Abstract
Respiratory syncytial virus (RSV) infects 99% of children by age 2 years and is a leading cause of serious lower respiratory tract infection (LRTI) and infant hospitalization in the United Kingdom. Identification of efficacious RSV therapeutics has been hindered by the lack of a robust and appropriate primary assay for high-throughput screening (HTS). Here we report an HTS cascade that identified inhibitors of RSV replication using a robust RSV replicon luminescence-reporter assay for the primary campaign. The performance of the assay was consistent and reliable at scale, with Z′ of 0.55 ± 0.08 across 150 assay plates and signal-to-background ratios >40. The HTS assay was used to screen the AstraZeneca compound library of 1 million compounds at a single concentration of 10 µM. Hits specifically targeting the RSV replicon were determined using a series of hit generation assays. Compounds nonspecifically causing cell toxicity were removed, and hits were confirmed in live viral inhibition assays exhibiting greater physiological relevance than the primary assay. In summary, we developed a robust screening cascade that identified hit molecules that specifically targeted RSV replication.
Introduction
Severe lower respiratory tract infections (LRTIs) such as pneumonia and acute bronchiolitis are the leading cause of infant mortality in both the developed and developing world. 1 Respiratory syncytial virus (RSV) is the most common viral cause of infant LRTIs, associated with around 75% of all bronchiolitis 2 and up to 40% of pneumonia cases. 3 In temperate climates such as the United Kingdom, RSV epidemics occur between late October and early March and peak over December and the New Year period (data from Health Protection Agency; www.hpa.org.uk). In England, around 3% of all hospital admissions in infants younger than 1 year are attributable to RSV, and it is this population that is most at risk from infection. Infants in this age group represent more than 50% of reported cases and are the most susceptible to forming severe LRTIs (data from Health Protection Agency; www.hpa.org.uk). Elderly patients with chronic health conditions are also at increased risk of developing a severe infection due to RSV. To highlight the impact to the National Health Service, it is estimated that RSV accounted for almost 90,000 occupied hospital bed days in the 2010–2011 RSV season. 4 The burden of RSV infection in the developing world is higher still, where additional risk factors such as malnutrition, human immunodeficiency virus (HIV) infection, crowding, and incomplete vaccination status make children more susceptible to severe LRTIs.3,5 Efficacious treatments for RSV would have a significant worldwide benefit to the patients described here and affect the current global economic burden of this virus.
There is one efficacious RSV antibody therapy, palivizumab (Synagis; MedImmune, Gaithersburg, MD), prescribed prophylactically to patient populations thought to be at high risk, but currently there are no effective small-molecule treatments for RSV infection. Ribavirin is a small molecule that was first marketed in the 1980s and is currently the only licensed drug for RSV. It is rarely prescribed due to demonstrating only marginal clinical benefit in patients with RSV and evidence of toxicity. 6 There are a number of antiviral RSV approaches, 7 including YM-53403, a small molecule that targets the RSV L (polymerase) protein, required for transcription and replication of the RSV genome. 8 Although there are a number of clinical vaccine studies currently taking place, 9 there are at present no licensed vaccines for RSV. The aim of this study was to screen the AstraZeneca chemical diversity collection to identify small-molecule inhibitors of RSV.
We have previously described the development of a high-throughput replicon assay for the identification of RSV inhibitors. 10 The use of an immortalized replicon cell line to prosecute multiple target classes was highly desirable. It gave us the advantages associated with a robust phenotypic cellular assay without the challenge of the biological safety requirements associated with pathogen screening at scale. An initial screening campaign of 158,000 compounds using this replicon cell line was performed with limited success. The campaign was detrimentally affected by significant variation in assay signal and noise, both within a screening run and on different screening occasions. The enrichment of the replicon-expressing cell population and use of cryopreserved assay-ready cells led to the development of a robust high-throughput screening (HTS) assay capable of screening at scale. Here we report the successful screening of 1 million compounds and the use of a validated hit generation cascade to identify selective and potent inhibitors of RSV replication. These molecules will be progressed further to potentially provide effective new small-molecule treatments for RSV infection.
Materials and Methods
Cells and Media
All cell lines were grown under standard culture conditions at 37 °C with 95% humidity in 5% CO2. RSV replicon cell line APC126, originating from Dr. Mark Peeples’s laboratory, was licensed from Apath, LLC (Brooklyn, NY). The cell line contains a stable and self-replicating RSV A2 subgenomic replicon encoding the viral proteins essential for RSV replication and the Renilla luciferase reporter.
9
The enriched RSV replicon cell line APC126-
RSV Propagation
Human RSV A2 strain was originally purchased from ATCC (VR-1450), in the form of an infected HeLa lysate. Virus was expanded using HEp2 cells in virus infection medium consisting of modified Eagle’s minimum essential medium (Opti-MEM; Invitrogen) supplemented with 5% heat-inactivated FBS, 1 U/mL penicillin, and 1 µg/mL streptomycin. RSV A2 seed stock was diluted in virus infection medium to infect cells at a multiplicity of infection (MOI) of 0.1. Supernatant was removed 24 h postinfection and replaced with Opti-MEM, and infection continued for a further 48 h. Viral supernatant and cell lysates postinfection were collected, lysed by freeze-thaw, and centrifuged at 2000 g for 5 min at 4 °C. Viral stocks were aliquoted and stored at −80 °C in the presence of virus stabilizer (HyClone, Logan, UT).
