Abstract
Artificial intelligence is the future of clinical practice and is increasingly utilized in medical management and clinical research. The release of ChatGPT3 in 2022 brought generative AI to the headlines and rekindled public interest in software agents that would complete repetitive tasks and save time. Artificial intelligence/machine learning underlies applications and devices which are assisting clinicians in the diagnosis, monitoring, formulation of prognosis, and treatment of patients with a spectrum of neuromuscular diseases. However, these applications have remained in the research sphere, and neurologists as a specialty are running the risk of falling behind other clinical specialties which are quicker to embrace these new technologies. While there are many comprehensive reviews on the use of artificial intelligence/machine learning in medicine, our aim is to provide a simple and practical primer to educate clinicians on the basics of machine learning. This will help clinicians specializing in neuromuscular and electrodiagnostic medicine to understand machine learning applications in nerve and muscle ultrasound, MRI imaging, electrical impendence myography, nerve conductions and electromyography and clinical cohort studies, and the limitations, pitfalls, regulatory and ethical concerns, and future directions. The question is not whether artificial intelligence/machine learning will change clinical practice, but when and how. How future neurologists will look back upon this period of transition will be determined not by how much changed or by how fast clinicians embraced this change but by how much patient outcomes were improved.
Keywords
Introduction
Artificial intelligence (AI) is rapidly changing the face of society and inevitably, clinical practice.1–7 While the history of AI stretches back decades, most recently, the release of ChatGPT3 in 2022 brought generative AI to the headlines and rekindled public interest in domestic robots, singularity and software agents that would complete repetitive tasks and enable more time for tasks which require dedicated human expertise. Since then, there has been a shift in public perception as industries have scrambled to add ‘genAI’ as a suffix or prefix to their products or work practices. However, this has not always been accompanied by an increase in understanding. The deluge of both tech industry hyperbole perpetuated by opportunists, mixed with genuine technology advancements and publications, has made it challenging for the average clinician to discern innovation from marketing. Particularly in this age of social media, there are a multitude of opinions which range across extremes, either magnifying the impact of the technology or foreshadowing dystopian outcomes. Some of us are privately drowning in a sea of unfamiliar terms, which provide abundant raw fodder for malapropisms in the workplace.
There is no shortage of comprehensive reviews of the use of AI in medicine.8–12 This primer will review the basics of machine learning (ML) and describe the utility of ML models in neuromuscular and electrodiagnostic medicine, with the hope that by providing an understanding of the fundamentals, opportunities, and pitfalls of ML, a decision to implement such tools can be made from a position of knowledge. In addition to the forthcoming discussion, a glossary of ML terms is provided for reference (Table 1).
Important terminology.

Key concepts in machine learning
As a grounding, it is worthwhile to first understand how the different terms seen in contemporary publications relate to each other (Figure 1). AI is used to describe the field of computer science determined to create machines that replicate human intelligence.6,13 Within the field of AI, machine learning refers to a large set of statistical and mathematical methods enabling computers to be trained to recognize patterns and relationships between clinical inputs and outcomes of interest using existing data, without explicit programming. Deep learning falls within ML and refers to multi-layer artificial neural networks that are trained on extremely complex patterns to make predictions. 13
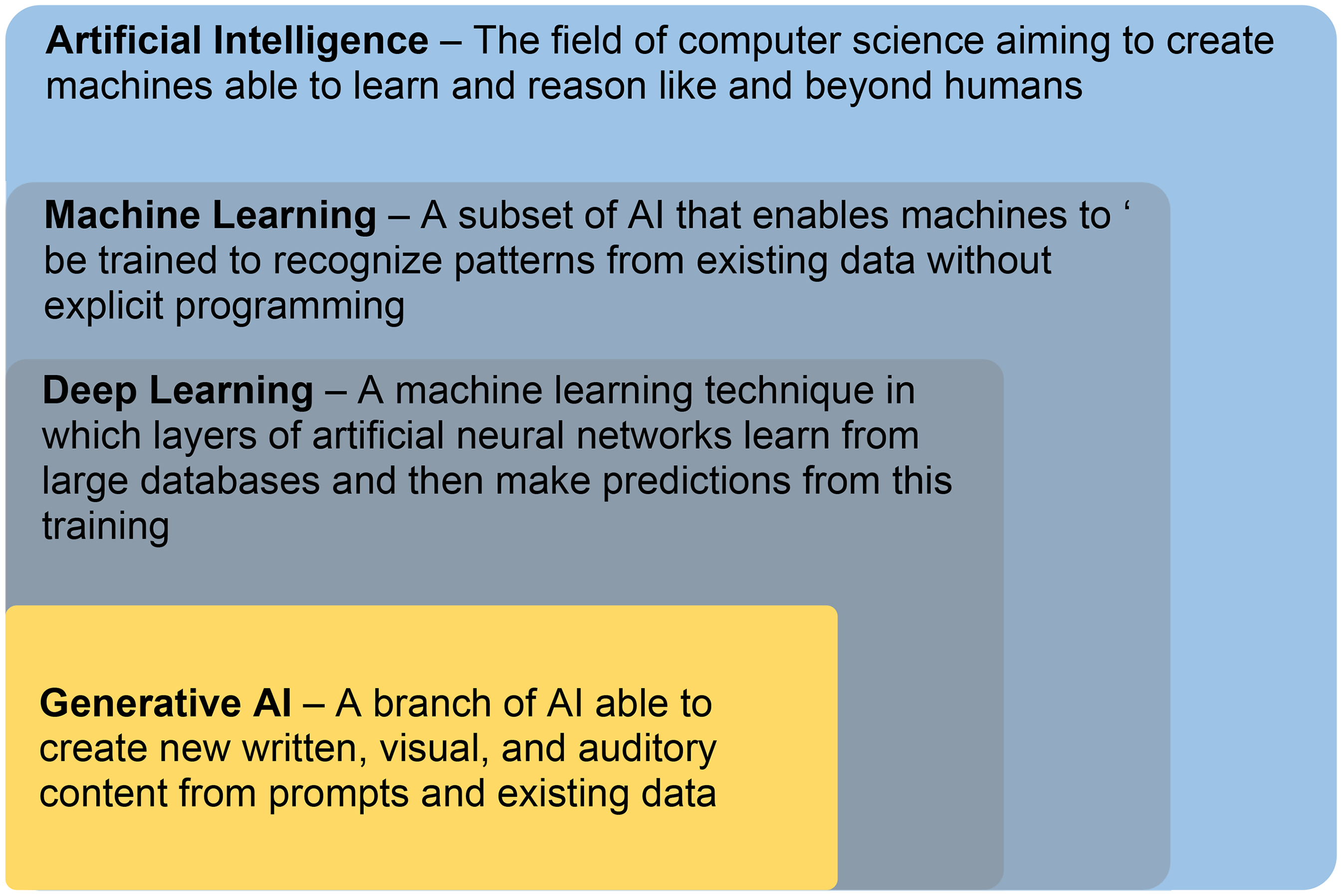
Relationship between AI, ML, DL, and generative AI.
