Abstract
In order to identify novel treatment principles specifically affecting cancer stem cells in triple-negative breast cancer, we have developed a high-throughput screening method based on the mammosphere and anoikis resistance assays allowing us to screen compounds using a functional readout. The assay was validated against manual protocols and through the use of positive controls, such as the response to hypoxia and treatment with the known cancer stem cell–targeting compound salinomycin. Manual and robotic procedures were compared and produced similar results in cell handling, cell cultures, and counting techniques, with no statistically significant difference produced from either method. The variance between samples processed manually versus robotically was no greater than 0.012, while Levene’s test of significance was 0.2, indicating no significant difference between mammosphere data produced manually or robotically. Through the screening of 989 FDA-approved drugs and a follow-up screen assessing the antineoplastic subgroup, we have identified three therapeutic compounds with the ability to modulate the breast cancer stem cell fraction in the triple-negative breast cancer cell line MDA-MB-231, highlighting their potential usage as stem cell–specific adjuvant treatments.
Introduction
Recent advances in gene expression profiling have led to the expansion of our understanding of the subclassifications of breast cancer. Indeed, the genomic landscape of breast cancer as a whole has been steadily advancing not only through studies outlining the genetic and mutational drivers behind the disease, 1 but also through our understanding of the degree of inter- and intra-tumor heterogeneity. 2 While our understanding of the subdivisions of breast cancer is advancing, and hence better informing clinical treatment options, certain breast cancer types still represent a clinical challenge. Among them are those tumors referred to as basal or triple-negative breast cancer (TNBC), characterized by low expression of estrogen receptor (ESR1), progesterone receptor (PGR), and human epidermal growth factor receptor 2 (HER2). 3 TNBC is often associated with poor prognosis and shorter periods of overall survival. 4 In addition, subpopulations of cells have been identified that possess the ability to drive tumor growth and recurrence. These populations share many of the characteristics of stem cells, including resistance to chemo- and radiation therapy. These populations have been termed cancer stem cells (CSCs). 5 Due to the overall poor prognosis linked to TNBC and a lack of sufficient treatment options to target CSCs, it is of vital importance that new treatment strategies are developed.
The tumorsphere assay was originally developed to evidence the existence of neural stem cells; 6 this method has since been adapted to a number of other cancer subtypes. The mammosphere assay is a widely used variation of this assay that has been established as a robust method for the isolation, propagation, and enumeration of breast cancer cells, allowing them to act with stem cell–like traits. 7 The methodology by which this assay is carried out can vary significantly between groups; however, there are a number of commonalities inherent to the assay. Chief among them is the labor and time-intensive nature of the assay, which can limit the number of conditions studied in any one experiment. Current methods also rely heavily on manual counting of spheres via microscope visualization. This can introduce a degree of variability between operators and individual experiments.
The application of high-throughput screening (HTS) technologies to the identification of chemical compounds that specifically target CSCs is of particular interest. However, standard currently available high-throughput cell viability assays applied to bulk populations of cancer cells cannot identify agents with CSC-specific toxicity. 5 Current methodologies include the generation of CSC-like populations of cells through the induction of epithelial–mesenchymal transition (EMT)8,9 or fluorescence-activated cell sorting (FACS) based on specific cell surface markers, that is, CD44high/CD24low, 10 followed by standard cytotoxicity assays to identify compounds that target these cells. Other studies rely on stem-associated phenotypes, such as the presentation of EMT markers that can be directly measured, 11 or enzymatic studies, including alkaline phosphatase. 12 Recently, great advances have been made through the usage of patient-derived tumor xenografts (PDTXs) in HTS methodologies. The work carried out by Bruna et al. represents a significant step forward in the area of HTS; by harnessing the clinical relevancy of PDTX material and combining this with HTS analysis, they have presented a platform for preclinical drug development. 13
In light of the current screening platforms available, we feel our automated HTS assay contributes a unique functional-based assay that uses mammosphere formation as an end point, exploiting the anoikis resistance (AR) of CSCs in order to identify compounds that specifically target these cells. This system has a specific advantage over previously mentioned assays, in that it does not require the exclusion of cells from the bulk population or require genetic modification prior to study, thus allowing us to examine the entire, highly heterogeneous population while only assaying for effects that target CSCs.
Materials and Methods
Overview
A comparative workflow is detailed in Figure 1 to highlight the main differences in procedure between the traditional low-throughput manual mammosphere assay and that of our novel high-throughput method. A traditional mammosphere assay is carried out by treating cells in a monolayer culture for an appropriate period of time, followed by enzymatically detaching cells in culture, diluting and syringing them into a single cell suspension, counting the cells using a cell counter or hemocytometer, and then diluting to a seeding density before plating into large low-adherence culture plates. Resultant mammospheres are counted manually using a microscope fitted with a graticule following 5 days of culture. We have detailed here for the first time an automated high-throughput alternative to this assay, which includes modification to the initial cell counting steps, processing methods, and sphere enumeration. Each step has been compared against the traditional method to ensure results are not artifacts of the modified procedure.
