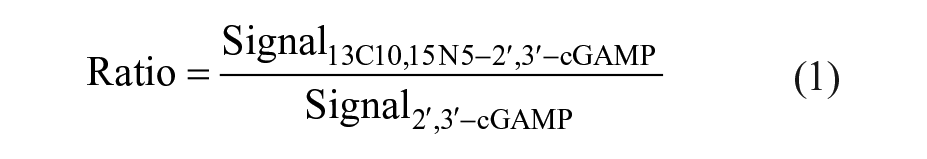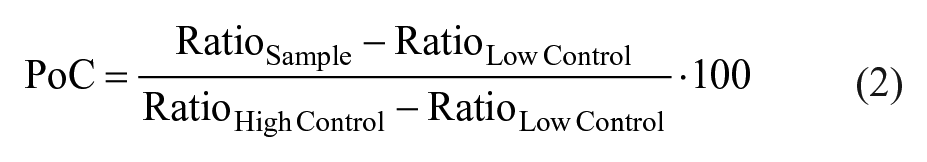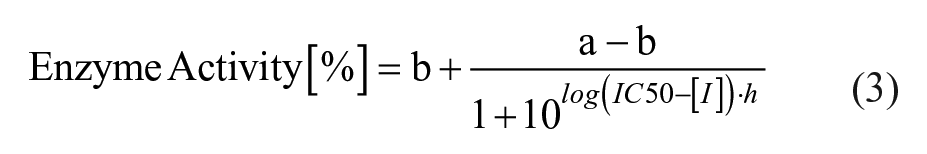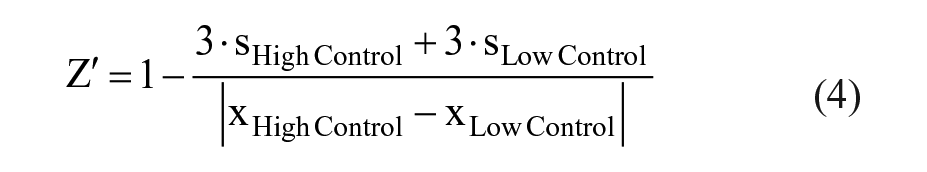Abstract
Comprehensive and unbiased detection methods are a prerequisite for high-throughput screening (HTS) campaigns within drug discovery research. Label-free matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry (MS) has been introduced as an HTS-compatible readout for biochemical test systems to support the drug discovery process. So far, reported HTS applications were based on surface-modified systems or proof-of-concept studies. We present the utilization of a MALDI-TOF-based screening platform to identify inhibitors of human cyclic GMP-AMP synthase (cGAS), a mediator of innate immune response whose aberration has been causally correlated to a number of inflammatory disorders. In this context, the development and validation of a MALDI-TOF-based activity assay is reported to demonstrate fast, robust, and accurate detection of chemical cGAS inhibition by direct quantification of the physiological reaction product cyclic GMP-ATP (cGAMP). Results from a screen of a diverse library of more than 1 million small molecules in 1536-well format against the catalytic cGAS activity are presented with excellent assay performance and data quality. Identified hits were qualified in dose–response experiments and confirmed by RapidFire-MS measurements. Conclusively, the presented data provide the first proof of applicability of direct automated MALDI-TOF MS as a readout strategy for large-scale drug discovery HTS campaigns.
Introduction
The cyclic GMP-AMP synthase (cGAS) is a mammalian cytosolic pattern recognition receptor for aberrantly modified, foreign, or mislocalized double-stranded DNA (dsDNA) and a main driver of innate immune response to infectious or cellular stress stimuli.1–5 Upon binding to dsDNA, cGAS forms a 2:2 cGAS/dsDNA complex and a conformational change within the catalytic core region is initiated that rearranges the active site of the enzyme and allows for binding of the two nucleotide substrates ATP and GTP. 6 The enzyme then exerts its catalytic activity in a two-step catalysis mechanism that involves the formation of a 2′,5′-GpA-linkage followed by a 3′,5′-ApG cyclization step of the linear dinucleotide at the same active site to produce the final product cyclic GMP-AMP [G(2′,5′)pA(3′,5 ′)p], further referred to as 2′,3′-cGAMP (cyclic GMP-ATP).7,8 This mammalian-specific isomer serves as an inflammatory second messenger by binding and activating stimulator of interferon genes (STING) at the endoplasmatic reticulum (ER) membrane.9,10 As a consequence of 2′,3′-cGAMP binding, STING translocates to the Golgi apparatus, where it serves as a signaling platform ultimately inducing the transcription of type 1 interferon (IFN) and proinflammatory cytokine genes.3,10–12 Perturbations and misregulation of the cGAS-STING pathway have been causally correlated with a number of inflammation-driven pathological phenotypes including Aicardi–Goutières syndrome (AGS), systemic lupus erythematosus (SLE), systemic sclerosis (SSc), and myocardial infarction (MI).13–21
The many implications of the cGAS-STING signaling axis in inflammatory disease pathogenesis suggest that chemical cGAS inhibition could be an effective strategy for therapeutic approaches. Recent advances regarding this topic have used in vitro screening assays or fragment-based approaches to identify small-molecule inhibitors in medium throughput.22–24 Despite showing submicromolar activity on the isolated enzyme under in vitro conditions, the further development of these substances is hampered by their limited cellular activity and considerable off-target effects. Therefore, novel lead structures are desired to be developed into more specific and cell-active small-molecule cGAS inhibitors. A promising approach to identify potential lead structures is unbiased screening of structurally diverse compound libraries at high throughput. One reason for the confined number of reported efforts to target the catalytic function of cGAS so far is the lack of sensitive and high-throughput-compatible assay systems.