While this publication focuses on machine learning, it would be remiss to not mention generative AI. As can be seen in Figure 1, generative AI is borne from machine learning. It recognizes patterns and makes predictions to generate content. Generative AI in the form it is today, whether it be OpenAI's ChatGPT, 14 Anthropic’s Claude, 15 Google’s Gemini, 1 or Meta’s Llama, 16 would not have been possible without prior work such as the landmark research paper from Google researchers in 2017, entitled “Attention Is All You Need”, 17 describing an augmentation to artificial neural networks known as a transformer. The transformer enables longer patterns of data to be analyzed and processed in parallel and is why large language models (LLMs) such as ChatGPT can ‘magically’ comprehend long passages of text, hold conversations, and generate equally long content. Yet, despite this impressive ability to manipulate language, current generative AI lacks genuine understanding of the meaning behind the text they generate. They are no more than stochastic parrots, 18 generating content by statistically stitching together sequences of linguistic forms observed in their training data, relying on probabilistic information about word combinations rather than semantic understanding. This is an important understanding as it explains in part their fundamental flaws, one of which are hallucinations, instances where the model generates text that is factually incorrect or nonsensical, despite appearing grammatically correct and superficially coherent. This phenomenon arises from the probabilistic nature of LLMs and their reliance on statistical patterns in the training data rather than deep semantic understanding of content.
LLMs prioritize fluency and coherence over factual accuracy, causing them to produce plausible-sounding but entirely fabricated statements especially when faced with ambiguous prompts or incomplete information. Hallucinations undermine the reliability of AI in any setting but are particularly dangerous in a clinical setting where factual accuracy drives decision-making.
There are ways to minimize this – prompt engineering, retrieval augmented generation, establishing safety guardrails within the software, but none are adequate solutions. For now, it is perhaps prudent to exercise caution and unwise to rely on LLMs in clinical practice, no matter how fluent they appear to be.
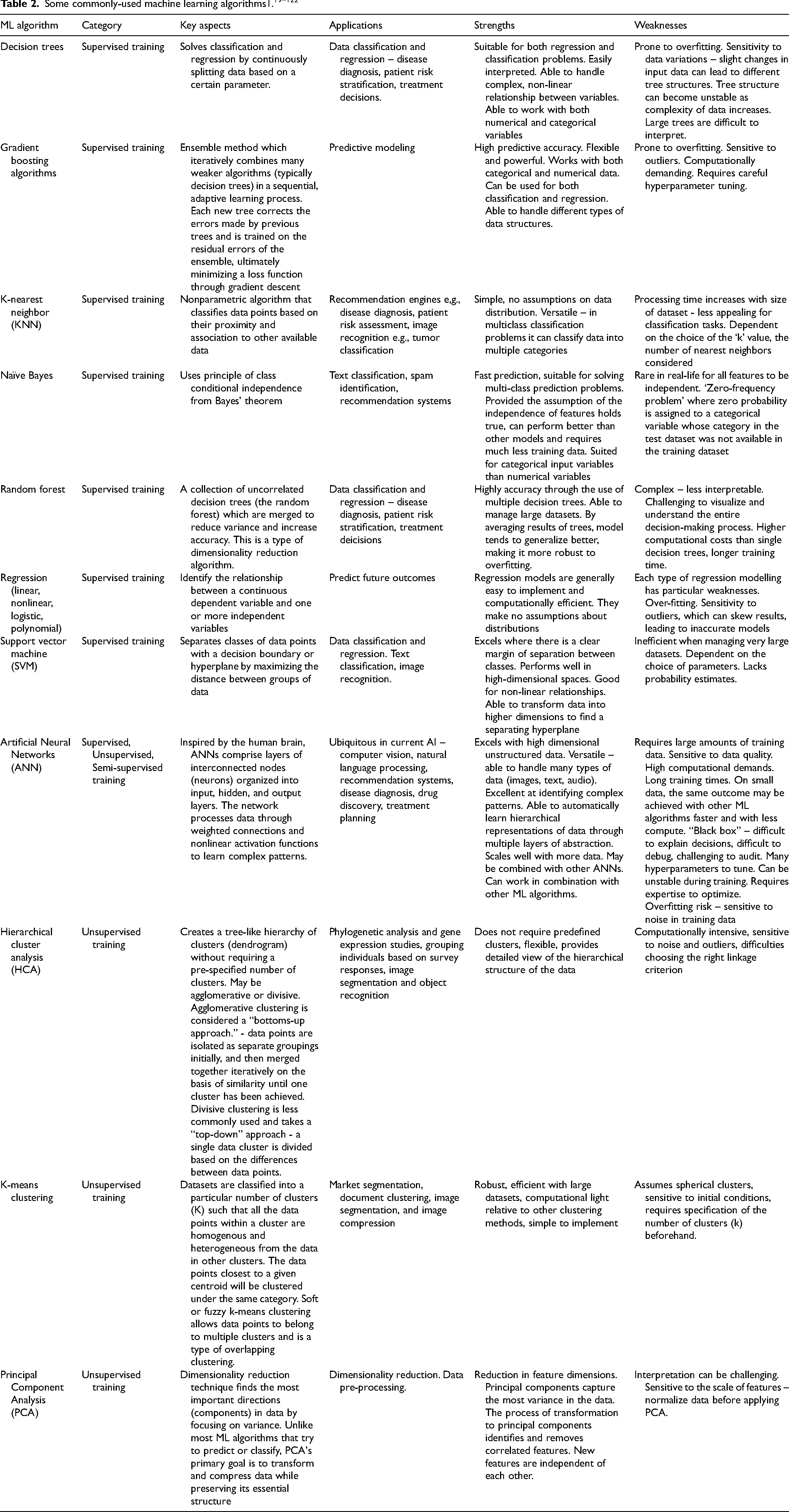
ML algorithms and models
Fundamental to ML are the algorithms - mathematical procedures and techniques that allow computers to learn from data, identify patterns, make predictions, or perform tasks without explicit programming. Which algorithm(s) to use for a particular problem is at the discretion of the data scientist who should be properly armed with knowledge of contemporary techniques. Some options are listed in Table 2. These are far from comprehensive. Many terms will be recognized by those who are familiar with statistical methodologies. In medical applications, it is common to employ several combinations of ML algorithms to enhance accuracy and reliability. Diagnostic imaging may use principal component analysis combined with a neural network for dimensionality reduction and pattern recognition. Disease prediction models may use a combination of gradient boosting and logistic regression, or support vector machine and K-nearest neighbor. Patient monitoring models could use ensemble methods to combine multiple classifiers.
ML algorithms can be broadly classified into supervised or unsupervised learning methods although there are additional categories such as semi-supervised, when labelling is incomplete, and reinforcement learning, where algorithms learn through trial and error by interacting with an environment to maximize rewards,
