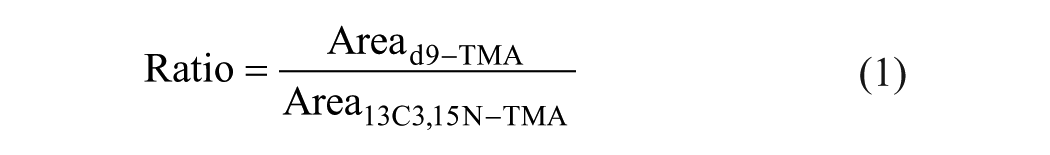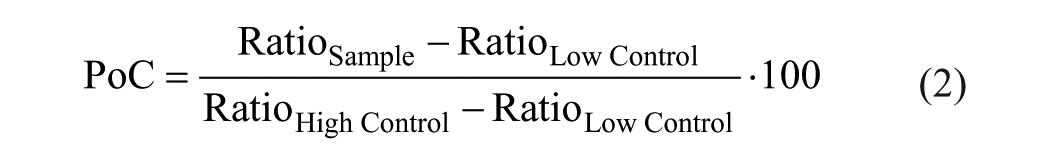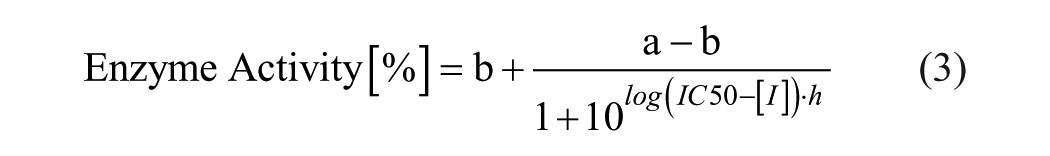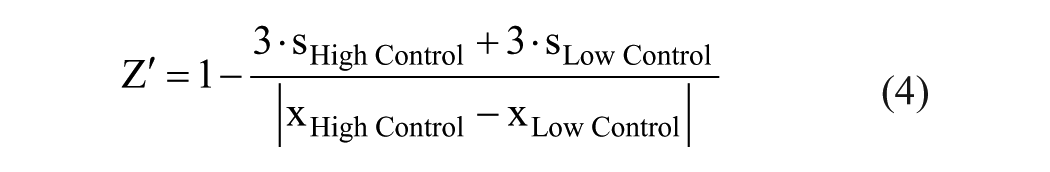Abstract
Microbial-dependent trimethylamine (TMA) generation from dietary precursors such as choline was recently linked to cardiovascular diseases (CVDs) as well as chronic kidney disease (CKD). Inhibition of TMA-generating enzymes in gut bacteria would be an innovative approach to treat these diseases. The potential to accurately quantify secreted TMA levels highlights the capacity of mass spectrometry (MS) for tracking microbial TMA-lyase activity. However, high-throughput screening (HTS) by conventional MS instrumentation is hampered by limited sample throughput. Recent advancement in liquid handling and instrumentation of matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) MS provides an HTS-compatible MS technology. The deciphering of enzymatic reactions using this label-free readout has been successfully applied but has thus far been limited to peptide/protein-centric activity assays. Here, we demonstrate the versatile applicability of MALDI-TOF by tracking a small molecule within a highly complex sample background. The key to success for this concept was chemical derivatization of the target molecule enabling quantitative assessment of microbial TMA formation. Further, its potential was demonstrated in a side-by-side comparison to RapidFire-MS in a primary screen and subsequent dose–response experiments. Overall, the established assay enables the screening for microbial TMA-lyase inhibitors and serves as a proof of concept for the applicability of MALDI-TOF for demanding assay concepts per se.
Introduction
Recent studies demonstrate a linkage between the gut microbiome and cardiovascular diseases (CVDs) as well as chronic kidney disease (CKD). 1 One important pathway is microbial-dependent trimethylamine N-oxide (TMAO) generation from dietary precursors such as choline, phosphatidylcholine, and carnitine being associated with increased risk for atherosclerosis, 1 as well as mortality risk in CKD. 2 Additionally, multiple studies have linked circulating TMAO levels with accelerated atherosclerosis in animal models3,4 and demonstrated its applicability as a biomarker for CVD events in humans.5,6 A two-step metaorganismal pathway is responsible for TMAO formation: (1) trimethylamine (TMA)-lyase expressed in gut microbes metabolizes dietary nutrients such as choline to TMA, 7 and (2) the host hepatic enzyme flavin monooxygenase 3 (FMO3) oxidizes TMA to TMAO ( Fig. 1A ). 4 The reduction of TMA generation by gut bacteria and the host hepatic metabolism of TMA into TMAO serve as potential targets to lower circulating TMAO levels and reduce the risk for CVD/CKD events. 7 Accordingly, it has been demonstrated that suppression of FMO3 activity significantly decreased circulating TMAO levels and consequently atherosclerosis.8,9 However, FMO3 suppression evokes unwanted side effects such as hepatic inflammation,9,10 as well as accumulation of TMA resulting in a noxious odor (fish odor syndrome) of the patient. 11 This diminishes the attractiveness of FMO3 as a potential therapeutic target to reduce TMAO levels. A novel innovative concept is the nonlethal treatment of the gut microbiome (“Drug the Bug!” 12 ) by targeting microbial TMA-generating enzymes. The potential of this concept has been recently demonstrated with the small-molecule 3,3-dimethyl-1-butanol (DMB); this structural analogue of choline decreased bacterial TMA production and reduced atherosclerotic plaque formation in a mouse model. 7
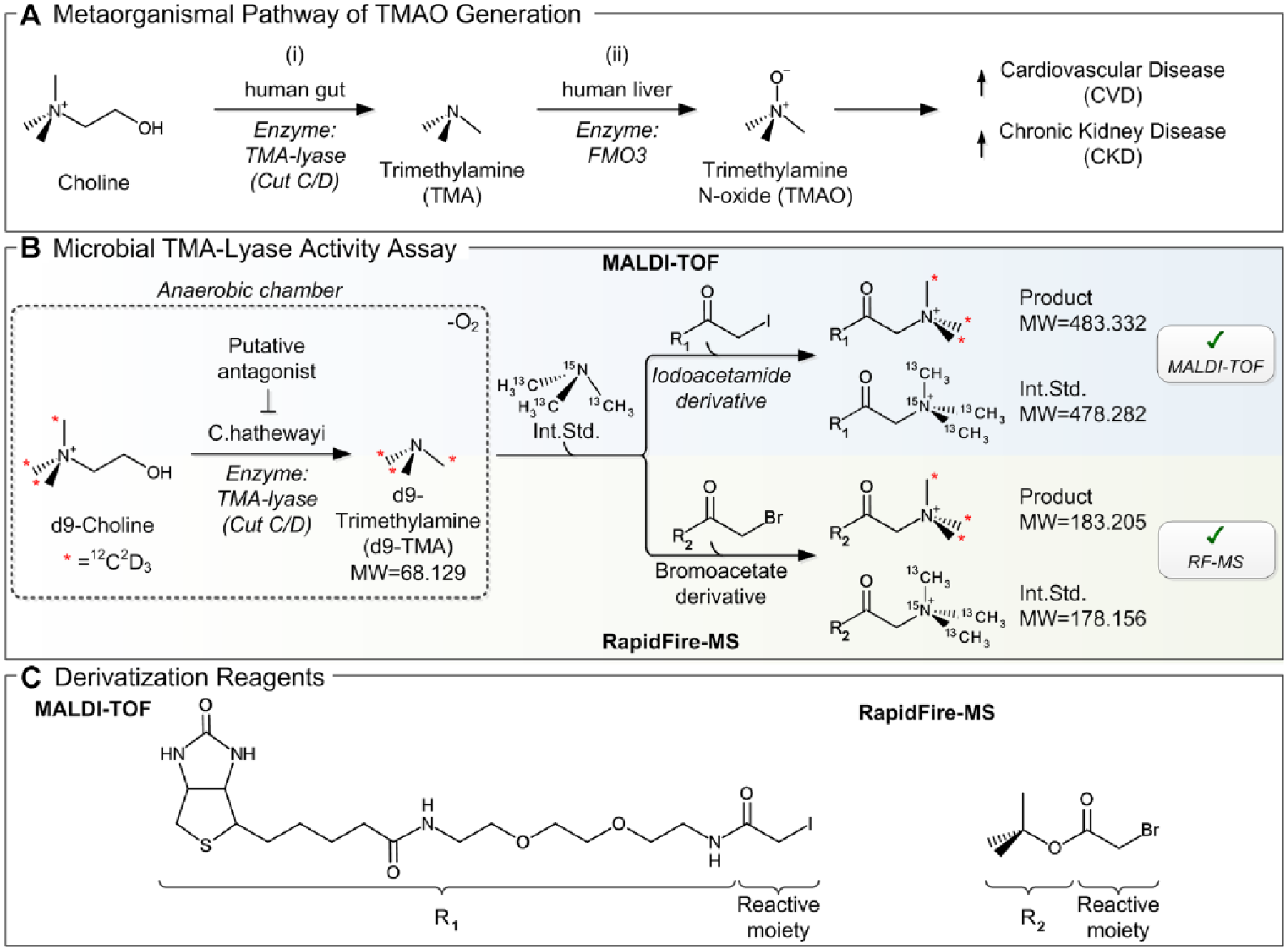
Descriptive representation of the MS-based microbial TMA-lyase inhibitor screening concept. (