Compound Preparation and Screening
All test compounds, including Ribavirin (Sigma) and YM-53403 (synthesized in house), were solubilized in 100% (v/v) DMSO to 10 mM. The RSV L inhibitor used in the HTS refers to a YM-53403 analogue synthesized in house. To generate compound transfer plates (CTPs) for campaign 1, compounds were acoustically dispensed using an Echo 555 (Labcyte, Sunnyvale, CA) into clear 384-well polypropylene microplates (Greiner Bio-One, Monroe, NC), and 15 µL APC126 assay media was added to create 10× compound stocks. For single-concentration screening, 3 µL of 10× compounds were transferred from CTPs using a proprietary tip-based dispenser. To generate assay-ready plates (ARPs) for campaign 2, compounds were acoustically dispensed into white 384-well tissue culture–treated microplates (Greiner Bio-One) for the luminescence-based reporter assay, black microclear 384-well tissue culture–treated microplates (Greiner Bio-One) for the cytotoxicity assays, and clear 384-well tissue culture–treated microplates (Corning, Oneonta, NY) for the enzyme-linked immunosorbent assay (ELISA). For single-concentration screening, 150 nL of 4-mM compounds was dispensed, and ARPs were thermally sealed and stored at room temperature. All single-concentration screening was at a final concentration of 0.25% (v/v) DMSO and a compound concentration of 10 µM. For concentration response (CR) screening, varying volumes of compound stocks were dispensed to generate 7-point, half-log CR curves, with a final highest concentration of 30 µM. Wells were backfilled with the required volume of DMSO to ensure a final concentration of 0.3% (v/v). Plates containing maximum (0% inhibition of assay response), minimum (100% inhibition of assay response), and reference (50% inhibition of assay response) compounds were used to determine assay quality and compound activity. For both campaigns, the maximum assay signal was defined with 0.25% v/v DMSO. Campaign 1 used a mixture of Ribavirin and YM-53403, and campaign 2 used RSV L inhibitor (YM-53403 analogue 11 ) as a minimum and reference control.
RSV Replicon HTS Assays
For screening assays using APC126 (campaign 1), cryopreserved cells were thawed and cultured for 7 days with two passages prior to being plated at a density of 7000 cells/well in a 15 µL volume. Plates were incubated under standard culture conditions for at least 4 h before a further 15 µL of assay media was added. Compounds were prepared as described earlier, transferred to assay plates at 3 µL/well, and incubated under standard culture conditions for 48 h. Replicon activity was measured via the Renilla luciferase reporter and subsequent generation of a luminescence signal following the addition of EnduRen Live Cell Substrate (Promega, Madison, WI). Briefly, 15 µL assay media was removed from the assay wells prior to the addition of 2 µL assay media containing 86 µM EnduRen substrate. Cells were incubated with EnduRen for 1.5 h under standard culture conditions, followed by a 30-min incubation at ambient temperature. Luminescence was measured using an Ultra plate reader (Tecan, Männedorf, Switzerland). Screening assays using cryopreserved APC126-
Cytotoxicity Assays
Cryopreserved cells were dispensed into ARPs using a Multidrop Combi (Thermo Scientific, Waltham, MA). APC126-
RSV ELISA
All reagent additions were performed using a Multidrop Combi and well contents aspirated using a Powerwasher (Tecan). All wash steps were performed three times using phosphate-buffered saline (PBS) containing 0.1% (v/v) Tween-20 (PBS-T). Unless stated, all reagent additions were at a volume of 30 µL, and incubations were performed at ambient temperature. Cryopreserved HEp-2 cells were dispensed into ARPs at a density of 5000 cells/well. After a 1 h incubation under standard culture conditions, 2500 pfu of RSV A2 in assay medium was added and plates incubated for 72 h. Well contents were then aspirated and cells fixed for 30 min at 4 °C using ice-cold 80% acetone. The acetone was removed and plates left to air dry for at least 15 min. Casein block solution (Thermofisher Scientific, Rockford, IL) was added, followed by a 45-min incubation before wells were washed. Mouse anti-RSV monoclonal antibody (Millipore, Billerica, MA) diluted 1:8000 in PBS-T was added, and following a 1 h incubation, wells were washed. Goat anti–mouse IgG horseradish peroxidase antibody (Millipore) was diluted 1:4000 in PBS-T and added for 1 h. After a further wash step, SureBlue TMB Microwell Peroxidase Substrate (Millipore) was added and microplates incubated for 30 min. To stop the enzyme reaction, 0.25 M sulfuric acid was added, before measuring absorbance at 450 nm on an EnVision plate reader (PerkinElmer, Waltham, MA).
Data Analysis
All screening data were analyzed using proprietary AstraZeneca software. The activity of each compound was calculated based on the Minimum control data for each run (% effect = 100 × [1 − (test compound − median Minimum control) / (test plate median − median Minimum control)]). Compound EC50 and CC50 were calculated using a four-parameter logistic model.
Results
Monitoring RSV Replicon Using a Luciferase Reporter Assay
In 2010, the screening of a subset library of 158,058 compounds was attempted (campaign 1) using the continuously cultured nonenriched APC126 RSV replicon cell line. Attempts to generate an assay-ready cryobank with this cell line were unsuccessful, as the cells were not functional upon thawing and required several passages to generate sufficient assay window, and quickly lost assay functionality when maintained in suspension prior to plate addition (data not shown). These issues were circumvented by generating multiple small batches of plates (up to 10 batches of 30 plates for each screening run). Due to high assay variability (20% coefficient of variation), there was a requirement that compounds were tested in duplicate. Figure 1A shows the control data from campaign 1 that were used to calculate assay Z′. 12 Throughout the screening campaign, an overall Z′ of 0.42 ± 0.09 (n = 6) was achieved. Based on internal AstraZeneca guidelines, this was on the borderline of acceptable tolerances for single-test primary screening. This, plus the requirement for multiple daily cell batches and duplicate compound testing, meant that it was impractical to use the replicon assay with APC126 cells to prosecute a full-collection HTS campaign.