Hyperparameters influence the learning trajectory and the final accuracy of the model and can be manually tuned by the model developer prior to training. Examples of hyperparameters can be found in Table 1. In contrast, parameters are defined as internal variables which are learned by the ML model during training and cannot be manually tuned. Regardless of the method category, each ML algorithm has a set of hyperparameters which control the learning of the model, and these are set before a model is trained.
In research and business meetings today, the phrase “neural network” is often uttered sagely, sometimes without true understanding of its meaning. A neurologist who understands the human biological neural network should be somewhat familiar with its computational equivalent, and if not, at least strongly curious. A primer on machine learning would therefore be remiss without discussing the artificial neural network (ANN). The life sciences equivalent of the ANN in terms of impact might resemble the polymerase chain reaction technique – both methods earned their developers a Nobel Prize – John Hopfield and Geoffrey Hinton were awarded theirs in 2024 for their work advancing the ANN. 123 From a history of fluctuating popularity and high computational power requirements, the ANN is now the foundation of global-scale AI tools today across all industries and domains. 124 The aforementioned transformer which generative AI today relies upon, is itself a type of artificial neural network.
How an ANN works is loosely analogous to the way biological neurons work together. An ANN consists of nodes arranged in layers – an input layer, one of more hidden layers not visible to the user, and an output layer. Each individual node can be viewed as its own linear regression model, composed of input data, weights, a bias (threshold), and an output. A node can be thought of as a thing which calculates a number, a function. Each node is connected to others and has its own associated weight and threshold for activation. A node only activates and transmits data to the next layer if the output reaches a specific threshold value. Thus, activations in one layer determine the activations in the next until they reach the output layer, where each node represents a particular outcome contains the probability of that outcome given the inputs. For example, each node could be a character in the case of an ANN designed for optical character recognition. This is similar to how neurons communicate with each other across neurological synapses. Chemical neurotransmitters bring signals from the terminal axons of preceding neurons across the synaptic gap to the dendrites of a neuron. These neurotransmitter signals may be “weighted” to activate or deactivate the neuron using electricity. The neuron is activated to transmit signal to the following neuron only when the summed signal from all its dendrites passes the electrical threshold for activation, triggering an electrical wave known as an action potential. The action potential of the activated neuron then propagates across the neuron down its axon, to the terminal axons which synapse with the dendrites of the following neurons in the next “layer”. The terminal axons then convert the electrical signal into neurotransmitter signals which cross the synapse and bind to the dendrites of the following neuron in the next “layer”. If the threshold is reached, then that layer is activated.
Backpropagation, short for “backward propagation of errors,” is the method used to train a neural network. It involves calculating the error (the difference between predicted and actual values) and propagating this error backward through the network to update the weights. During training, initial weights and biases are set, then for each input data, a guess is made as to the output. This is compared with the known answer and if incorrect, the weights and biases are updated and the process repeated.