High-throughput screening (HTS) is a well-established concept to identify novel small-molecule inhibitors with superior potency and druglikeness. Mass spectrometric quantification of TMA levels represents a direct measure of enzymatic activity and has been successfully demonstrated using liquid chromatography tandem mass spectrometry (LC-MS/MS) instrumentation; however, this procedure requires cycle times incompatible with HTS (>10 min/sample).3,13 Considerably faster cycle times can be achieved by instruments comprising online solid-phase extraction (SPE) and electrospray ionization (ESI) MS, such as the RapidFire-MS (RF-MS) system (Agilent Technologies, Santa Clara, CA). This MS technology enables cycle times of less than 10 s 14 and was successfully applied for the analysis of small molecules in complex samples. 15 However, for comprehensive HTS campaigns RF-MS applicability is still limited. An emerging HTS-compatible alternative is provided by recent advancement in liquid handling and instrumentation of matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS). Successful establishment of MALDI-TOF as an ultrafast label-free readout for enzymatic assays has been demonstrated in several studies16–21 and marks, together with the proven HTS capabilities of automated MALDI target preparation, 22 the upcoming era of label-free MS-based readouts within HTS campaigns.
In this study, we aimed to set up a MALDI-TOF-based assay for accurate quantification of secreted TMA levels within bacteria cultures to assess microbial TMA-lyase activity. This approach is hampered by low MALDI-TOF compatibility with (1) analytes in a highly complex sample background and (2) the analysis of small molecules (<100 Da) per se. The analysis of complex samples (e.g., whole-cell lysates) by MALDI-TOF is limited due to the lack of enrichment/purification steps. Hence, the application of MALDI-TOF for complex samples is mainly focused on mass spectrometric fingerprinting in order to distinguish cellular phenotypes23,24 or identify microorganisms 25 instead of quantifying specific analytes. The analysis of specific target molecules within complex cellular samples has been demonstrated for proteins in high abundance (e.g., histones 26 ) but is rarely done for small molecules. The capability of MALDI-TOF to detect compounds in the low mass range is generally considered limited due to interference peaks induced by the matrix. 27 Several strategies exist to enable the analysis of small molecules by MALDI-TOF, such as matrix-free laser desorption/ionization, 28 usage of matrix additives to suppress background signals,29,30 or chemical derivatization of analyte molecules. 31 Our strategy, to enable MALDI-TOF-based quantification of TMA levels in complex microbial cell culture-derived samples, comprised sample dilution for complexity reduction and chemical derivatization to increase the analyte’s molecular weight. This concept enabled accurate quantification of secreted TMA levels with an HTS-compatible platform that provides the potential to uncover novel inhibitors of bacterial TMA-lyases for the treatment of CVD as well as CKD.
Materials and Methods
Materials
HTS MALDI disposable target plates (cat. 1847006) and HTS MALDI Adapter (cat. 8283496) were purchased from Bruker Daltonics (Bremen, Germany). 13C3,15N-TMA (trimethylamine-13C3,15N1 hydrochloride, cat. 730092), biotin polyethyleneoxide iodoacetamide (BPI; cat. B2059), tert-butyl bromoacetate (TBBA; cat. 124230), α-cyano-4-hydroxycinnamic acid (HCCA; cat. 70990), ammonia solution (cat. 09859), acetonitrile (cat. 34851), DMSO (cat. D5879), glycerol (cat. G9012), and isopropanol (cat. 34863) were obtained from Sigma Aldrich (St. Louis, MO). Trifluoroacetic acid (TFA; cat. 6957) was obtained from Carl Roth (Karlsruhe, Germany). Sodium hydroxide solution (cat. 35256) was purchased from Honeywell (Seelze, Germany). Assay plates in 384-well (cat. 784201/cat. 781270) and 1536-well (cat. 782101) format were purchased from Greiner Bio-One (Frickenhausen, Germany). Further chemicals were purchased from different suppliers: formic acid (cat. A117-50, Thermo Scientific, Waltham, MA), reinforced clostridial medium (RCM; cat. 215192, BD Biosciences, San Jose, CA), Clostridium hathewayi (cat. 13479, DMSZ, Braunschweig, Germany), d9-choline (choline chloride-[trimethyl-d9]; cat. DLM-549-PK, Cambridge Isotope, Tewksbury, MA), and d9-TMA (d9-trimethylamine; cat. D-0764, CDN Isotopes, Quebec, Canada).
