Abstract
DNA double-strand breaks (DSBs) are repaired primarily by homologous recombination (HR) or nonhomologous end joining (NHEJ). Compounds that modulate HR have shown promise as cancer therapeutics. The V(D)J recombination reaction, which assembles antigen receptor genes in lymphocytes, is initiated by the introduction of DNA DSBs at two recombining gene segments by the RAG endonuclease, followed by the NHEJ-mediated repair of these DSBs. Here, using HyperCyt automated flow cytometry, we develop a robust high-throughput screening (HTS) assay for NHEJ that utilizes engineered pre-B-cell lines where the V(D)J recombination reaction can be induced and monitored at a single-cell level. This approach, novel in processing four 384-well plates at a time in parallel, was used to screen the National Cancer Institute NeXT library to identify compounds that inhibit V(D)J recombination and NHEJ. Assessment of cell light scattering characteristics at the primary HTS stage (83,536 compounds) enabled elimination of 60% of apparent hits as false positives. Although all the active compounds that we identified had an inhibitory effect on RAG cleavage, we have established this as an approach that could identify compounds that inhibit RAG cleavage or NHEJ using new chemical libraries.
Introduction
DNA double-strand breaks (DSBs) are generated as intermediates of physiologic processes and upon exposure to genotoxic agents, such as cancer therapeutics. 1 The two main pathways of DSB repair are homologous recombination (HR) and nonhomologous end joining (NHEJ), which use functionally distinct sets of proteins.2,3 NHEJ is active in all phases of the cell cycle and is the main pathway of DSB repair in G1-phase cells. NHEJ functions to rejoin broken DNA ends, frequently in an imprecise manner. In contrast, HR functions to repair DSBs in cells in the S and G2 phases of the cell cycle using the sister chromatid as a template for precise DSB repair.
Novel agents that target DNA repair have been used to treat cancers. One of the most promising recent advances has been the use of the synthetic lethal relationship between BRCA-1 or BRCA-2 deficiency and inhibitors of the poly(ADP-ribose) polymerase (PARP).4,5 BRCA-1 and BRCA-2 are required for HR-mediated DNA DSB repair. Inhibition of PARP leads to the formation of DNA DSBs that cannot be repaired by HR in BRCA-deficient cells, resulting in cell death. While this new therapy is promising, it has a limited scope, as only a small fraction of tumors are BRCA deficient, and many cancer cells, including cancer stem cells, are in G0–G1, where NHEJ, and not HR, is the major pathway of DSB repair. 6
Opportunities may exist to develop therapeutic approaches to treat tumors by targeting NHEJ. 7 NHEJ relies on the core factors Ku70, Ku80, DNA ligase IV, and XRCC4. 3 Deficiency in any of these factors leads to a complete block in NHEJ in all cell types analyzed. In addition to core NHEJ factors, there are other proteins that function during NHEJ, but do not appear to be essential for DSB repair in all settings. For example, deficiency in the NHEJ factor XLF leads to defects in DSB repair in fibroblasts, but not in lymphocytes. 8 Loss of the histone H2A variant H2AX does not lead to NHEJ defects in any cell lines examined to date; however, loss of H2AX and XLF in lymphocytes leads to a severe block in NHEJ. 9 Thus, chemical inhibitors of XLF, or proteins that function in the same pathway, could cripple NHEJ in H2AX-deficient tumors, resulting in increased sensitivity to DNA DSB-inducing therapies, such as ionizing radiation.
Developing lymphocytes assemble antigen receptor genes through the process of V(D)J recombination. 10 This reaction is initiated when the RAG-1 and RAG-2 proteins, which together form the RAG endonuclease, introduce DNA DSBs at the border of two recombining gene segments and their flanking RAG recognition sequences (RSs). RAG cleavage occurs only in G1-phase developing lymphocytes and generates two hairpin-sealed coding ends (CEs) and two blunt signal ends (SEs). NHEJ is required to process and join the CE pairs and SE pairs to generate a coding join (CJ) and signal join (SJ), respectively. 11
We have exploited the requirement for NHEJ to repair RAG DSBs to develop a cell line–based approach to identify compounds that can inhibit NHEJ. Murine bone marrow pre-B cells, transformed by the Abelson murine leukemia virus, hereafter referred to as abl pre-B cells, grow in single-cell suspension cultures. 12 Treatment of abl pre-B cells with the Abl kinase inhibitor imatinib leads to G1 arrest, RAG induction, and V(D)J recombination at endogenous antigen receptor genes and at chromosomally introduced retroviral recombination substrates.13,14 We have developed retroviral recombination substrates, such as pMX-INV, in which successful V(D)J recombination leads to inversion of an antisense green fluorescent protein (GFP) cDNA and GFP expression ( Fig. 1A ). 14 Thus, after treatment with imatinib, abl pre-B cells that express GFP have generated RAG DSBs at pMX-INV and successfully repaired them by NHEJ, whereas those that are GFP negative have not generated a RAG DSB or have a RAG DSB but are unable to repair it due to a defect in NHEJ.