This back-and-forth process is guided by a loss function, and an optimization algorithm which is used to minimize the loss function. The loss function quantifies how well the predictions match the answer. It measures the error between the predicted and true outputs and provides a signal to guide weight updates. Examples of loss functions include mean squared error and cross-entropy loss. Gradient descent, the optimization algorithm commonly used in neural networks, iteratively adjusts a model's parameters in the direction of the steepest descent (negative gradient) of the loss function to minimize error. An analogy would that of a ball rolling around a skateboard park, trying to settle in the deepest bowl (global minimum). The ball naturally rolls downhill (following the gradient), picking up speed on steep ramps (larger gradients) and slowing down on flatter surfaces. If the ball gets stuck in a shallow bowl (local minimum), it might need a push or adjustment to move toward the deepest bowl, ensuring it finds the lowest possible point. At this point the weights and biases for each node are optimal, the network has ‘learned’. This is a topic where informal video125–128 is more effective for teaching than formal literature.129,130 Both are provided in the references.
There are many possible augmentations to the neural network – Transformers, the Convolution Neural Network, the Recurrent Neural Network, Generative Adversarial Networks, Graph Neural Networks – these are only a few of the options available. Each advances the traditional ANN in a particular way to produce better results for certain applications.
While variants and augmentations of the classical ANN dominate AI today, it has not rendered other ML algorithms obsolete any more than the tractor eliminated the need for picks and shovels. Sometimes, a simpler algorithm can achieve the same result in less time and computing power. This is especially the case when data is small, or there is enough prior knowledge that deep learning is unnecessary, or a situation requires a method that is explainable and interpretable in its process. It is a question of determining the best tool for the problem.
ML model training
An ML model must be properly trained before it can be deployed. This process is divided into training and testing and shown as a data flow diagram in Figure 2.
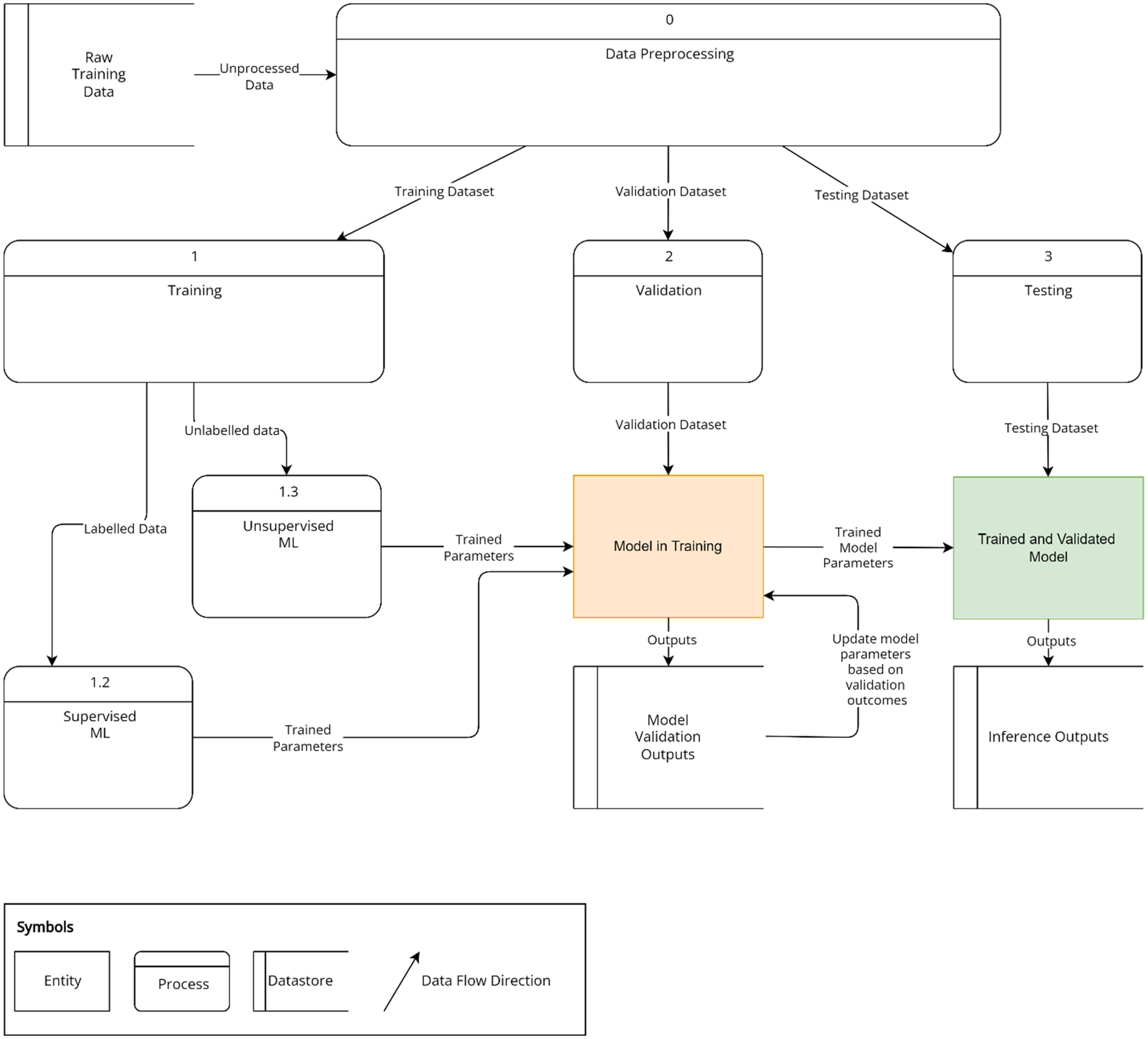
Overview of machine learning model training. Pre-processed data is divided into independent training, validation and testing datasets. Processes 1, 2, and 3 are sequential and dependent upon the completion of the previous process. Labelled training data is run through a supervised machine learning model (1.2). Unlabelled data is run through an unsupervised machine learning model (1.3). Validation data (2) is used to ascertain stopping criteria for training. The training (1) and validation processes (2) run through multiple iterations until satisfactory accuracy as assessed by performance metrics is achieved. The trained model is then tested using the third dataset (3) and the outputs are referred to as inferences.
