-CoV-2 variants from patients for quality control of sequencing and antiviral assays for manuscripts in preparation.
Recipes
10% FBS DMEM
500 mL DMEM. 50 mL FBS (It has been previously decomplemented for 30 min at 56°C and filtered with a 0.22 µm filter). 5 mL P/S. 500 mL DMEM. 12.5 mL FBS decomplemented and filtered. 5 mL P/S. 7.2 g Avicel. 300 mL water.
2.5% FBS DMEM
2.4% Avicel
Dissolve with a magnetic bar for 10 min and autoclave.
Crystal violet solution
0.5 g crystal violet. 100 mL ethanol absolute. 400 mL water. 108.1 mL formaldehyde 37%. 891.9 mL PBS.
4% Formaldehyde
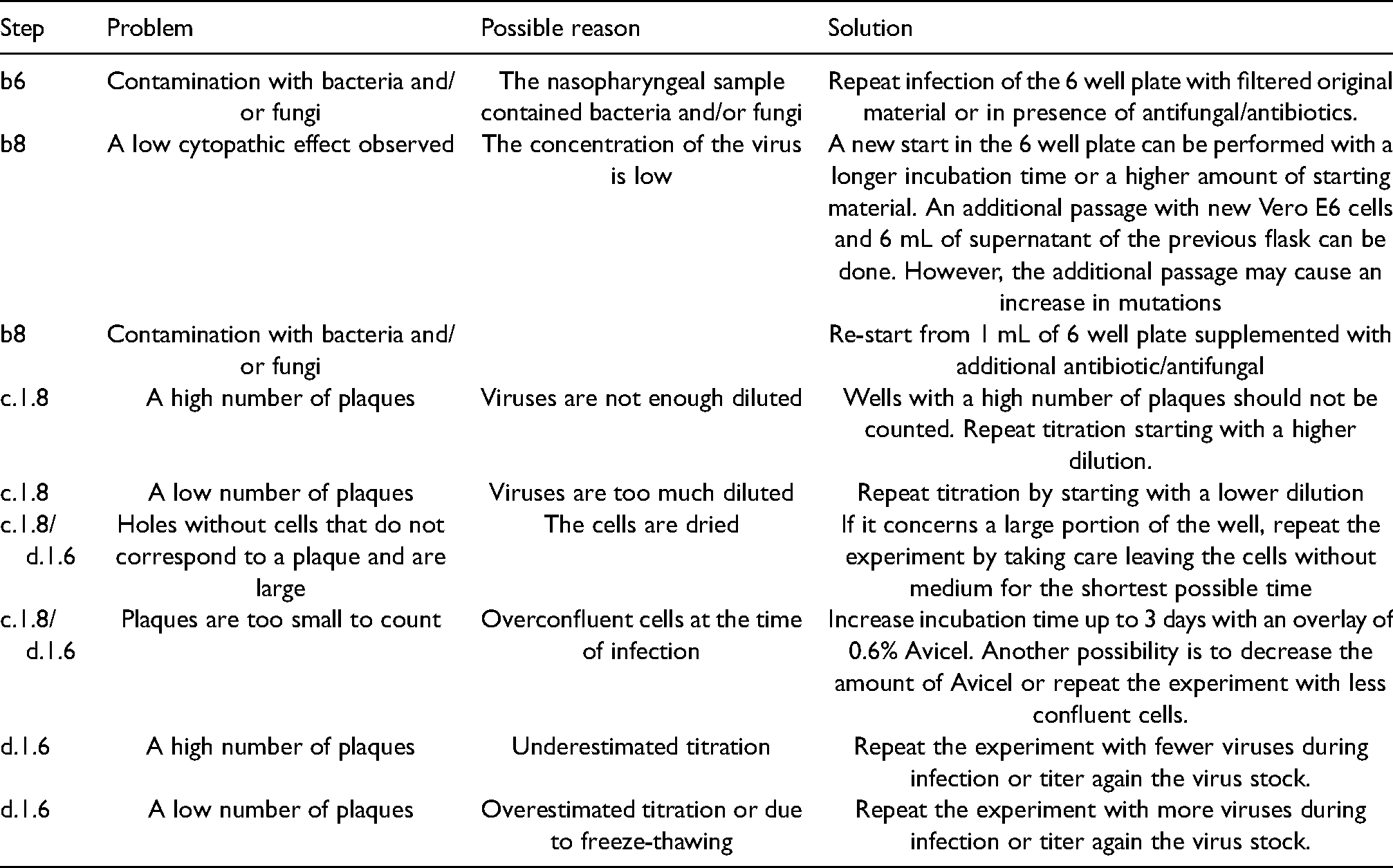
Troubleshooting