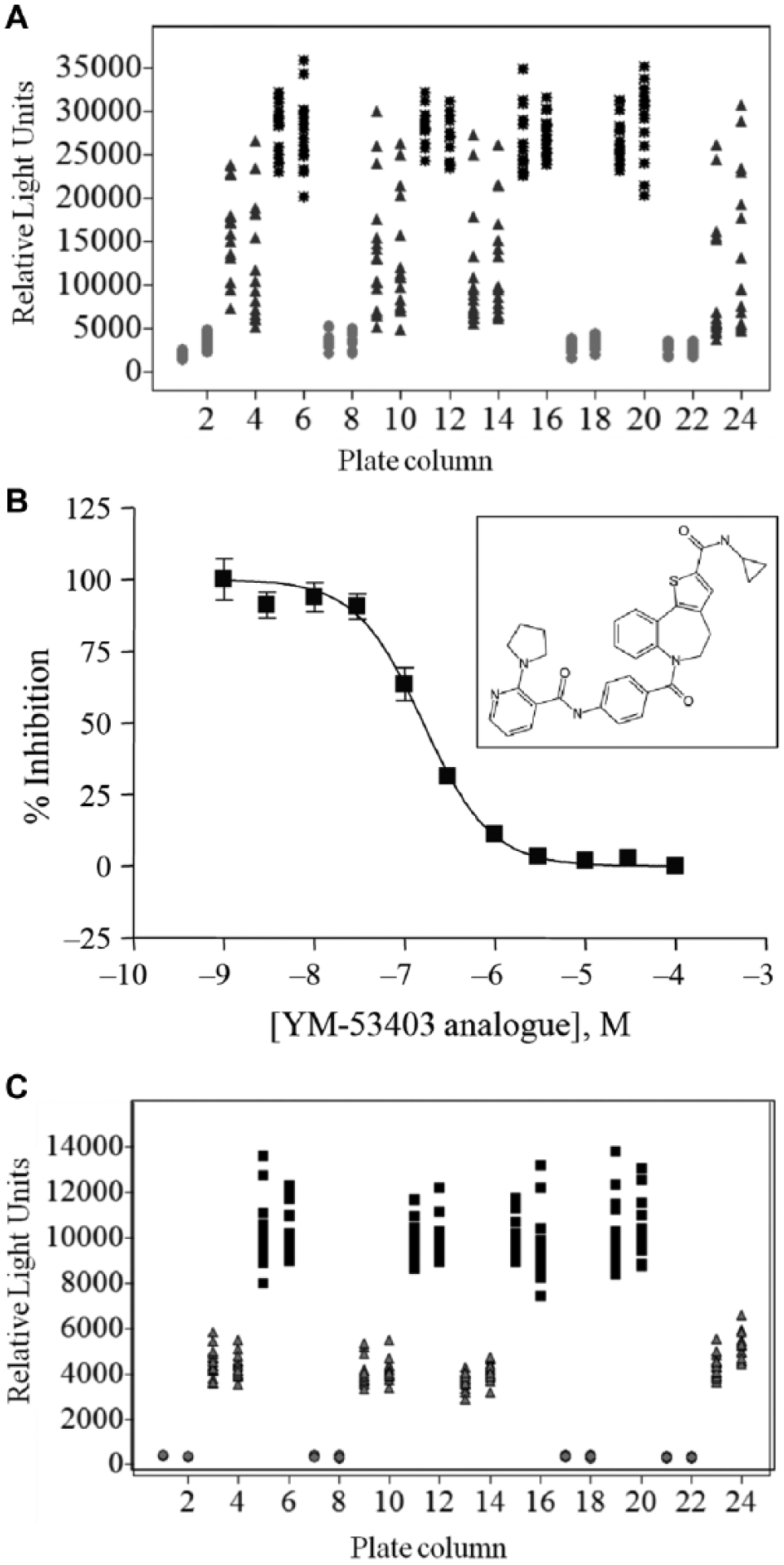
Data from respiratory syncytial virus replicon high-throughput screening assays. (
RSV Replicon Primary HTS
In 2012, we completed the development of an RSV replicon HTS assay using a single batch of cryopreserved APC126-
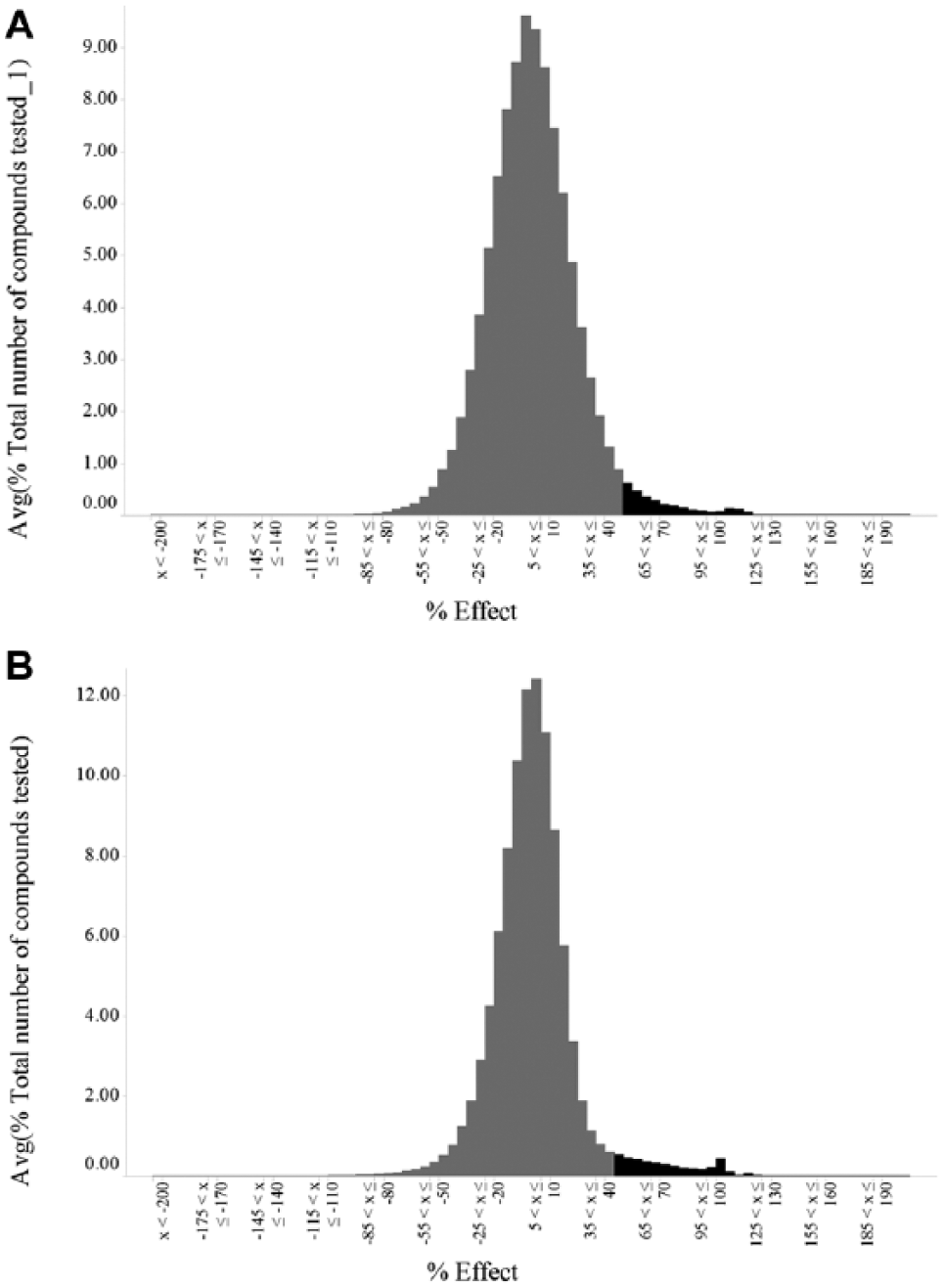
Single-concentration screening data from respiratory syncytial virus replicon high-throughput screening assays. (
Confirmation Screening Phase
All compounds identified as active in the APC126-
EC50 Screening Phase
To test the validity of the replicon assay in CR format, we used the validation data 10 to select a set of 47 reference compounds with a range of potencies. These were tested twice in CR format in the replicon assay. We then examined the reproducibility of the EC50 estimation 14 by plotting the arithmetic ratio of the EC50 between tests against the geometric mean EC50 ( Fig. 3 ). No obvious trends were seen in this analysis, indicating that the replicate EC50 measurements gave values that were reproducible and that the RSV replicon assay was suitable for EC50 determination. Testing the 8321 compounds from the confirmation screening phase of the cascade on two separate occasions in both the replicon