Abstract
Histidine decarboxylase (HDC) is the primary enzyme that catalyzes the conversion of histidine to histamine. HDC contributes to many physiological responses as histamine plays important roles in allergic reaction, neurological response, gastric acid secretion, and cell proliferation and differentiation. Small-molecule modulation of HDC represents a potential therapeutic strategy for a range of histamine-associated diseases, including inflammatory disease, neurological disorders, gastric ulcers, and select cancers. High-throughput screening (HTS) methods for measuring HDC activity are currently limited. Here, we report the development of a time-resolved fluorescence resonance energy transfer (TR-FRET) assay for monitoring HDC activity. The assay is based on competition between HDC-generated histamine and fluorophore-labeled histamine for binding to a Europium cryptate (EuK)-labeled anti-histamine antibody. We demonstrated that the assay is highly sensitive and simple to develop. Assay validation experiments were performed using low-volume 384-well plates and resulted in good statistical parameters. A pilot HTS screen gave a Z′ score > 0.5 and a hit rate of 1.1%, and led to the identification of a validated hit series. Overall, the presented assay should facilitate the discovery of therapeutic HDC inhibitors by acting as a novel tool suitable for large-scale HTS and subsequent interrogation of compound structure–activity relationships.
Introduction
Histidine decarboxylase (HDC) is a fold type I pyridoxal 5′-phosphate (PLP)-dependent enzyme that catalyzes the α-decarboxylation of
The discovery of therapeutic small inhibitors of HDC would be accelerated by a biochemical high-throughput assay. Although several HDC assays have been proposed, none are available for high-throughput screening (HTS). The most commonly used assay to determine HDC activity measures the enzymatic liberation of 14CO2 using 14C-carboxyl-labeled substrate.
9
Although the assay is sensitive, it does not lend itself to HTS as it requires specialized equipment and presents special safety concerns related to the generation of radiochemical gas. An alternative nonradioactive approach uses phosphoenolpyruvate carboxylase and malate dehydrogenase to couple the generation of CO2 to convert NADH to NAD, a step that can be followed spectrometrically.
10
This system is also low throughput due to a required initial degassing of all reagents and the need to work with great haste to prevent atmospheric CO2 from diffusing back into the samples. In addition, a fluorometric assay has been developed involving
Here, we describe a time-resolved fluorescence resonance energy transfer (TR-FRET) method to assay HDC. The method is based on detecting the histamine release that occurs as the HDC-catalyzed reaction proceeds. We repurposed a commercially available TR-FRET histamine kit that was designed to detect histamine release from cells to an in vitro assay for measuring HDC activity. 13 The major advantage of TR-FRET is that the long emission half-lives of lanthanide-based fluorophores lead to fewer fluorescent artifacts from library compounds. Enzyme kinetics and inhibitor experiments indicate that this assay has a good signal-to-background ratio (S/B) and exhibits full inhibition. Assay validation in low-volume 384-well plates indicated a robust assay well suited to HTS campaigns for the discovery of small-molecule HDC inhibitors. Finally, the utility of the assay for surveying chemical libraries was probed through a pilot screen of ~7000 small molecules, an effort that gave good assay performance and a hit rate of 1.1%. Hit follow-up activities identified a series of confirmed inhibitors with IC50 values ranging from 2.4 to 18.9 µM. One such molecule was identified as a histidine analog that was confirmed as a bona fide HDC binder.
Materials and Methods
Reagents
The histamine TR-FRET kit was purchased from Cisbio Bioassays (Codolet, France; cat. 62HTMPEC).