ML models typically require large amounts of data for modelling the data to draw meaningful inferences from them. Both supervised and unsupervised approaches require input data with sufficient parameters that are descriptive of the problem under study. 131 Examples of parameters include demographic data, laboratory values, drug prescriptions, and imaging data. This data consists of continuous values, such as grayscale units for imaging, or categorical values, such as gender and ethnicity. Complex data such as images or -omics datasets can be used. Simpler machine learning systems use limited input variables, such as age, gender, and other demographic features, and have less information to predict an outcome.
Initially, data used for ML needs to be preprocessed, generating parameters which are the input variables. Parameters can be further processed by feature engineering. Supervised learning methods train the ML algorithms to more accurately “label” or classify data into categories that are determined by experts or gold standard tests. Supervised learning methods require data which is already “labelled” into categories. For example, patients may be labelled with better or worse prognosis,132–137 as responders or non-responders to treatment, 138 or with longer or shorter duration of disease. 139 Other examples of clinical data which may be labelled for supervised learning include labelling imaging findings with their clinical diagnoses7,19–22,140–142 and labelling EMG audio or visual data with various EMG waveforms.7,23 This is the more commonly used form of ML, which requires less data than unsupervised learning (see below), and can achieve human-level performance.
When labelling of data is not possible or impractical due to the volume of data, unsupervised learning approaches are used. The unsupervised ML algorithm processes the unlabelled data to cluster patients with similar characteristics together or learn the distributions of most of the data to identify outliers or patterns. Applications of clustering approaches include clustering similar clinical phenotypes of diseases,24,136 clustering similar imaging findings, 25 and clustering associated gene expression networks.26,27
Data are always divided into a training dataset, which is used to train the model, and an independent testing set, which is used to assess the accuracy of the model (Figure 1). The training set is used to learn parameters and tune hyperparameters to achieve the best ML accuracy on the independent test set. A small subset of the training set (referred to as the validation set) is used to validate the model during the training process to make decisions on the number of training iterations (consecutive training events) and tuning of hyperparameters. If the dataset is small, a technique known as cross-validation can be used to improve reproducibility. In a cross-validation study, the whole dataset is divided into n-subsets, with each model trained on (n-1) subsets and the remaining subset is used for testing. In this way, each subset will be a test set in at least one model.
One of several issues which may manifest during the training process is overfitting in which the model perfectly fits the data used in training but also mistakenly learns irrelevant features or random fluctuations in that particular dataset as significant concepts influencing the outcomes, when there is actually no logical relationship between them. This could happen when too many parameters are available to fit the data. For example, a model trained too well on a dataset from patients in Europe may be very accurate in answering questions on this particular set of European patients but have very poor performance on a North American database of patients. Regularization techniques can be applied to prevent overfitting which stop the training process early, before the model overfits to the dataset, or increase the effective dataset size by augmenting the model with synthetic data that have been artificially modified so that the model is exposed to a diversity of data distribution and can generalize well. Alternatively, the number of parameters may be reduced.
After training the independent testing dataset is run through the model and results assessed against known outcomes. If the model meets predefined requirements, it is deployed.