and cytotoxicity CR assays allowed us to use CC50 data during the analysis of the replicon data. Removal of cytotoxic compounds from the data set left a total of 3969 active compounds with an EC50 of less than 30 µM (or where CC50/EC50 ≥ 10). This equated to a 0.4% active rate from the original primary screen of 1 million compounds.
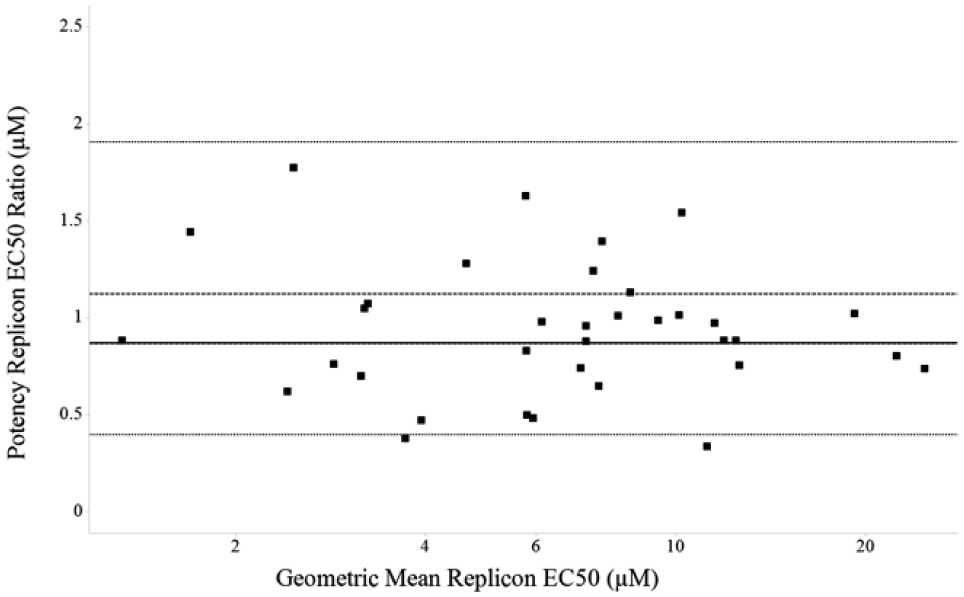
Analysis of respiratory syncytial virus (RSV) replicon EC50 assay performance. To test the validity of the replicon assay in concentration response (CR) format, we used validation data generated previously to select a set of 47 reference compounds with a range of potencies. These were tested twice in CR format in the assay. The graph shows the arithmetic EC50 ratio versus the geometric mean potency for these compounds. Mean EC50 ratio (solid line) is 0.87, the minimum significant ratio (MSR) is 2.19, and the limits of agreement (LsA) (0.4–1.91) are indicated by short dashed lines. The 95% confidence limits (0.87–1.12) are shown as long dashed lines (data from two separate occasions are shown). General acceptance criteria of an MSR <3 and LsA between 0.33 and 3.0 indicated that the RSV replicon assay was suitable for EC50 determination.