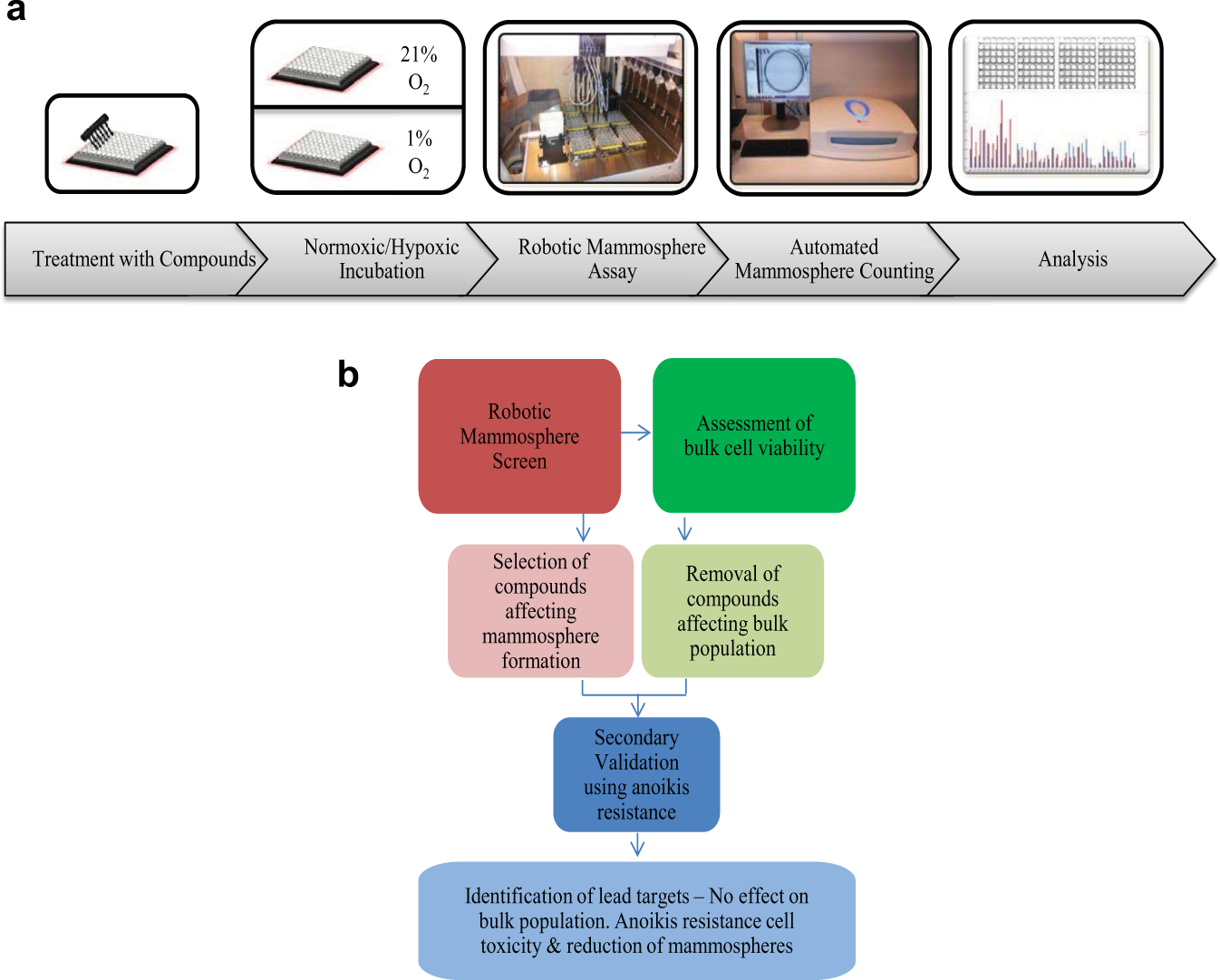
Robotic mammosphere and lead target validation steps workflow. (
In brief, cells are cultured in 96-well plates, which allows for the testing of a high volume of compounds. Cells in this monolayer culture are then treated for an appropriate period of time under standard culturing conditions. Luminescence readings are taken immediately following the addition of luciferin. Cell number is estimated from a standard curve of luminescence versus cell number. Using Microsoft Excel, required dilution volumes are calculated. A VBA algorithm within Microsoft Excel then automatically generates CSV files containing these volumes in a format that is compatible with the robotic platform. Culture plates of cells, deep-well dilution plates containing mammosphere media, polyHema-coated 48-well plates, and reservoirs of phosphate-buffered saline (PBS) and Accutase are added to the Janus robotic platform. A custom-designed program then automatically progresses each 96-well plate through the steps of cell detachment, dilution, and seeding. Following 5 days of nonadherent three-dimensional culture, resultant mammospheres are stained using MTT. Following staining, mammosphere plates are scanned using a high-resolution plate scanner and mammospheres enumerated using predefined conditions.
Cell Culture
MDA-MB-231/Luc (Cell Biolabs, Inc., San Diego, CA), a luciferase-expressing epithelial breast cancer cell line, was cultured in Dulbecco’s modified Eagle’s medium (DMEM) complete media (supplemented with 10% fetal bovine serum, 100 U/mL penicillin, and 100 μg/mL streptomycin). Cells were grown in a humidified atmosphere of 5% CO2/95% air at 37 °C. For hypoxia experiments, cells were incubated for 48 h in the SCI-tiveN hypoxic workstation (Ruskinn, Bridgend, UK) in 1% O2, 5% CO2, and 94% N2 in a humidified environment at 37 °C. To generate anoikis-resistant cells, T150 flasks were coated with polyHema and allowed to dry. Cells were then seeded at an appropriate concentration and allowed to grow for 24 h, at which point the surviving stemlike cells were harvested.
Cell Counting
Cells were cultured for 48 h in clear-bottomed, black-walled 96-well plates. Two random wells were chosen for counting, and both methods of counting were applied to the same wells. Luminescent-based cell counting was carried out through the addition of D-Luciferin (PerkinElmer, Waltham, MA) to culture media, and the resultant luminescent signal immediately read using a CentroPRO Microplate Luminometer (Berthold Technologies, Bad Wildbad, Germany). The cell number was calculated from a standard curve of known cell concentrations versus luminescent readings. Traditional cell counting was carried out using a Moxi Z cell counter (ORFLO Technologies, Ketchum, ID) after cells had been detached and syringed into a single cell suspension. Statistical analysis was carried out to measure the degree of difference between each method of counting per sample, and also the degree of difference between samples in each method.
