Abstract
The inherent ability of nucleic acids to recognize a complementary pair has gained wide popularity in DNA sensor applications. DNA molecules can be produced in bulk and easily incorporated with various nanomaterials for sensing applications. More complex designs and sophisticated DNA sensors have been reported over the years to allow DNA detection in a faster, cheaper, and more convenient manner. Here, we report a DNA sensor designed to function like a switch to turn “on” silver nanocluster (AgNC) generation in the presence of a specific DNA target. By defining the probe region sequence, we are able to tune the color of the AgNC generated in direct relation to the different targets. As a proof of concept, we used dengue RNA-dependent RNA polymerase conserved sequences from all four serotypes as targets. This method was able to distinguish each dengue serotype by generating the serotype-respective AgNCs. The DNA switch was also able to identify and amplify the correct target in a mixture of targets with good specificity. This strategy has a detection limit of between 1.5 and 2.0 µM depending on the sequence of AgNC. The DNA switch approach provides an attractive alternative for single-target or multiplex DNA detection.
Introduction
Dengue is a prevalent viral disease worldwide, transmitted mostly by Aedes aegypti mosquitos. 1 The principal vector (Aedes aegypti) is a day-biting mosquito that is able to breed in natural and artificial water sources. 2 Approximately 390 million dengue infections are reported annually in 125 countries, and more than 2.5 billion people living in the tropic and subtropic regions are at risk of infection.3,4 Dengue infection appears to be one of the chief reasons for morbidity and mortality worldwide because of the growing virulence over the past half-century. 5 Approximately 12,000 deaths worldwide are associated with dengue infections every year. 6 The first ever dengue vaccine, Dengvaxia®, has been successfully developed by Sanofi Pasteur for dengue prevention and approved to be used in some highly dengue-endemic regions. 7 Even so, recent investigations have raised concerns about the potential for antibody-dependent enhancement (ADE) to hinder the development of dengue vaccination.8,9 This highlights the importance and need for dengue serotyping in endemic regions.
There are four infective serotypes (DENV-1 to -4), with ⩾30% differences in their overall amino acid sequences responsible for dengue infections. 10 Dengue infection usually causes acute febrile illness known as dengue fever. However, certain cases will develop into dengue hemorrhagic fever (DHF) and dengue shock syndrome (DSS), which is life threatening.1,11 It is crucial to distinguish the dengue serotype during the infection to determine primary or secondary infections. Infection with one dengue serotype confers a lifelong immunity against that specific serotype yet affords transient cross-protection against other serotypes. Secondary infections with different dengue serotypes results in more severe risk than primary infections due to ADE. Antibodies produced during the primary infection are unable to neutralize the virus from a secondary infection. Instead, the antibodies form immune complexes with the virus and have high affinity toward Fcγ receptors. The binding subsequently allows the virus to enter the cells and replicate. 12
Aside from infection by a single dengue serotype, reports have shown that concurrent infection with multiple dengue serotypes is on the rise recently since the first report made in 1982 that occurred in Puerto Rico. 13 The correlation between concurrent infection and severity of diseases is still not fully understood. 14 Early prognosis of dengue concurrent infections is crucial to avoid complications caused by severe forms of the disease such as DSS or DHF. The gold standard assay for dengue serotyping is usually performed with virus isolation or detection of antibodies in blood. However, to identify dengue concurrent infections, these methods are too tedious and time-consuming, 15 and antibody levels are undetectable during the early phase of infection. 16 Nucleic acid sensing appears to be a more promising alternative in the determination of dengue virus infection as it would be faster, more sensitive, and more specific.17–19 The specificity of nucleic acid sensing is mainly attributed to the specificity of DNA strands to find a matching complementary DNA sequence by hybridization. Recently, we have shown the feasibility of nucleic acid detection of dengue RNA by the generation of silver nanoclusters (AgNCs) in either single-tube format or probe cocktail assay. 20 The performance of nucleic acid sensing over the past decade has highlighted the potential of this technique to be expanded for wider applications. Various nucleic acid sensing techniques have been reported for dengue detection such as reverse-transcriptase polymerase chain reaction,19,21 nucleic acid sequence–based amplification, 17 and transcription-mediated amplification assays. 22
Here, we propose the application of a DNA switch sensor to detect the presence of dengue DNA mediated by a toehold mechanism as a derivative from our previous work. 20 The previous work used a single color for serotyping, whereas this work allows the utilization of a two-color system to carry out the serotyping. The mechanism of the DNA switch sensor is illustrated in Figure 1 . The proposed DNA switch sensor is designed with one main probe (MP) that hybridizes with two dengue priming strands (DPSs). The design of both MP and DPSs is shown in Supplemental Figure S1 . Hybridization of MP with DPSs was done using a temperature-cascading process. Detection of dengue DNA by the DNA switch involves three stages. First, the presence of the desired dengue target will displace the hybridized DPS. The displaced DPS is later isolated by magnetic separation via the interaction between the biotin group on the MP with the streptavidin magnetic beads. Then, a common primer is introduced to bind with the 6-bp spacer and nicking site of DPS to initiate the amplification and nicking process. At this stage, AgNC strands will be produced as a result of the toehold displacement. Lastly, AgNC strands are reduced to form fluorescent AgNCs. We are able to distinguish between dengue serotypes by the resulting fluorescent emission peaks. This method also revealed a detection limit of at least 1.5 to 2.0 µM. This approach provides the possibility of regulating a fluorescent switch using the DNA toehold mechanism.