Protein Expression
HDC protein expression and purification were carried out by the Nanyang Technological University Protein Production Platform. His6-tagged human HDC residues 1–477 corresponding to the catalytic domain were expressed in
Histidine Decarboxylase Assay
All enzymatic assays were performed in a 9 µL reaction mixture containing 5 nM HDC,
Assay Validation and High-Throughput Screening
The reaction was carried out with a final concentration of 5 nM HDC, 50 µM histidine, 68 nM PLP, and 2% DMSO. The assay volume was 9 µL in a 384-well plate. Enzyme stocks were diluted with buffer consisting of 25 mM HEPES, pH 7.4, 150 mM NaCl, and 0.3 mM DTT. Inhibitors/DMSO vehicle were dispensed to a dry plate using an Echo 555 acoustic liquid handler (Labcyte, Sunnyvale, CA); subsequently, enzyme solution was added using a BRAVO automated liquid handler (Agilent, Santa Clara, CA) to the plate and sealed. Sealed plates were incubated at room temperature for 30 min prior to substrate addition via BRAVO. After 30 min at room temperature, the reaction was quenched with acylation reagent and acylation buffer. Acylation was carried out for 1 h, followed by 5 h incubation of histamine-XL665 and anti-histamine-EuK antibody. The amount of acylation reagent, acylation buffer, histamine-XL665, and anti-histamine antibody added in the assay was done according to the Cisbio product manual. TR-FRET signal was detected at 665/615 using an EnVision plate reader. Resulting data were analyzed and evaluated for robustness through determination of the signal window and Z′. The Z′ factor was determined using the following equation: Z′ = 1 – (3σmax + 3σmin)/(|µmax – µmin|), where σmax and σmin represent the standard deviations (SDs) of the positive and negative controls and µmax and µmin represent the mean values of the positive and negative controls.
Thermal Shift Assay
Thermal shift assay was performed on a QuantStudio 12K flex system (Thermo Fisher Scientific) with a ramp speed of 0.015 °C per second. Solution containing 5 μM HDC, 100 μM compound, and 5X SYPRO Orange dye (Sigma-Aldrich) was dispensed into 384-well PCR plates (10 μL per well). The plates were sealed and heated in the instrument across a temperature range of 25–95 °C.
Statistics
Assay Z′ values were calculated using the standard equation Z′ = 1 – 3(σp + σn)/|µp – µn|, where σp and σn are the SDs of the positive and negative controls and µp and µn are the arithmetic means of the positive and negative controls.
Results and Discussion
Here, we have addressed a gap in screening tools by developing a high-throughput enzymatic assay to search for inhibitors of HDC. By repurposing a commercial kit that measures histamine concentration, we have optimized and validated an assay that is compatible with 384-well plates and has good assay statistics (Z′ > 0.5). Furthermore, we have applied this assay to a pilot screen, an exercise that (1) further validated the assay as suitable for HTS and (2) identified several hit compounds. One of these, a histidine analog, was confirmed as giving concentration-dependent inhibition and was validated as a bona fide binder using a thermal shift assay. Finally, we present some initial structure–activity relationships (SARs) that should inform future medicinal chemistry efforts.
HDC Assay Development Using a TR-FRET Histamine Detection Strategy
HDC catalyzes the decarboxylation of histidine, yielding histamine and CO2. Hence, one way to measure HDC activity is to quantify the amount of histamine generated during the reaction. Cisbio has developed a TR-FRET-based histamine assay that measures histamine release from cells. The assay is based on the competition between sample histamine and allophycocyanine (XL665)-labeled histamine for binding to a EuK-labeled antibody ( Fig. 1A ). The assay was reported to have a detection range of 1.56–400 nM histamine. 13 To assess whether the Cisbio TR-FRET histamine kit could be adopted to measure HDC activity, we first examined if the readout antibody can discriminate between the histidine substrate and histamine product. Various concentrations of histidine and histamine were tested in the presence of a fixed amount of XL665-labeled histamine and EuK-labeled antibody ( Fig. 1B ). Histamine was able to compete with XL665-histamine for antibody binding at concentrations as low as 13 nM. Histidine, however, did not show significant competition below 200 µM, suggesting that the assay has a > 10,000-fold selectivity for histamine over histidine.