Because of its unselective spectrophotometric properties and the lack of suitable label attachment sites, cGAMP is a rather challenging analyte to track with conventional in vitro screening methods. Published test systems use either fluorescence polarization, 23 luminescence, 22 or mass spectrometry (MS)-based readouts22,24 to quantify biochemical cGAS activity, all of which have individual hurdles regarding their high-throughput screening (HTS) application. The use of fluorescence polarization relies on competitive displacement of a labeled cGAMP derivative from a receptor. Elaborative production of the chemical tracer molecule and a specific monoclonal antibody are therefore required, both of which are not commercially available to date. The adaptation of available luminescence-based assays to track the ATP consumption by cGAS is straightforward but does not provide the desired sensitivity, which hampers assay miniaturization and increases protein consumption. MS-enabled direct detection of substrate mass shifts as a consequence of enzymatic activity provides a label-free and unbiased approach, granting access to analytes challenging to address with conventional methods. So far, commonly used electrospray ionization (ESI) systems have been employed for cGAS inhibitor screening.22,23 However, these methods require analysis times of more than 10 s/sample, which is not compatible with cycle times required for the HTS of large compound libraries.
The conjunction of matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) MS with chemically modified target surfaces has convincingly demonstrated the feasibility of the comprehensive MS-based assay readout for HTS applications.25–27 Immobilization and enrichment of analytes at the MALDI target plate surface enables the use of stringent washing procedures prior MALDI-TOF analysis, which results in increased sensitivity while maintaining HTS-compatible capacity. 28 Recent innovations in MALDI-TOF MS and liquid handling instrumentation have provided significant advancements in sample preparation and throughput of this technology. Triggered by these progressions, the feasibility of direct MALDI-TOF MS as an assay readout strategy, independent from target surface chemistry, has been evaluated in a number of studies.29–33 We have recently demonstrated the successful integration of fast direct MALDI-TOF MS into the HTS environment and its adaptation as a versatile and label-free readout strategy for in vitro enzyme assays.34–36
In this work, we advance beyond the proof of concept and demonstrate its application in a large-scale full-diversity HTS campaign against the catalytic activity of human cGAS. MS-based direct and label-free quantification of cGAMP was used to track compound-dependent enzyme inhibition at high speed. Besides the optimization of relevant assay parameters and the assessment of critical assay performance characteristics, the results from a HTS library screen of more than 1 million small molecules in 1536-well format are presented and discussed. Robust and sensitive analyte detection and high system stability, as proposed by proof-of-concept studies, were confirmed. Conclusively, our study demonstrates the first proof of applicability of MALDI-TOF MS as an in vitro assay readout strategy in extensive maximum-diversity HTS campaigns.
Materials and Methods
Materials
HTS MALDI disposable target plates (no. 1847006) and an HTS MALDI adapter (no. 8283496) were purchased from Bruker Daltonics (Billerica, MA). Recombinant human cGAS (cGAS-2-522-N-His-SUMO-BV) was produced in High Five insect cells in-house at Boehringer Ingelheim (Biberach an der Riß, Germany). α-Cyano-4-hydroxycin-namic acid (4-HCCA; no. 70990), isotope-labeled 13C10, 15 N5-GTP substrate (no. 645680), 45 bp DNA oligos (custom synthesis; for exact sequence see Supplemental Information), 2′,3′-cGAMP (no. SML1229) internal standard, acetonitrile (no. 34851), DMSO (no. D5879), formic acid (no. 33015), and isopropanol (no. 34863) were obtained from Sigma Aldrich (St. Louis, MO). ATP (no. V915B) substrate was purchased from Promega (Madison, WI). Tween 20 (no. 9127.1) and trifluoroacetic acid (TFA; no. 6957) were purchased from Carl Roth (Karlsruhe, Germany). Bovine serum albumin (BSA; no. 11945.03) and dithiothreitol (DTT; no. 20710.04) were obtained from Serva (Heidelberg, Germany). Tris (no. 108382) and MgCl2*6 H2O (no. 105833) were obtained from Merck Millipore (Burlington, MA). KCl 2 M solution (no. AM9640G) was purchased from Thermo Scientific (Waltham, MA).