Preparation of Compound Plates
For single-dose screening, 1 µL of DMSO containing 0.5 mg/mL compound was transferred into 384-well microplates using a CyBio Well vario (Analytik Jena, Jena, Germany) liquid handling unit equipped with a capillary head. This resulted in a final screening concentration of 5 µg/mL for each compound. In dose–response experiments ascending compound concentrations (final screening concentration between 0.01 and 100 µM) were likewise transferred in 1 µL of DMSO. Each prepared 384-well plate comprised 352 compound-containing wells, while 1 µL of DMSO was added to the last two columns (23/24) for high/low controls.
Bacteria Culture in Anaerobic Chamber
In vitro TMA formation was tracked using a bacteria cell culture of C. hathewayi. This species is an obligate anaerobe and all relevant assay steps were performed under strict oxygen-free conditions in a Bactron900 anaerobic chamber (Sheldon Manufacturing, Cornelius, OR). 32 To reduce oxygen content in the RCM, it was pretreated by exposure to the anaerobic atmosphere for 18–24 h. Stocks were frozen at OD600 = 0.4 in 1 mL aliquots containing 15% glycerol at –80 °C. In order to initiate bacteria culture, C. hathewayi cells were thawed on ice and suspended in 50 mL of RCM. Cells were grown for 6 h at 37 °C in a humidified incubator and subsequently divided, diluted (1:1000 in RCM), and grown overnight under equal conditions. The prepared compound plates were incubated with 45 µL of RCM for 30 min in the anaerobic chamber. Subsequently, the bacteria culture was diluted once again with RCM (1:25) and 45 µL of this solution was transferred to columns 1–24 prior to an additional incubation step for 1 h at 37 °C in a humidified incubator. The substrate d9-choline was prepared at 30 mM in 0.9% sodium chloride prior to a dilution to 12 mM in RCM. Ten microliters of this substrate dilution was added to columns 1–23, resulting in a final concentration of 1.2 mM substrate per well, and 10 µL of pure RCM was added to column 24 of the compound plate (= low control). This solution was incubated for 3 h at 37 °C in a humidified incubator. Prior to MS analysis, the assay plates were centrifuged for 5 min at 5000 rpm to pellet the bacterial cells and only the supernatant was used for subsequent sample preparation.
Derivatization Protocol: MALDI-TOF MS
Samples generated in the microbial assay were derivatized using BPI to enable MALDI-TOF-based TMA quantification. Initially, 20 µL of bacterial cell culture supernatant was transferred to new 384-well assay plates using an Agilent Bravo (Agilent Technologies). Subsequently, the bacteria solution was diluted 1:10 by transferring 2 µL of bacteria solution and 18 µL of ddH2O to new 384-well assay plates using a CyBio Well vario pipettor. After brief centrifugation (2 s at 1000 rpm) and vortexing (30 s at 1000 rpm), 2 µL was transferred into 1536-well plates and prepared for derivatization by adding 2 µL of an internal standard (13C3,15N-TMA, at 30 µM) containing 0.1 M ammonium hydroxide solution. For derivatization, 2 µL of 37 mM BPI was added and subsequently centrifuged (2 s at 1000 rpm) and vortexed (30 s at 1000 rpm). The mixture was incubated in the dark at room temperature for 30 min and finally quenched by adding 1 µL of 0.7% TFA. The mixture was briefly centrifuged (2 s at 1000 rpm) and the assay plates were sealed for subsequent MALDI target preparation.
MALDI Target Preparation
MALDI target preparation was performed as described previously. 22 Briefly, a saturated solution of α-cyano-4-hydroxycinnamic acid (HCCA) was prepared in 50% acetonitrile and 50% water/0.05% TFA (v/v). The CyBio Disk was equipped with a 1536-channel ceramic tip head to transfer the 1536-well plate within one step. For highly reproducible and homogenous spot shapes, we applied double-layer spotting. Here, 100 nL of saturated matrix solution was spotted onto plain steel plates and dried with the aid of vacuum. Subsequently, 100 nL of matrix and 100 nL of sample was aspirated successively and dispensed together onto the dried matrix spots prior to a second vacuum drying step. In order to prevent carryover, ceramic tips were thoroughly washed with 70% isopropanol and 30% water/0.1% TFA (v/v).
MALDI-TOF MS Acquisition
Mass spectra were acquired with a rapifleX MALDI TOF/TOF PharmaPulse instrument from Bruker Daltonics equipped with a Smartbeam 3D laser. FlexControl (v4.0), flexAnalysis (v4.0), and MALDI Pharma Pulse (v2.0) were used for MS acquisition as well as data processing and analysis. Mass spectra were acquired in the mass range of m/z 400–600 to cover product and the internal standard of the biochemical assay. A pulsed ion extraction of 160 ns was used and the digitizer was set to 5.00 GS/s. The laser was operated at a 10 kHz frequency applying the “Single” Smartbeam parameter at a 50 µm × 50 µm scan range. Laser power was adjusted prior to a batch process to acquire sufficient peak intensity for quantification (≈104 cts intensity thresholds for the internal standard peak). Sample spots were analyzed in the automatic mode acquiring 10,000shots per spot composed of 20 steps at the raster spot. The acquired mass spectra were processed with the following parameters: centroid algorithm for peak detection, Gaussian smoothing of spectra (m/z 0.02, one cycle), and subsequent TopHat baseline subtraction. Prior to each campaign, external calibration was performed once with Peptide Calibration Standard II (Bruker Daltonics) containing seven peptides with corresponding [M+H]+ masses: angiotensin II = 1046.5418, angiotensin I = 1296.6848, substance P = 1347.7454, bombesin = 1619.8223, adrenocorticotropic hormone (ACTH) clips 1–17 = 2093.0962, ACTH clips 18–39 = 2465.1983, and somatostatin 28 = 3147.10. Additionally, internal calibration was performed using the monoisotopic peak of product M+ = 483.3315 and internal standard M+ = 478.2816.