RSV A2 Live Virus ELISA Screening
To determine whether compound activity in the surrogate replicon assay used in the primary screening assay translated to a biological effect in a more physiological live virus system, we adapted an ELISA to measure RSV A2 levels in HEp2 cells. 15 To pharmacologically validate the ELISA, we tested an analogue of YM-53403, obtaining an EC50 of 0.11 µM, in line with published data 11 ( Fig 4A ). Figure 4B represents the data generated using the 47 reference compound set described earlier. We examined the reproducibility of the EC50 estimation 14 by plotting the arithmetic ratio of the EC50 between tests against the geometric mean EC50. No obvious trends were seen in this analysis, indicating that the replicate EC50 measurements gave values that were reproducible and that the RSV ELISA assay was suitable for EC50 determination. This assay was screened using a batch size of 30 plates and demonstrated a mean Z′ of 0.37 ± 0.12. All 3969 active compounds identified from the replicon CR screening phase of the campaign were tested on two separate occasions using the ELISA. They were also all screened in CR format in the HEp2 cytotoxicity assay. Of these compounds, 1144 were deprioritized based on a CC50 potency threshold of ≤30 µM.
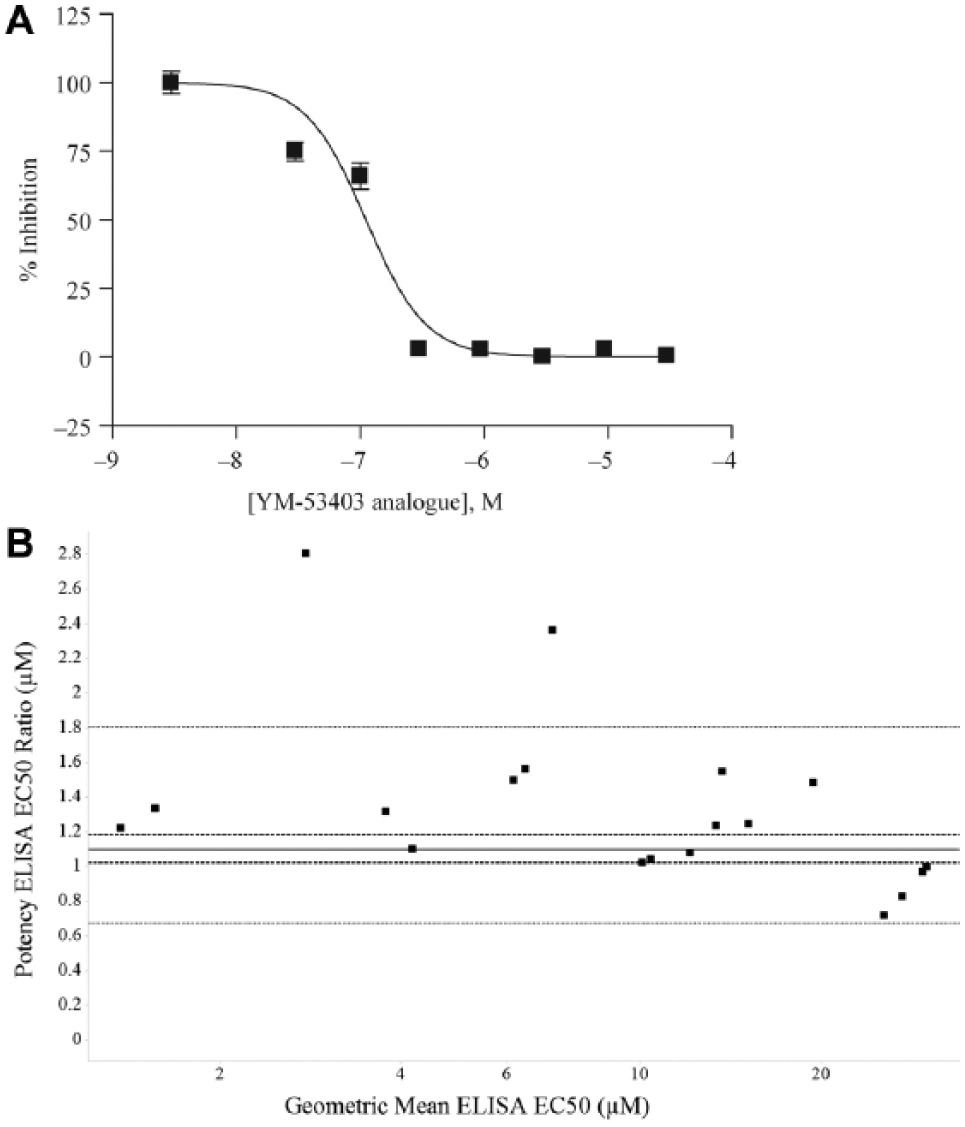
Analysis of respiratory syncytial virus (RSV) enzyme-linked immunosorbent assay (ELISA) EC50 assay performance. (
Figure 5 shows a comparison of mean pEC50 data (tested on two separate occasions) for 47 compounds tested in both RSV replicon and ELISA assays. The majority of reference compounds identified in the replicon assay had activity that translated into the live virus HEp2 ELISA ( Fig. 5 ), validating the replicon technology as a good hit-finding approach. These data suggest that the top five most attractive, highest potency compounds are equipotent in the two assays within the statistical limits of agreement. For some compounds, there was a drop-off in potency against the viral strain tested: this is not unexpected and may in part be attributed to the two assays probably measuring different levels of cellular viral protein. In contrast, four compounds exhibited a higher potency against the native ELISA rather than recombinant replicon system.