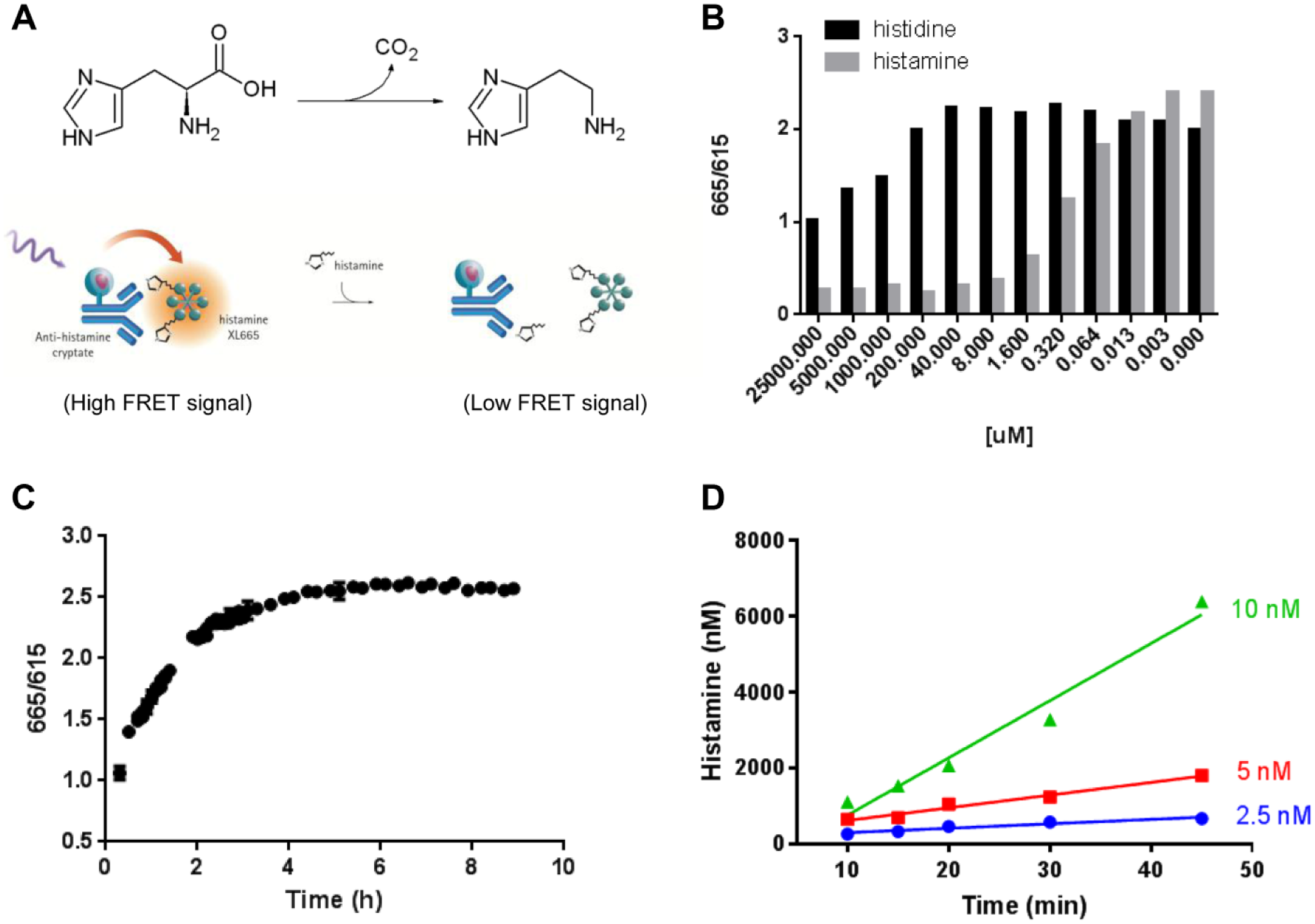
Development of TR-FRET-based HDC assay. (
As the anti-histamine antibody used in the assay was raised against carrier protein conjugated to histamine via an acyl linker, acylation of sample histamine is required to achieve high-affinity binding to the readout antibody. Thus, the effect of acylation time was examined. We did not observe significant signal differences between 15 and 180 min (data not shown), suggesting that acylation is complete by 15 min. Next, to understand the antibody/histamine binding kinetics, an incubation time-course study was carried out. As the TR-FRET signal reached an equilibrium plateau at 5 h ( Fig. 1C ), a 5 h readout incubation was chosen as our standard condition to maximize the S/B window.