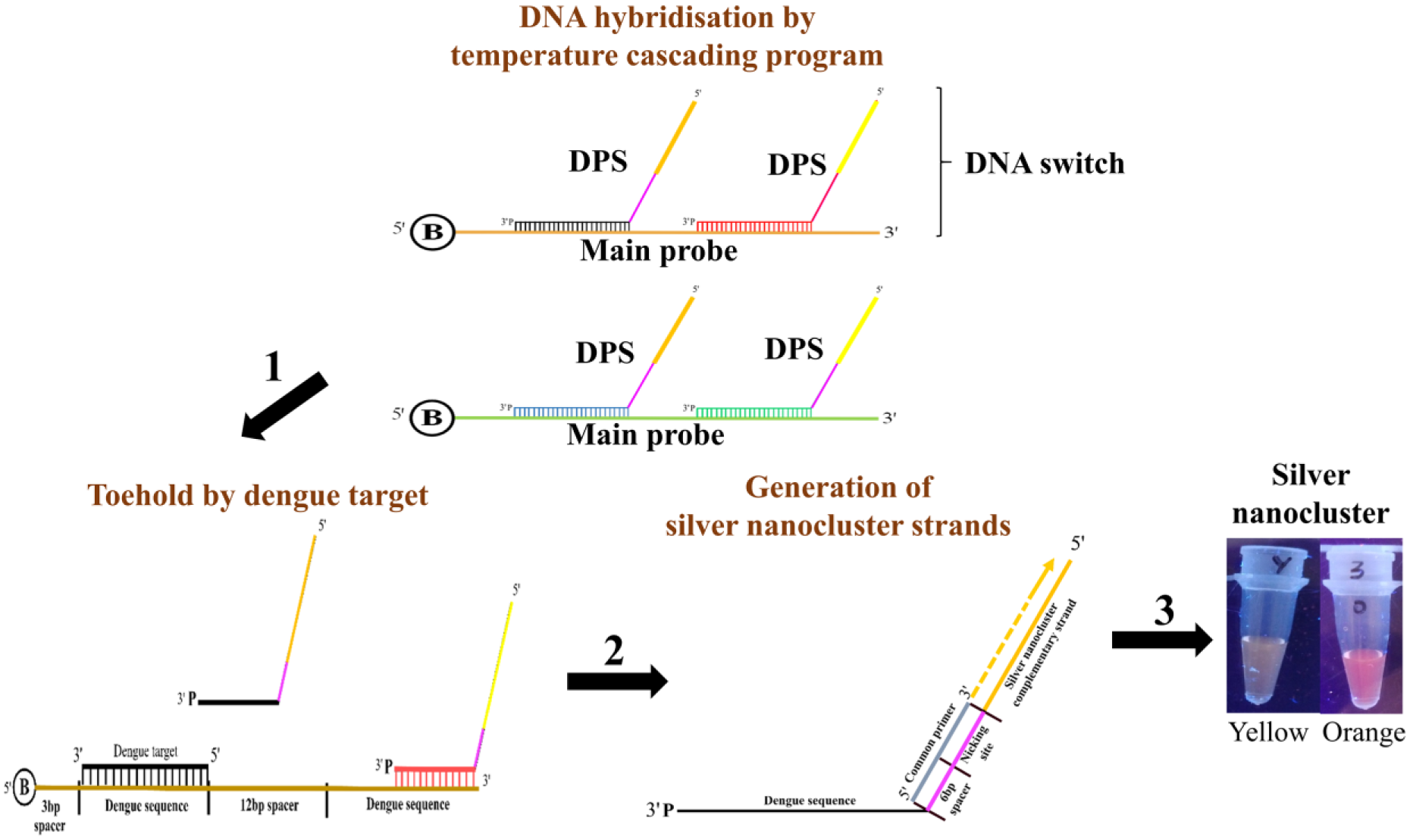
Schematic diagram of the overall DNA switch sensor workflow. The DNA switch sensor is composed of two main components: main probe (MP) and the dengue priming strand (DPS). Fabrication of the DNA switch commences with the hybridization of DPSs to MP by a temperature-cascading program. Then, addition of a target to displace the hybridized DPS is initiated via a toehold-mediated strand exchange mechanism (1). Binding of a common primer to the displaced DPS is required to initiate the polymerization and nicking stage to generate silver nanocluster (AgNC) strands (2). Finally, formation of AgNCs is achieved via reduction of silver ions with the nanocluster oligonucleotides (3).
Materials and Methods
Reagents and Materials
All oligonucleotides were synthesized by Integrated DNA Technologies (Lowa, US). The sequences of all synthesized oligonucleotides are shown in Supplemental Table S1 . Vent (exo-) DNA polymerase and Nt.BstNBI nicking enzyme were purchased from New England Biolabs (Ipswich, MA). Sodium borohydride (NaBH4) and silver nitrate (AgNO3) were purchased from Fisher Scientific (Waltham, MA). Streptavidin magnetic beads (1-µm hydrodynamic diameter) was purchased from Chemicell (Berlin, Germany).
Fabrication of DNA Switch
A total of 3 µM of each DPS was first hybridized with 3 µM of MP in 2 µL of buffer A (25 mM Tris-HNO3 [pH 7.9], 50 mM NaNO3, 5 mM Mg [NO3]2, 0.5 mM dithiothreitol). Buffer A must be prepared fresh before each run. The mixture was incubated with a cascading program from 95 °C to 25 °C (95 °C, 80 °C, 70 °C, 50 °C, 30 °C, 25 °C) with each temperature point maintained for 5 min. Then, the mixture was incubated with 50 µL of streptavidin magnetic beads. The streptavidin beads were washed thrice with 500 µL of distilled water before and after incubation with mixture. DNA switch can be stored in distilled water at 4 °C.
