Abstract
Label-free, mass spectrometric (MS) deciphering of enzymatic reactions by direct analysis of substrate-to-product conversion provides the next step toward more physiological relevant assays within drug discovery campaigns. Reduced risk of suffering from compound interference combined with diminished necessity for tailored signal mediators emphasizes the valuable role of label-free readouts. However, MS-based detection has not hitherto met high-throughput screening (HTS) requirements because of the lack of HTS-compatible sample introduction. In the present study, we report on a fully automated liquid-handling concept built in-house to concatenate biochemical assays with matrix-assisted laser desorption/ionization time-of-flight closing this technological gap. The integrated reformatting from 384- to 1536-well format enables cycle times of 0.6 s/sample for automated spotting and 0.4 s/sample for MS analysis, matching the requirements of HTS compatibility. In-depth examination of spotting quality, quantification accuracy, and instrument robustness together with the implementation of a protein tyrosine phosphatase 1B (PTP1B) inhibitor screening (4896 compounds) demonstrate the potential of the heavily inquired HTS integration of the label-free MS readout. Overall, the presented data demonstrate that the introduced automation concept makes label-free MS-based readouts accessible for HTS within drug discovery campaigns but also in other research areas requiring ultrafast MS-based detection.
Introduction
High-throughput screening (HTS) is a key technology within lead identification in early stages of pharmaceutical drug discovery. Common HTS screening approaches are largely driven by fluorescence- and chemiluminescence-based readouts providing joint features including sensitive, reproducible, and fast analysis of enzymatic reactions combined with near universal applicability. In turn, they suffer from compound-dependent screening artifacts leading to false-positives/negatives 1 and require time-consuming and costly assay development including the generation of artificial, nonphysiological probes and/or specific antibodies. 2 An emerging alternative to conventional label-based screening technologies is the use of mass spectrometry (MS) exemplified by the growing landscape of MS-based screening approaches in the field of drug discovery research.3–5 These offer the possibility to track physiological relevant molecules, without prior labeling steps.
MS-based readouts in the field of drug discovery were up to now largely dominated by instruments concatenating online solid-phase extraction and electrospray ionization (ESI) MS. One representative of these hyphenated instruments is the RapidFire (RF-MS) system of Agilent (Santa Clara, CA) providing cycle times of less than 10 s, which vastly outperforms conventional high-performance liquid chromatography–MS instrumentation. 6 However, implementation of HTS or even ultra-HTS campaigns is still elusive for this technology. Here, cycle times of less than 1 s per sample are required as provisioned by fluorescence/luminescence–based readouts. Because of this, RF-MS is almost exclusively used for lead optimization campaigns, 5 compound profiling, 7 and ADME assays.6,8
Novel opportunities to provide HTS-compatible MS readouts involve the use of acoustic droplet ejection as well as an open-port probe that delivers samples directly from multiwell plates into the mass spectrometer.3,9,10 However, the lack of commercial availability has hampered their broad applicability so far. An alternative to enable label-free HTS campaigns is provided by recent advancements in matrix-assisted laser desorption/ionization (MALDI) that offer speed and robustness of conventional label-based technologies accompanied by the advantages of an MS-based readout (e.g., label-free, interference-resistant, comprehensive assay assessment). This, together with the evolution of sophisticated liquid-handling concepts to transfer samples onto MALDI target plates, raised the potential to enable MS-based full-diversity HTS campaigns and pushed the efforts to integrate MALDI time-of-flight (TOF) into the HTS environment.11–15
Key challenges that remain within this effort include assay buffer adaption to match MS compatibility while enzymatic activity remains unaffected 11 as well as tailored integration into the automated liquid-handling process with a focus on the challenging sample transfer onto MALDI target plates. Several concepts exist for the MALDI-TOF integration, such as the acoustic liquid handler Echo 550 (Labcyte, Sunnyvale, CA) 12 and the Mosquito (TTP Labtech, Hertfordshire, UK) nanoliter dispenser.13,14 Here, we present a concept for the utilization of the multichannel pipettor CyBio Disk (Analytik Jena, Jena, Germany) as a connecting link between the biochemical assay and the MALDI-TOF readout. We aimed to provide greatest flexibility by establishment of a fully automated standalone spotting station. This station is capable of handling 384- or 1536-well plates originating from biological assays, merging four 384-well plates into 1536-well format if necessary, and finally transferring the samples onto MALDI target plates.
Overall, we established a concept that is suitable for MALDI-TOF HTS integration. Within this study, we demonstrate high accuracy and reproducibility of the introduced spotting process accompanied by convincing data for the quantification of enzymatic activity by MALDI-TOF. Moreover, we provide evidence that the system can indeed run a 1-million-compound screen without the need for major maintenance of the spotting station or MALDI-TOF, respectively. The chosen design is composed of separated spotting and MS units, which enables the versatile application of MALDI-TOF within different stages of the drug discovery process, starting with the accomplishment of full-diversity screening campaigns to support lead qualification and profiling projects. Finally, we demonstrate performance of this overall concept using a protein tyrosine phosphatase 1B (PTP1B) inhibitor screen to approve the entire spotting/analysis concept as a valuable tool for drug discovery campaigns.
Materials and Methods
Materials
HTS MALDI disposable target plates (No. 1847006) and HTS MALDI Adapter (No. 8283496) were purchased from Bruker Daltonics (Bremen, Germany). Insulin receptor substrate (ETDpYYRKG-amidated) and product peptide (ETDYYRKG-amidated) as well as the labeled AlphaScreen peptide (biotin-Ahx-ETDpYYRKG-amidated) were purchased from Biosyntan (Berlin, Germany). Internal standard peptide ETDYY
Preparation of Assay Plates
Assay plates were prepared in 384-well format containing ingredients mimicking an actual biological assay in order to assess quality parameters of the entire automation concept. For this purpose, 0.25 µM ETDpYYRKG-amidated (substrate peptide;
PTP1B Inhibitor Screen
Label-based as well as label-free detection of PTP1B inhibition were conducted by AlphaScreen and MALDI-TOF, respectively, as previously published with minor modifications. 16 Briefly, for single-dose concentration screening, 100 nL DMSO containing 0.5 mg/mL compound was transferred into 384-well microplates (No. 781075, Greiner, Frickenhausen, Germany) using the CyBio Well vario (Analytik Jena, Jena, Germany) liquid-handling unit equipped with a capillary head. This resulted in a final screening concentration of 5 µg/mL for each compound. To assess performance characteristics and compare label-free and label-based assay technologies, a set of 4986 compounds were examined. These were composed of a selected subset of structural class representatives of the full-deck screening pool.