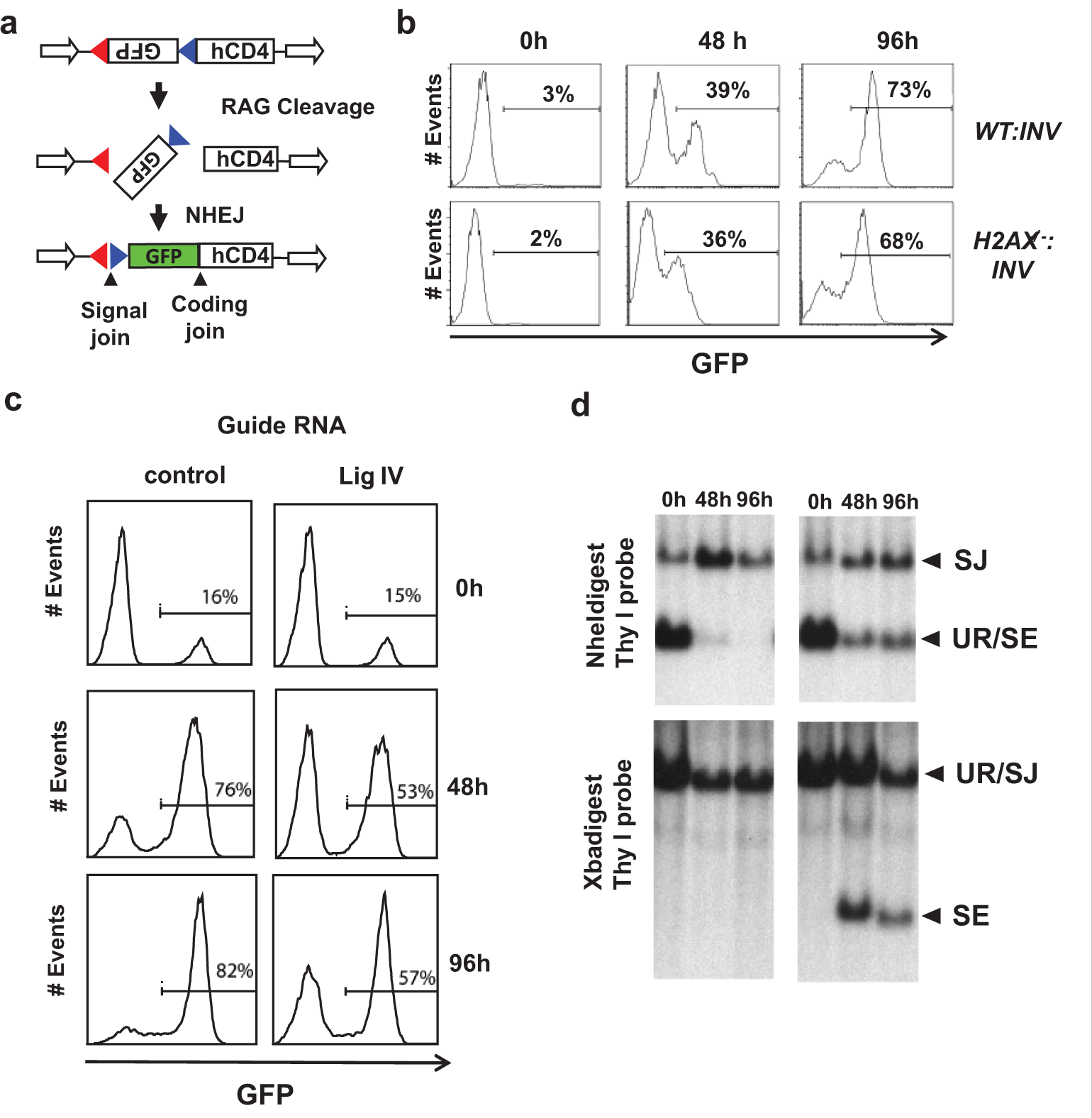
Retroviral V(D)J recombination substrate. (
Modern drug discovery strategies employ high-throughput screening (HTS) approaches where libraries of chemical compounds can be tested in a variety of cellular assays. 15 Flow cytometry is capable of simultaneous analysis of numerous biomarkers associated with physiological and biochemical cellular responses. HyperCyt (IntelliCyt, Albuquerque, NM) is a high-throughput flow cytometry (HTFC) platform where individual microplate wells with volumes as small as 1 µL are sampled in a time-resolved manner.16,17 This approach has been validated using cell- and bead-based end-point assays for ligand binding, surface antigen exposure, and GFP expression.18,19
Here we report the development of an HTFC assay to identify compounds that inhibit NHEJ in H2AX-deficient abl pre-B cells containing pMX-INV retroviral V(D)J recombination substrates. Compounds that inhibit XLF, or proteins that function in the same pathway as XLF, would inhibit V(D)J recombination and NHEJ in H2AX-deficient, but not wild-type, abl pre-B cells. Such compounds could be used to selectively inhibit NHEJ in H2AX-deficient tumor cells but not normal tissues, thus sensitizing the tumor cells to DSB-inducing therapies, such as ionizing radiation. The optimized HTFC assay was used to screen the National Cancer Institute (NCI) NeXT library containing more than 80,000 small molecules. This library was derived from the Molecular Libraries Small Molecule Repository (MLSMR) library, with the addition of some commercial chemical libraries and the removal of compounds that have chemical liabilities that would prohibit advancing them therapeutically.