Target-Induced Toehold-Mediated Displacement of DPS
The distilled water with the DNA switch was first removed. Then, 3 µM of target with buffer A was introduced to the DNA switch. The beads were incubated at 37 °C with shaking at 800 rpm for 5 min followed by a final incubation at 37 °C without shaking for 15 min. The whole incubation process was repeated for 2 h. Separation of beads and solution was done with a magnetic separator (DynaMag-2 Magnet, Thermo Fisher Scientific, Waltham, MA). The solution was pipetted out for downstream process.
Isothermal Amplification of AgNC Strands
A total of 3 µM of common primer was introduced to 5 µL of toehold product from the previous step in buffer A. Hybridization of common primer to the toehold product was performed with the cascading program mentioned earlier. Then, 3 µL of buffer B (10 mM NaNO3, 20 mM NH4NO3, 20 mM Tris-HNO3, pH 8.8, 2 mM Mg[NO3]2, 0.1 % [v/v] Triton X-100), 1 µM dNTPs, 0.13 U Vent (exo-) DNA polymerase, and 0.40 U of Nt.BstNBI nicking enzyme were added to the solution. Distilled water was added to the mixture to a final volume of 20 µL and incubated at 55 °C for 5 h. The solution can be stored at 4 °C.
Formation of AgNCs
The resulting isothermal amplification mixture was mixed with 200 µM of AgNO3 and topped up to a final volume of 50 µL with distilled water. The mixture was centrifuged at 12,000 rpm for 5 min. The supernatant was collected and incubated on ice in the dark for 15 min. Then, 60 µM of NaBH4 was freshly prepared and added into the mixture. The solution was incubated in the dark at room temperature for 1 h. The formation of AgNCs upon reduction by NaBH4 was determined using a fluorescent spectrophotometer (Varian Cary Ecplise, Agilent, Santa Clara, CA). The excitation wavelength was set at 512 nm and 525 nm, respectively, for yellow and orange AgNCs. The emission was expected to be in the range of 572 nm and 612 nm, respectively. The excitation slit was set at 10 nm and 5 nm for emission slit.
Results
Construction of DNA Switch
The proposed DNA switch sensor consists of two main parts: MP and DPS. One MP could accommodate two DPSs, and each DPS could generate either yellow- or orange-emitting AgNC strands after polymerization and nicking. In this work, two sets of DNA switch sensors were constructed to detect all four dengue serotypes, as shown in Figure 1 .
Validation of the proposed system was done step by step to ensure the feasibility of the concept. Fabrication of the DNA switch commenced with the hybridization of DPSs to MPs using a temperature-cascading program, which was validated by agarose gel. All DPSs were successfully hybridized to their respective MPs, as a higher band was observed when DPS was introduced ( Fig. 2A , B ). Upon successful hybridization, the DNA switch was bound to streptavidin magnetic beads to validate the optimized conditions. Supplemental Figure S4 shows that the optimized condition for the assay was determined to be 3 µM of DNA switch binding to streptavidin magnetic beads at 20 °C for 2 h. The two sets of DNA switch sensors successfully showed displacement of their respective DPS upon introduction of appropriate dengue target strands ( Fig. 2C ). In contrast, no displacement of DPS was observed when target dengue strand was absent ( Fig. 2D ). Figure 2E shows the final DNA switch construct was successfully applied for dengue DNA target detection by producing a faint band (lane 2 of Fig. 2E ) that has a similar size to the AgNC (lane 4 of Fig. 2E ) after the polymerization and nicking process. No similar band was observed when the target DNA was not introduced into the reaction (lane 3 of Fig. 2E ).