To optimize enzyme concentration and define the kinetic linear range, we next performed an enzymatic reaction time course at various concentrations of HDC. As full-length hHDC protein did not overexpress in
To define optimal substrate concentrations for inhibitor profiling, we sought to determine the Km of histidine and PLP against HDC. For the former, we performed the reaction at various concentrations of the
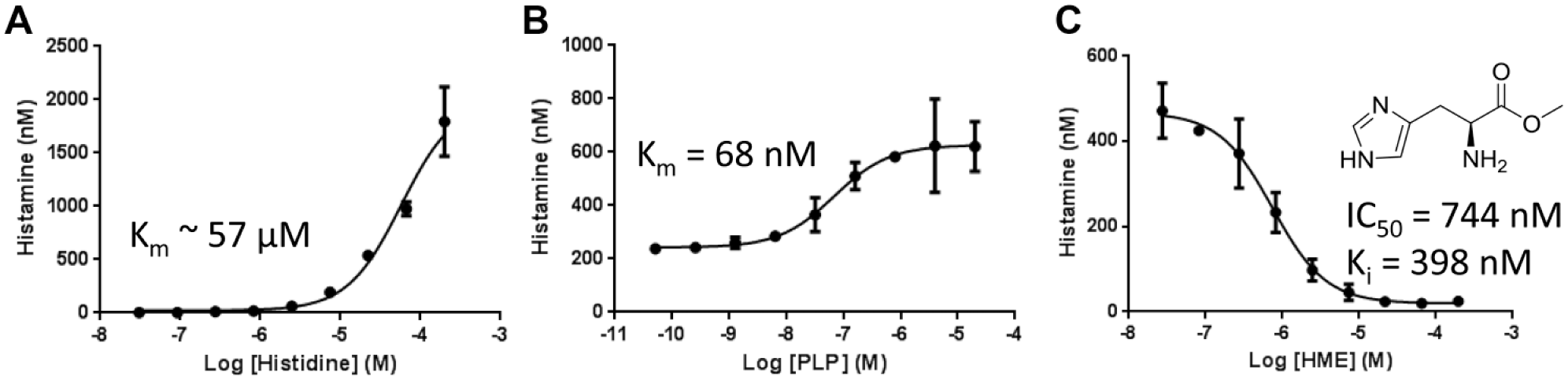
TR-FRET-based HDC assay can be used to determine EC50 values of substrates and inhibitors. Enzymatic reactions were run using the indicated concentrations of
We further validated our assay using a known HDC inhibitor, HME, a structural analog of histidine that inhibits HDC with a Ki of 460 nM. 16 HME was incubated with enzyme for 30 min, followed by addition of histidine and PLP. The IC50 value generated was 744 nM ( Fig. 2C ). The Ki calculated based on the Cheng–Prusoff equation (Ki = IC50/(1 + [S]/Km) was 398 nM, which is within a twofold difference from the value reported previously. The good correlation between our assay and the published results also suggested that the signal seen in our assay is specific to HDC rather than potential contamination. To assess the robustness of the assay, an intra- and interday experiment was performed. The IC50 values generated were 772 ± 196 nM with intra- and interday shifts less than threefold, suggesting that the assay developed here is highly reproducible.
Assay Validation and High-Throughput Screening
Following the establishment of the standard conditions, the assay was validated for HTS using 384-well plates and whole-plate analysis to access the assay window (signal-to-noise ratio), reproducibility, and intraplate variability. The negative control used consisted of 5 nM HDC, 50 µM histidine, and 68 nM PLP, and the positive control contained 5 nM HDC, 50 µM histidine, 68 nM PLP, and 196 µM HME. In addition, a concentration of 2% DMSO was added to prevent compound precipitation. No significant effects on enzymatic activity were observed between 0% and 2.5% DMSO (data not shown). The reactions were initiated by the addition of histidine and PLP, and all wells were quenched with acylation reagent after 30 min. Whole-plate analysis revealed the coefficient of variation for the negative and positive controls to be 6.36% and 5.34%, respectively. Further analysis showed the Z′ factor and S/B to be 0.46 and 1.94 (respectively), within acceptable ranges ( Fig. 3A ). Overall, the result suggested that the assay is suitable for high-throughput application.