Materials and Methods
Materials
Dulbecco’s modified Eagle’s medium (DMEM, Sigma-Aldrich, St. Louis, MO) was used as a base for complete medium and was supplemented with standard concentrations of
Generation of Abl Pre-B Cell Lines
Southern Blotting
Native Southern blot analyses of V(D)J recombination of the pMX-INV and pMGINV retroviral recombination substrates were carried out as previously described.14,21
HTS Procedure
On day 1, 10 μL complete medium with or without 6 µM imatinib was added to columns 2–24 of a sterile 384-well culture plate (Greiner Bio-One, Monroe, NC, no. 781186) using a BioTek (Winooski, VT) MultiFlo liquid dispenser. Column 1 contained medium without cells. Following this initial addition and prior to adding cells, a Beckman Coulter (Brea, CA) FX Liquid Handler equipped with 200 nL pin tools (V&P Scientific, San Diego, CA) was used to add library compounds (columns 3–22) and DMSO (columns 23 and 24). Plates were mixed before cells were added (10 μL, 2 × 106/mL) to columns 2–24. Empty wells in column 1 were used during analysis to orient well and row identification and to serve as a wash well between the two types of control wells. Cells in column 2 were exposed to both imatinib and dequalinium. Dequalinium was identified during a preliminary validation screen of the PCL as a small molecule that reduced GFP expression in the presence of imatinib. It served as a positive control for inhibition of NHEJ in the screen of the NCI NeXT collection. Cells that were only exposed to 1% DMSO served as an alternate positive control for cell growth and GFP expression in the absence of imatinib and test compounds. Final concentrations in the assay wells were 106/ml abl pre-B cells (20,000 in 20 µL), 10 μM library compounds, 1% DMSO, 3 μM imatinib, and 10 μM dequalinium. Assay plates were incubated for 4 days (37 °C/5%CO2) and were harvested on day 5 using the HTFC platform.
Dose–response plates were assembled similarly. Test compounds were serially diluted 1:3 prior to pin tool addition, resulting in a final concentration range of 100 μM to 15 nM. Data points were fitted by Prism software (GraphPad Software Inc., San Diego, CA) using nonlinear least-squares regression in a sigmoidal dose–response model with variable slope, also known as the four-parameter logistic equation. Curve-fit statistics were used to determine the concentration of test compound that resulted in 50% of the maximal effect (EC50), the confidence interval of the EC50 estimate, the Hill slope, and the curve-fit correlation coefficient.