Cell Viability Staining
Cells were processed into a single cell suspension by manual syringing and by robotic pipetting in parallel. Cell viability was assessed using the LIVE/DEAD Cell Viability kit (Life Technologies, Carlsbad, CA). Viable cells stain green with calcein-AM, indicating intracellular esterase activity; dead cells stain red with ethidium homodimer-1, indicating loss of plasma membrane integrity. Cells were imaged using a fluorescent microscope, and the number of red and green cells counted.
Mammosphere Assay
Cells were cultured for 48 h in normoxic and hypoxic conditions in the presence and absence of normal culture media, DMSO vehicle, and salinomycin, a known stem cell inhibitor. Following incubation, the mammosphere assay was carried out manually and robotically on identically treated samples.
Manual assay
Mammosphere culture was carried out as previously described, and spheres were counted on day 5 at 4× magnification using a graticule-containing microscope. 14
High-throughput assay
Cell number is estimated using the luminescence-based cell counting method described above. Using a Microsoft Excel template spreadsheet, cell number per well and dilution volumes are calculated. A VBA algorithm within Microsoft Excel then automatically generates CSV files containing these volumes in a format that is compatible with the robotic platform. Following cell number calculations, culture plates of cells, deep-well dilution plates containing mammosphere media, polyHema-coated 48-well plates, and reservoirs of PBS and Accutase are added to the Janus robotic platform. The Janus platform automatically adds Accutase to all wells of the culture plate in order to detach cells. Following detachment, the platform then adds sterile PBS to each well in order to raise the liquid volume to a level required for the subsequent processing step. Each column of wells in the culture plate is then processed individually in order to prevent cell reattachment. Each well is robotically pipetted at high speed to produce a single cell suspension. An individual volume (provided by the CSV files) is then removed from each well and added to the deep-well dilution plates to ensure each well of the plate contains an equal cell number. An equal volume is then removed from the dilution plates and added to the polyHEMA-coated 48-well plates for incubation. Following 5 days of culture, resultant mammospheres are stained using MTT. Mammosphere plates are scanned using the GelCount high-resolution plate scanner (Oxford Optronix, Oxford, UK). The associated software using a CHARM II (Compact Hough and Radial Map) image processing algorithm then enumerates all mammospheres in each well.
Statistical Analysis to Determine Effect Size, Required Sample Sizes, and Assay Variance
In order to accurately determine the degree of assay variance in response to similar treatment conditions, first the effect size was calculated. A positive control pilot mammosphere assay was carried out in triplicate with both manual and robotic procedures, measuring the response to hypoxia and salinomycin. Using the effect seen in response to hypoxia, the effect size (Cohen’s d) was calculated. Second, a power analysis was then carried out to determine the number of replicates that would be required to ensure sufficient statistical power for our results in a larger-scale variance assay. The Cohen’s d value of the hypoxia effect, although much greater, was reduced to the standard 0.8 value assigned by Cohen to large effects, in an effort to be as conservative as possible when interpreting the data. Using a confidence interval of 0.95, Cohen’s d of 0.8 thus increased our replicate number to 42. The mammosphere assay was carried out both manually and robotically on normoxic and hypoxic samples with 42 replicates of each. The degree of variance between these replicates was assessed via basic statistical calculations, as well as with Levene’s test for homogeneity of variances using SPSS (IBM, version 20). Effect size and power calculations were carried out using R statistical software version 3.2.4.
FDA-Approved Drug Screen Using Established Platform
A library of Food and Drug Administration (FDA)–approved drugs was kindly gifted from the Drug Discovery Unit, CRUK Manchester Institute, Manchester, UK. Ninety-six-well plates were precoated with compound such that the final concentration upon the addition of media would be 10 μM. Cells were added to these plates and allowed to grow for 72 h. Following incubation, plates were processed as per the HTS method outlined above.
Gene Expression Analysis
Quantitative real-time PCR (qPCR)–based gene expression analysis was conducted in accordance with the MIQE guidelines, 15 as described previously. 16 In short, cells grown as adherent monolayer or in anoikis-resistant conditions were harvested and RNA was isolated using the RNeasy Mini Kit (Qiagen, Valencia, CA) according to the manufacturer’s instructions. RNA was reverse transcribed into cDNA, followed by qPCR using gene-specific oligonucleotides. qPCR data analyses were performed with GenEx (MultiD version 5.4.3). Cq values larger than 35 were replaced with a cutoff value of 35. Missing data were replaced with the cutoff Cq value + 1, reflecting 50% of the lowest target concentration detected. Cq values were interpolate calibrated, and technical replicates were averaged and normalized to reference genes. Suitable reference genes were identified with the Normfinder software. 17 Cq values were then converted into relative copy numbers and log 2 transformed for statistical analysis. All applied primer sequences have been published previously. 16
Secondary Validation of Lead Target Antineoplastic Drugs
Drugs for follow-up validation were identified from the subset of antineoplastics. These compounds were prepared in an identical manner to the primary screen. Anoikis-resistant cells were cultured for 24 h in nonadherent conditions, harvested, and subsequently plated at a concentration of 9000 cells per well and allowed to adhere and grow for 72 h. Following incubation, cell viability was assessed using the Alamar Blue assay. By combining data from the primary screen and secondary validation assay, lead compounds for further study were identified as those that had no effect on the bulk population of monolayer cells, as determined by the luminescence cell counts and reduced mammosphere formation in the primary screen and induced cell death in the follow-up secondary validation using anoikis-resistant cells.