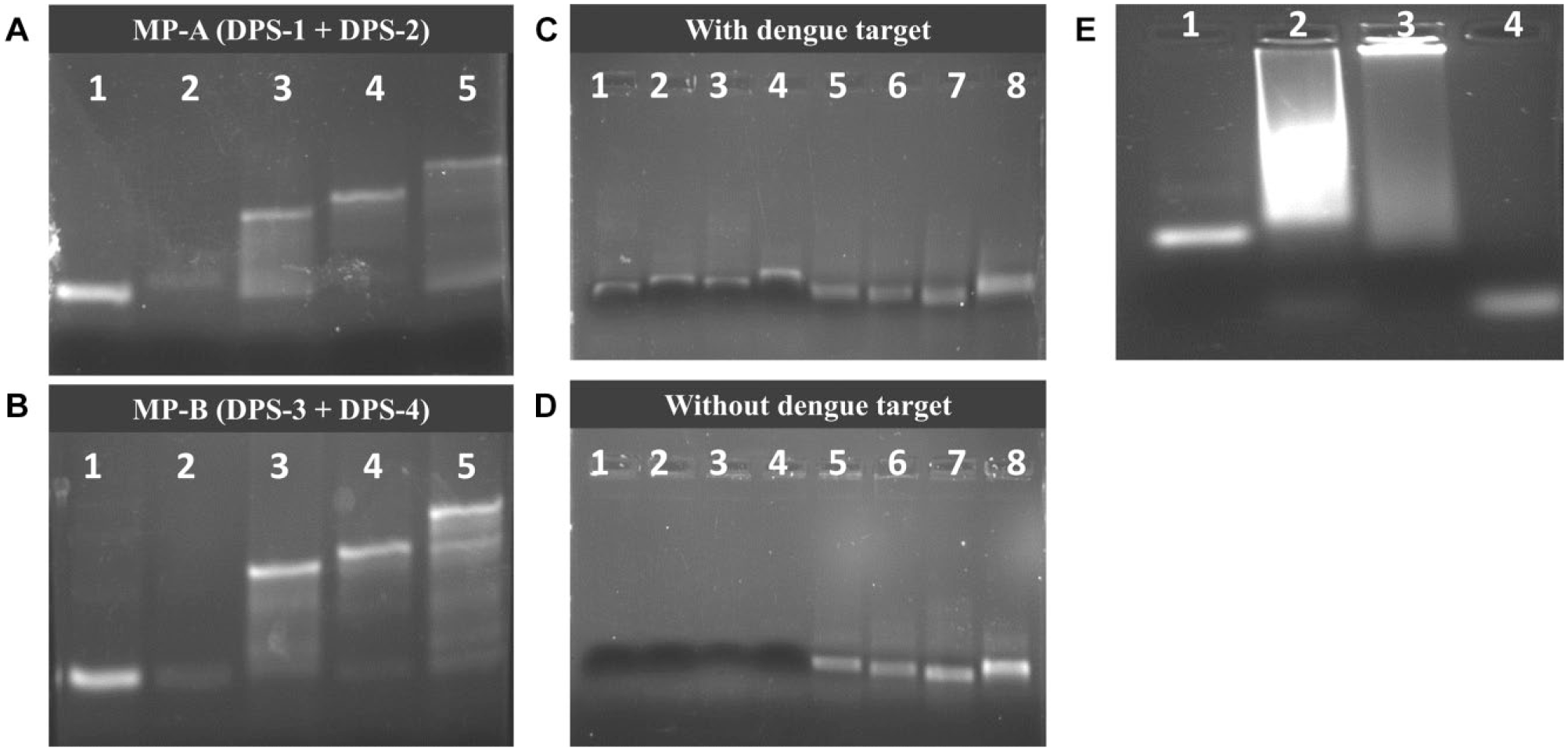
Validation of DNA switch construct. Hybridization of dengue priming strands (DPSs) to main probe (MP) via temperature-cascading program for (
Excitation and Emission Spectra of AgNCs
The excitation wavelengths of the yellow and orange AgNC were determined to be 512 nm and 525 nm, respectively ( Fig. 3A ). The emission wavelengths of the yellow and orange AgNC were found to be very close. The yellow and orange AgNCs gave emission wavelengths of 572 nm and 612 nm, respectively. The two AgNCs were then mixed together and excited at their respective wavelengths to validate whether there would be any cross-interference in the emission spectra. Figure 3B shows that two peaks were observed in the emission spectra when excited at 512 nm, where one peak was about 570 nm and the other peak was about 610 nm. Only one emission peak at about 610 nm was observed when excited at 525 nm. A clearer picture was seen when the AgNCs were excited separately at the same wavelength. Both yellow and orange AgNCs when excited at 512 nm produced a peak at 570 nm and 610 nm, respectively ( Fig. 3C ). Only the orange AgNC yielded a peak at 610 nm when excited at 525 nm ( Fig. 3D ).