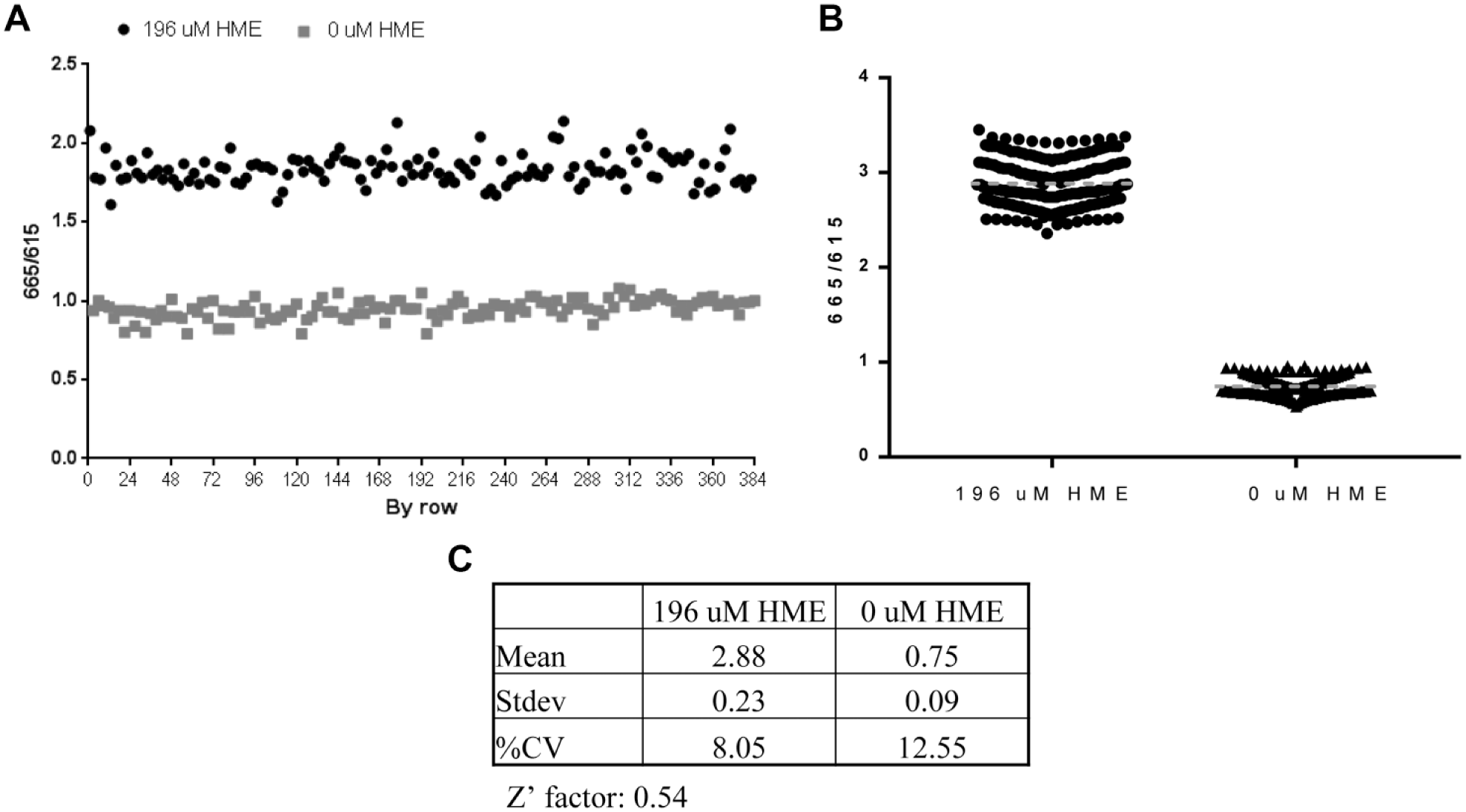
The TR-FRET-based HDC assay can be adapted as a robust screening assay. (
To identify initial hit compounds and further validate the assay for HTS campaigns, we next conducted a pilot chemical library screen using the developed assay. Specifically, approximately 7000 compounds from the MSD HTS diversity set were screened at a final concentration of 10 µM. The HTS plate map included 16 wells of each our negative control (complete reaction mixture) and our positive control (complete reaction mixture plus 196 µM HME). Average values across the 21 screening plates for Z′ factors (0.54) and S/B (3.87) ( Fig. 3B , C ), further confirmed the assay as robust for HTS. To select compounds for follow-up, we set a cutoff at 3SDs from the mean of the negative control, resulting in 74 hit compounds and hit rate of 1.1%, within the desired range. To remove compounds with fluorescent artifacts, we inspected the raw data to find compounds with high reference channel signals, as these compounds were likely false positives. Accordingly, 17 inhibitors with reference signals that were > 3SDs from the mean of control wells were removed, yielding 57 compounds in the hit list. Of these 57 hits, 15 were confirmed in 10-point dose–response studies. The results for three representative compounds are shown in Figure 4A , which shows IC50 values of 2.4, 9.5, and 18.9 µM, respectively.
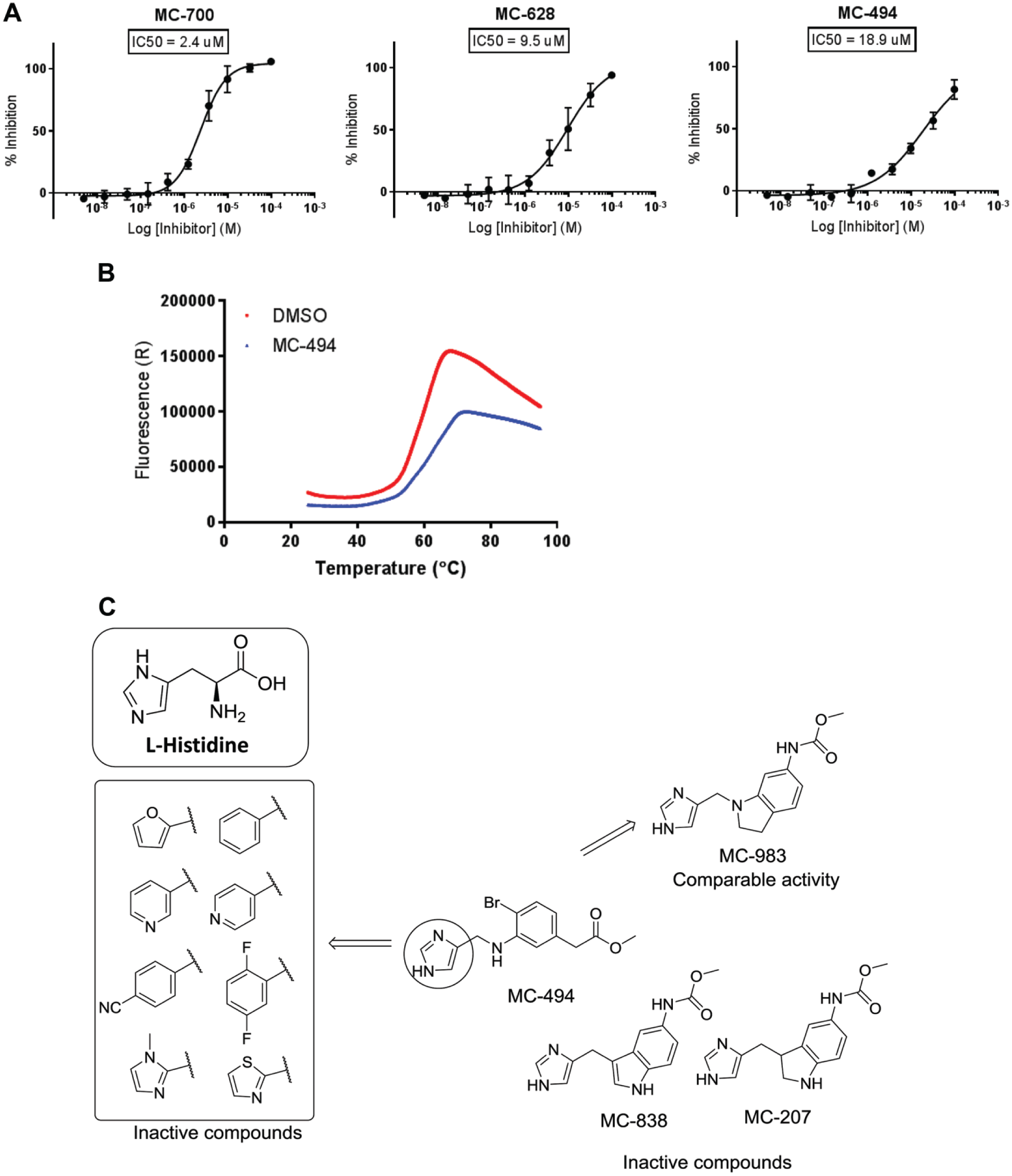
(