HyperCyt High-Throughput Flow Cytometry
HyperCyt was recently adapted for the screening of samples displayed in high-density, 1536-well plate format. This was achieved using a Cluster Cytometry platform consisting of four networked BD Accuri C6 flow cytometers that was capable of processing individual plates in less than 15 min and routine screening of up to 60,000 wells/day. 19 In the current study, the Cluster Cytometry platform was modified for processing four 384-well plates in parallel to extend the range of assays amenable to this advanced level of HTFC screening ( Fig. 2 ).
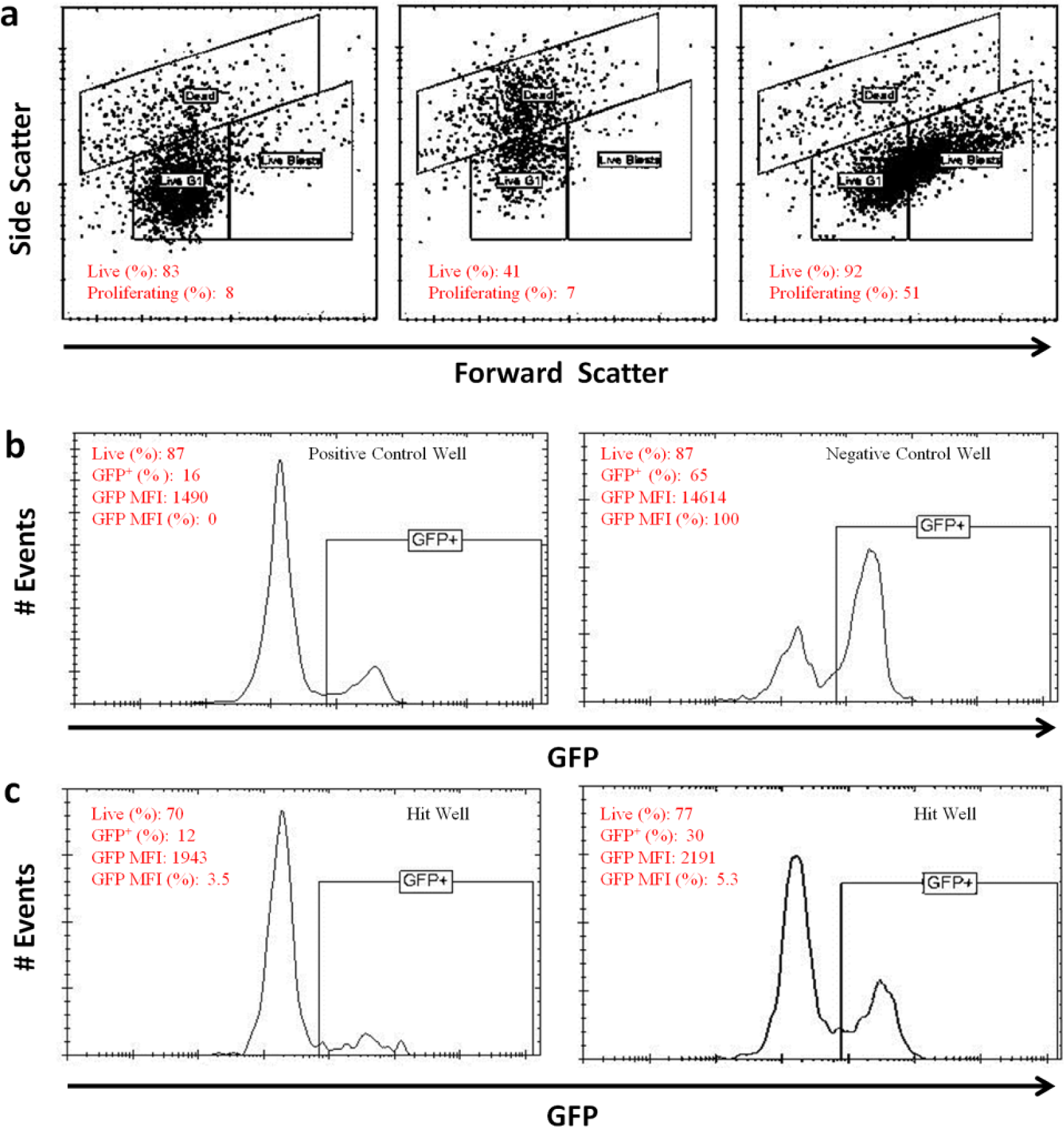
Gating configuration for flow cytometry analysis. (
The fluorescence of GFP was excited at 488 nm and detected with a 533/30 nm optical bandpass filter. Custom-modified HyperView software and ForeCyt software (IntelliCyt) were used to analyze data. First, flow cytometer data files were parsed by software-based well identification algorithms to segregate data from individual wells for separate analysis. Next, dead cells were identified and gated based on forward and side light scatter profiles elicited by the 488 nm laser (dead, Fig. 3a ). Light scatter profiles were also used to discriminate two subpopulations of live cells: nonproliferating G1-arrested cells (live G1) and proliferating blast cells (live blasts, Fig. 3a ).
The percent of live cells was calculated as
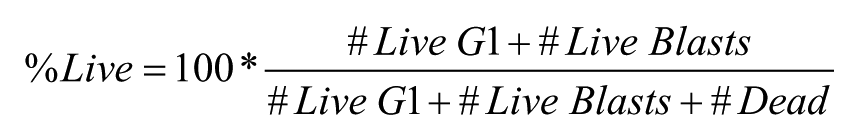
The percent of proliferating cells was calculated as
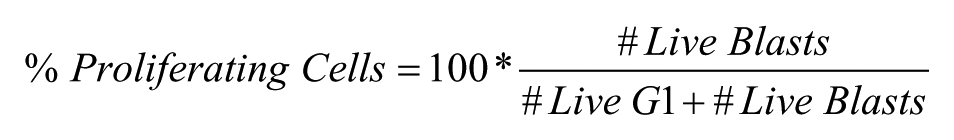
The response range for the assay was defined as the difference between the averages of the GFP median fluorescence intensity (MFI) of cells in the negative (NCntrl) and positive control (PCntrl) wells, respectively ( Fig. 3b ). Normalized GFP expression in the presence of compounds in sample wells was designated as % GFP MFI and calculated as
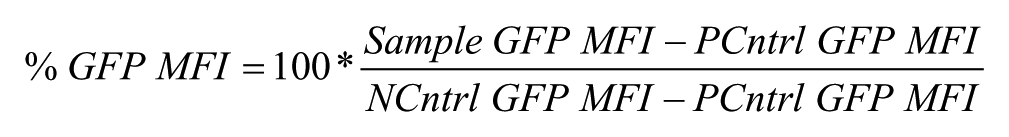
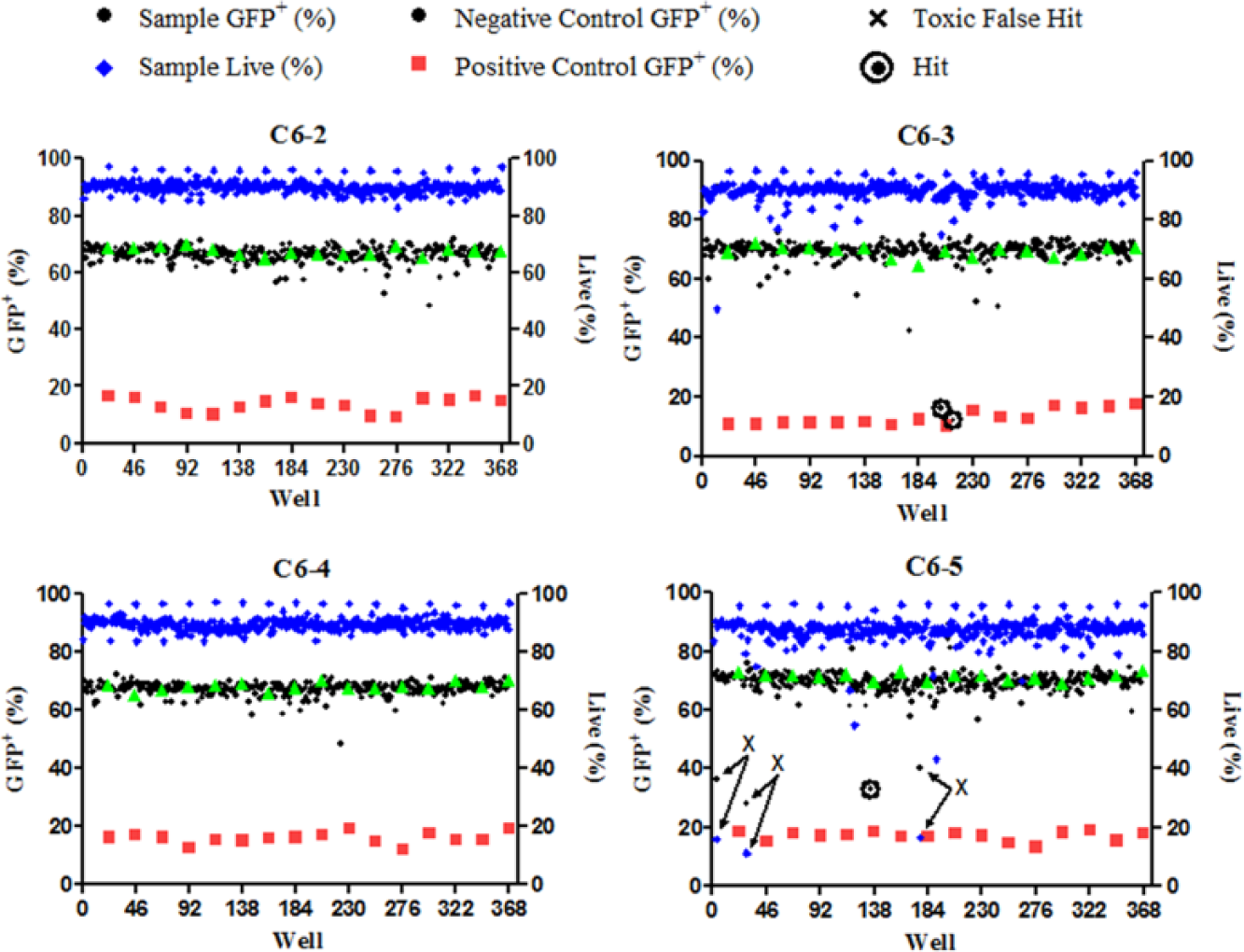