Derivatization Protocol: RF-MS
For RF-MS quantification of TMA-lyase activity we derivatized TMA using TBBA. For this purpose, 4 µL of bacterial cell culture supernatant were diluted with RCM 1:100 (v/v) and 4 µL of this solution was transferred to new 384-well assay plates using an Agilent Bravo device. To prepare the solution for derivatization, 8 µL of an internal standard (13C3,15N-TMA, at 1 µM) containing 0.1 M ammonium hydroxide solution was added. Subsequently, 6 µL of 3 mM TBBA was added and incubated for 30 min at room temperature. To quench the derivatization reaction, 180 µL of water/acetonitrile/formic acid (50/50/0.025) was added. Remaining bacteria cells were spun down for 5 min at 4000g and 140 µL of supernatant was transferred into a new 384-well plate, sealed, and stored at –4 °C until mass spectrometric analysis.
RF-MS Acquisition
An ABSciex API 6500 mass spectrometer (ABSciex, Toronto, ON) was connected to a RapidFire sample injector (Agilent Technologies), a plate hotel (in-house solution), and a binary pump (Agilent 1290). This setup was operated by an in-house-developed LabView-based master software. Ten microliters of this sample was aspirated from the 384-well sample plate by negative pressure at –50 kPa. Samples were aspirated for 250 ms and subsequently flushed onto a H5-12µL HILIC cartridge with 70/30 isopropanol/hexane (eluent A). Subsequently, the analytes were back flushed toward the MS analyzer with methanol supplemented with 0.1% formic acid (eluent B). The mass spectrometer was operated in positive mode recording the following transitions: (1) derivatized d9-TMA 183.1/127.3 and (2) derivatized internal standard 13C3,15N-TMA 178.2/122.0, both at a declustering potential (DP) of 46 V and collision energy (CE) of 20 V. Pump 1 was operating with a flow rate of 1.1 mL/min delivering eluent A, pump 2 with 1.25 mL/min, and pump 3 with 0.5 mL/min, which was delivering eluent B.
Data Analysis
MALDI-TOF data, processed with flexAnalysis (v4.0) or MALDI Pharma Pulse (v2.0), were exported as a comma delimited (.csv) file. RF-MS data were recorded by Analyst 1.6.2 (ABSciex) and DiscoveryQuant 2.3 (ABSciex) and exported to Excel (Microsoft, Redmond, WA). Both datasets were further processed with either GraphPad Prism (v7.03; GraphPad Software, La Jolla, CA) or in-house software MegaLab.
TMA-lyase activity was tracked by analyzing the enzymatic product (d9-TMA) as well as the corresponding stable isotope labeled internal standard (13C3,15N-TMA). Their ratio was calculated in order to diminish variations ascribed to the MS readouts according to following equation:
Generated data were normalized and expressed as the percentage of control (PoC) according to following equation:
The IC50 calculation was performed in MegaLab by fitting the data to the following four-parameter logistical equation:
where a is the uninhibited value, b is the completely inhibited one, [I] is the inhibitor concentration, h is the Hill slope, and IC50 is the half maximal inhibitory concentration. GraphPad Prism was utilized for graphical representation of the data.
Basic appraisal of the assays was reviewed by the statistical quality parameter Z′ 33 with the following equation:
where s is the standard deviation and x are the mean readouts of the mass spectrometric assays.
Results and Discussion
Setup of the Microbial TMA-Lyase Activity Assay
In order to identify small molecular inhibitors of bacterial TMA production for the treatment of CVD and CKD patients, we established a high-throughput-compatible TMA-lyase activity assay. The setup of this strategy is outlined in
Figure 1B
. An enzymatic TMA-lyase activity assay composed of the unique glycyl radical enzyme CutC and the activating protein CutD was reported previously.
34
However, in order to identify druglike compounds that (1) inhibit TMA-lyase and (2) are able to penetrate bacterial cell walls, we decided to apply a microbial activity assay as the primary screen. While there are multiple bacteria species within the human gut that harbor the genes encoding for the choline TMA-lyase enzyme, C. hathewayi was selected for our initial screening campaign due to its relatively high abundance within human CKD stool and its high capacity to convert choline into TMA (data not shown). Due to C. hathewayi oxygen sensitivity, all experiments had to be performed under strict anaerobic conditions in an anaerobic chamber (
MS was selected as the readout to assess TMA-lyase activity due to its ability to provide a label-free and direct quantification of secreted TMA levels. Initially, RF-MS was established to characterize the microbial assay (data not shown). For this instance, d9-TMA was derivatized using TBBA (shown in Fig. 1C ) for increased lipophilicity and molecular mass (+115.075 Da), as previously suggested. 36 As outlined in Figure 1B , the assay mixture was supplemented with isotope-labeled TMA (13C3,15N1-TMA) as the internal standard prior to derivatization enabling accurate quantification of d9-TMA.
To provide an HTS-compatible, MS-based readout, we subsequently aimed to transfer the assay to a MALDI-TOF format. The usability of this technology for drug discovery screening has recently been proven in several studies assessing the activity of kinases, 18 histone demethylases, 19 acetylcholinesterases, 19 phosphatases,21,22 and histone lysine methyltransferases, 37 as well as the ubiquitin system.38,39 Together, all presented studies employed peptide/protein-centric activity assays that are well suited for MALDI-TOF analysis. MALDI-TOF compatibility for small molecules (<100 Da), however, is generally considered limited due to the high number of interfering matrix signals in the low mass region. This could be overcome by derivatization of the target molecule to increase its molecular weight. First, the derivatization strategy established for RF-MS was tested in this context. Unfortunately, derivatization by TBBA did not lead to MALDI-TOF-compatible analytes due the minor molecular weight increase of 115.076 Da (MWProduct = 183.2; MWInt.Std. = 178.2). For this reason, we replaced TBBA with BPI (shown in Fig. 1C ). This derivatization reagent increased the molecular weight of the target molecules by 415.203 Da, providing analytes with improved MALDI-TOF compatibility (MWProduct = 483.3; MWInt.Std. = 478.3). Of note, both derivatization strategies not only increase the molecular weight of d9-TMA but also introduce a permanent charge onto the analytes, which is expected to improve ionization efficiency and consequently mass spectrometric sensitivity. The presented derivatization strategy was hampered by the high complexity of samples derived from the microbial assay. Hence, we identified that 1:10 (v/v) dilution with water, prior to the derivatization step, was crucial for successful derivatization. Not only the target analyte but also other biomolecules containing amines and thiols are derivatized likewise, for which reason we titrated the concentration of the derivatization reagent to provide sufficient excess for efficient TMA derivatization. The mass-selective detection of derivatized TMA enables differentiation between other biomolecules derivatized in this procedure and the target analyte, enabling quantitative assessment of TMA formation in the bacterial samples.