Data quality
While an understanding of ML algorithms is important as it helps anticipate the shortcomings of a particular model, it equally important for a clinician to be able to critically assess the quality of the data used to train and validate a model. If possible, it is worthwhile spending time reviewing and discussing the training data with the responsible data scientist – determine whether the training data is from a reputable source, how is it quantified, how was it collected, how was data quality and accuracy of content verified, whether the data been used for other models – this information will impact data quality and subsequent model quality. Check if there is any reason for bias – such as missing data, inaccurate labelling, not including patients of a particular ethnicity. Even if presented with a trained model, these questions are still relevant.
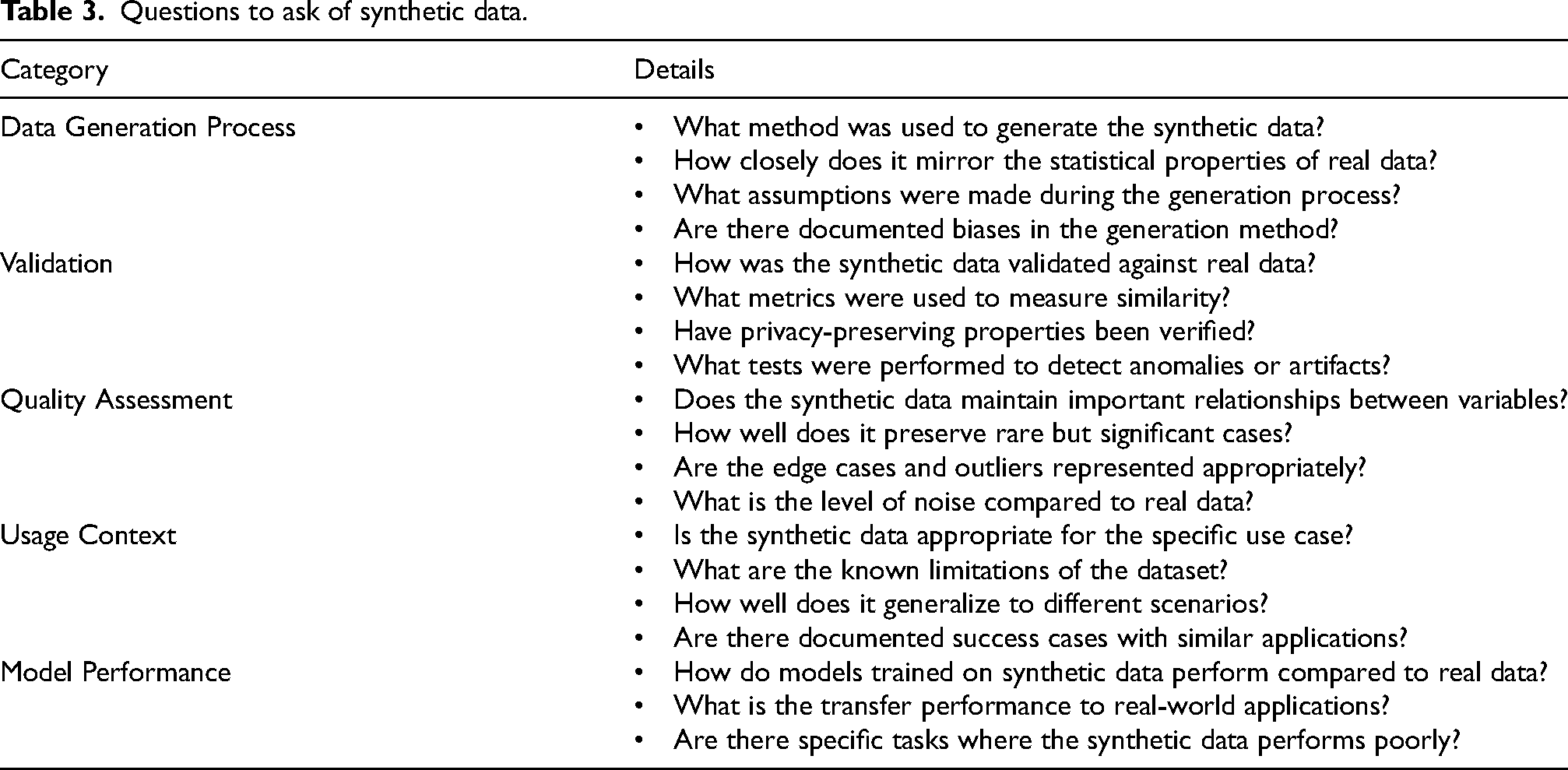
The tech industry claims to have exhausted the global supply of quality training material for AI28,29 and in certain areas of medicine, big data is inaccurate term – sparse data, fragmented data, poorly documented data is more accurate. Consequently, many advocate and are using synthetic data for training.30,31 In this context, synthetic data can be defined as data generated by AI for the purpose of training new AI. Synthetic data addresses the challenges associated with using real world data for training – namely sparse data, fragmented data, data privacy regulations, data security, and resource constraints. Examples include - autonomous vehicle training, financial fraud detection models trained using synthetic data, and even health records have been generated for training AI to avoid using patient records. To what extent is this acceptable? Currently there is no regulation around the use of synthetic data, and it is in the interests of the tech industry to foster this practice – it is cheaper and faster to use synthetic data. Synthetic data contains all the biases and flaws of the AI which generated it and these negative qualities are transferred to the next AI. When considering synthetic data is it particularly important to understand how the data was generated, how it was validated, the quality assessment, how the data will be used and how model performance will be assessed. Example questions for each of these areas are presented in Table 3.
Questions to ask of synthetic data.
AI applications in clinical medicine
The progression of AI into healthcare applications has been predicated on quality data for training such as high-resolution medical imaging, biosensors with continuous output of physiologic metrics, -omics (epigenomics, genomics, transcriptomics, proteomics), and electronic medical records, as well as advancements in computer processing and cloud computing. At present AI processing power is greatest though cloud computing and enables moderately powerful devices with a good Internet connection to access the power of AI inferencing servers in a distant location.
AI has been applied in clinical decision support tools to improve diagnostic and prognostic accuracy. AI tools developed by Zebra Medical Vision and Aidoc are FDA-approved to triage or enhance radiological diagnoses in neurology, pulmonology and cardiology from X-rays, mammograms and CT scans. Google’s DeepMind for eye diseases operates under CE marking for European Union usage and is utilized at Moorfields Eye hospital to analyze optical coherence tomography (OCT) for the detection of macular disease. AI tools also have good potential for treatment optimization to make more cost-effective treatment decisions, optimization of resource utilization, and improvement in quality of life. For example, the AI-based Sepsis Immunoscore is FDA-approved to identify patients at risk of sepsis. The MiniMed 670G system is FDA-approved as the first hybrid closed-loop insulin delivery system. Watson for Oncology (WFO) is an artificial intelligence assistant decision system developed by IBM with Memorial Sloan Kettering Cancer Center (MSK) to recommend appropriate chemotherapy regimens for specific cancer patients.32,33 While it is not FDA-approved, it is widely deployed internationally.