Representative results from parallel HTS analysis. Each panel represents analysis of a separate 384-well plate by each of the four flow cytometers (C6-2, C6-3, C6-4, and C6-5) in the Cluster Cytometry platform. Illustrated is GFP expression in positive control wells (red squares), negative control wells (green triangles), and wells containing test compounds (black circles). The percentage of live cells in test compound wells was also quantified on the basis of light scatter profiles (blue triangles). False-positive results are indicated by significantly diminished GFP expression associated with low viability (Xs with arrows indicating decreases in both percent of live and percent of GFP+ cells in the wells). Active hit compounds are indicated by diminished GFP expression without diminished viability (circled black circles).
Compounds resulting in a normalized cell GFP MFI of 40% or less (≥60% inhibition, Fig. 3c ) were considered active.
Results
Cell-Based V(D)J Recombination Assay
We developed H2AX-deficient (
High-Throughput Flow Cytometric Screening Platform
The Cluster Cytometry platform was reconfigured to process four 384-well plates with 20 µL/well in parallel. A 3-D printer-fabricated custom positioning assembly was secured to the autosampler positioning arm to enable positioning of the four sample uptake probes in matching locations of the four 384-well plates. Four plate positioning frames were fabricated and mounted on the autosampler deck in vertically offset locations amenable to access by an Agilent plate-handling robot. Custom software was developed to enable Agilent automation software to communicate with and control the Cluster Cytometry platform.
HTFC Screen of H2AX-Deficient Abl Pre-B Cells
In the test wells, GFP expression indicative of RAG DSB generation and NHEJ repair was detected as the percentage of live cells with green fluorescence intensity (excitation 488 nm, emission 533/30 nm) exceeding a threshold set by the separation of GFP-expressing and nonexpressing MFI peaks in the positive and negative control wells ( Fig. 2b ). A compound was considered active if it produced a greater than 60% decrease in the MFI of GFP-expressing cells relative to the negative control cells ( Fig. 2c ). Compounds that were considered active by this criterion, but exhibited increased forward or side scatter, were eliminated from further analysis, as these compounds nonspecifically led to increased cell death or proliferation, respectively.