Configuration of the MALDI Spotting Station
Arrangement of the in-house–built MALDI spotting station is schematically outlined in Figure 1A . The core of this station was a centralized robotic system (Stäubli TX60L, Stäubli International AG, Pfäffikon, Switzerland) that orchestrated the entire process and moved assay and MALDI target plates to their designated destination within the automation unit. Together, four stackers (3x CyBio QuadStacks L/1x CyBio Well vario stacker, Analytik Jena, Jena, Germany) were included to store up to 60 MALDI target plates (stacker 1) and 480 assay plates in 384- or 1536-well format (stacker 2+3) as well as up to 224 assay plates in 1536-well format in the CyBio Well vario stacker. Sealer (PlateLoc Typ G5402A, Agilent, Palo Alto, CA), peeler (XPeel, Brooks Automation, Chelmsford, MA), shaker (BioShake 5000, Quantifoil Instruments, Jena, Germany), and two barcode readers (Opticon NLV3101, Dietzenbach, Germany) were enclosed. Merging of the four 384-well plates onto one 1536-well plate was conducted by the multichannel pipettor CyBio Well vario (Analytik Jena) supplied with disposable 384-channel tips (CyBi TipTray 384-25 µL, Analytik Jena). Potential air bubbles were removed by centrifugation (Bionex Automated Centrifuge, BioNex Solutions, San Jose, CA). The transfer of samples as well as matrix solution onto MALDI target plates was conducted using the CyBio Disk (Analytik Jena) robot system supplied with a 1536-well ceramic tip head.
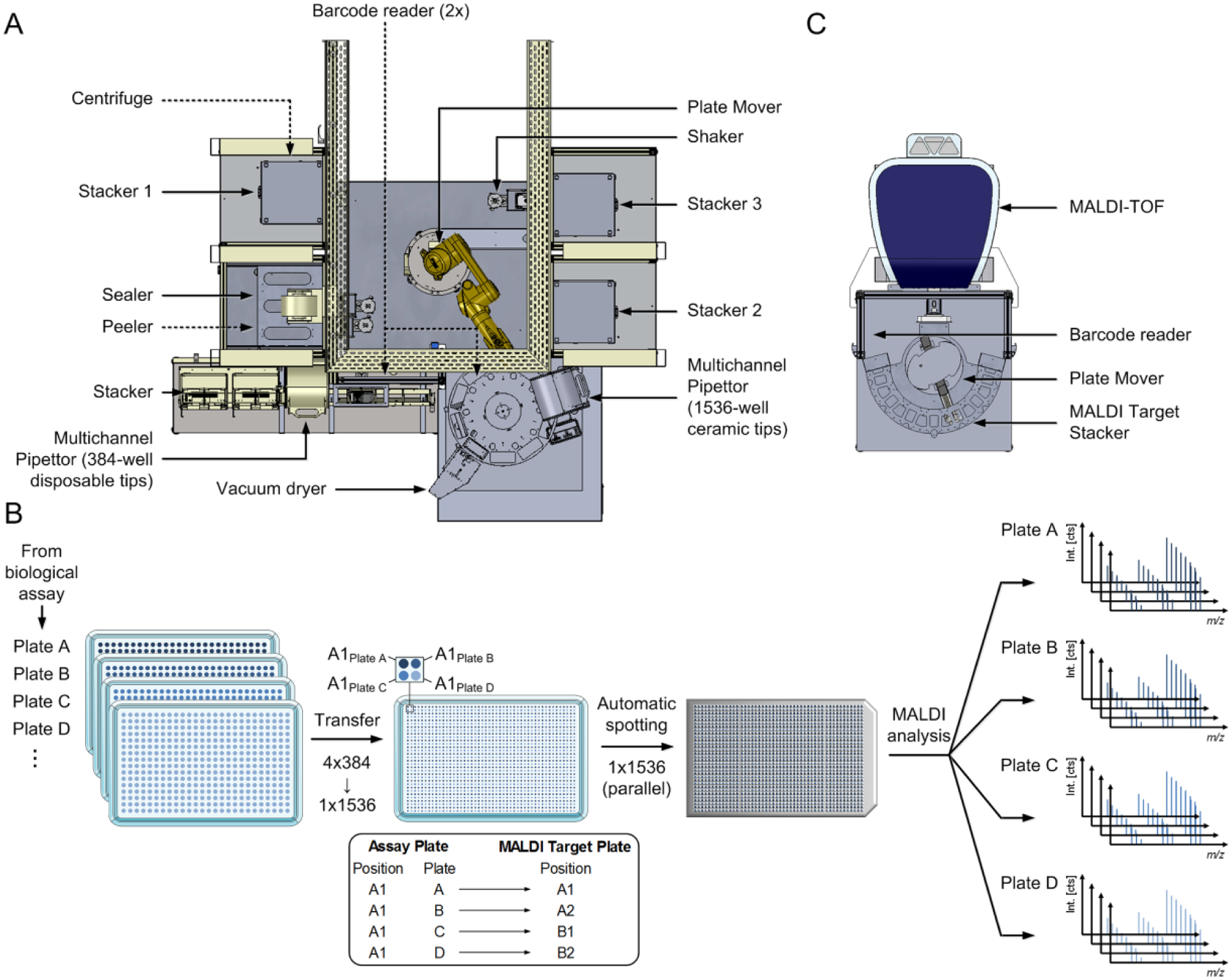
Schematic illustration of the automation concept for concise matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) high-throughput screening (HTS) integration. (
Plate Fusion: 384-Well to 1536-Well Format
Merging of four 384-well plates into one 1536-well plate is briefly sketched in
Figure 1B
. Assay plates derived from the biochemical assay were loaded into stacker 2, and empty 1536-well target plates were loaded into the CyBio Well vario stacker. The first assay plate was transported to the shaker to provide a homogenous assay solution by brief mixing (20 s at 1000 rpm) of the plate. Subsequently, the seal of the assay plate was removed by the peeler, and 5 µL of each well was transferred into the first quadrant of the empty 1536-well target plate. For storage, the assay plate was sealed and finally transferred to stacker 3 while the multichannel pipettor tips were washed three times with 23 µL pure water. This procedure was conducted for four consecutive assay plates, which were concatenated in the 1536-well plate as illustrated in
Figure 1B
(exemplified for position A1 of the four respective 384-well plates). After transfer completion, the 1536-well plate was centrifuged for 30 s at 1000 rpm to remove air bubbles and was subsequently moved to the CyBio Disk to execute the spotting procedure. During the corresponding spotting procedure, transfer of the next assay plate was initiated. Precise scheduling of the automation steps enabled acceleration of the entire procedure and is exemplarily illustrated in
MALDI Target Spotting Using the CyBio Disk Pipetting System
α-Cyano-4-hydroxycinnamic acid (4-HCCA) was diluted to a final concentration of 10 mg/mL in 50% acetonitrile and 50% water/0.05% TFA (v/v) prior to vigorous vortexing to provide a saturated matrix solution. The CyBio Disk was equipped with a 1536-channel ceramic tip head to transfer the 1536-well plate within one step. MALDI target plates were placed onto a spring adapter to ensure planarity and enable contact-based matrix/sample transfer, facilitating simultaneous and reproducible displacement of 1536 spots. For highly reproducible and homogenous spot shapes, we applied double-layer spotting. Here, 100 nL saturated matrix solution was spotted onto plain steel plates and dried with the aid of a vacuum in the active dryer for 120 s. Subsequently, 100 nL matrix and 100 nL sample were aspirated successively and dispensed together onto the dried matrix spots prior to a second vacuum drying process for 300 s. To prevent carry over, ceramic tips were washed with 70% isopropanol and 30% water/0.1% TFA (v/v). Here, we applied a prewash using 0.15 µL followed by three consecutive steps with 0.3 µL within sample processing, and after completion of an entire spotting procedure, a prewash using 0.4 µL followed by three steps with 0.6 µL was performed. Crucial was the usage of a tip-wash station, which aided the removal of the remaining solution in the pipette tips by vacuum suction. Selected samples were washed on target by transferring 0.3 µL of 0.1% TFA (v/v)/10 mM ammonium dihydrogenphosphate onto each spot with subsequent removal after 1 s of incubation. After completion of the spotting procedure, the 1536-well plate was moved to stacker 3, while the MALDI target plate was moved to stacker 1. To maintain spotting quality and prevent matrix-induced clogging of the ceramic tips, they were rinsed three times with 0.1 M sodium hydroxide after the last spotting operation in a batch process followed by a tip-washing procedure as described above to remove the remaining sodium hydroxide.