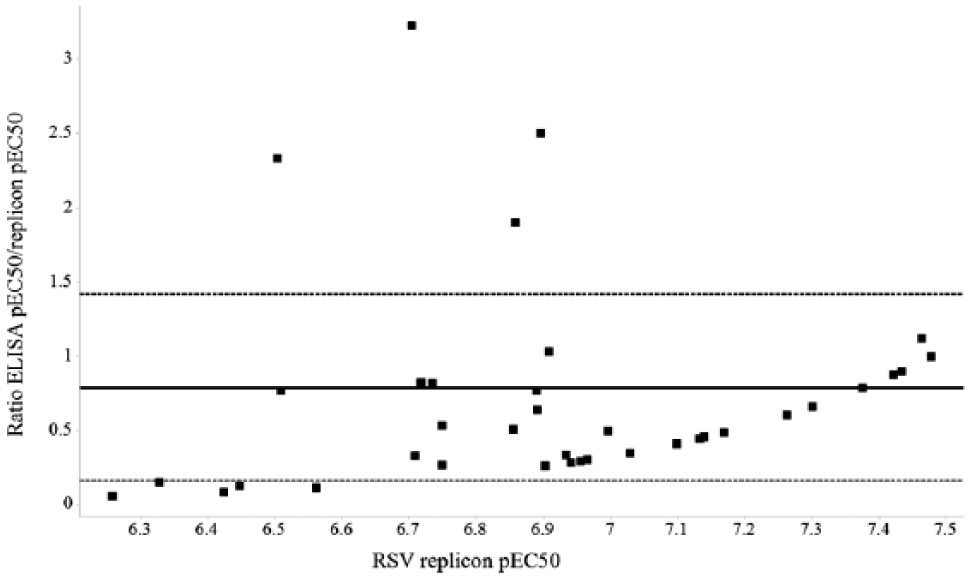
Comparison of respiratory syncytial virus (RSV) replicon and enzyme-linked immunosorbent assay (ELISA) EC50 assays. Graph of pEC50 ELISA/replicon ratio for the set of 47 reference compounds tested in both assays versus the mean RSV replicon pEC50 calculated from two separate occasions for each assay. Solid line is the mean pEC50 ratio, and short dashed lines indicate mean pEC50 ratio ± 1.0 × standard deviation.
Further Triage Assays
Figure 6 shows the overall screening cascade for campaign 2, with numbers of compounds tested at each stage of the cascade and the hit rate from each assay calculated as a percentage of the original compound collection. The RSV replicon active compounds validated by the ELISA and cytotoxicity assays were filtered to prioritize hits for further spectrum and selectivity testing. In total, 379 compounds were selected based on the criteria of (a) EC50 ≤ 10 µM, no cytotoxicity, or CC50/EC50 ratio ≥10 in replicon cells; (b) EC50 ≤ 10 µM, no cytotoxicity, or CC50/ EC50 ratio ≥10 in HEp2 cells in the RSV ELISA assay; and (c) no replicon or ELISA technology artifacts or known RSV inhibitors. The final pool of 379 prioritized compounds was then tested against representative strains from RSV A (A2 and Along) and B (Washington) subtypes in an RSV cytopathic effect (CPE) assay in HEp2 cells. A cytotoxicity assay was run in parallel, and all data were assessed to determine compound potency and spectrum coverage, as well as filter out compounds that were cytotoxic to proliferating cells. Compounds were also tested for selectivity for RSV in cell-based infection assays against influenza A virus (IAV) and herpes simplex virus (HSV). Compounds showing activity against these distinct viruses were deprioritized to filter out potential pan-antivirals targeting essential cellular pathways. This screening cascade identified four compound series that were potent and selective. These were progressed into an internal chemistry program for further characterization and optimization.