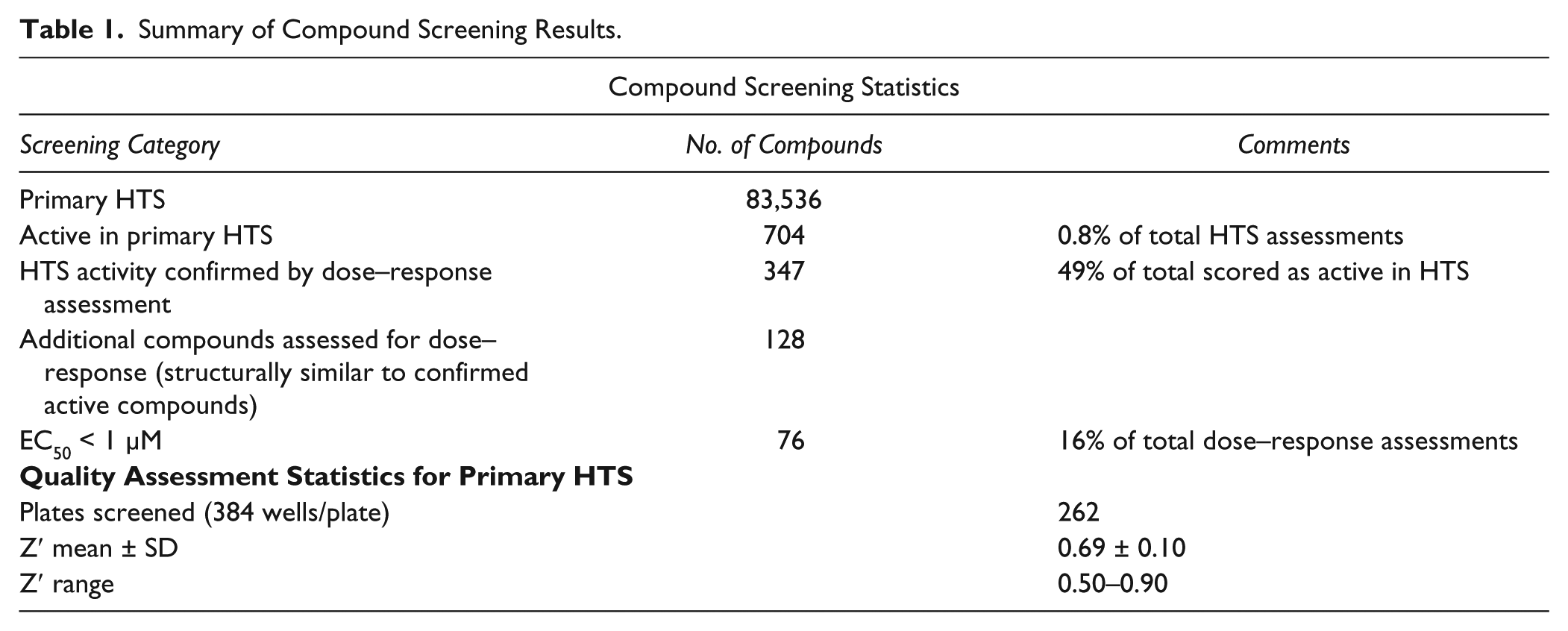
A representative example of results from four of the two hundred sixty-two 384-well plates screened by independent cytometers in parallel is shown ( Fig. 3 ). The values from positive (red squares) and negative (green triangles) control wells produced Z′ scores of 0.77–0.84, from the four Accuri C6 cytometers used for analysis, indicating uniformly sensitive and robust screening results from these cytometers. Examples of the positive hits are indicated by the circled data points, and those indicated by arrows were excluded due to unfavorable, cell death–associated side scatter parameters ( Fig. 3 ). Of the 83,536 compounds analyzed, there were 704 active compounds, a hit rate of 0.8% ( Table 1 ). Z′ scores were 0.5 or better for all the plates processed in the primary HTS.
Summary of Compound Screening Results.
Evaluation of Positive Hits and Related Compounds
Subsequent concentration–response analyses of the active compounds, together with an additional 128 that were structurally similar, identified 83 compounds that lead to a 50% (EC50) or greater reduction in the MFI of GFP-expressing cells at concentrations of 1 µM or less ( Table 1 ). Representative dose–response curves from three structurally different compounds are shown ( Fig. 4a–d ). Of the confirmed active compounds, 46 structurally distinct compounds were acquired as powder and retested in dose–response. Of these 46 compounds, 21 were reconfirmed as exhibiting an EC50 of less than 1 µM.