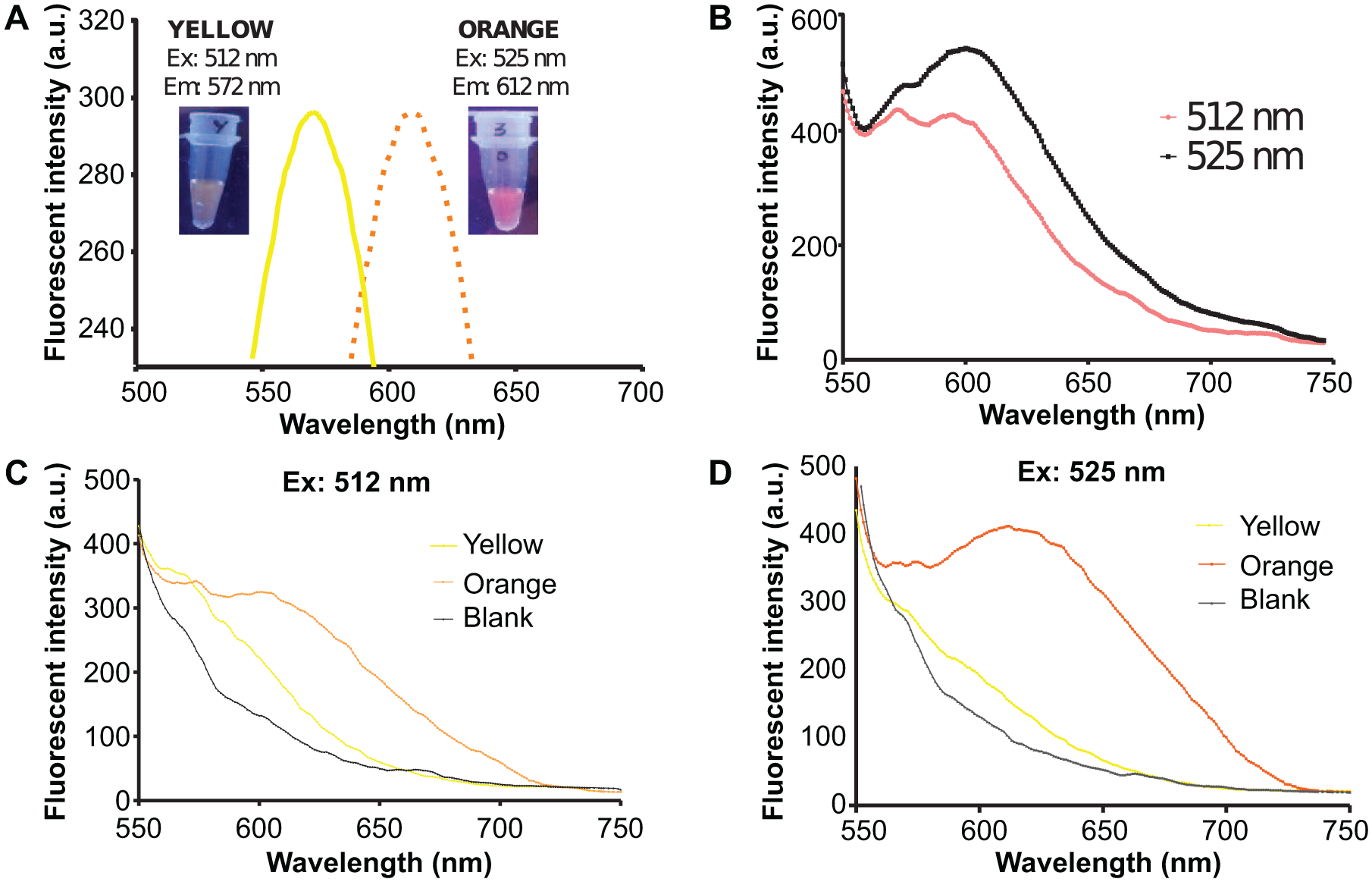
Emission and excitation spectra of silver nanoclusters (AgNCs). (
Cross-Reactivity Assays
Specificity of the proposed DNA switch was investigated by performing a series of cross-reactivity assays and using a mixture of target DNAs as demonstrated in Figure 4 . Both DNA switches (MP-A and MP-B) were able to detect and distinguish between single dengue serotypes as shown in Figure 4a to d . The correct emission of the AgNC was generated upon addition of the respective dengue target to the reaction. These results also show the specificity of the DNA switch sensor in detecting the correct target.
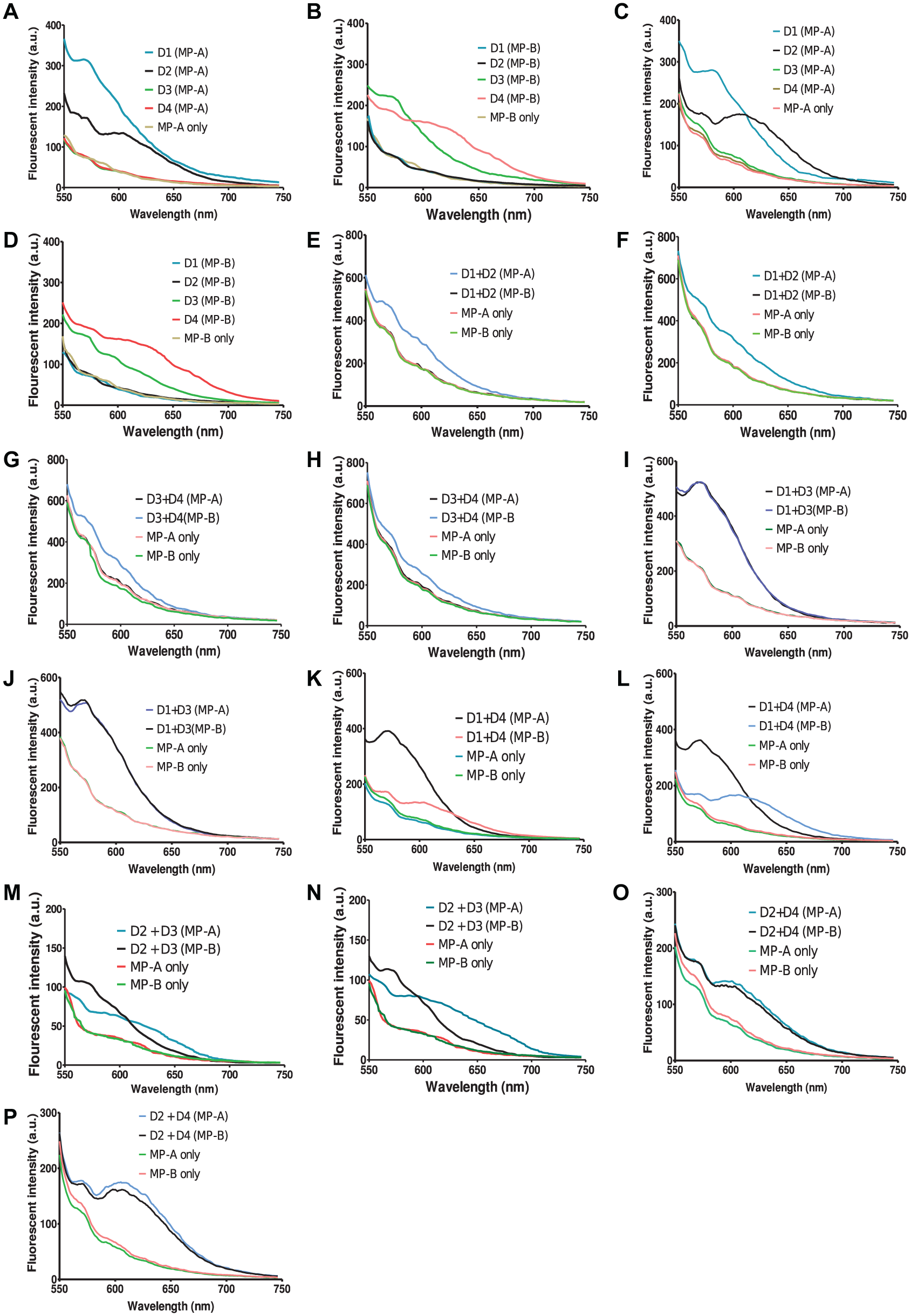
Emission spectra for cross-reactivity assays. Spiked dengue DNA targets were used to perform the cross-reactivity assays with excitations at 512 nm (yellow DNA switch) and 525 nm (orange DNA switch), respectively. D1: dengue target serotype 1; D2: dengue target serotype 2; D3: dengue target serotype 3; D4: dengue target serotype 4. Samples a, b, e, g, i, k, m, and o were excited at 512 nm. Samples c, d, f, h, j, l, n, and p were excited at 525 nm.