Initial hit lists from HTS campaigns typically include a significant percentage of promiscuous inhibitors, usually in the form of colloidal aggregators. To remove these undesired compounds, the hits were tested for specific binding using a thermal shift assay. Five compounds showed positive shift in the melting temperature of > 2 °C, suggesting these as bona fide binders (
Fig. 4B
). Interestingly, one of the confirmed hits (MC-494) shared a common imidazole moiety to that of histidine (
Fig. 4C
), giving us further confidence that it was a true binder. This compound was thus selected for some initial SAR studies. Specifically, we searched the MSD compound repository for highly similar compounds using a 2D similarity method based on atom pairs and topological torsions, and selected 15 compounds for IC50 determination in the HDC assay. Although none of the similarity search compounds had a potency greater than that of MC-494, some preliminary SARs did emerge. In particular, the imidazole moiety is essential for activity. Specifically, replacing the imidazole moiety with aryl or heteroaryl functionalities, such as 2-furyl, phenyl, 3-pyridyl, 4-pyridyl, 4-CN-phenyl, 2,4-difluorophenyl, 2-thiazolyl, or
In conclusion, we have repurposed a previously developed cell-based TR-FRET histamine detection kit to an in vitro HDC activity assay. The assay is simple to develop and is highly sensitive. We first demonstrated that the antibody used is highly selective for histamine product but not histidine substrate. Next, enzyme kinetics and inhibitor experiments exhibited a good S/B. Moreover, the determined EC50 values of substrate and inhibitor are in good agreement with published results, suggesting that this TR-FRET assay can be a replacement for the traditional HDC enzymatic assays. Assay validation was performed in 384-well plates and resulted in good statistical parameters (Z′ factor and S/B), which positioned it for a pilot HTS campaign. Application of the assay to a ~7000-member chemically diverse library resulted in an initial hit rate of 1.1% and ultimately the identification of five compounds (including MC-494, a histidine analog), which was reconfirmed via dose titration, and also gave a > 2 °C shift in a thermal shift assay. Initial SAR studies on MC-494 suggest that it could provide a valuable starting point to develop therapeutically effective compounds that inhibit HDC activity. In addition, the assay developed is primed for application to large small-molecule libraries to identify additional hit matter. Indeed, the simplicity and robustness of the assay provide a new tool to identify and characterize novel inhibitors of HDC and will contribute significantly to the development of therapeutics targeting HDC.
Footnotes
Declaration of Conflicting Interests
The authors declared potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors are employed by Merck & Co. (USA based employees)/MSD (Singapore based employees) and their research and authorship of this article was completed within the scope of this employment.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