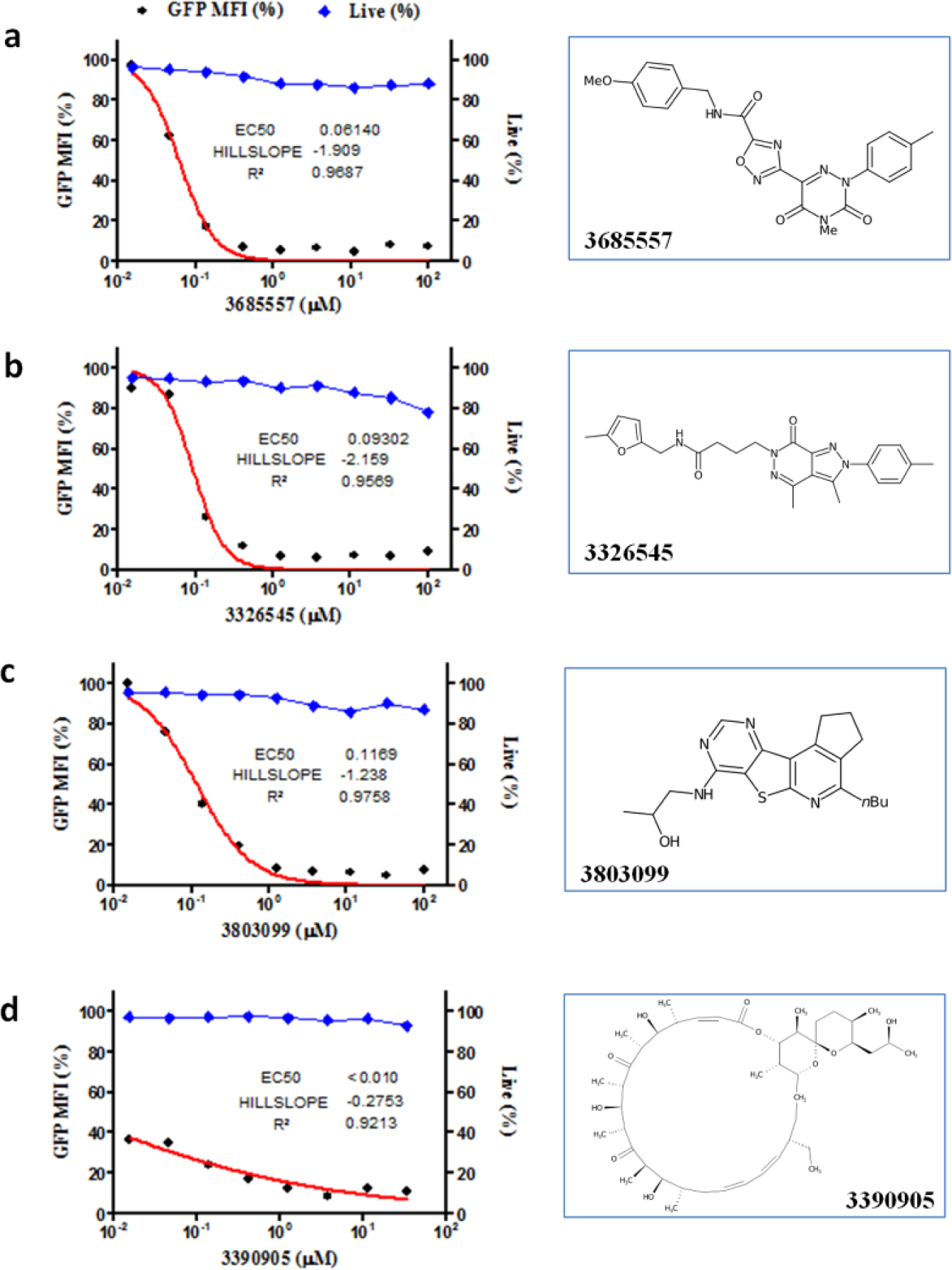
Dose–response profiles: Examples of compounds with good viability and dose–response kinetics (
Effect of Positive Hits on V(D)J Recombination
Compounds that inhibit V(D)J recombination in
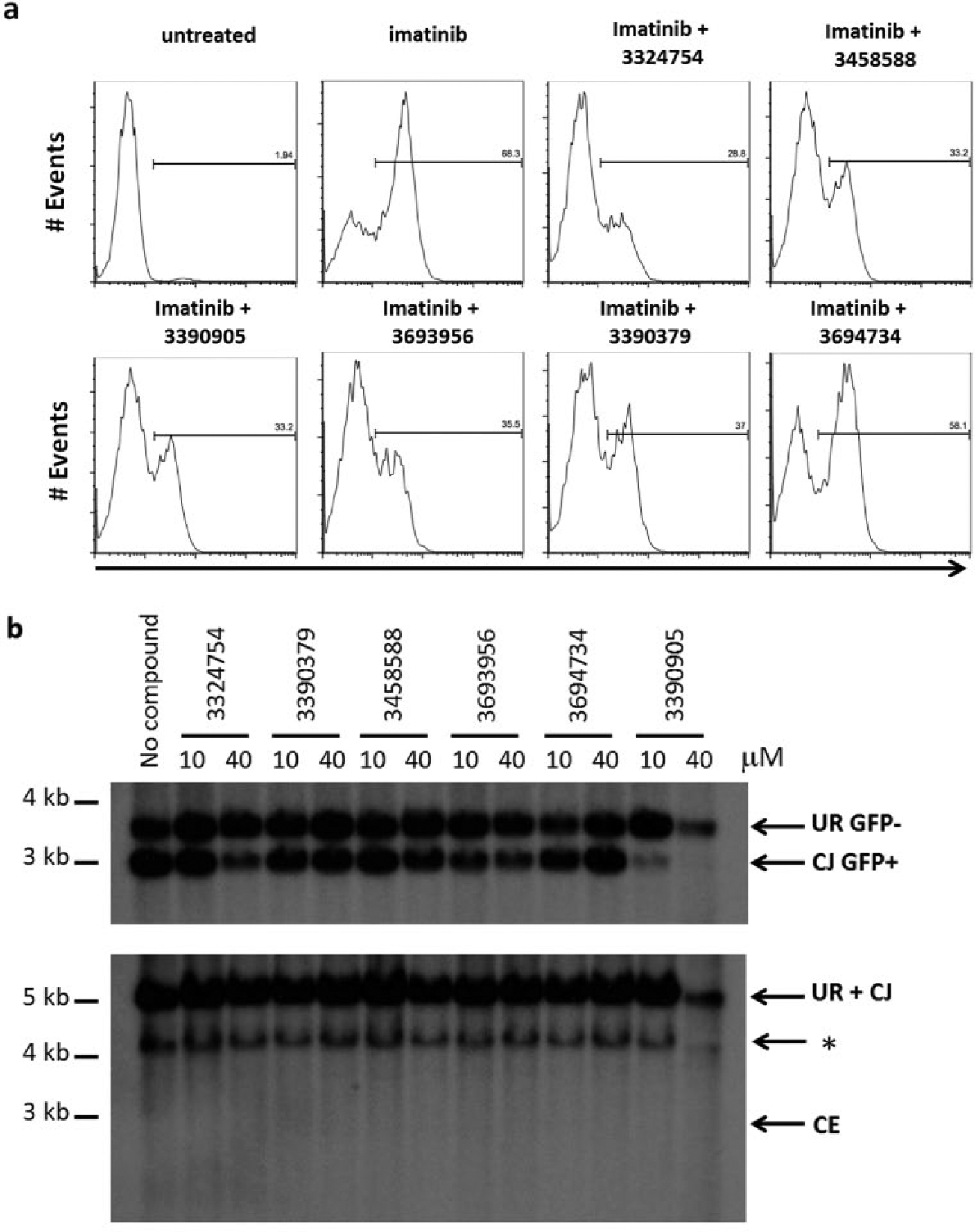
Assessment of active compounds. (
Discussion
Here we have developed a single-cell-based assay for successful V(D)J recombination. This assay was adapted to an HTFC compound screening platform to identify compounds that inhibit V(D)J recombination. Active compounds could prevent RAG cleavage or inhibit the repair of RAG DSBs by NHEJ. As RAG DSBs can only be repaired by NHEJ, this approach could identify compounds that inhibit DNA DSB repair by NHEJ.
There are several important features of this new HTFC screening approach. We have shown that we can induce and follow V(D)J recombination in small culture volumes (20 µL) in a robust and reproducible manner with Z′ scores in excess of 0.5, and usually the range of 0.7 is easily achievable. Abl pre-B cells have been engineered through the ectopic expression of bcl-2 so that they do not die upon activation of DNA damage responses by unrepaired DNA DSBs. 14 As a result, compounds that prevent the repair of RAG DSBs will not lead to cell death. Thus, any compounds that cause cell death can be excluded from further analysis, as they effect processes that are essential for cell viability other than the generation or repair of RAG DSBs. Importantly, cell viability can be assessed directly from the primary HTFC screening data by analyzing side scatter. In the screen carried out here, this allowed us to eliminate 50% of the compounds (907), which prevented V(D)J recombination due to a nonspecific effect on cell viability. RAG DSBs are generated and repaired in G1-phase cells. RAG-2 is degraded upon entry into the S phase; thus, any compounds that prevented G1-cell-cycle arrest by imatinib would indirectly inhibit V(D)J recombination. 22 Such compounds can also be excluded by analyzing forward scatter data from the primary HTFC screen. In this regard, we eliminated an additional 10% (187) of the initial positive hits due to their ability to promote proliferation in the presence of imatinib. Thus, a very powerful feature of this screening approach is the ability to eliminate compounds that have a nonspecific effect on V(D)J recombination by either promoting cell death or proliferation. In our primary screen of the 1798 compounds that inhibited V(D)J recombination, we were able to eliminate 1094 based on forward and side scatter analysis, saving considerable effort in validating these compounds.