The final assay protocol includes anaerobic cultivation of C. hathewayi, addition of the isotope-labeled internal standard (13C3,15N-TMA), and finally chemical derivatization. This provides an HTS-compatible MALDI-TOF assay for TMA quantification in microbial samples. A representative mass spectrum of derivatized d9-TMA is illustrated in
Figure 2A
for high (no compound) and low (no substrate) control. In order to assess TMA-lyase activity of C. hathewayi within this experimental setup, we aimed to quantify the amount of d9-choline being transformed into d9-TMA. For this instance, we prepared a d9-TMA standard curve in the range of 1 µM to 1 mM (exemplary mass spectra sections are illustrated in
Suppl. Fig. S1
). As presented in
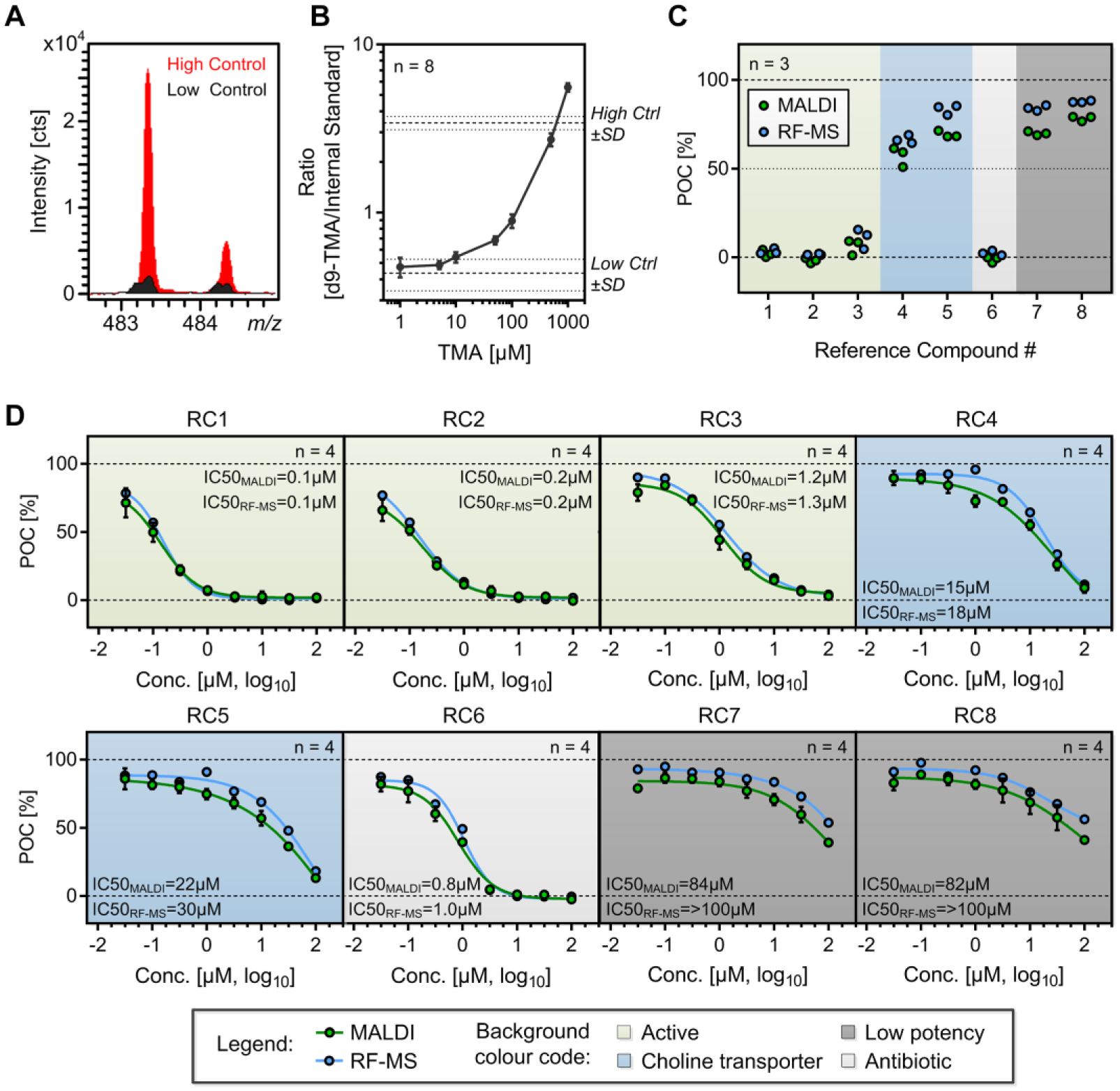
Assessment of general microbial assay characteristics and potency of reference compounds. (
In order to further validate the chemical derivatization and MALDI-TOF analysis, we added a constant concentration of 500 µM d9-TMA in growth medium to a 384-well assay plate representing technical high controls (no d9-TMA in column 24 representing low controls) and analyzed the plate according to the established protocol. Obtained results are illustrated in Supplemental Figure S2 . The coefficient of variation of 7% together with a Z′ value of 0.73 demonstrated the general suitability of the established workflow for tracking the microbial TMA formation in an HTS campaign.
Evaluation of Assay Performance Characteristics by Reference Compounds
After demonstrating successful d9-TMA quantification in complex growth medium by MALDI-TOF, we aimed to perform a proof of concept using known reference compounds. We selected five compounds with known alteration of TMA-lyase activity from two recent patent applications representing different structural choline analogues.40,41 Additionally, two known inhibitors of choline transport 42 as well as one compound with antibiotic activity 43 were included to demonstrate potential alternative modes of TMA inhibition. A list of these eight compounds, including chemical structures as well as description of their mode of action, can be found in Supplemental Table S1 .
Initially, single-dose experiments were performed by MALDI-TOF and RF-MS. As presented in Figure 2C , three of the structural choline analogues were found to be highly active (RC1–3) while two of them induced only minor alterations of TMA-lyase activity (RC7+8). The two compounds reported to be inhibitors of choline transport (RC4+5) induced only slight reduction of TMA formation, while reference compound 6 (ornidazole), an antibiotic agent with known activity against anaerobic bacteria, 43 induced an intense decrease of TMA formation. This emphasizes the necessity for counterscreens that rule out potential antibiotic activity of drug candidates.
Subsequent dose–response experiments of all eight reference compounds confirmed the findings of the single-dose experiments ( Fig. 2D ). Further, all experiments performed in this context show high consistency between the two MS-based readouts. A minor, though not significant, shift to increased activity is observable in the MALDI-TOF readout compared with RF-MS. This was not observed in subsequent single-dose and dose–response experiments, leading to the assumption that the shift might be caused by minor experimental variations in these preliminary experiments.