MALDI-MS Acquisition
Mass spectra were acquired with a rapifleX MALDI TOF/TOF PharmaPulse instrument from Bruker Daltonics comprised of a Smartbeam 3D laser. FlexControl (v4.0), flexAnalysis (v4.0), as well as MALDI Pharma Pulse (v2.0) were used for MS acquisition as well as data processing and analysis. Measurement of multiple target plates was facilitated by automated handling of target plates using the robot system Orbitor RS (Thermo Scientific) including a barcode reader as depicted in
Figure 1C
. Mass spectra were acquired in the mass range of
Prior to each campaign, external calibration was performed once with Peptide Calibration Standard II (Bruker Daltonics) containing seven peptides with corresponding [M+H]+ masses: angiotensin II = 1046.5418, angiotensin I = 1296.6848, substance P = 1347.7454, bombesin = 1619.8223, ACTH clip 1–17 = 2093.0962, ACTH clip 18–39 = 2465.1983, somatostatin 28 = 3147.10. In addition, internal calibration was performed using the monoisotopic peak of substrate [M+H]+ = 1110.4616, product [M+H]+ = 1030.4952, and internal standard [M+H]+ = 1040.5035 for the PTP1B enzymatic reaction.
Software Tools and Data Analysis
Instrument control and process scheduling of the entire system was provided by the Momentum laboratory automation software (v4.2.3; Thermo Scientific). This master control software enabled setup of the entire system, definition of processes, controlling of devices, and implementation of assay/MALDI target plate barcode tracking (concept illustrated in
MALDI-TOF data, processed with flexAnalysis (v4.0) and MALDI Pharma Pulse (v2.0), were exported as a tab delimited (.txt) file and further processed with Microsoft Excel (Microsoft, Redmond, WA), GraphPad Prism (v7.03; GraphPad Software, La Jolla, CA), or in-house software MegaLab.
PTP1B activity was tracked by analyzing dephosphorylated product peptide and the corresponding heavy labeled internal standard peptide. Their ratio was calculated in order to diminish variations ascribed to the MS measurement according to following equation:

The AlphaScreen signal was transformed into substrate concentration with the substrate standard curve by fitting the data to a four-parameter logistical calibration curve implemented in MegaLab.
Both transformed AlphaScreen and MALDI-TOF data were normalized likewise and expressed as percentage of control (POC) according to following equations:
MALDI-TOF data:

AlphaScreen:
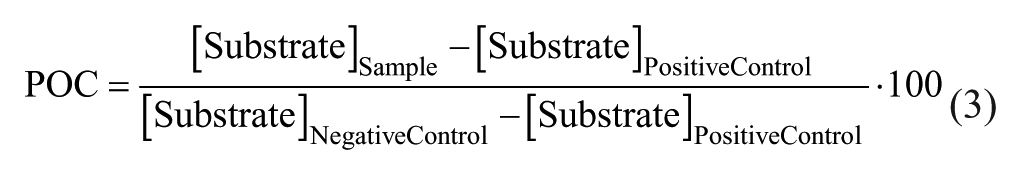
Basic appraisal of the assays was reviewed by the statistical quality parameter Z′, 18 with the following equation:
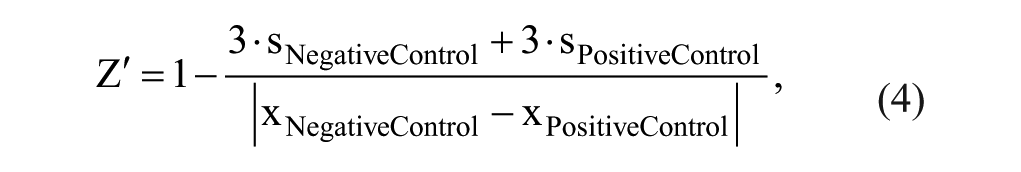
where s is the standard deviation and x is the mean readout of the AlphaScreen and MALDI-TOF assays, respectively.
Results and Discussion
MALDI-TOF Automation Concept
Robust, fast, and miniaturized assays are a prerequisite for HTS campaigns. So far, HTS implementation by label-free MS-based readouts was hampered by the lack of speed largely induced by the transfer of solubilized analytes into ESI mass spectrometers.6,8 This issue has recently been overcome by the introduction of fast and precise MALDI-TOF instruments concatenating the label-free MS readout with speed and robustness of conventional label-based readouts. Several publications in this context demonstrate the extensive efforts in the drug discovery community to integrate MALDI-TOF into the HTS environment.11–14,16,19 However, they largely lack a concept of providing a robust and reproducible transfer of samples from assay plates as well as matrix solution onto the MALDI target plates in an HTS-compatible manner.
We demonstrate a fully automated spotting concept to transfer samples derived from biochemical assays onto MALDI target plates. The structure of the compiled automation units is schematically illustrated in
Figure 1A
Verification of Reproducible Spotting Quality
Crucial for high-throughput analysis of biochemical assays by MALDI-TOF is the simultaneous and accurate deposition of both sample and matrix solutions onto MALDI target plates. The presented spotting station employs a multichannel pipettor equipped with 1536 ceramic tips. This enables simultaneous deposition of an entire 1536-well plate onto the MALDI target plate. In comparison to alternative liquid-handling systems that have been reported previously (Echo and Mosquito), the CyBio Well vario offers advantages such as (1) simultaneous deposition of sample spots in 384- or 1536-well format, (2) diminished consumable consumption, and (3) providing the possibility to implement on-target washing of sample spots. One potential limitation of the multichannel pipettor is the risk for carryover, which has to be reviewed carefully for each biochemical assay and analyte, respectively.
To provide homogenous spot shapes in a reproducible fashion, we decided to apply double-layer spotting. 20 Here, the matrix solution (100 nL) is transferred onto plain steel plates. After desiccation, this prespotted matrix serves as an anchor system for the following transfer of a matrix/sample (100 nL each) mixture being aspirated successively and dispensed together onto the dried matrix spots. Drying of transferred sample and matrix solution is aided by a vacuum dryer, which provides fast and reproducible drying times crucial for fully automated target plate preparation. Between each spotting step, ceramic tips are washed three times with 70% isopropanol and 30% water/0.1% TFA (v/v) to prevent carryover, which has been shown to be sufficient for the present peptide-based assay (data not shown).