Expression and Purification of Recombinant cGAS
Human cGAS (amino acids 2–522) with an N-terminal 6× His-tag followed by a TEV cleavage site was expressed in High Five insect cells. Cells were lysed in buffer containing 20 mM HEPES (pH 7.5), 400 mM NaCl, 10 mM imidazole, 10 % glycerol, 1 mM TCEP, protease inhibitor cocktail (cOmplete, EDTA-free, Roche, Basel, Switzerland), and DNase (10 µg/mL). The cGAS protein was isolated by affinity chromatography on Ni-NTA agarose resin and further purified on a HiTrap Heparin HP (GE Healthcare, Chicago, IL) column, followed by size exclusion chromatography using a Superdex 200 column (GE Healthcare) equilibrated in 20 mM HEPES (pH 7.4), 250 mM KCl, 10 % glycerol, and 1 mM TCEP. Purified protein was concentrated to 2 mg/mL and stored at −80 °C.
Biochemical cGAS Assay
Single-stranded DNA oligos were mixed in equimolar ratio and subjected to a standard annealing procedure to yield dsDNA. Enzymatic reactions were set up in assay buffer containing 10 mM Tris (pH 7.5), 10 mM KCl, 5 mM MgCl2, 1 mM DTT, 0.01% BSA, and 0.001% Tween 20. First, 50 nL of test compound (5 µg/mL final) or DMSO was placed into the wells of a 1536-well plate (Greiner Bio-One, Kremsmuenster, Austria; no. 782101) using a CyBio Well vario (Analytik Jena, Jena, Germany) liquid handling unit equipped with a capillary head. Therefore, four 384-well substance plates were combined into one 1536-well assay plate. Next, 2 µL of a 2.5× concentrated intermediate mixture of cGAS enzyme and dsDNA (12 nM and 0.32 µM final concentrations, respectively) was added. The plates were then incubated for 10 min in a humidified chamber at 24 °C. Subsequently, 3 µL of an equimolar ATP/13C10, 15 N5-GTP mixture (60 µM final concentration each) was added to each well and the plates were incubated for 1.5 h at 24 °C. After incubation, the enzymatic reaction was stopped by adding 1 µL of 0.6% formic acid (v/v) containing the 2′,3′-cGAMP internal standard (1 µM final concentration). Dispensing steps were executed with the aid of a Certus Flex Micro Dispenser (Gyger, Oensingen, Switzerland). The plates were then sealed with an adhesive foil and stored at room temperature until preparation of the MALDI target plates. Each 1536-well assay plate contained high (no compound; columns 45–46) and low (no compound; assay buffer instead of enzyme/dsDNA mixture; columns 47–48) controls to follow the enzymatic reaction. For dose–response experiments, eightfold dilution series of compound solutions were prepared in DMSO in 1:5 or 1:3.16 dilution steps starting from 10 mM or 5 mg/mL stock solutions, respectively.
MALDI Target Preparation
4-HCCA was dissolved in a 1:1 mixture of acetonitrile and 0.05% TFA (v/v) with the aid of vigorous vortexing until a saturated solution was obtained. The CyBio Well vario liquid handling system (Analytik Jena) equipped with ceramic tips and operated in 1536-well format was employed to conduct double-layer spotting providing highly homogeneous spot shapes. Here, 100 nL of matrix solution was spotted onto plain steel MALDI target plates and dried in a vacuum chamber. Subsequently, assay plates were centrifuged at 1000 rpm for 60 s and the seals were removed before 100 nL of matrix solution and 100 nL of sample were aspirated successively from the matrix reservoir and the assay plate, respectively, and dispensed together onto the dried matrix spots. The MALDI target plate was then dried under vacuum and stored until analysis. Between the first and second spotting step, ceramic tips were washed three times with 70% isopropanol and 30% water/0.1% TFA (v/v). After successful transfer of the matrix–analyte–mixture, three repetitive washing cycles using 0.1 M NH4OH followed by three cycles of 70% isopropanol and 30% water/0.1% TFA (v/v) were carried out in order to prevent carryover and clogging of the tips.
MALDI-TOF-Based HTS Assay Readout
Mass spectra were acquired with a rapifleX MALDI-TOF/TOF instrument from Bruker Daltonics including a Smartbeam 3D laser. FlexControl (v4.0), FlexAnalysis (v4.0), and MALDI PharmaPulse (v2.0) were used for MS acquisition and data analysis. Target plates were loaded onto an Orbitor RS (Thermo Scientific) robotic system controlled by the laboratory automation software Momentum (v4.2.3, Thermo Scientific) and automatically inserted into the MALDI-TOF device. Gantt charts of the automated plate handling for the biochemical assay, as well as the spotting procedure and MALDI TOF measurements, are depicted in
Data Analysis
MALDI-TOF data, processed with flexAnalysis or MALDI Pharma Pulse, were exported as a comma delimited (.csv) file. Data sets were further processed with either GraphPad Prism (v7.03; GraphPad Software, La Jolla, CA) or in-house software MegaLab. cGAS activity was tracked by analyzing the measured area or intensity (signal) for the enzymatic product (13C10, 15 N5-2′,3′-cGAMP) as well as for the corresponding unlabeled internal standard (2′,3′-cGAMP). During the primary screening campaign against cGAS activity, compound-containing wells where no internal standard could be detected were automatically excluded from further analysis and the respective compounds were retested at a different occasion. Control wells without detectable internal standard were excluded from analysis. The ratio of the signals of the reaction product and the internal standard was calculated in order to diminish variations ascribed to the MS readouts according to following equation:
The average of ratio values was calculated for high controls (RatioHigh Control) and low controls (RatioLow Control), respectively, for every substance plate. Generated sample data were then normalized and expressed as the percentage of control (PoC) according to the following equation:
IC50 calculation was performed in MegaLab by fitting the data to the following four-parameter logistical equation:
where a is the uninhibited value, b is the completely inhibited one, [I] is the inhibitor concentration, h is the Hill slope, and IC50 is the half maximal inhibitory concentration. GraphPad Prism was utilized for graphical representation of the data.