Results
High-Throughput Cell Counting Based on Luminescence Is Comparable to Standard Cell Counting
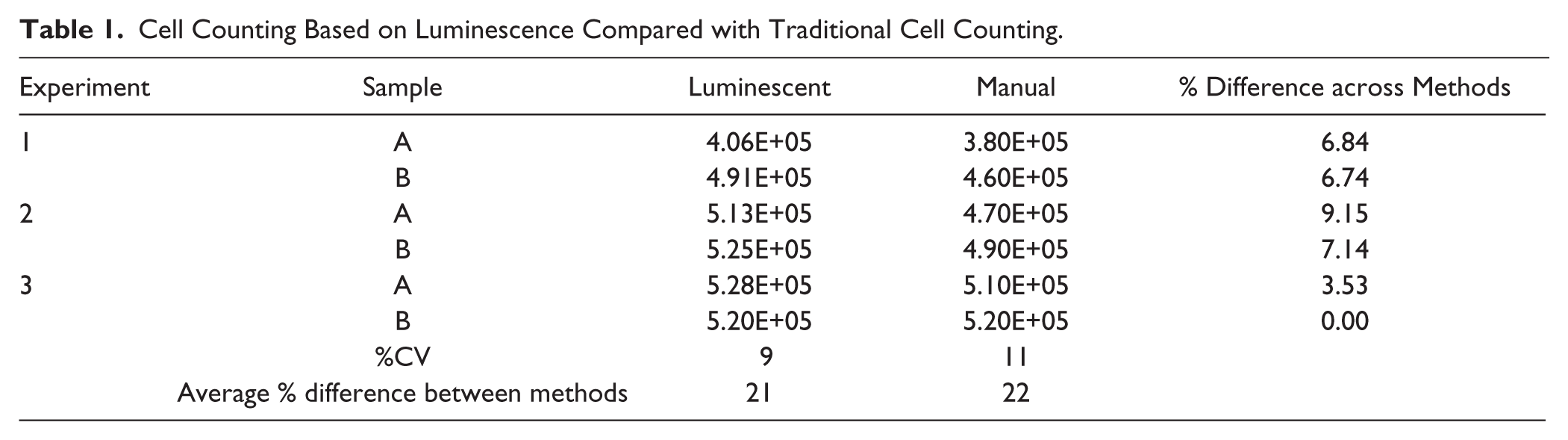
Of pivotal importance to the mammosphere assay is that differences in cell seeding densities between samples lie within an acceptable range; this can be somewhat standardized, but due to differing methods of cell counting, it must be experimentally determined for each research group. Mammosphere-forming capacity (MFC) relative to the control can only be determined if the cell number seeded into anchorage-independent growth is also relative to the control and within an acceptable range. In addition, cell seeding densities can influence mammosphere growth; if the seeding density is too high, cells can clump, forming false positives. In order to determine that the degree of difference between the high-throughput luminescence-based cell counting method used in our screen and the standard methods of cell counting was within acceptable limits, cell samples were counted in duplicate using both methods. Cells were counted using a cell counter, referred to here as manual counting; cell number for the same sample was also calculated through the use of a luminescence–cell number standard curve ( Suppl. Fig. S1 ). Counts were compared for percent difference and for percent coefficient of variation (%CV) (see Table 1 ). The degree of variation between methods was within 10%, which we consider acceptable for variance between cell counts. The %CV was calculated between experiments and was found to be lower using the luminescence-based method than manual counting, but not significantly different. These data indicate that the high-throughput cell counting method used is acceptable and does not contribute to subsequent differences in mammosphere numbers.
Cell Counting Based on Luminescence Compared with Traditional Cell Counting.
Robotic Processing to Single Cell Suspension Does Not Decrease Cell Viability
The mammosphere assay is particularly sensitive to the opposing influences of cell aggregation, causing false positives, and overmanipulation, causing false negatives. It is therefore of critical importance that cells are processed to a single cell suspension without overmanipulation, which could result in cell death. In order to compare the viability of cells post manual syringing and with that of robotically pipetted cells, we used the LIVE/DEAD viability kit. Calcein-AM stains viable cells green due to intracellular esterase activity, and ethidium homodimer-1 stains dead cells red, indicating loss of plasma membrane integrity. Cells were imaged using a fluorescent microscope and the number of red and green cells counted using ImageJ software. As indicated in Supplementary Figure S2 , following viability staining, there was no difference in the ratio of viable to nonviable cells. Robotic handling therefore does not contribute to any loss of cell viability during the assay and is comparable to manual syringing for the production of a single cell suspension.
Manual and High-Throughput Mammosphere Assays Produce Similar Results in a Pilot Study Using Known Controls
In order to establish that the novel high-throughput version of the mammosphere assay developed here could accurately reproduce results similar to those of the standard manual method, both methods were carried out side by side in a pilot study and results compared. Results from this pilot study were used in effect size and power calculations for assay variance and to establish cutoff limits for the FDA screen. Data previously published by our research group have shown that estrogen receptor–negative breast cancer cells, following hypoxic pretreatment, have a lower MFC than their normoxic counterparts. 18 It has also been previously shown that salinomycin has the ability to selectively target CSCs. 5 We exploited both of these chemical and physical MFC suppressors in order to validate that both the manual and newly developed robotic assays produced similar results. Cells were cultured for 48 h in normoxic and hypoxic conditions in the presence and absence of normal culture media, DMSO vehicle, and salinomycin. Following incubation, the robotic mammosphere assay was started and allowed to proceed unsupervised, while the manual procedure was carried out on identically treated samples. Results in Figure 2a indicate that the high-throughput robotic and standard methods produce comparable outputs, but at different throughputs. Hypoxia induced a 62% and 51% drop in MFC in the manual and robotic methods, respectively, while salinomycin induced a 73% and 68% drop. There was also no change in sphere morphology based on smaller culturing conditions, as seen in Figure 2b . The effect size Cohen’s d value was calculated as 7 for both manual and robotic methods when measuring the response to hypoxia.