Generated matrix/sample spots are of highly homogenous shape, as presented in Figure 2A . This constitutes a generic target plate composed of 1536 sample spots along with the zoom onto two neighboring spots in order to improve visibility of the spot shape. Homogenous spot shapes are prerequisite for equally distributed analytes enabling sample analysis using only one centric laser position per spot, which in turn provides fast cycle times. To confirm the ability to provide homogenous spot shapes over time, we performed the entire spotting procedure for forty 384-well assay plates, mimicking the actual composition of the previously published PTP1B inhibitor screen assay. 16
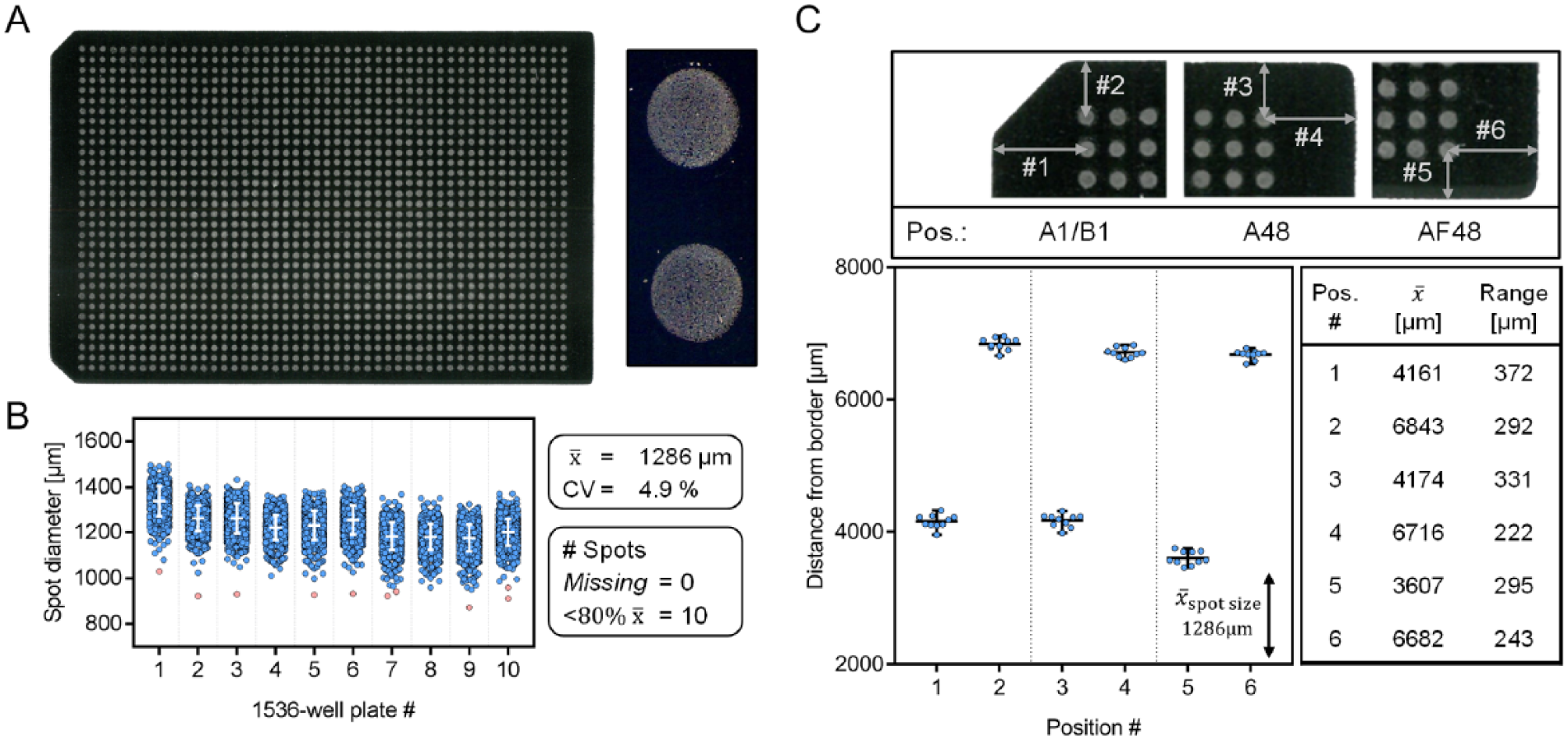
Assessment of the fully automated matrix-assisted laser desorption/ionization (MALDI) target plate preparation. (
A direct measure for reproducible and accurate spotting is provided by the analysis of the spot diameter for each of the 15,360 spots in this experiment. In Figure 2B , the spot diameter is plotted as a function of the 10 × 1536 target plates to uncover trends in spot size. Overall, spot sizes with high precision (coefficient of variation [CV] <5%) and a low rate of outliers (10/15,360 spots <80% of mean diameter) were generated. This, combined with no missing spot on the tested sample plates, implies high accuracy and reliability of the presented procedure.
Furthermore, for automated MALDI-TOF analysis, it is crucial to ensure reproducible positioning of sample spots on the target plate. Positioning of sample spots is taught to the MALDI-TOF for the first target plate only. The instrument expects that each of the subsequent target plates contains identical spot placement. Hence, the automatic spotting procedure is required to fulfil this assumption. For this purpose, we decided to check the positioning of the spot grid as outlined in Figure 2C . Distances between the centers of selected spots to the border of the target plates were determined at six positions, precisely defining the positioning of the entire spot grid. The compiled data for the 10 × 1536 target plates demonstrate that the border distance varies by less than 400 µm, which is much smaller than the mean spot size within this experiment (1286 µm). This demonstrates reproducible positioning of the 1536-spot grid and entails a negligible risk for spots to move out of the MALDI laser focus, which would result in missing data points.
To verify these findings, we reproduced the entire experiment on another day. Data of equal quality were generated and are presented in
MALDI-TOF Analysis for HTS Applications
After confirming the fundamental prerequisite of reproducible and accurate spotting, we intended to examine the suitability of the mass spectrometric readout for HTS campaigns. For this purpose, target plates being prepared to verify quality parameters of the spotting procedure contained a mixture mimicking the actual composition of a biochemical assay. Three peptides representing substrate, product, and internal standard of the PTP1B enzymatic reaction were enclosed at constant concentration levels, which were adapted from our previous publication assessing MALDI-TOF applicability as versatile readout within drug discovery campaigns.
16
A representative mass spectrum of this assay imitation is outlined in
Scatter plots representing the peak area of the internal standard (
Fig. 3A
) and product (
Fig. 3B
) illustrate the variability of the intrinsic fluctuating MALDI-TOF signal. However, the mean peak area for the 1536 spots per plate remains stable, confirming the persisting sensitivity of the MALDI-TOF instrument for the analyzed 15,360 spots without intermediate instrument maintenance (e.g., ion source cleaning or adjustment of laser power). To provide accurate and reproducible quantification by MALDI-TOF, it is necessary to calculate the ratio between the analyte and a reference. In our concept study, we used the isotope-labeled counterpart of the product peptide for this purpose. However, calculation of the product/substrate ratio provides a measure of enzymatic activity with equal accuracy and could be applied instead as suggested previously.