We then challenged the sensor with mixtures of serotypes of dengue targets. There are three sets of mixtures: (1) both serotypes are targets, (2) one is a target serotype while the other is not, and (3) both serotypes are not targets. Set (1) and (3) are represented by (e), (f), (g), and (h) of Figure 4 . The presence of two targets gave a combination of the emission spectra from both the AgNCs, which is similar to that demonstrated in Figure 3B . However, we did observe some reduction in fluorescent emission in the assay. The presence of two dengue serotypes that are not targets gave no emission as no toehold-mediated displacement of DPS was expected. For set (2), only one of the serotypes is present in the sample as a target, which is represented from panel (i) to (p). From Figure 4 , we can clearly observe the emission spectra of the respective DNA AgNC upon introduction of the respective target. The presence of other serotype targets that are not complementary to the MP would not affect the efficiency of the toehold displacement mechanism. This highlights the ability of the toehold mechanism to correctly displace complementary DNA targets based on their sequence to yield a specific sensor. This was possible even with the high variability of the displacement sequence region. In this case, a region with 13-bp similarity was sufficient to provide a specific displacement by toehold mechanism.
Detection Limit of DNA Switch Sensors
Figure 5 demonstrates the detection limit of the proposed system for both AgNCs at different concentrations of spiked dengue targets from 0 to 3 µM. The yellow AgNC DNA switch shows a relatively good linear correlation (regression coefficient R2 = 0.9911) between the dengue target concentration and fluorescent intensity ( Fig. 5A ). The detection limit of the yellow DNA switch was determined to be 1.0 µM by visual estimation from the graph ( Fig. 5A ). However, the detection limit of the yellow DNA switch according to the definition of 3σb/slope (σb, standard deviation of the blank) was determined to be higher at 1.5 µM. Hence, the detection limit of the yellow DNA switch was determined to be 1.5 µM. On the other hand, the orange AgNC DNA switch did not produce a satisfying regression coefficient value (R2 = 0.8971) together with a higher detection limit at approximately 2.0 µM by visual estimation ( Fig. 5B ) and about 1.7 µM according to the definition of 3σb/slope. The detection limit of the orange DNA switch is 2.0 µM. Fluorescent intensities close to the background value were seen at low concentrations of target concentration.
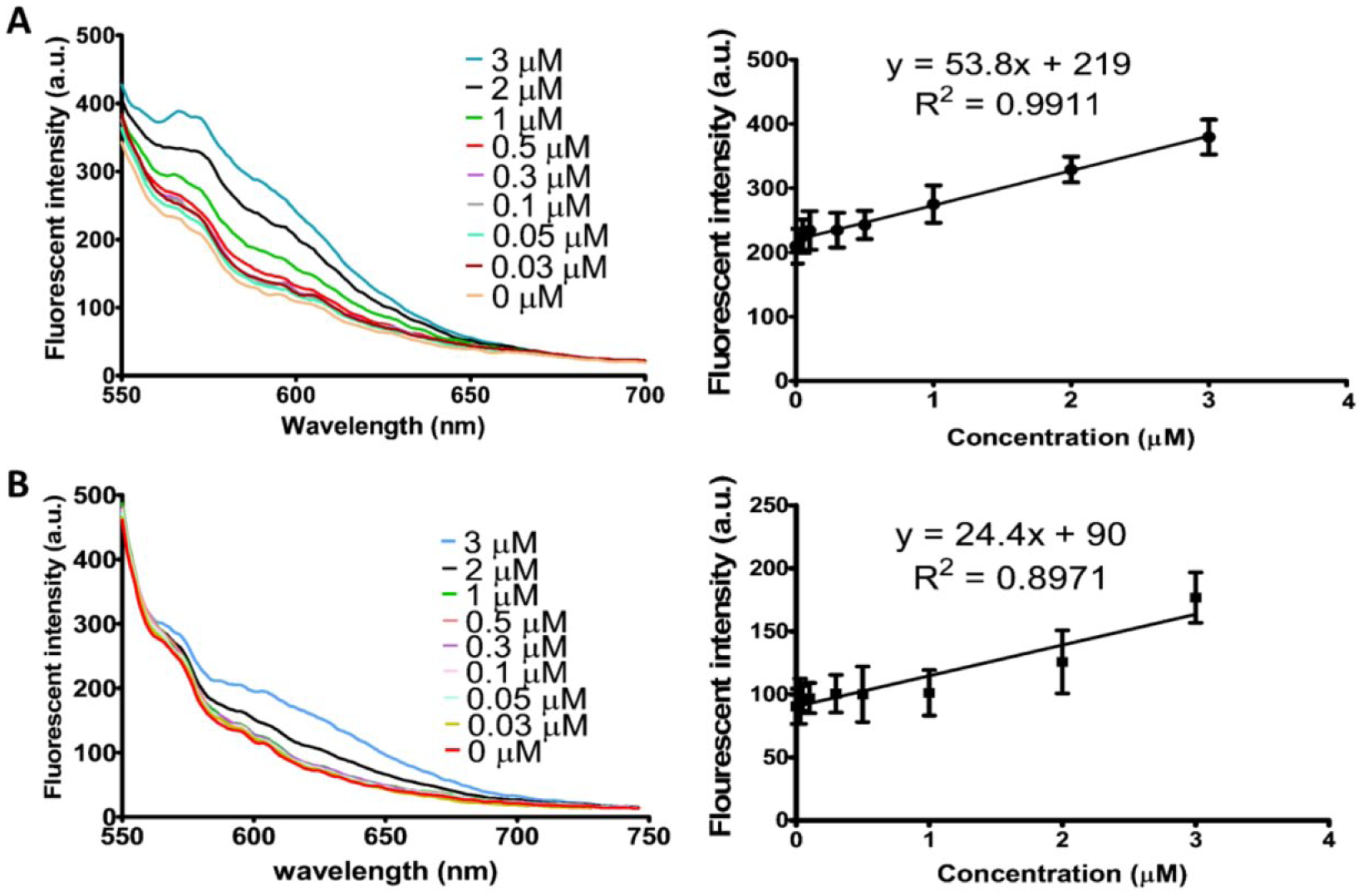
Limit of detection of spiked dengue target at a range from 0 to 3 µM. (
The detection limits for both yellow and orange AgNC DNA switches to detect dengue DNA target in the range of 1.5 to 2.0 µM could be considered as acceptable. In addition, it should be taken into account that potential loss of material during the transferring process of DPSs after toehold-mediated displacement could have a negative impact on the detection limit. Even so, the detection limits achievable by the system could be considered reasonable.
Discussion