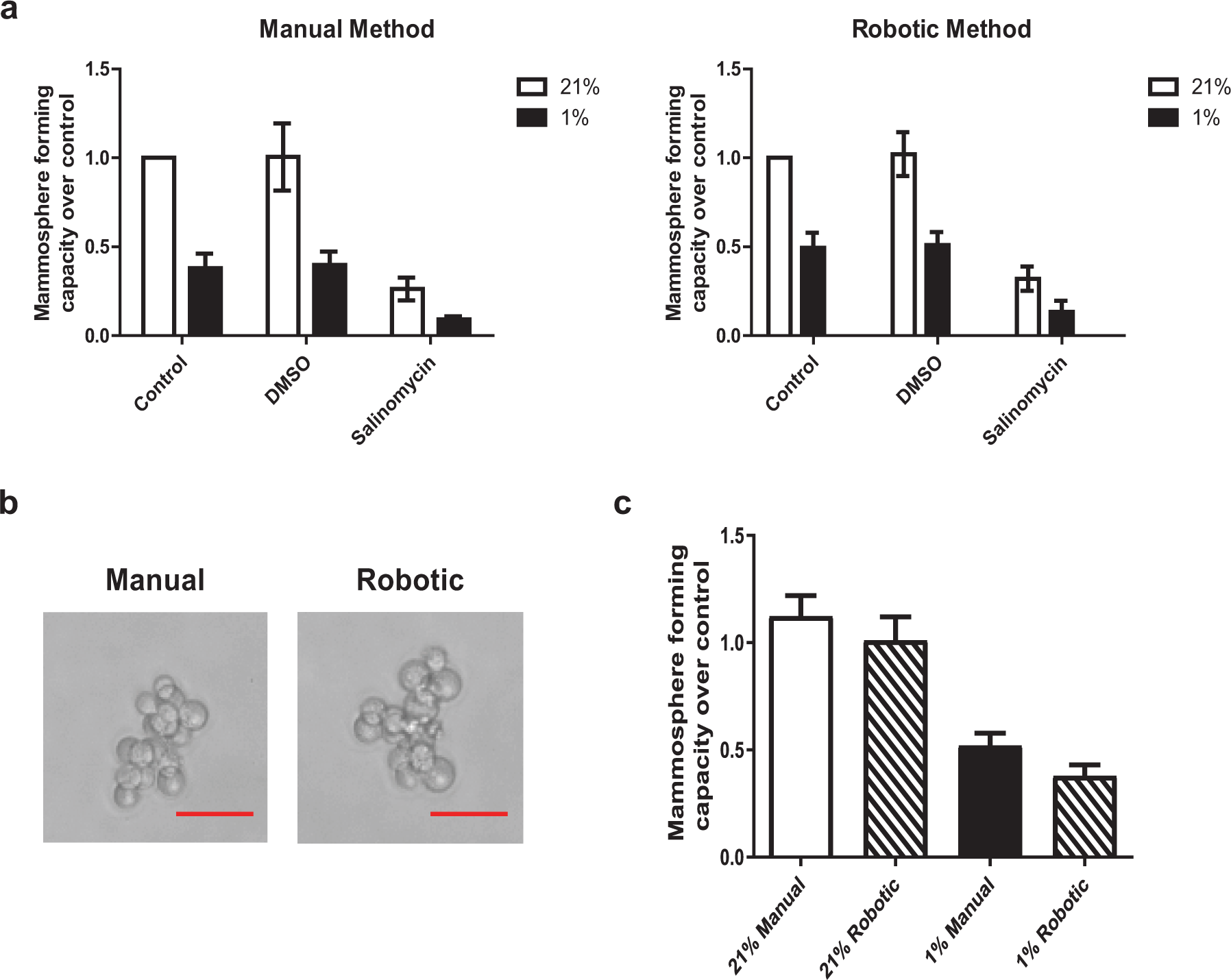
The MFC in response to known controls is similar in the manual and robotic pilot study. (
High-Throughput Robotic Mammosphere Assay Has a Similar Degree of Variance as Standard Mammosphere Assay
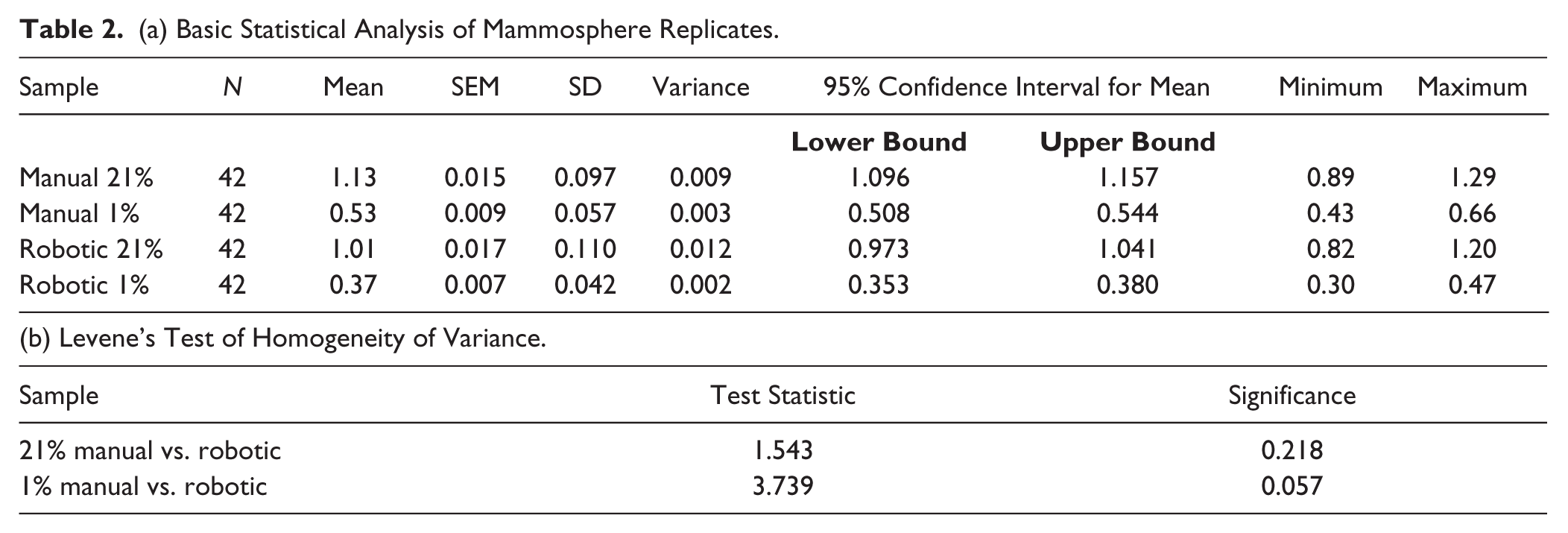
The degree of variance between the standard manual and the robotic approach was assessed as follows. MDA-MB-231 cells were cultured in normoxia and hypoxia for 48 h prior to seeding the cells into the mammosphere assay. Cohen’s d tables were used to determine a sample size with sufficient statistical power to evaluate variation between replicates within each assay. A confidence interval of 0.95 and a conservative limit for Cohen’s d of 0.8 required a sample size of 42 replicates of each condition. Calculations were carried out using the R statistical software environment and double-checked using an online calculator. 19 The degree of variance between replicates was assessed via basic statistical calculations, as well as with Levene’s test for homogeneity of variances using SPSS (IBM, version 20). For both systems, we observed a robust 0.6-fold downregulation of MFC following hypoxia ( Fig. 2c ). Applying basic statistic calculations and Levene’s test, we found that the variances between the two systems did not differ significantly ( Table 2a , b ). These data indicate that the degree of variance is not significantly different between the methods.
(a) Basic Statistical Analysis of Mammosphere Replicates.
High-Throughput Robotic Mammosphere Screen Identifies Potential Lead Compounds from Previously FDA-Approved Drug Library