13
This ultimately leads to reduced expenses for HTS campaigns. Nevertheless, we observed diminished ionization efficiency of the phosphorylated substrate peptide, which may hamper data evaluation due to limited presence of substrate peaks. This is in line with previous knowledge that phosphate groups reduce the ionization efficiency of peptides.
21
Consequently, for this study, it is favorable to apply an isotope-labeled internal standard of the product peptide. This entails identical desorption/ionization characteristics because of its equivalent chemical nature and hence provides ideal properties for accurate measurement of product quantity. Calculation of the product/internal standard ratio reduced the mean coefficient of variation of the MS signal from 96% (internal standard) and 92% (product) respectively, to 4.0% (see
Fig. 3C
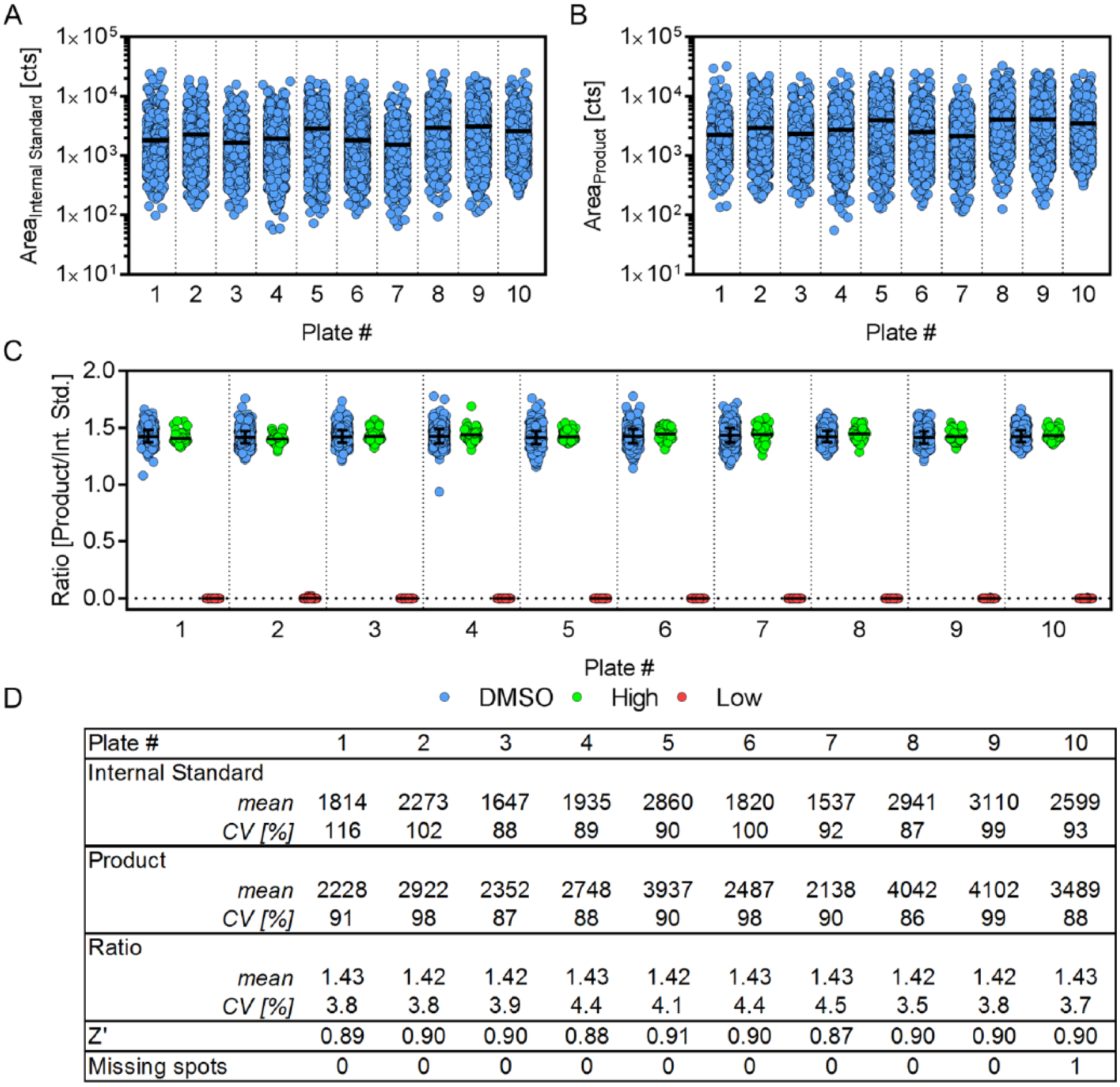
Verification of accurate and robust matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) analysis. (
To verify these findings, we performed two confirmation experiments: (1) reanalysis of the target plates to prove the repeatability of MALDI-TOF MS analysis as well as (2) repeated implementation of the entire procedure (spotting and MALDI-TOF analysis) to ensure general technical repeatability of the established procedure. Both reanalysis (
Implementation of MALDI-TOF for HTS
In previous sections, we demonstrated robust and reproducible spotting as well as competitive data quality provided by the label-free MS readout. Another major aspect of MALDI-TOF HTS integration is the achieved speed of the proposed workstation setup. For this purpose, we determined the time required for the spotting procedure and the MALDI-TOF analysis along with the time required for the biochemical assay, which was also conducted on a fully automated system. Table 1 encompasses the obtained process times depending on the number of acquired data points.
Overview of Cycle Times for Biochemical Assay, Automated Spotting, and MALDI-TOF Analysis in Dependency of Acquired Data Points.
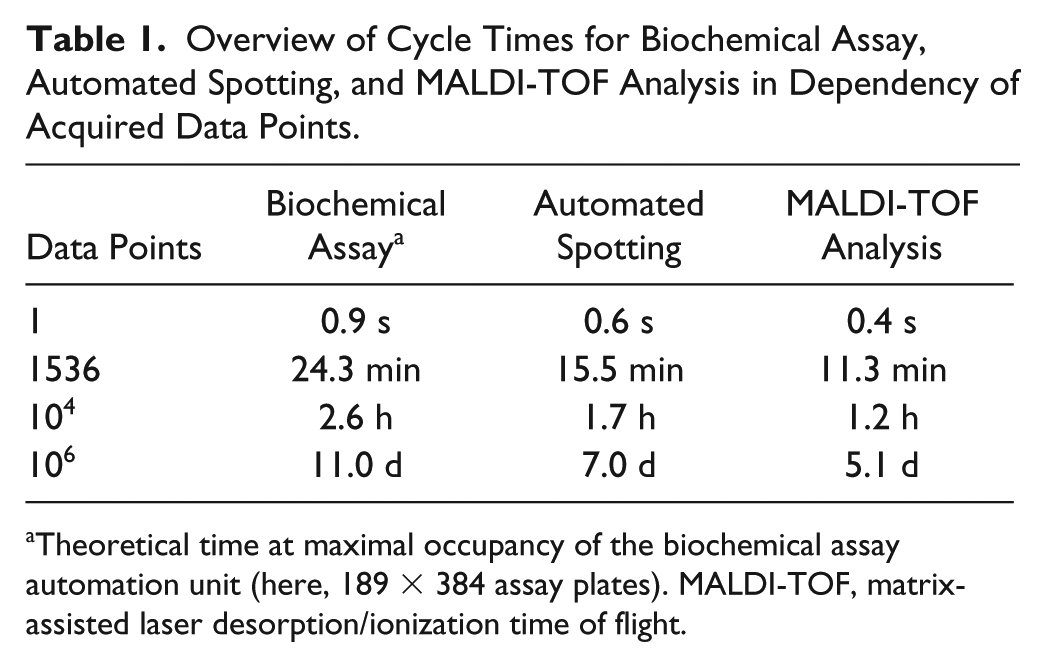
Theoretical time at maximal occupancy of the biochemical assay automation unit (here, 189 × 384 assay plates). MALDI-TOF, matrix-assisted laser desorption/ionization time of flight.