The assignment of compounds to the corresponding measurements was achieved by software-aided deconvolution of every 1536-well assay plate to the corresponding 384-well substance plates in the course of data analysis, as we have described before. 34
Basic appraisal of the assays was reviewed by the statistical quality parameter Z′ with the following equation: 37
where s is standard deviation and x is the mean readouts of the mass spectrometric assays of individual 384-well plates.
RapidFire-MS-Based Measurement of cGAMP
An API6500 (AB Sciex, Darmstadt, Germany) was equipped with a RapidFire (Agilent, Waldbronn, Germany) and a liquid chromatography pump of the 1290 series (Agilent). Solvent A consisted of 10 mM ammonium acetate (pH 7.4), which was delivered by pumps 1 and 2. Solvent B consisted of 47.5% acetonitrile, 47.5% methanol, and 5% water supplemented with 10 mM ammonium acetate, which was delivered by pump 3. Pump 1 delivered 1.5 mL/min and pumps 2 and 3 delivered 1.25 mL/min, respectively. The sample was aspirated for 250 ms into the 10 µL sample loop and trapped onto a C18 trap column (Agilent Technologies, Santa Clara, CA, no. G9526-80000) with 12 µL of bed volume for 3000 ms with solvent A. Subsequently, solvent B was used to elute the analytes by a 4000 ms step. The MS was operating in the positive electrospray mode with a curtain gas of 35, a spray voltage of 4200 V, a source temperature of 550 °C, and an ion source gas 1/2 of 65/80 psi at unit resolution. The following transitions were monitored: 13C10, 15 N5-cGAMP at Q1: 690 m/z, Q3: 529 m/z, DP: 130 V, CE: 30 V; and cGAMP at Q1: 675 m/z, Q3: 524 m/z, DP: 130 V, CE: 30 V.
Test for Isobaric Compound Interference
For the analysis of potential compound interference with the internal standard signal, eight replicates of each candidate compound were first transferred to wells of an assay plate, as described for the biochemical cGAS assay. Then, 2 µL of assay buffer, 3 µL of an equimolar ATP/13C10, 15 N5-GTP mixture (60 µM final concentration each), and 1 µL of 0.6% formic acid (v/v) were added subsequently to every well. For control wells (n = 16), DMSO was transferred instead of compounds and the formic acid solution was supplemented with 6 µM 2′,3′-cGAMP (1 µM final concentration). The samples were spotted onto MALDI target plates and measured as described above. The average of signal intensities at m/z = 673.1 was calculated for every candidate compound and referenced against the average of internal standard signals from control wells. A 5% threshold was applied for assigning candidates to isobaric interfering compounds.
Results
Reliable, fast, and unbiased readout strategies are a prerequisite for HTS-capable assay workflows within the hit discovery process. We used a well-established MALDI-TOF screening platform 35 for the development of an in vitro activity assay for the direct detection and quantification of 2′,3′-cGAMP, the enzymatic reaction product of the NTase cGAS activity, to initiate a screening campaign against this promising drug target.