The assay validation data showed that the robotic miniaturized mammosphere assay was comparable to the standard manual method; thus, a proof-of-concept screen was carried out using 989 FDA-approved drugs. MFC was assessed for each compound following incubation. Potential compounds for follow-up were identified as those giving a positive or negative response greater than 50% difference in MFC over the control. This is illustrated in Figure 3 as data points falling above or below the top and bottom red lines, respectively. Of the compounds tested, 78% were found to fall outside of our requirements as a robust hit and were therefore excluded from further analysis; 5% were found to increase MFC, while 17% lead to a decrease ( Fig. 3 ). The compounds were then subdivided based on clinical usage to establish if one particular type of drug had a more significant impact on the stem cell compartment (data not shown). Subdivision of compounds based on clinical usage did not, however, yield any data that would suggest that one particular drug type was more or less effective at targeting breast CSCs. This is potentially due to the fact that while these drugs may have similar clinical usage in the treatment of specific medical conditions, their usage is not limited to these conditions; rather, some compounds can be indicated in multiple disease states. Furthermore, even though they may target similar pathological conditions, they have greatly dissimilar molecular targets. We did, however, find that the antineoplastic drugs presented a 27% positive hit rate, with 12 of 44 compounds showing a reduction in MFC greater than 50% ( Fig. 4b ). The antineoplastics were therefore selected for a secondary screen to support and validate the primary screen data and to further refine the potential lead target list.
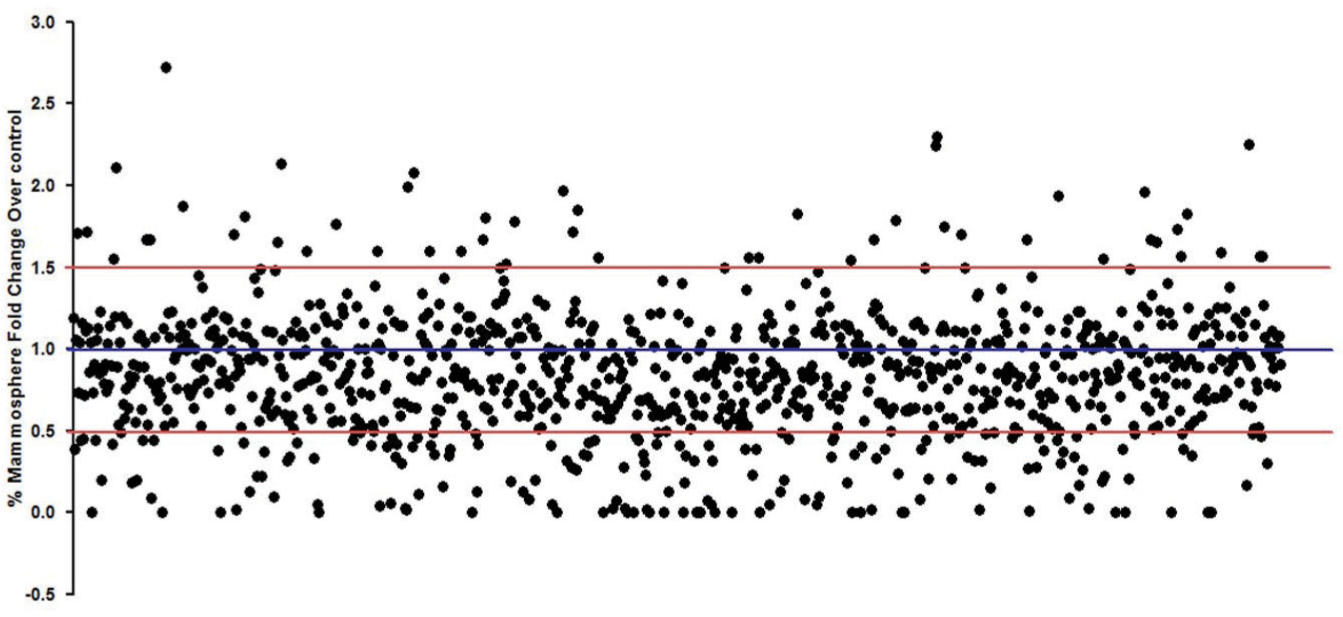
HTS of FDA-approved drug library (989 compounds). Each dot indicates an individual drug’s mammosphere response. Blue line indicates control level, while red lines indicate the level of a robust alteration in MFC.