The idea of the DNA switch sensor was inspired by the programmable DNA biosensor that allows for the correct target to switch a biosensor “on” or “off.” Hybridization of the correct target to the MP will initiate the toehold displacement of the respective annealed DPS. This particular setup was designed to turn on upon target presence and remains off in the absence of target. The “on” mode of the DNA switch allows for the hybridization of the common primer to serve as a primer for DNA polymerization. The subsequent nicking process is done to generate AgNC strands that are then reduced with silver ions to form fluorescent DNA AgNCs ( Suppl. Fig. S2 ). All DPSs have the same binding region for the common primer at a 6-bp spacer and nicking site. By identifying the color emitted by the DNA AgNC, we are then able to determine the dengue serotype present. The components of the DNA switch are made up of one MP with two DPSs. The MP consists of two dengue sequence regions for attachment to the respective DPS. Hence, two sets of DNA switches were designed to detect all four dengue serotypes ( Suppl. Fig. S2 ). Main probe A (MP-A) hybridizes with DPS serotype 1 and 2, whereas MP-B hybridizes with DPS serotype 3 and 4. DPS serotypes 1 and 3 will generate yellow-colored AgNC, while DPS serotypes 2 and 4 generate orange-colored AgNC ( Suppl. Fig. S3 ).
Construction of the DNA switch sensor was done using a temperature-cascading program from 95 °C to 25 °C, which allows for the DNA strands to come into proximity to improve hybridization. At 95 °C, the secondary structures of DNA that may interrupt DPSs binding were denatured to form linear strands. As the temperature is lowered, the kinetics of the DNA molecules are lowered, thus allowing for efficient hybridization of DPSs to MPs. Successful hybridization of DPS to MP increases the total size of DNA, which results in a reduced migration rate on agarose gel. Later, the displacement of DPS by dengue target was successfully performed by toehold-mediated mechanism. Both the dengue targets and DPSs are complementary to the dengue sequence region of MP. However, the binding regions of DPSs are nine bases shorter than the dengue targets. Hence, the binding affinity of dengue targets to MPs are stronger than DPSs, which causes the DPSs to be released from MPs when the dengue target is bound. In the presence of a dengue target, displacement of DPSs will occur, allowing it to bind with the common primer to give a band on agarose gel. On the other hand, in the absence of the dengue target, no invading strands were able to displace the bound DPSs. Hence, common primers were unable to bind to any DNA strands, thus resulting in no visible bands on agarose gel analysis. Separation of free DPSs from bound DPSs could be easily done via pipetting the free DPSs out from the solution.
Common primers were later introduced to bind with the free DPSs to serve as the primer to start the DNA amplification and nicking process by using DNA polymerase and the Nt.BstNBI nicking enzyme to generate AgNC strands. The common primer hybridized to DPSs would serve as the primer to initiate the process. As all DPSs have a similar binding site to the common primers, the common primers would be able to bind to DPSs to initiate the amplification and nicking process regardless of which DPSs is being displaced by the dengue targets. Generation of AgNC strands via DNA amplification and nicking is similar to the previous publication by our lab. 20 However, in our current proposed dengue serotyping system, two different-colored AgNCs were incorporated simultaneously. DPS serotypes 1 and 3 were designed to generate yellow-color AgNCs, whereas DPS serotypes 2 and 4 are able to generate orange-colored AgNCs. Hence, by separating MP-A and MP-B into two different tubes, we were able to recognize all four specific dengue serotypes via the color generated by the AgNCs.