Setup of the MALDI-TOF-Based In Vitro cGAS Activity Assay
In the presence of dsDNA, cGAS links the two substrates ATP and GTP to generate the cyclic dinucleotide 2′,3′-cGAMP. Because of the lack of commercially available isotope-labeled 2′,3′-cGAMP to be used as an internal standard, we decided to employ stable isotope-labeled GTP (13C10,
15
N5-GTP) in combination with ATP as substrates, which yields 13C10,
15
N5-cGAMP ([M-H]– = 688.1103 Da) as the enzymatic reaction product (
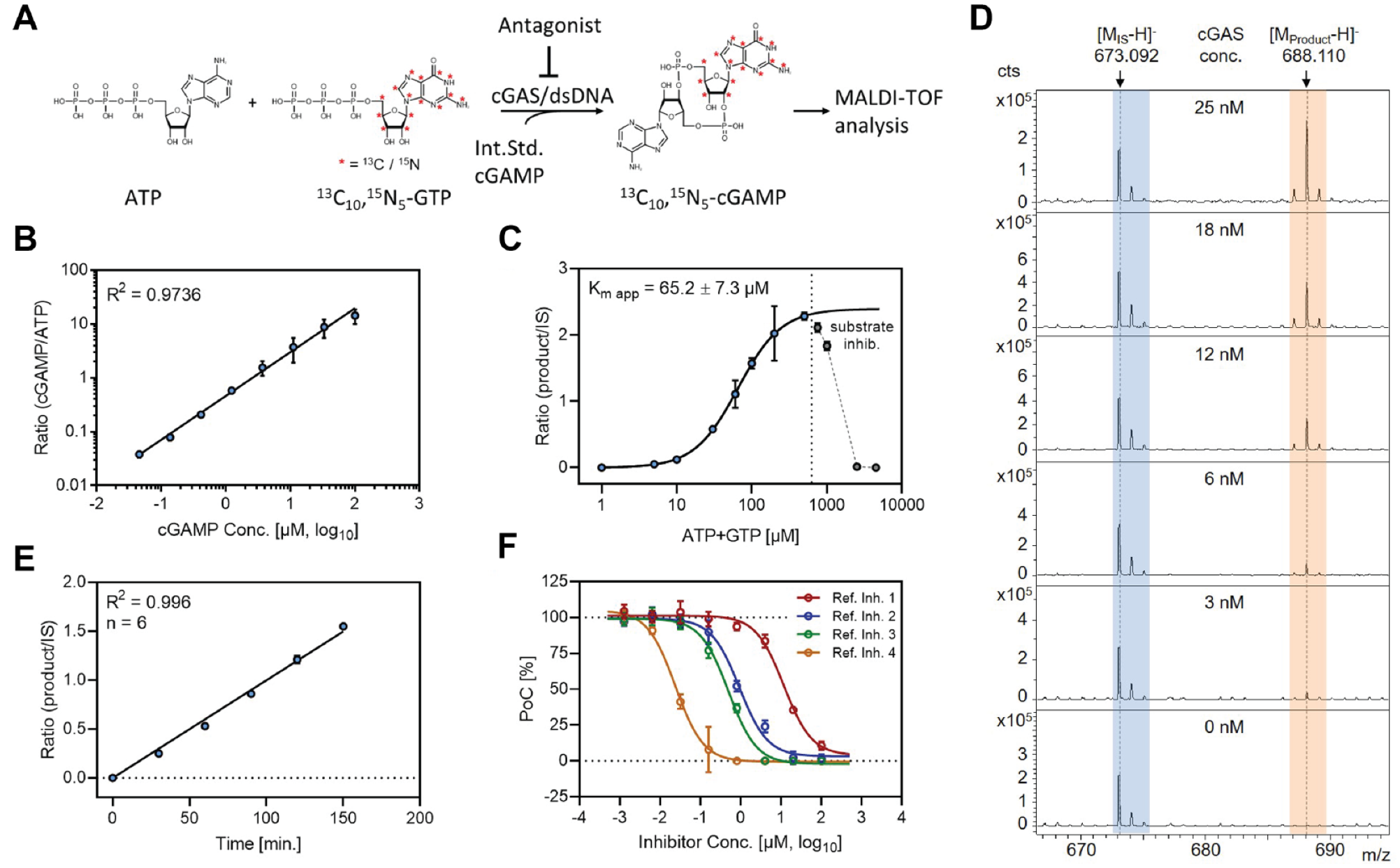
Setup of the MALDI-TOF-based in vitro cGAS activity assay. (
The enzyme displayed an increase in 13C10,
15
N5-cGAMP production that was dependent on the presence of dsDNA in the incubation mixture (
Evaluation of Assay Performance Characteristics
Based on defined assay parameters, basic assay statistics to validate the reproducibility, robustness, and stability of the MALDI-TOF-based cGAS assay were determined. Therefore, a representative number of DMSO controls were dispensed in 1536-well plates and processed by the automated MALDI target spotting system and the MALDI-TOF MS device. During the analysis of the acquired data, we observed random variations in the baseline signal that caused minor deviations in the integration procedure for a small subset of sample spots. We found that it was possible to circumvent these effects via data evaluation based on the intensity values (
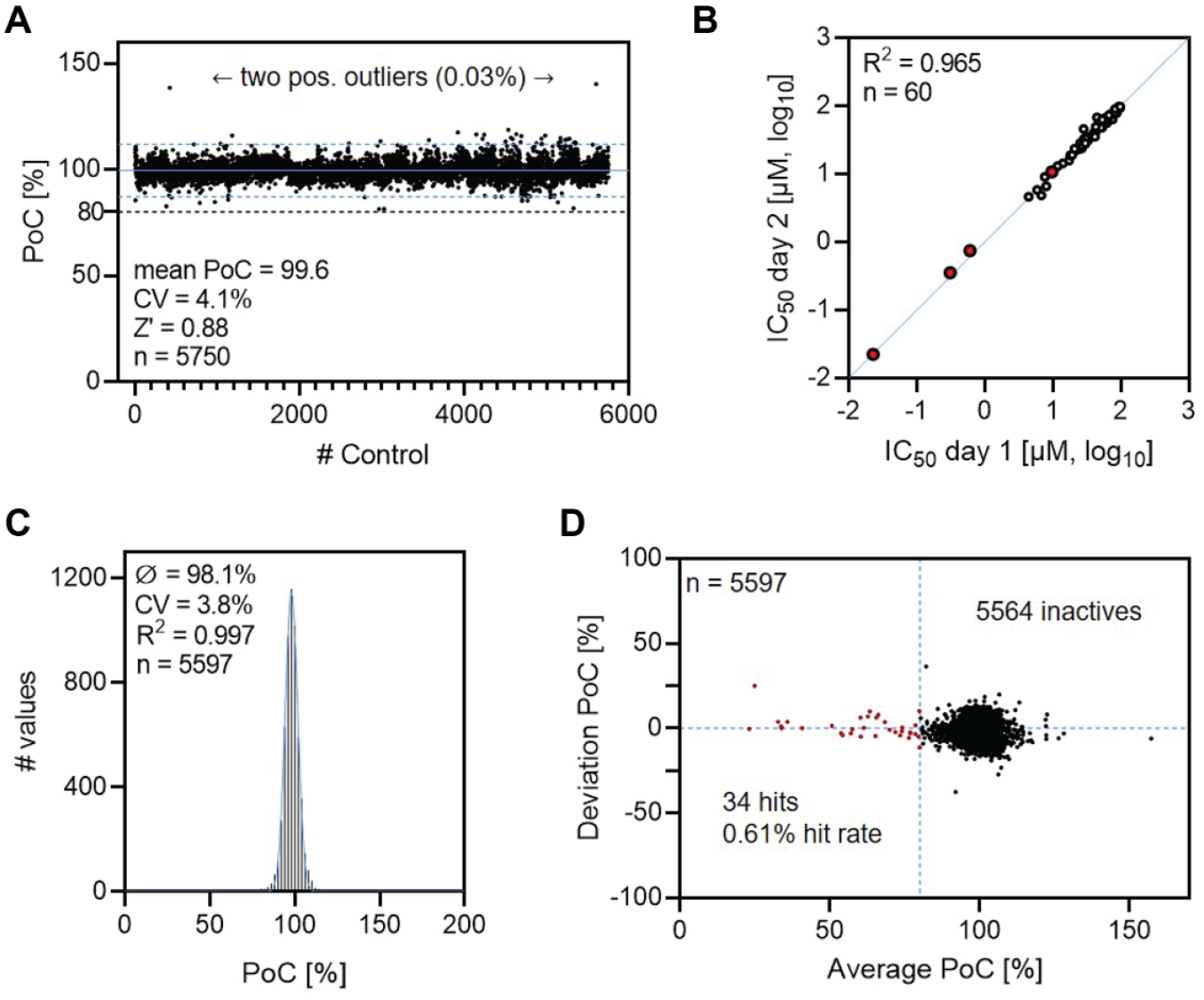
Results of the assay characterization and validation procedure. (
Following the confirmation of assay stability, dilution series of reference compounds and a set of 58 promiscuous enzyme inhibitors were tested against cGAS in two individual experiments to determine the reproducibility of the assay. Comparing the results of IC50 determinations from different assay days revealed a high interday reproducibility (R
2
= 0.965), further emphasizing the robustness of the test system (
To provide evidence that our system is suitable for screening cGAS inhibitors under HTS conditions, a validation set of 5597 chemically diverse compounds were selected from the in-house library and subjected to the assay procedure. Following the biochemical assay, MALDI target preparation, measurement, and data analysis, the scattering pattern of measured signals exhibited a good fit to a Gaussian distribution (R2 = 0.997), as illustrated in
MALDI-Based High-Throughput cGAS Inhibitor Screening
The main library screen was conducted in fully automated fashion with spatial separation of the biochemical assay, the MALDI target spotting system, and the MALDI-TOF MS device. An average of ~60,000 samples were tested per day by parallelized execution of the biochemical assay, MALDI target spotting, and MALDI-TOF MS analysis for batch sizes up to 45 assay plates. DMSO control plates and dilution series of reference compounds were included in every screening batch to ensure consistent assay performance.