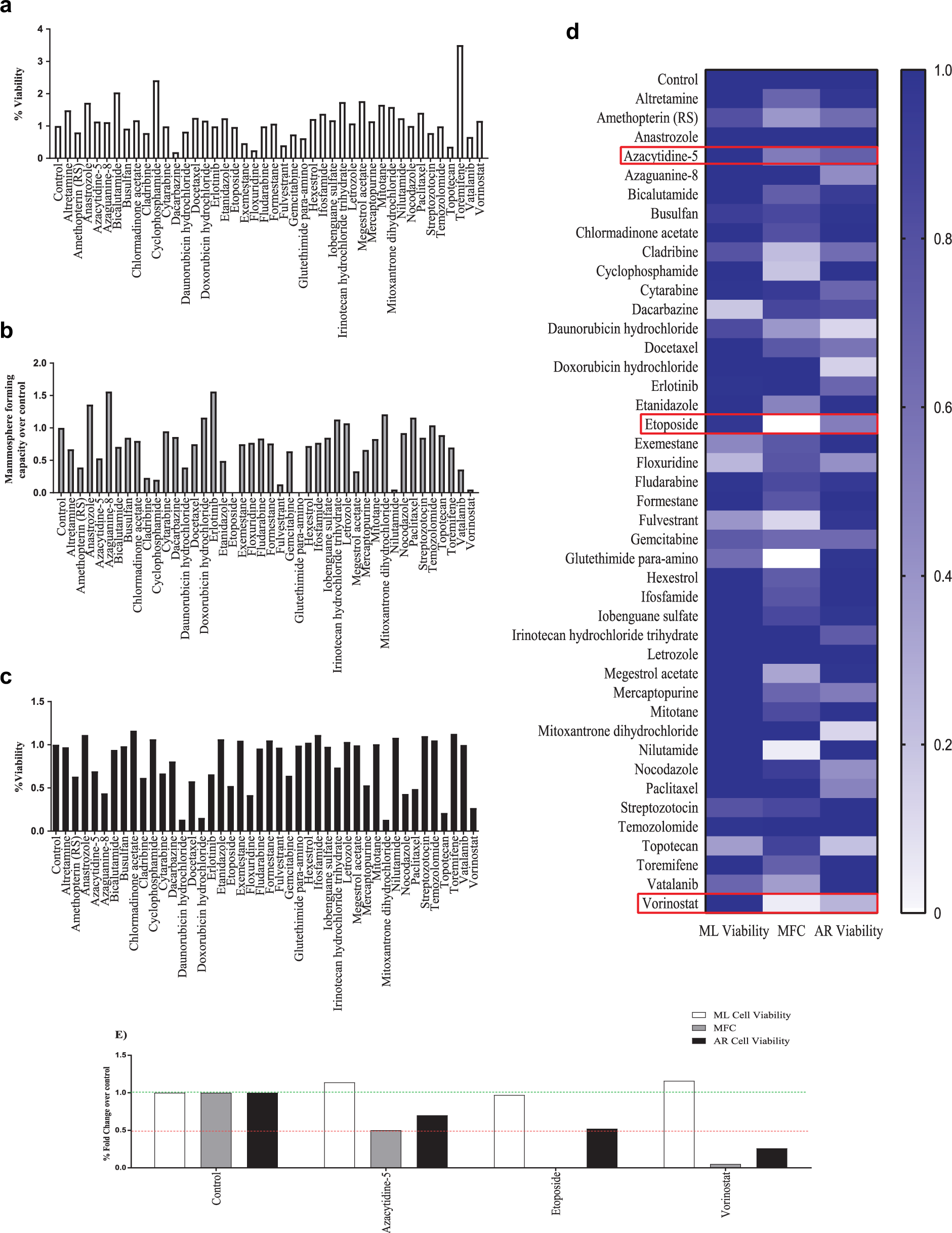
Examination of antineoplastic drugs from the FDA library. (
Anoikis-Resistant Growth Conditions Enrich Breast Cancer Cells with a Stemlike Phenotype
qPCR-based gene expression analysis was conducted in order to establish that cells surviving anoikis-resistant growth conditions represented a more stemlike fraction of the total cell population. Expression levels of pluripotency-associated genes, POU5F1 and NANOG, were increased in the anoikis-resistant cells one- and twofold, respectively, with a minor increase of 0.5-fold in SOX2, while differentiation markers CDH1 and CD24 were decreased ( Suppl. Fig. S3 ). This indicates that the short-term culture method is an appropriate enrichment technique to select for and isolate stemlike cells.
Validation Testing Using Anoikis-Resistant Cell Viability Further Refines Primary Screening Data to Identify Three Potential Lead Compounds
Following the primary robotic screening, we sought to further validate our list of hit compounds by performing a follow-up viability assay using anoikis-resistant cells. The secondary assay was designed to measure which compounds had a toxic effect on the stemlike population of cells. To this end, anoikis-resistant culture was used to enrich for this population of cells, which were then harvested, counted, and plated in the presence of the antineoplastic compounds. The Alamar Blue assay was used to measure the anoikis-resistant cell-specific cytotoxicity of these compounds after 72 h of incubation ( Fig. 4c ). Compounds were then assessed for the following criteria: (1) monocytotoxicity to the bulk population of cells, as there is no significant impact on the bulk monolayer cells, and hence they are stem cell specific; (2) reduction in mammosphere formation; (3) and toxicity toward anoikis-resistant cells. Of the antineoplastic compounds tested, three were found to robustly meet this criteria: 5-azacytidine, etoposide, and vorinostat ( Fig. 4d , e ).