The detection limit of both AgNCs used in this assay was determined by a fluorescence spectrophotometer using a range of spiked synthetic AgNC oligonucleotides at different concentrations (0–3 µM; Suppl. Fig. S5 ). The fluorescence intensity of the AgNCs showed a linear correlation with the concentration of AgNC, where the R2 value was more than 0.97 for both yellow and orange AgNCs, respectively. The detection limit for both yellow and orange AgNCs was determined to be 0.3 µM. This limit was determined as lower-concentration readings were not able to provide a good differentiation between the sample and background. Therefore, the fluorescence emission of AgNC with a concentration lower than 0.3 µM will be difficult to be detected by the fluorescence spectrophotometer. Excitation of the mixture of yellow and orange AgNCs showed that we were able to distinguish and identify the AgNC even when mixing both AgNCs together ( Fig. 3B ). This is crucial when it comes to the determination of concurrent infections, as there will be multiple dengue DNA present in the sample. Yellow and orange AgNCs have close excitation wavelengths. Hence, the emission peak of orange AgNC was still observed at a lower intensity even when not excited at the expected wavelength, making it easier to apply in an assay ( Fig. 3C ).
The cross-reactivity assays show the ability of our proposed DNA switch sensor to detect and differentiate dengue serotypes when one single target or even two targets are present in one sample. By identifying the emission spectra of the sample after polymerization and nicking, we were able to conclude the presence and differentiate the type of targets present. However, we did observe some reduction in fluorescent emission in the presence of two targets. This might be due to the splitting of dNTPs and enzymes for the production of two sets of AgNCs at the same time. Even so, further optimization of dNTPs and enzymes for production of two sets of AgNCs might be useful to enhance the yields. The specificity of the assay is good, as the presence of two dengue serotypes that are not targets gave no emission. This turn “off” mechanism occurs as there is no toehold-mediated displacement of DPS, resulting in no DNA AgNC strand formation. This highlights the suitability of the designed hybridization region for sample differentiation with good specificity.
In the context of dengue infection, we envision this method to be used for dengue serotyping after reverse transcription of dengue RNA. Dengue infection with a single serotype or concurrent infections with two serotypes could be identified via the proposed method. Despite the fact that concurrent infection with more than one serotype is very rare, 23 concurrent infections of multiple dengue serotypes has been reported in certain geographic areas where occurrence of multiple dengue serotypes are found to be co-circulating.23–25 This is mainly critical for use in endemic countries such as Malaysia.
Detection limits of the yellow and orange DNA switch sensors were at least 1.5 µM and 2.0 µM of dengue target, respectively. The high detection limit might be due to inefficient transferring of displaced DPSs during pipetting, unoptimized polymerization and nicking process, or reduced formation of AgNC due to the remaining salt impurities from the previous process. Also, the detection cutoff point of the fluorescence spectrophotometer at 0.3 µM might be another factor in the high detection limit when the dengue target concentration was low. Although other DNA template sequences for AgNC formation were incorporated in this proposed system, unfortunately not all AgNCs were well formed under similar conditions or exhibited lower emission intensities, especially blue and green fluorescent emitting AgNCs.26,27 This proved to be a bottleneck in generating a true multiplex (AgNC with four colors) system for DNA sensing.
Here, we described the successful development of a toehold-mediated strand exchange DNA switch sensor to generate AgNCs isothermally for the detection of dengue DNA targets. The proposed DNA sensor design is based on a switch mechanism whereby AgNCs are produced when the target is present and vice versa. The detection limit achieved by the DNA switches was within the range of 1.5 to 2.0 µM depending on the fluorescent strength of the AgNC used. Both the hybridized DPSs on MP can be recognized by the target and displaced for subsequent applications. The development of the DNA switch is universal as it allows for future expansion to other diseases with the change of the dengue region sequence to other desirable target sequences. Different types of AgNC fluorescence could also be formed via changing the AgNC sequence. However, the challenge is to identify a combination of AgNC that is able to form under one similar condition. Both DNA switches showed reproducible results and is a feasible concept for DNA multiplexing. The DNA switch sensor could also potentially be applied in a quantitative manner. The proposed method offers a cost-effective, easy, and simple alternative to standard DNA detection systems.
Supplemental Material
Supplemental_material_for_DNA_Switch_Toehold_Mediated_DNA_Isothermal_Amplification_for_Dengue_Serotyping_by_Chan,_et_al – Supplemental material for DNA Switch: Toehold-Mediated DNA Isothermal Amplification for Dengue Serotyping
Supplemental material, Supplemental_material_for_DNA_Switch_Toehold_Mediated_DNA_Isothermal_Amplification_for_Dengue_Serotyping_by_Chan,_et_al for DNA Switch: Toehold-Mediated DNA Isothermal Amplification for Dengue Serotyping by Soo Khim Chan, Akinori Kuzuya, Yee Siew Choong and Theam Soon Lim in SLAS Discovery
Footnotes
Acknowledgements
The authors would like to thank the members of the Antibody Engineering Group at INFORMM, USM for their kind assistance and support.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors acknowledge financial support from the Malaysian Ministry of Education through the Fundamental Research Grant Scheme (203/CIPPM/6711473) and Higher Institutions Centre of Excellence (HICoE) Grant Scheme (311/CIPPM/4401005).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