In this screen, all the compounds that we identified had an inhibitory effect on RAG cleavage and not on NHEJ. There are many ways that compounds could inhibit RAG cleavage. It is conceivable that some may directly inhibit the expression of RAG-1 or RAG-2, possibly through interactions with transcription factors required for Rag expression. The generation of RAG DSBs also depends on transcription and accessibility of the pMX-INV retroviral recombination substrate and any inhibition of transcription or accessibility would lead to a decrease in RAG cleavage. In addition, specific histone modifications are required for RAG recruitment and activity. Compounds that inhibit histone-modifying enzymes that regulate these modifications would also influence RAG cleavage. Finally, it is possible that some of these compounds directly inhibit RAG nuclease activity.
While the screen of the NeXT library did not reveal any compounds with activity in inhibiting NHEJ in H2AX-deficient cells, the establishment of this approach will provide a powerful tool for the future analysis of NHEJ. In this regard, it can potentially be used with different chemical libraries to identify compounds that inhibit NHEJ. Moreover, testing compounds on backgrounds of different genetic deficiencies is facilitated in abl pre-B cells given the ease with which genes can be inactivated using guide RNA and Cas9 approaches. Finally, this HTFC screening approach can be easily adapted for genetic screens using either inhibitory RNA or guide RNA (with Cas9) libraries to identify proteins that are required for NHEJ or RAG cleavage. Indeed, we have shown here that expression of a DNA ligase IV guide RNA and Cas9 can block NHEJ in abl pre-B cells. Such approaches are already underway to identify new proteins that function during NHEJ.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by NIH grants R01CA177759 (B.P.S. and L.A.S.), R01 AI074953 (B.P.S.), R01HG005066 (B.S.E.), U54MH084690 (L.A.S.), U54 MH074425 (L.A.S.), and CCSG P30CA118100.