Discussion
In order to improve the speed and accuracy of the classic mammosphere assay, we have developed a miniaturized version that is compatible with existing robotic platforms, thus allowing for automation of the process. Through the data outlined in this study, we have demonstrated that the miniaturized and automated assay developed here is comparable to standard methodology.
We have compared the component steps of each procedure and have found no operational difference between them, other than their throughput rate. Both cell counting techniques were found to yield similar numbers and were consistent across experiments. Single cell suspensions produced by both methods had analogous viability postprocessing, and the resulting mammospheres following treatment were also reproducible using the new platform. Finally, the robotic platform was also found not to introduce any additional statistically significant variance between replicates. The main advantage of this robotic method over the manual procedure was that of throughput rate. On average, it takes 7.5 min per sample when carrying out the mammosphere assay by hand using the method employed by our group. The robotic platform, however, can process six individual samples simultaneously, and thus operate at a rate of 45 s per sample. Mammosphere counting time is also vastly decreased from 9 min per sample to 56 s. This is approximately a 10-fold decrease in overall operating time without the introduction of additional variance.
In comparison with existing drug screening methodologies, such as those employed by Germain et al. 8 and Gupta et al., 5 which rely entirely on a preselection of cells prior to experimental treatment, our novel methodology does not require any pretreatment or preselection of cells, such as shRNA-induced knockdown or FACS. Given the heterogenic 20 and highly plastic nature of cancer cell populations, 21 any premodification or preselection steps will inherently introduce an experimental bias and thus should only be used as secondary screening methods. CSCs already represent a minimal subfraction of the total population; further selection steps reduce the number of available target cells to increasingly small numbers with an increasingly refined phenotype. This could account for the low positive hit rates observed in some studies. It should be noted that the method outlined in this study does require that a luciferase-expressing cell line is used in order to allow for cell estimation in a 96-well plate. The cells used in this study were a commercially available and validated luciferase cell line. Other luciferase-expressing breast cancer cell lines are also commercially available, one of which, an estrogen receptor–positive variant, is currently under validation in our laboratory for use in future studies. HTSs are inherently prone to noise and false-positive or -negative results; hence, an approach that combines more than one screening platform and lead target validation is a prerequisite for an efficient system. 22 The system employed by our group does not exclude cells from the bulk population, thus allowing us to potentially target all subgroups present. In addition, because our end-point measurement is mammosphere formation and is not specifically limited to cytotoxicity, this method also allows us to study compounds that could affect the (de-)differentiation machinery at play within the tumor environment. Through the usage of a secondary validation method, we can reduce the false-positive hit rate and select for only the most robust of candidates to analyze further. This dual protocol of screening and validation provides high-throughput, accurate information on test compounds’ abilities to impact the stemlike population of cells within breast cancer.
Our proof-of-concept screen using FDA-approved drugs indicates the successful application of this platform to the search for CSC-specific therapies. Of the 989 compounds screened, we found a significant number within the antineoplastics to be effective in reducing the MFC. This is not surprising given that the intended nature of these compounds is cytotoxicity against cancer cells. In this instance, reduction in MFC is a likely result of overall cytotoxicity, which could mask a true stem cell–specific targeting ability. 23 It is for this reason that we also measured the viability of the monolayer cells after treatment and excluded compounds that had a significant impact on the cell population as a whole. By assessing only those compounds that had no impact on the bulk of the cell population but reduced the MFC as seen in the primary screen, we could effectively identify those compounds with a stem cell–specific action. Furthermore, our validation assay refined our potential hit list by elucidating those compounds that not only cause a reduction in mammosphere formation, but also possess toxicity toward anoikis-resistant cells. Of the antineoplastic compounds tested, three were found to robustly meet our criteria, 5-azacytidine, etoposide, and vorinostat. Indeed, one of these three, etoposide, has also been identified in previous screening platforms by other groups,5,24 for potential usage against the stem cell subpopulation of breast cancer. These studies used vastly different methodologies from both each other and our currently outlined assay; however, both studies used a number of validation techniques to add weight to their screen findings for etoposide, which, when taken in combination with our own study, points significantly toward the usage of etoposide as a potential anti-CSC therapeutic. In addition, a recent study by Pathania et al. identified 5-azacytidine as a compound capable of blocking the tumorigenicity of cancer stemlike cells; their work also points toward a TNBC-specific action for this compound through the targeting of RAD51AP1, which they found to be highly expressed in estrogen receptor–negative tissue types. 25 Finally, vorinostat has been highlighted previously 26 to have the ability to modulate EMT in MDA-MB-231 cells through the increasing of E-cadherin levels. Indeed, this study pointed toward the requirement to combine vorinostat with additional inhibitors for effects in vivo to mimic in vitro measurements. This may indicate why this compound displayed the lowest efficacy of the three compounds identified in our screen. We feel these data, taken together with our screening data, indicate that our screening method is a high-throughput, stable, and effective platform for the identification of compounds with potential therapeutic usage against the stem cell compartment of TNBCs. In addition, TNBCs could be uniquely targeted through the usage of currently available antineoplastics as a stem cell–specific adjuvant treatment.
Footnotes
Supplementary material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants from BioCARE, Cancerfonden, Sahlgrenska Universitetssjukhuset, and Vetenskapsrådet.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