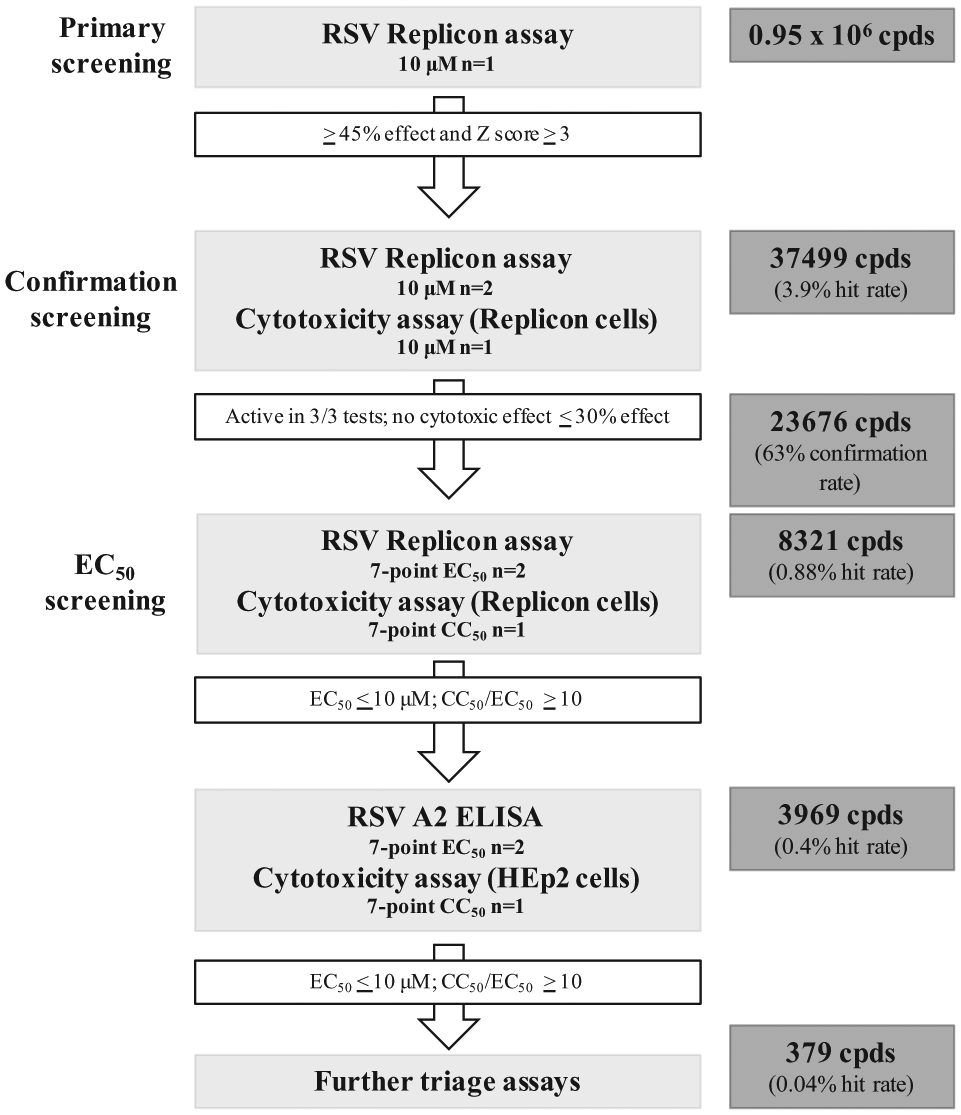
Screening cascade for campaign 2. Numbers of compounds tested in each step are shown, and the hit rate from each assay is calculated as a percentage of the original compound collection of 0.95 million. RSV, respiratory syncytial virus.
Discussion
The identification of efficacious RSV therapeutics has been hindered by the lack of a robust and appropriate primary assay for ultra-HTS. To enable screening of large compound libraries, an assay should ideally be simple, with the minimal number of operations, and using automation where possible. The length of time to perform the assay is also of importance, as lengthy incubations result in long lead times to generate data. The use of assay-ready plates separates the compound delivery from the assay assembly, which simplifies and streamlines the screening process. The availability of a cryopreserved cell bank reduces the resource requirements for screening at scale and can also enhance standardization of assay signal and reproducibility. The cost of an HTS is a further consideration, as expensive reagents can prohibit the screening of large numbers of compounds. Finally, the biosafety level requirements should be carefully considered, since higher biological containment increases the complexity of screening.
The RSV replicon assay does not require the use of live virus, reducing the biocontainment requirements for screening. The assay is also amenable to automation, enabling screening of large numbers of compounds. During our first screening campaign using the replicon assay, we encountered several challenges, including a variable assay with multiple steps, a requirement for the use of continuously cultured cells, and a limit to the compound throughput due to the cost of EnduRen substrate. Through further assay optimization and improvements in cryobank generation techniques, we addressed these challenges and prosecuted a successful screening campaign, delivering high-quality data for 1 million compounds on an automated platform. We selected to screen at 10 µM to try to balance on-target activity against nonspecific cytotoxic effects and to avoid solubility issues that can occur at higher concentrations. The validation data 10 also showed a sufficiently high active rate to allay fears that suitable equity would be found at this concentration.
There are a number of potential ways to screen for RSV inhibitors at scale. Advances in our knowledge around the mechanism of viral life cycles have led to a number of viral proteins being identified as potential targets for drug therapy. Targeting isolated viral proteins such as RNA-dependent RNA polymerase 16 and the viral F-protein 17 using biochemical assays at scale have been reported in the literature. These types of screens are readily amenable to scale and do not pose the biological safety hazards associated with live viral assays. However, they have the disadvantage of targeting only one viral mechanism, whereas the replicon assay targets several.
Several HTS campaigns have been performed to identify RSV inhibitors. Functional cell CPE assays have been used to screen large numbers of compounds.18,19 Whole-cell phenotypic assays are not limited to targeting a single mechanism and have the additional advantage of affecting either viral or host systems. They are also generally more physiologically relevant than isolated enzyme assays, but such assays require lengthy incubation times of several days. This can present challenges, including edge effects due to evaporation, storage of assay plates containing functional virus at the appropriate biocontainment level, and assays more prone to issues such as cytotoxicity. Since the replicon assay has a shorter 48-h incubation period, the impact of these factors is reduced. The mechanism of action of inhibitors identified in phenotypic screening can also be challenging to deconvolute. Cellular antiviral screens can also have biological safety and viral instability 20 considerations if live RSV is employed. The cell line used in the replicon assay does not release infective viral particles, and the deletion of some cytopathic proteins means cells can be cultured over multiple passages without compromise to their viability.