The complete screening campaign took 20 working days during which we processed and analyzed 1,132,365 samples (1,080,879 compounds, 51,486 controls) with high data quality. An average Z′ value of 0.90 with low interplate variability (3.8% CV) was obtained across 3164 tested 384-well substance plates (
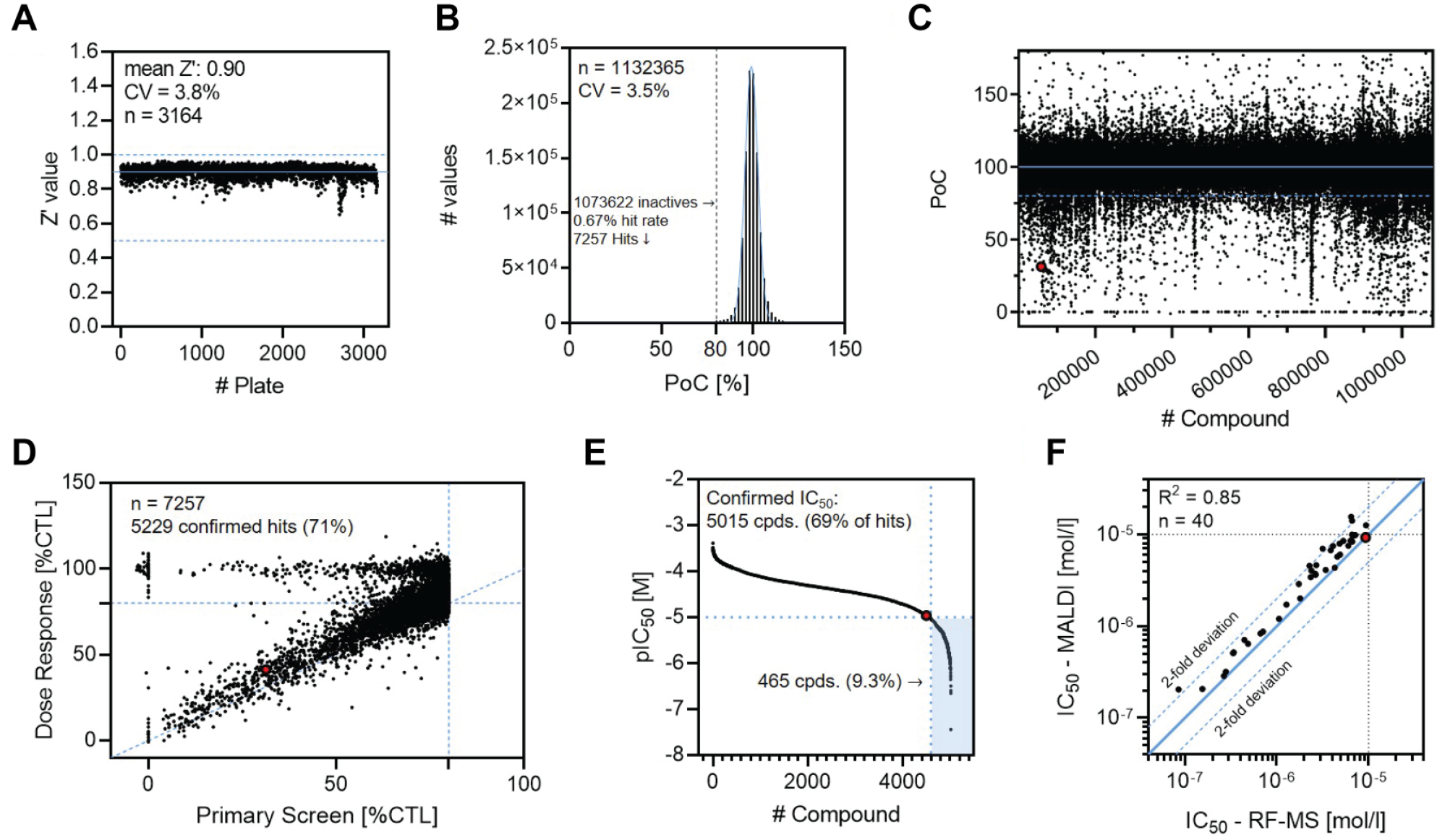
Results from the cGAS HTS assay campaign and hit triaging. (
Hit Triaging and Confirmation
A total of 7257 compounds (0.67% hit rate) reducing cGAS activity by ≥20% were identified, a number that is well in line with the approximations based on the validation run (
MS-based detection of cGAS activity using RapidFire-MS (RF-MS) is a well-established method described for inhibitor screening and profiling approaches with medium throughput.22,24 We repeated the dose–response experiments for the 40 selected compounds with RF-MS-based detection and compared the resulting IC50 values with MALDI-TOF data. The compound profiles showed a good correlation (R2 = 0.85) between the two individual methods with a slight tendency to higher half inhibitory concentrations for the MALDI-TOF readout (4 compounds outside two-fold deviation;
Potential False Positives and Assay Interference
Although the MALDI-TOF-based readout is largely unbiased and interference tolerant, we observed a compound population that did not reproduce their inhibitory effects from the primary screening during dose–response experiments (see
First, during the primary screening campaign, we obtained an average ratio of 0.64 (intensity of signals [product/internal standard]) for high control wells with a variability of 3.5% CV for ratio values and >30% CV for the individual signals. The high intrinsic signal variability during MALDI-TOF analysis is the result of fluctuating ionization efficiencies on different sample spots and is efficiently compensated by the use of an internal standard. We found that in presence of an excess internal standard compared with the analyte, low ionization efficiencies could reduce the signal intensity of the analyte below the detection limit while the internal standard remains measurable, therefore producing a false-positive assay hit. This effect was the presumable cause for a small pool of compounds (102 compounds, 0.009% of samples, 1.4% of hits) that showed around 100% inhibition in the primary cGAS screen but was completely inactive in the dose–response experiments (
In the course of data analysis, we observed a second mechanism of potential assay interference. The molecular weight of 2′,3′-cGAMP and the tracked isotope-labeled analyte are well within the mass range of the tested screening library. Isobaric signals arising from test substances or one of their isotopes could therefore produce false-positive (if isobaric with internal standard) or false-negative (if isobaric with product) results. To evaluate the proportion of this interference mechanism on our set of screening hits, we calculated the exact mass values for the first three isotopes of each screening hit. Candidates showing values within the defined peak picking delta of 0.3 (m/z) with respect to the internal standard were considered as potentially interfering compounds (36 compounds; 0.5% of hits) and reordered for separate analysis. The substances were then incubated under low control conditions and measured without addition of the internal standard. Four compounds (interferences 1–4) were detected under the applied conditions (11% of potential interfering compounds), which produced average MALDI-TOF signals of 7%–65% compared with the internal standard. A plot of the normalized signal intensities of interfering compounds and representative mass spectra is included in
Discussion