Single-Dose Screen of a Representative Compound Subset
After confirming performance characteristics of both the MALDI-TOF- and the RF-MS-based microbial TMA-lyase activity assays, we screened a selected small-compound library subset of 10,229 compounds. This library subset comprised diverse chemical structures representing the major chemical classes of the full-deck Boehringer Ingelheim corporate library (~1,100,000), allowing an estimate of the hit rate of a projected full-deck screen. The tested 10,229 compounds were spread across 32 assay plates in 384-well format. High and low controls were included on each of these assay plates and the corresponding results are visualized in Figure 3A for improved assessment of assay robustness. Additionally, the statistical quality parameter Z′ is plotted in Figure 3B for all consecutive assay plates. Mean Z′ values of 0.73 for MALDI-TOF and 0.84 for RF-MS were obtained in these experiments, which is competitive with previously published data for label-free MS readouts (0.87/0.86, 21 0.88/0.82, 19 and 0.77 44 ). This emphasizes the suitability of both technologies to track TMA formation in this complex microbial assay.
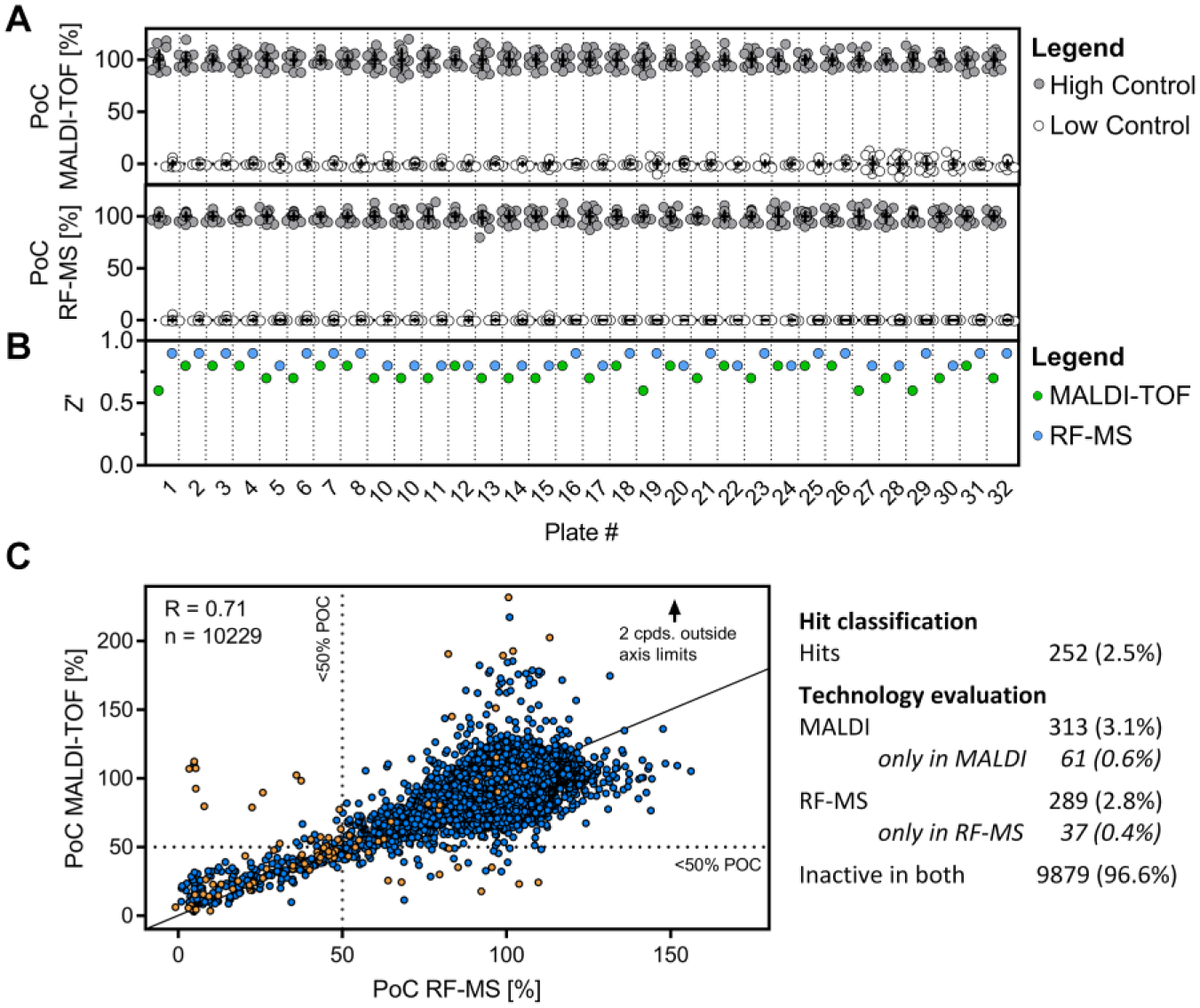
Single-dose screen for microbial TMA-lyase inhibitors using MALDI-TOF and RF-MS. (
Sample complexity led to hampered signal intensity by ion suppression, which consequently required an elevated number of laser shots and automated moves on each sample spot. Thus, MALDI-TOF cycle time was slightly elevated to approximately 1.5 s/sample compared with the previously published cycle times of 0.4 s/sample in a peptide-based MALDI-TOF assay. 22 However, it still outperforms cycle times of common RF-MS assays (6–12 s/sample14,45) and enables MS-based HTS campaigns for the discovery of novel microbial TMA-lyase inhibitors.
The PoC of the 10,229 compounds tested in this single-dose screen is plotted in Figure 3C . One technical replicate was performed on each of the established MS technologies. As expected for a single-dose experiment performed by two different readouts, the results were not identical. However, they did correlate (R = 0.71) and are consistent with previously published correlation coefficients of single-dose experiments performed by two distinct enzyme activity readouts (0.73 22 and 0.66 18 ).
The mean PoC observed within this assay was 92.7% (MALDI-TOF) and 94.7% (RF-MS). The standard deviation of this single-dose experiment was comparable between the two datasets, with 17.8% for MALDI-TOF and 16.2% for RF-MS (visualized in Suppl. Fig. S3 ). Compounds significantly reducing microbial TMA formation were defined as those producing a PoC smaller than 50%. A majority of tested compounds were identified to be inactive within both technologies (9879 compounds; 96.6%), while 252 (2.5%) compounds were identified as hits when both readouts are required to exceed the selected hit criteria. Only a small fraction of compounds exceed the hit criteria for only one of the MS readouts (MALDI: 61 [0.6%]; RF-MS: 37 [0.4%]). Interestingly, some compounds in the MALDI-TOF assay were shifted to elevated PoC values (PoC > 150%), which is not present in the RF-MS readout. However, this finding was not confirmed in subsequent dose–response experiments and hence attributed to experimental variations. The compiled hit rates for the two assay technologies are summarized in Figure 3C , and detailed results of all compounds are listed in Supplemental Table S2 .