For the presented assay, we observed fast analysis times by the MALDI-TOF instrument (0.4 s/sample), which were superior to the cycle times provided by the automated spotting procedure (0.6 s/sample) as well as the biochemical assay (0.9 s/sample). These distinct cycle times make the separated workstation layout advantageous to generate degrees of freedom as compared with an integrated MALDI-TOF in-line reader. Vacant instrument time could be used for alternative vital processes such as instrument maintenance, development/testing of novel assays, or even running another assay prespotted on another device. Of note, the presented MALDI-TOF analysis time holds true only for the presented analytes. The MALDI-TOF cycle time is vastly dependent on analyte characteristics, such as (1) chemical characteristics of the analyte (e.g., polarity, proton affinity), (2) stoichiometry, (3) buffer components (e.g., ion suppression, adduct formation), and (4) data quality requirements (single-dose screening, profiling campaigns). MALDI-TOF instrument settings (e.g., number of shots, shot steps, laser power) should be aligned with these aspects to provide sufficient sensitivity to fulfil the requirements for accurate quantification.
We demonstrate the potential applicability of the presented concept to screen more than 1 million compounds for the PTP1B enzymatic assay in a short time frame (see Table 1 ). Nevertheless, persistent data quality within full-deck screening campaigns using MALDI-TOF needs to be determined. To assess the robustness of the concept, we again performed the analysis of 10 × 1536 assay plates mimicking the biochemical assay composition. Five of these plates were analyzed prior to and five after the analysis of 1 million spots (containing a mixture of matrix/assay buffer) on the MALDI-TOF instrument, reflecting the analysis of 660 × 1536 assay plates.
We intended to perform this experiment without any interim maintenance of the instrument (e.g., ion source cleaning) to evaluate its ruggedness. Only laser power was adjusted prior to the final measurements to produce appropriate signal intensity. In an actual screening campaign, this would be performed daily to ensure sufficient peak intensity for accurate quantification. Here, we required a laser power increase of 8% (44%
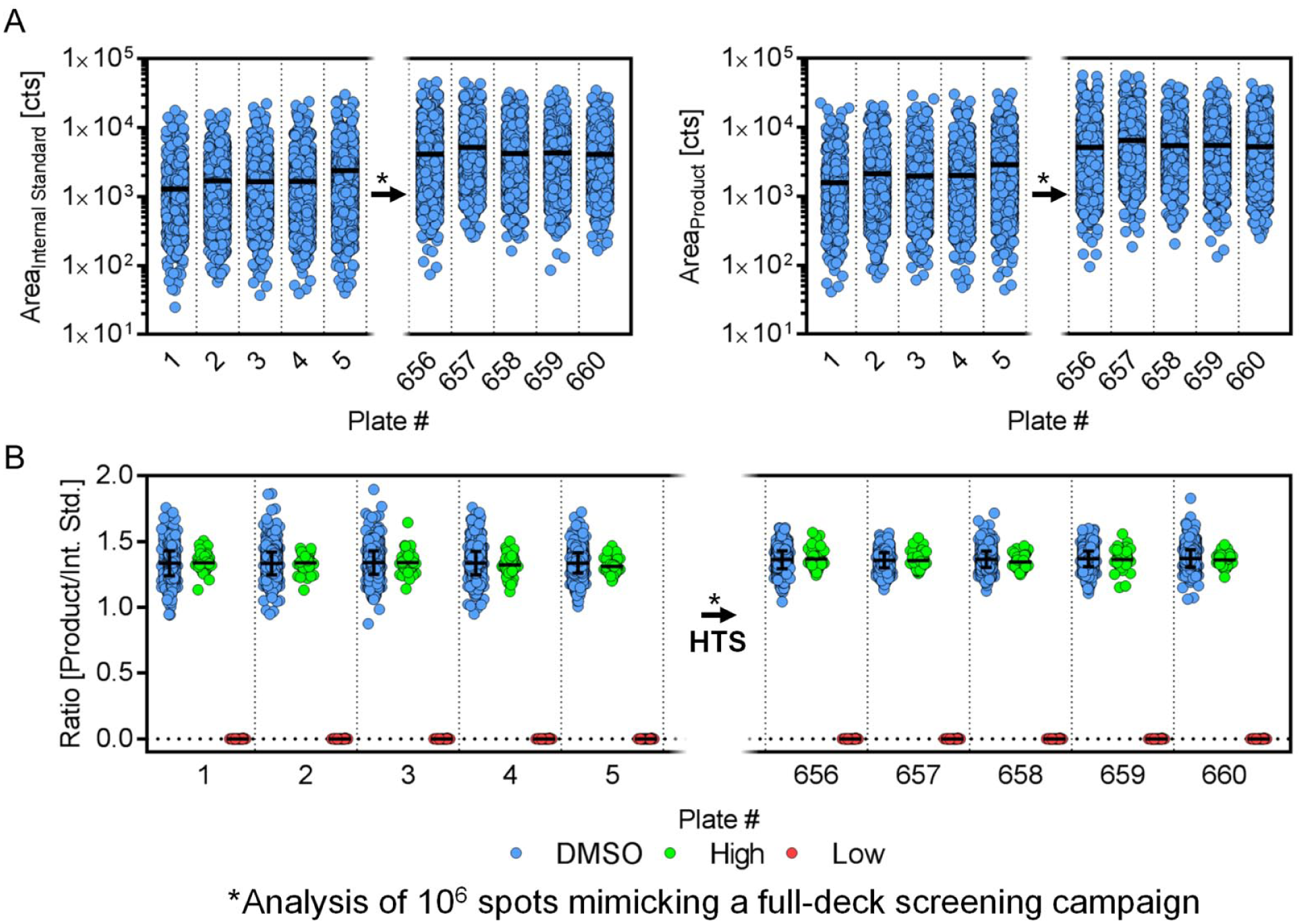
Examination of matrix-assisted laser desorption/ionization time-of-flight ruggedness within high-throughput screening campaigns. (
Optional: On-Target Washing Procedure to Increase Sensitivity
A beneficial ability of the applied multichannel pipettor is the on-target washing procedure, which expands the repertoire of strategies within assay development. On-target washing provides the potential to remove buffer components, such as sodium chloride, that are frequently present within assay buffers in order to preserve enzyme conformation and activity. Reduction of these salts and other components in the assay buffer increases analyte ionization efficiency due to diminished ion suppression as well as adduct formation (e.g., sodium adducts). The automated on-target washing procedure is visualized in the supplemented video clip (2:21–2:40 min in