Uses of machine learning in clinical practice and research in neuromuscular and electrodiagnostic medicine
In neuromuscular and electrodiagnostic medicine, there are many opportunities to use ML - in analyzing clinical data gathered from subspecialty tests such as nerve conduction studies (NCS),34–36 electromyography (EMG),23,37–39 electrical impendence myography (EIM),40,41 muscle MRI,19,25,42–47,139–142 nerve ultrasound,129–132 muscle ultrasound,22,37,48,140 muscle biopsies,49–51 patient biorepository, and from electronic medical records (EMR).1,24,27,52–65,132–136
EMG, NCS, neuromuscular ultrasound, muscle MRIs and muscle biopsies are sub-specialty tests which are routinely used by neuromuscular specialists in clinical practice for diagnosis and monitoring. It is well-known that EMG, which is currently manually performed and interpreted by trained electromyographers, is labor intensive and has significant inter-operator variability. The accuracy of analysis of the EMG waveform data depends heavily on the training and experience of the electromyographer. Consistency may become an issue when there is a need for repeat studies for monitoring or diagnosis, and the same electromyographer is not available to perform the study. ML tools have the potential to fill this gap by enhancing the accuracy and consistency of EMG.23,38 Unlike complex and unstructured EMG data, which is a time-series signal with high frequency and changes in amplitudes, collected in various muscles decided by the electromyographer based on clinical acumen, at rest and on voluntary movement, routine NCS data is structured in the form of discrete values relating to parameters like amplitude and conduction velocity, usually collected from routine nerves in the upper and lower limbs. This lends itself well to using ML to automatically detect common conditions such as polyneuropathy 34 or entrapment neuropathies and ensure they are not missed out on diagnosis or reporting. Neuromuscular ultrasound is another tool used in clinical practice by trained neuromuscular sonographers in the diagnosis of nerve or muscle diseases. It is a newer subspecialty compared to NCS/EMG and there are fewer widely accepted guidelines on diagnostic parameters such as nerve cross sectional area (CSA) and diameter, and echogeneity of muscle and nerve fascicles. ML tools, which are already in clinical practice for other forms of imaging such as MRI and CT diagnosis for other diseases, can address this gap and provide clinical insights for neuromuscular specialists to develop diagnostic criteria for various neuropathies and myopathies. ML can automate the precise drawing of boundaries of nerves and muscles in images or videos for correct analysis,46,66 and correctly identify common entrapment neuropathies.20–22,67,68 Muscle MRIs are generally used to distinguish between inherited and autoimmune muscle diseases and monitor disease activity; the use of ML to objectively screen and analyze multiple muscles may provide a practical and accurate way to diagnose and monitor muscle diseases which often affect the whole body, and eschew the need for muscle biopsies which are invasive and prone to sampling bias.19,42–44,141,142 None of the neuromuscular specialty tests currently have FDA-approved devices in clinical practice, but have shown promise in research.
ML can also aid clinicians in the diagnosis and monitoring, clinical prognostication, personalized treatment, classifying patients and understanding diseases mechanisms of neuromuscular diseases. Again, the field has not embraced these new technologies in clinical practice but research is promising. These are summarized in Figure 3 and elaborated in the sections below. Key studies are summarized in the Appendix.
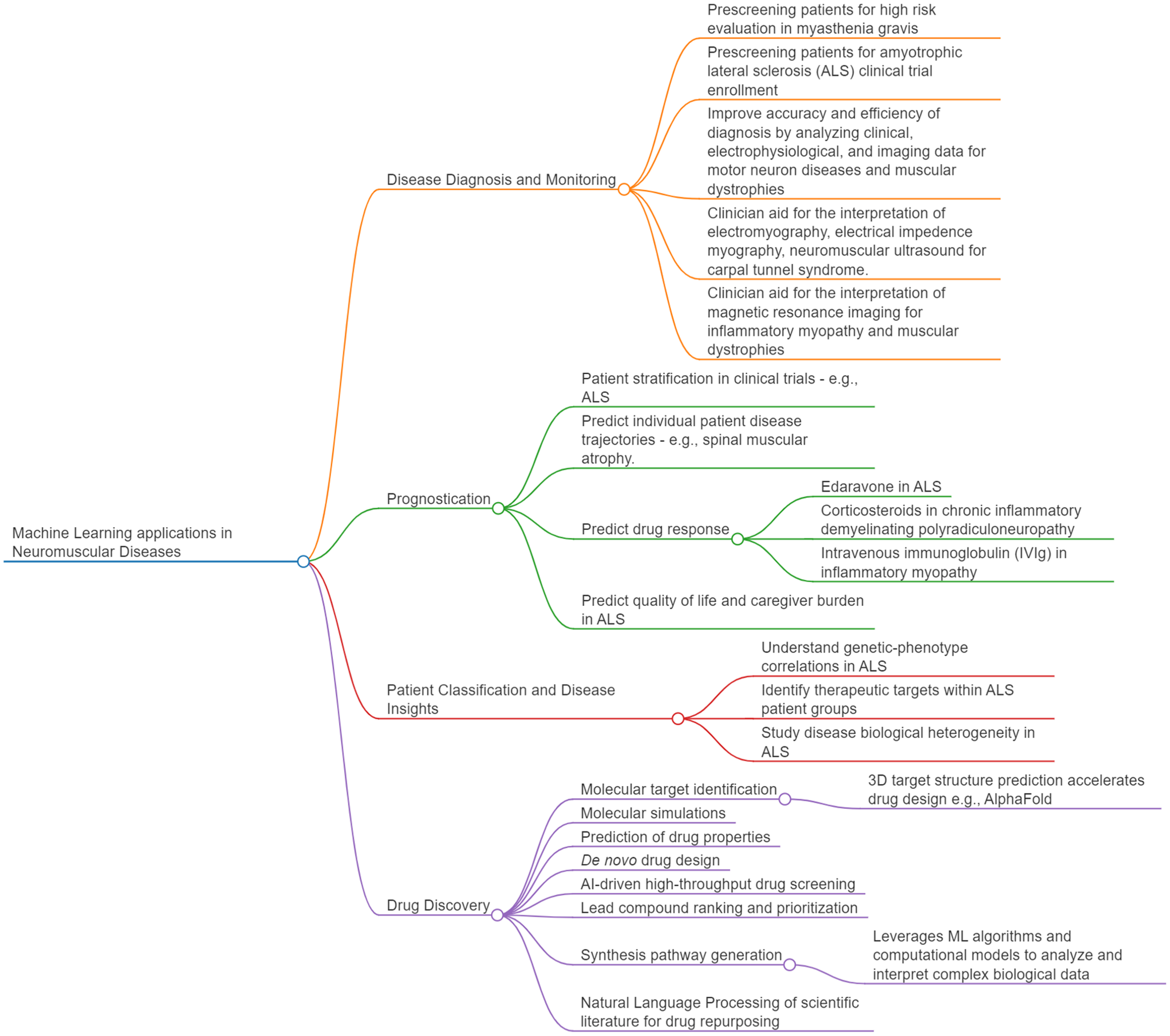
Machine learning applications in neuromuscular diseases.