Hit Qualification by Dose–Response Experiments
To confirm findings obtained by the single-dose screen, we selected a set of compounds with different characteristics. In total, 99 compounds were selected to represent four distinct categories: (1) inactive in both readouts (22 compounds), (2) active in both readouts (45 compounds), (3) active only in the MALDI-TOF readout (13 compounds), and (4) active only in the RF-MS readout (19 compounds). Additionally, 10 compounds with known alteration of microbial TMA formation were enclosed (including reference compounds 1–4 and 6 from previous experiments) and subjected to dose–response experiments. In order to reveal the accuracy and reproducibility of the assay, we performed the dose–response experiments in two independent biological replicates on two separate days on each MS platform. High correlation between these replicates was obtained for both technologies (RMALDI-TOF = 0.92; RRF-MS = 0.96), demonstrating high intratechnology and interday correlation of the established assays (see
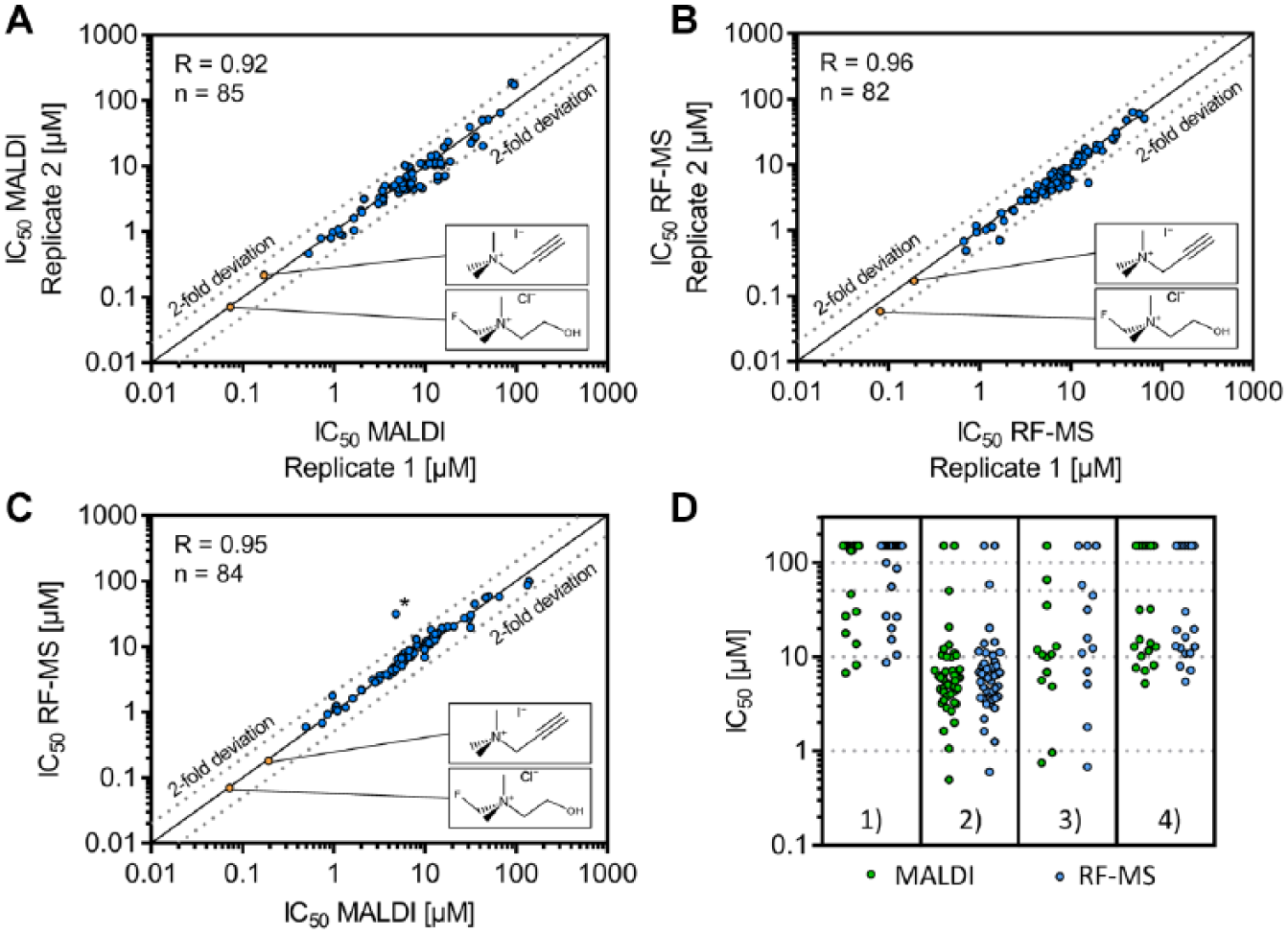
Dose–response confirmation of identified compound characteristics in the single-dose experiment. Overall, 109 compounds were examined in this approach. Of note, compounds triggering no enzyme inhibition at the highest concentration level (>100 µM) were excluded from the graphs. Details of this potency determination can be found in
Supplemental Table S3
. (
Despite the high correlation of MALDI-TOF and RF-MS readouts, three compounds were identified with a more than sixfold deviation in IC50 values determined by the two technologies. Dose–response curves of these compounds (IC50 - MALDI = 4.8/6.8/11.8 µM; IC50 - RF-MS = 31.5/>100/>100 µM) are illustrated in Supplemental Figure S4A . The origin of this potency variation was identified to be induced by the interference of compounds with the internal standard signal. In these samples, the intensity of the internal standard signal is elevated in dependency on the present compound concentration, as demonstrated in Supplemental Figure S4B,C . Two of these compounds have a small mass difference from the internal standard (Δm = −0.026 Da) that cannot be resolved by the applied MALDI-TOF instrument. The third interfering compound has a higher mass difference (Δm = 41.841 Da), and thus we assume a laser-induced neutral loss of the corresponding compound or a contamination within the compound solution being responsible for the interfering signal (for details see Suppl. Fig. S4D ). A solution for this issue would be retesting of interfering compounds with an alternative internal standard; calculating the product/substrate ratio as suggested previously is not applicable as substrate is not derivatized in this assay. 18 Alternatively, one could establish a fragmentation-based MALDI-TOF/TOF analysis similar to the selected reaction monitoring (SRM) performed by the RF-MS readout. Analysis of product ions instead of precursors increases selectivity, which might enable distinction between the analyte of interest and isobaric contaminations/compounds.