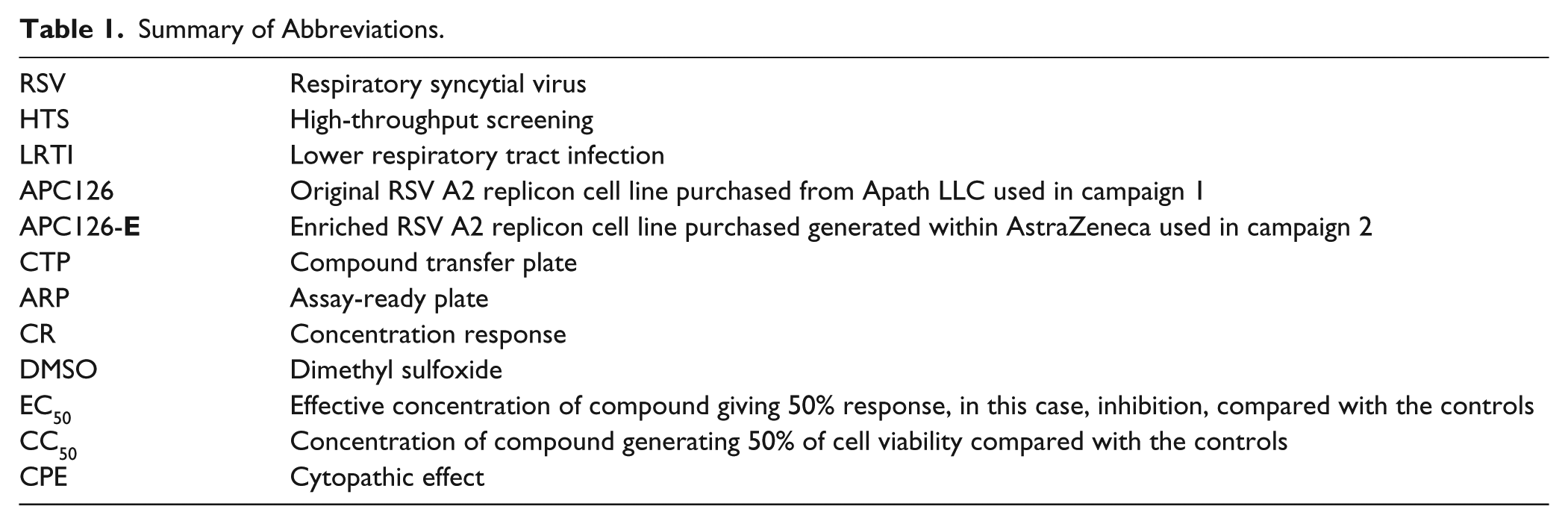
Summary of Abbreviations.

RSV messenger RNA transcription and replication are driven by cooperative interactions between the viral nucleocapsid (N) protein, large viral polymerase (L) protein, phosphoprotein (P), and M2-1 protein. 21 The replicon cell line used in this screen uses a modified viral genome encoding these proteins essential for viral replication. Viral proteins associated with viral entry and RSV-induced cytotoxicity such as the viral fusion (F) protein, attachment glycoprotein (G), and small hydrophobic (SH) glycoprotein are removed and replaced with a luciferase reporter gene. This assay will therefore identify inhibitors of viral replication that target either the pathogen or the host, without targeting viral entry or assembly mechanisms, a potential drawback of the system. However, given that the vast majority of successful antivirals are targeting viral replication compared with viral entry inhibitors,15,22 as well as the robustness and convenience of the replicon assay, we believe that the advantages of this screening approach outweigh this drawback and that the use of the replicon assay may be more desirable to identify attractive chemical start points. We expect the antivirals identified by this method to work against multiple strains of RSV within the same subtype (A or B), as the viral targets involved in viral replication are quite conserved within the same subtype. Whether the inhibitors work on strains from different subtypes would depend on their mechanisms of action and how conserved the compound binding pockets are between the two subtypes.
The risk to using a recombinant system within a rodent cell line is that we identify active compounds that do not translate across into functional viral assays. To demonstrate that the screening cascade was able to identify a bona fide RSV inhibitor, we tested an analogue of YM-53403 in all assays used and obtained EC50 values that were in line with published data. 11 Through use of a medium-throughput live virus ELISA as part of the screening cascade, we identified four novel compound series of interest. The potential of screening with this system was further endorsed as it resulted in the identification of several nucleoside-like compounds from within the AstraZeneca collection that were flagged as active. Nucleosides are a common class of antivirals, and there are currently over 25 approved analogues that are used to treat several viral infections such as hepatitis, HIV, and herpes. 23 These data together validate the use of recombinant replicon-based cell systems in HTS and highlight the potential to successfully employ this system in drug discovery for viruses other than RSV and dengue. 24 In conclusion, the development of a replicon-targeted assay along with the deployment of a robust hit generation strategy has led to the successful identification of new tractable RSV inhibitors with the potential to become new antiviral therapies.
Footnotes
Acknowledgements
We thank Paul Fleming and Pamela Hill for their evaluation of the output of the HTS screening campaign. We also thank Ruben Tommasi and Mark Wigglesworth for their comments and scientific input during the preparation of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