In summary, we developed a stable, sensitive, and robust MALDI-TOF-based in vitro assay, which was used to perform an HTS campaign against the NTase activity of human cGAS. This was achieved by employing a fully automated MS-based screening platform that was integrated into an HTS environment. The MALDI-TOF readout is capable of providing sensitivity, speed, and robustness that allow for HTS-compatible analysis times and low consumption of assay components, thereby omitting the limitations of alternative cGAS assay technologies for HTS applications. Although initial financial expenses for such systems are higher than for conventional biochemical methods, significant cost reductions can be achieved during screening campaigns due to assay miniaturization and omission of detection reagents or antibodies. Additionally, the direct analyte detection by MS-based methods is independent of functional labels and artificial signal mediators, which has the potential to reduce time and costs for assay development compared with conventional methods. The label-free concept further provides a more physiologically relevant readout and grants access to target and analyte classes that are challenging or unsuitable to address with label-based methods. While recent developments in the field of ESI-based MS suggest promising potential for HTS, MALDI-TOF is the only commercially available MS-based and high-throughput-compatible technology to date.38,39
The developed cGAS assay was profoundly validated and subsequently used to screen a full-diversity compound library with excellent assay statistics. To the best of our knowledge, this is the first report of a large-scale (>106) HTS campaign using a MALDI-TOF-based readout strategy. The throughput of ~60,000 samples per day was achieved by fully automated execution on three distinct workstations for the biochemical assay, MALDI target plate preparation, and MALDI-TOF analysis. This setup allowed us to optimize parameters of each part individually and enabled independent equipment maintenance. Consistent assay performance throughout the screen emphasized the sustainable potential of MALDI-TOF MS as a readout strategy for HTS applications. Further, identified assay hits were efficiently triaged and confirmed by RF-MS, demonstrating the accuracy and versatility of the MALDI-TOF-based readout. In ensuing steps, the activity of freshly synthesized material, cellular potency, target engagement, and potential DNA intercalating mechanisms should be investigated in downstream assays to further confirm and profile the biochemical compound activities.
Finally, we identified low ionization efficiency and isobaric compound interference as two potential mechanistic sources of false-positive/negative assay hits. The overall contributions of these effects to the screening results were low (<2% of hits) and possible corrective measures were provided. Altogether, the presented data demonstrate the combination of sensitivity, speed, and robustness provided by MALDI-TOF MS under true HTS conditions and establish the described screening platform as a valuable and versatile tool for drug discovery campaigns targeting the NTase activity of cGAS.
Supplemental Material
Supplemental_Material_for_cGAS_HTS_Simon_et_al – Supplemental material for MALDI-TOF Mass Spectrometry-Based High-Throughput Screening for Inhibitors of the Cytosolic DNA Sensor cGAS
Supplemental material, Supplemental_Material_for_cGAS_HTS_Simon_et_al for MALDI-TOF Mass Spectrometry-Based High-Throughput Screening for Inhibitors of the Cytosolic DNA Sensor cGAS by Roman P. Simon, Martin Winter, Carola Kleiner, Robert Ries, Gisela Schnapp, Annekatrin Heimann, Jun Li, Ljiljana Zuvela-Jelaska, Tom Bretschneider, Andreas H. Luippold, Wolfgang Reindl, Daniel Bischoff and Frank H. Büttner in SLAS Discovery
Footnotes
Acknowledgements
We thank Helmut Romig, Julian Rasch, and Markus Holstein for their support with RapidFire-MS experiments. Further, we thank Cedrickx Godbout, Patrick Groß, Christoph Hoenke, Sandra Handschuh, and Dirk Reinert for helpful discussion. We also thank Helga Bronner and Ursula Schmid for their support of the HTS campaign.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors are employees of Boehringer Ingelheim Pharma GmbH & Co. KG and completed this work within the scope of their employment.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: All work was completed as work-for-hire for the employer Boehringer Ingelheim Pharma GmbH & Co. KG.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.