In order to assess the confirmation rate of the established assay, compounds were separated according to their results obtained in the single-dose experiments (see
Tabular Results of the Dose–Response Experiments of Compounds Divided into Five Different Categories according to Their Activity in the Single-Dose Experiment.
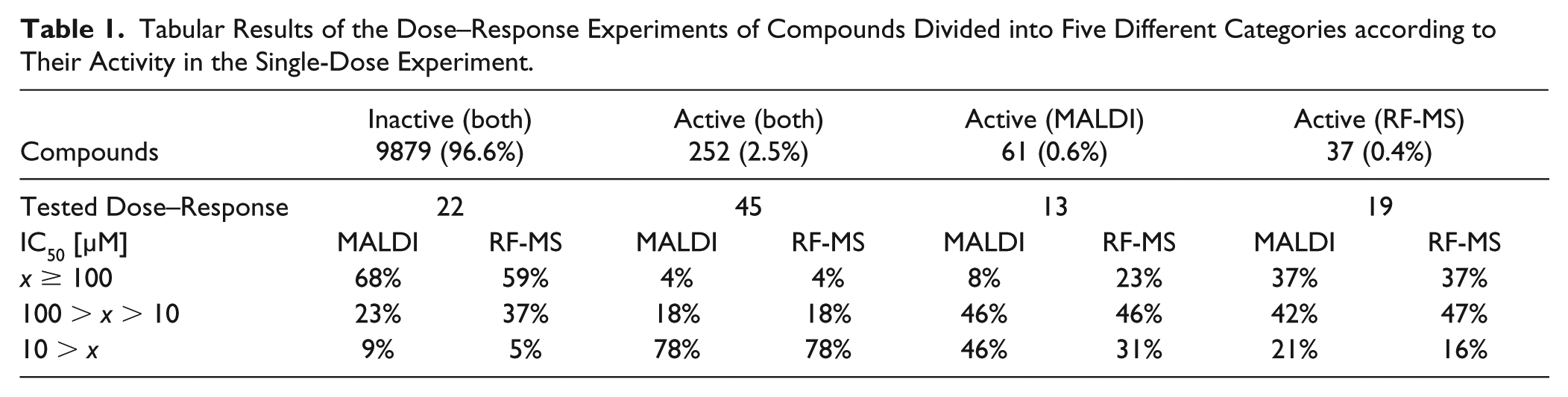
In summary, we applied MALDI-TOF as a label-free readout for the microbial choline TMA-lyase activity assay in single-dose (10,229 compounds) and subsequent dose–response (109 compounds) experiments. Side-by-side comparison to the RF-MS readout demonstrated high concordance while providing significantly reduced cycle times. This study revealed the promising prospects of MALDI-TOF to be used in HTS campaigns for the discovery of novel lead structures with choline TMA-lyase inhibitory properties.
The obtained hit rate of 2.5% is slightly elevated compared to a plain biochemical MS-based potency assay, which might be induced by potential off-target effects of the compounds but may also be target related. Also, the selected hit threshold of 50% PoC is an adjusted threshold that might cause false-positive results. To account for the assay variation in a full-deck screening campaign, we would select a lower hit threshold according to the three-sigma rule. In the presented data, this would lead to hit thresholds of 39.3% for MALDI-TOF and 46.1% for RF-MS, which indicates that the 50% cutoff will lead to a number of false-positive hits.
To rule out potential off-target effects of the tested compounds, important follow-up experiments are required to unravel the compounds’ mode of action. The development of nonlethal TMA-lyase inhibitors is less likely to induce bacteria to evolve resistance mechanisms than traditional antibiotics. 7 Hence, compounds with antibiotic activity should be ruled out by microbial viability assays. Further, to confirm the validity of identified hit candidates it would be crucial to perform direct on-target experiments. This should employ (1) the isolated recombinant CutC/D enzyme within an enzymatic activity assay, as previously described by Craciun et al., 34 as well as (2) biophysical methods to confirm direct target engagement. These combined results should decipher the compounds’ mode of action and rule out the majority of potential off-target effects. Finally, the investigation of different microbial species would demonstrate species-independent inhibition of TMA-lyase activity, which would elevate the expected treatment efficacy. Both biochemical assay and testing of different microbial species could be analyzed by the newly developed MALDI-TOF assay. With these follow-up assays, the overall hit rate (2.5%) obtained in the microbial assay should be vastly reduced.
Conclusively, we established an HTS-compatible MALDI-TOF-based concept to track microbial TMA formation enabling the screening of choline TMA-lyase inhibitors within large compound libraries. Chemical derivatization converts d9-TMA into MALDI-TOF-compatible conjugates that serve as a direct quantitative measure for TMA-lyase activity, highlighting the promising potential of derivatization strategies in general to give access to novel target classes within complex samples. To our knowledge, this is the first MS-based, HTS-compatible microbial assay tracking the generation of a small molecule such as d9-TMA by MALDI-TOF.
This serves as a proof of concept that not only peptide-based biochemical assays can be adapted to MALDI-TOF for HTS approaches as presented previously,18,19,21,37–39 but also assays tracking small molecules within a complex sample background. Overall, the presented data highlight MALDI-TOF as a versatile technology for HTS campaigns and further emphasize its promising perspective as integral technology within drug discovery research.
Supplemental Material
Supplemental_Material_for_MALDI-TOF_HTS_Assay_for_TMA-lyase_by_Winter_et_al – Supplemental material for Chemical Derivatization Enables MALDI-TOF-Based High-Throughput Screening for Microbial Trimethylamine (TMA)-Lyase Inhibitors
Supplemental material, Supplemental_Material_for_MALDI-TOF_HTS_Assay_for_TMA-lyase_by_Winter_et_al for Chemical Derivatization Enables MALDI-TOF-Based High-Throughput Screening for Microbial Trimethylamine (TMA)-Lyase Inhibitors by Martin Winter, Tom Bretschneider, Sven Thamm, Carola Kleiner, Daniel Grabowski, Sarah Chandler, Robert Ries, Jörg T. Kley, Danielle Fowler, Christina Bartlett, Ralph Binetti, John Broadwater, Andreas H. Luippold, Daniel Bischoff and Frank H. Büttner in SLAS Discovery
Footnotes
Acknowledgements
We thank Markus Holstein for the RapidFire-MS measurements. Further, we thank Marina Ilg for providing the MALDI derivatization reagent.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors disclosed the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors are employees of Boehringer Ingelheim Pharma GmbH & Co. KG and completed this work within the scope of their employment.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.