To assess the potential of this procedure, we spotted two 1536-well plates mimicking the assay conditions of the PTP1B inhibitor screen. One plate was spotted with and one without subsequent on-target washing using 0.3 µL of 10 mM NH4H2PO4/0.1% TFA. The peak area of internal standard, product, and substrate peptide is illustrated in
Figure 5A
, demonstrating a significant peak area increase for all three (fold change: FCInt.Std. = 2.6, FCProduct = 2.4, FCSubstrate = 2.0), whereas the ratio between product and internal standard remains unaffected (see
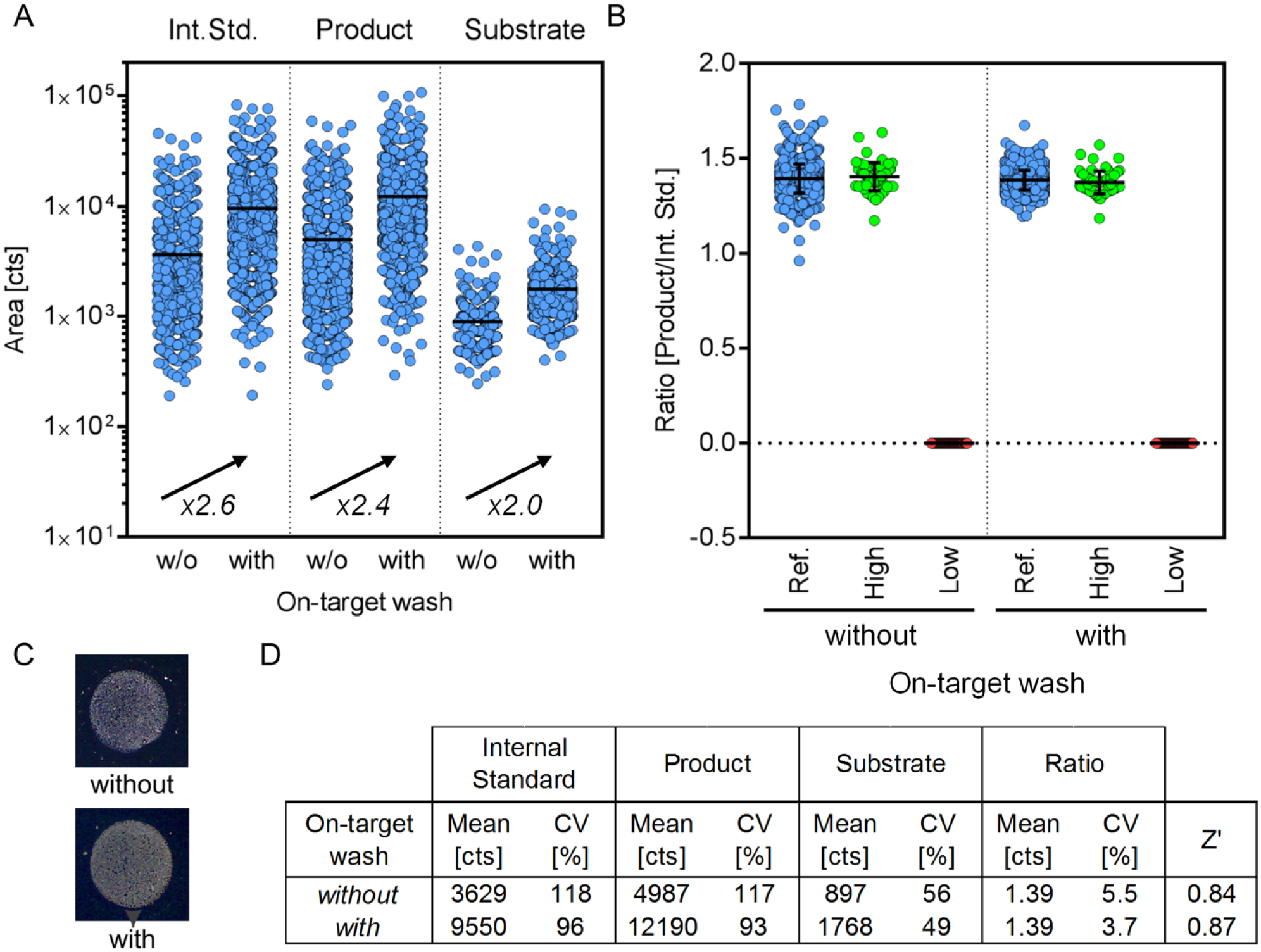
Assessment of the optional on-target washing concept. (
Overall, the data demonstrate the high quality of the on-target washing procedure by preserving spot characteristics as well as data quality measures (CV, Z′) while significantly elevating the signal intensity (see Fig. 5D ). This provides a valuable opportunity for fully automated assay development of upcoming target classes. We estimate that on-target washing concepts will be crucial to make MALDI-TOF analysis accessible for several analyte/buffer compositions. However, its applicability should be reviewed carefully for new analyte/buffer compositions to verify assets and drawbacks (e.g., dissolution of the analyte itself). If successful, this concept will have a significant impact on the MALDI-TOF analysis by (1) facilitating the analysis of formerly impractical buffer compositions, (2) diminishing the required number of laser shots and laser power to achieve a sufficient peak area for quantification, (3) reducing ion source contamination, and (4) boosting analysis time. Of note, among the commercially available automation equipment for MALDI target plate preparation used in the field of HTS, an efficient on-target washing procedure is solely provided by the CyBio Disk multichannel pipettor being induced by their mode of action (acoustic liquid handler Echo [Labcyte] and the nanoliter dispenser Mosquito [TTP Labtech]). For the CyBio Disk, it might be activated or inactivated depending on the respective project in a fully automated fashion. It has to be mentioned that the presented procedure elongates the spotting procedure by approximately 2.2 min (spotting time: twith OTW = 17.7 min/1536 plate, 0.7 s/spot; twithout OTW = 15.5 min/1536, 0.6 s/spot). However, the potential to make hitherto inaccessible analyte/buffer compositions MS compatible outweighs this limitation.
Fully Automated Analysis of Selected Structure Representatives
Finally, to validate the entire concept composed of biochemical assay, fully automated spotting, and MALDI-TOF analysis, we performed a single-concentration screening for PTP1B antagonists using 4986 compounds. Experiments were performed in duplicate on 2 separate days using the presented MALDI concept and a previously published AlphaScreen protocol 16 at a concentration of 5 µg/mL for each compound. A complete 384-well plate containing DMSO only was enclosed to assess the intrinsic assay variability. Here, we observed a comparable CV for the two orthogonal assays (CVAlphaScreen = 6.7%/7.0%, CVMALDI-TOF = 4.4%/4.9%). This, combined with Z′ values of 0.83/0.81 (Replicate 1/2, AlphaScreen) and 0.87/0.85 (Replicate 1/2, MALDI-TOF), demonstrates robust and reproducible data quality.
Results of the single-dose experiment are illustrated as a scatter plot in
Figure 6A
. Statistically significant inhibition of PTP1B induced by compound incubation is defined by a POC smaller than or equal to the mean POC ±3 SD being calculated for the whole screening data set. In addition, the number of compounds inducing a POC smaller than 50% was determined. The compiled number of hits and the associated hit rates for the two assay technologies are summarized in
Figure 6B
and
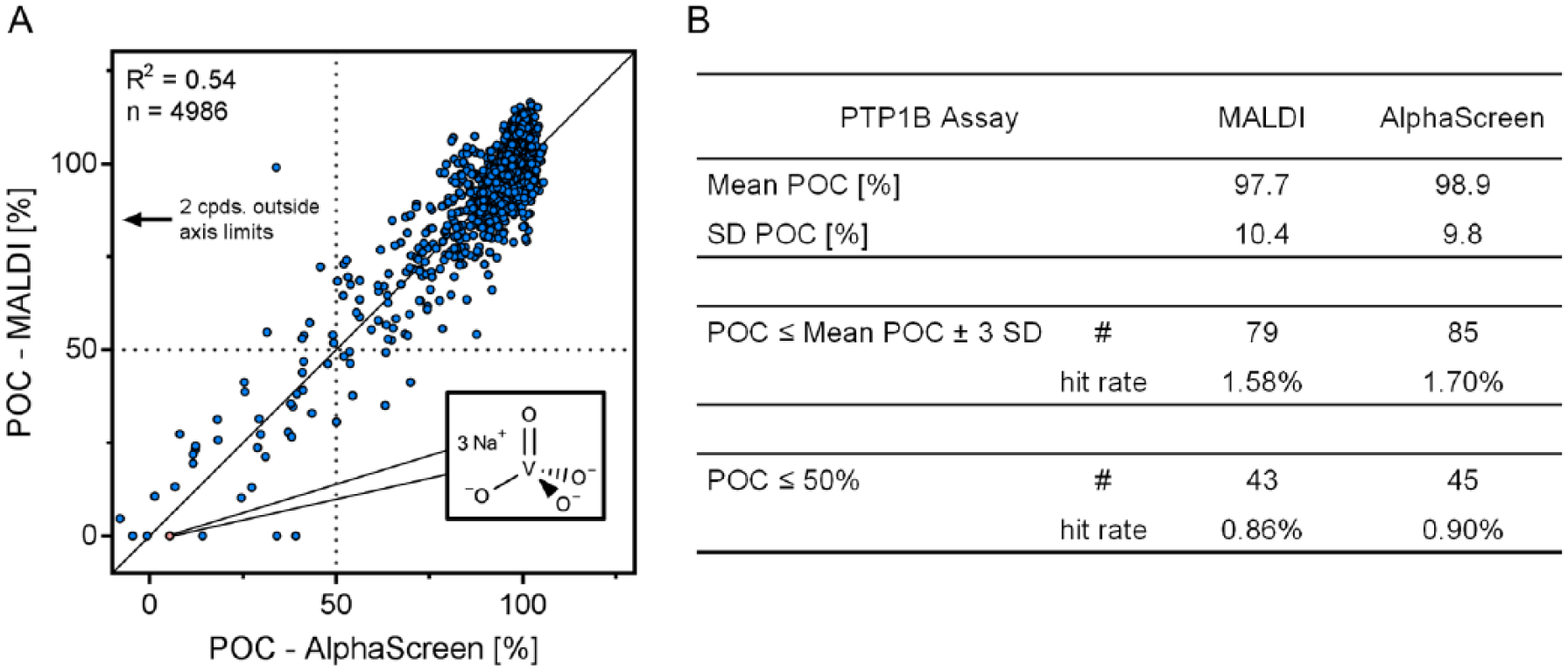
Comparison of label-free and label-based readouts for the protein tyrosine phosphatase 1B (PTP1B) inhibitor screen. Single-dose experiment (4986 compounds) for the discovery of PTP1B antagonists by AlphaScreen and matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF). (
Overall, 98 compounds were identified as hits within this experiment (POC < mean POC ±3 SD) composed of 67% (66 compounds) that were identified by both technologies (19 compounds only by AlphaScreen, 13 by MALDI-TOF). Applying the threshold of POC <50% leads to 53 hits containing 35 common ones (10 identified only by AlphaScreen, 8 by only MALDI-TOF). This confirms the comparability of the two orthogonal assay technologies. Interestingly, only three compounds in this experiment showed substantial difference between the assay technologies, which could be induced by compound interference of the luminescence-based readout. Compound interference within the AlphaScreen assay is generally known 24 but could also be present in the MALDI-TOF assay (e.g., covalent substrate/product modification by compounds). This, combined with the general assay variability (SD ≈ 10%), might be the reason for differences in the set of identified hits by the two technologies. Future work is required to characterize the exact origin of differences between the orthogonal assay technologies. However, the presented data being generated by MALDI-TOF together with reduced consumables, necessity of labeled substrates and/or specific antibodies, as well as reduced risk for compound interference marks the upcoming age of label-free MALDI-TOF readout within drug discovery.
In conclusion, we present the establishment of an in-house–built fully automated spotting system that concatenates the biochemical assays with the label-free MALDI-TOF MS readout. This empowers the hitherto elusive MALDI-TOF HTS integration and was concisely demonstrated by (1) highly reliable preparation of sample/matrix spots (accurate size/positioning), (2) excellent data quality of the MALDI-TOF readout (CV <5%, Z′ >0.85), (3) micro-HTS–compatible cycle times for spotting and MS analysis, as well as (4) instrumental and analytical robustness enabling the implementation of a 1-million-compound screen without intermediate instrument maintenance. Lastly, implementation of a PTP1B inhibitor screen for 4986 compounds revealed consistent data between label-free (MALDI-TOF) and label-based (AlphaScreen) assay technologies, confirming its applicability in actual screening efforts. This gives access to more physiological relevant assays, diminishing the necessity for follow-up screening concepts (counterscreens) to identify false-positive primary hits and ultimately provide the potential to reduce costs and time of drug discovery campaigns. Overall, the presented data fill in the blanks of label-free MALDI-TOF integration into HTS campaigns and further emphasize its viable and versatile role for drug discovery research per se.
Supplemental Material
Supplemental_Material – Supplemental material for Automated MALDI Target Preparation Concept
Supplemental material, Supplemental_Material for Automated MALDI Target Preparation Concept by Martin Winter, Robert Ries, Carola Kleiner, Daniel Bischoff, Andreas H. Luippold, Tom Bretschneider and Frank H. Büttner in SLAS Technology
Supplemental Material
Supplemental_Table_S1 – Supplemental material for Automated MALDI Target Preparation Concept
Supplemental material, Supplemental_Table_S1 for Automated MALDI Target Preparation Concept by Martin Winter, Robert Ries, Carola Kleiner, Daniel Bischoff, Andreas H. Luippold, Tom Bretschneider and Frank H. Büttner in SLAS Technology
Footnotes
Acknowledgements
We thank Margit Bauer for providing the enzyme PTP1B. Further, we thank Meike Hamester, Franz J. Mayer, Miriam Denkert, and Astrid Erdmann from Bruker Daltonics (Bremen, Germany) as well as Adrian Siemers and Swen Tyrasa from Analytik Jena (Jena, Germany) for their valuable input and helpful discussion.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. All work was completed as work-for-hire for the employer Boehringer Ingelheim Pharma GmbH & Co. KG.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