ML can be used to improve diagnosis and monitoring of neuromuscular diseases
ML may be useful for prediagnosis (pre-screening patients), peridiagnosis (improving the accuracy and efficiency of diagnosis by real-time assistance at the time of diagnosis), and postdiagnosis (acting as quality control to detect diagnostic errors before patients are affected), and may complement clinician evaluations while reducing clinical workload. ML and clinician evaluation can be more accurate and efficient in combination.
Pre-diagnosis may be used to screen highest risk patients for further evaluation and management at a higher level of care. It can also identify new patient risk factors for clinicians to take note of. Supervised ML on clinical data has been used in Taiwan to detect patients admitted with myasthenia gravis who are more likely to have prolonged and severe disease and require intensive care interventions with high accuracies,63,137 suggesting that these models have the potential to be developed into clinical predictive tools. In these studies, the presence of thymoma predicted ICU admission and prolonged hospitalization stay. Interestingly, the use of rescue treatments such as intravenous steroids and immunoglobulin (IVIG) was also associated with prolonged hospitalization stay. These models have to be validated in other international patient cohorts as the Asian myasthenia gravis disease phenotype may differ from patients in Europe or the United States. Supervised machine learning has also been used on ALS clinical data to predict survival and progression rate with good accuracy,53,69,70,134 which opens avenues for patients to be screened remotely for inclusion in clinical trials. It also enables the enrolment in clinical trials of fast progressors or short survivors – patients who are most likely to show treatment response in the shortest period of time.
During diagnosis (peri-diagnosis), a ML model has potential to improve the accuracy or efficiency of diagnosis by assisting clinicians in real time to detect abnormalities more quickly and consistently. In diseases where early diagnosis is challenging due to heterogeneous clinical presentations and etiology, and routine clinical diagnosis requires clinical history and examination by a neuromuscular specialist, usually accompanied by electrophysiological tests, supervised ML can facilitate early diagnosis and access to treatment, by the automatic analysis of routine data ranging from clinical to blood analytes to electrophysiological to imaging either alone or in combination.57,60,70,71,132 It can also distinguish between subsets of similar diseases to optimize clinical management – supervised ML has been used to distinguish between ALS and lower motor neuron disease using patient blood analytes with good accuracy, and identified immunological markers as important discriminators of these two diseases, 132 which suggests that they should be managed as two diseases and not two spectrums of one disease as these diseases are commonly viewed.
Genomics, transcriptomics, proteomics and metabolomics, and molecular networks, have also been used as data for ML, which give insights into the molecular complexity of diseases and propel the discovery of network biomarkers and new therapeutic targets.51,54,55,58,64,134 Supervised ML identified diagnostic biomarkers with good accuracy from proteomics performed in patient-derived stem-cell differentiated motor neurons, 134 ALS-related genes taken from brain biopsies 55 and RNA taken from post-partum samples. 58 Although patient sample sizes were invariably small, as expected for rare neuromuscular diseases, with some datasets being imbalanced with much more or fewer disease cases, 58 it is promising that ML was able to extract important disease associated features even from highly heterogenous -omics data.
ML image analysis lends itself well to peri-diagnosis and post-diagnosis. Ultrasound has been used to assess for the common entrapment neuropathy, CTS, through the evaluation of median nerve morphology, but operator dependency and lack of standard protocols limit its widespread use. Supervised ML can be used in ultrasound analysis to detect CTS20,21,36,66,67 by detecting median nerve cross sectional area 21 – commonly used in clinical practice, or other variables not commonly used in clinical practice such as the volume, 67 echogenicity and thickness of the epineurium and surrounding tissues. 20 The median nerve can be effectively delineated by ML, 70 with automatic cross sectional area measurement aligning well with manual measurement, implying the potential of reducing dependence on sonographer and catching human errors. Supervised ML models have identified variables in predicting CTS severity, 36 which can assist clinical decision making about surgical decompression.
Muscle imaging, together with detailed clinical examination and muscle biopsy, is one of the main tools for deep phenotyping and diagnosis of neuromuscular disorders. It can give clues to the underlying pathogenicity of variants of unknown significance and facilitate diagnosis in cases with inconclusive genomics, but again is limited by clinician/operator experience and hardware, and it is time and labor-intensive to analyze large amounts of imaging data.68,140 Limitations with visual image interpretation by human operators can be mitigated with ML. Supervised ML has been used to support quantitative whole-body MRI analysis for diagnosis of myopathies, 42 sonographic diagnosis of inherited and acquired myopathies 37 MRI analysis of muscle involvement patterns and muscle imaging texture to distinguish between inherited and acquired myopathies such as muscular dystrophies, congenital myopathies and idiopathic inflammatory myopathies,42,44,139,141,142 and to correlate with disease duration and disability.42,71,139 In some studies, disease rarity contributed to a low sample size, 37 which did not cover all the various stages of disease, or few myopathies were covered, which compromised the practical utility of the model unless the clinical suspicion of the particular rare myopathy was already high. 42 Other studies were limited by the data – when images of only certain muscles 42 or parts of muscles were available. 141 Accuracies ranged from moderate to good, suggesting potential for improvement by including more homogenous and whole body data from more patients across international neuromuscular centres. Interestingly, one study used unsupervised ML on MRI to discriminate between muscles with fatty replacement, edema or neither in patients with STIM1 tubular aggregate myopathy with good accuracy, 25 and the model was able to identify alterations in muscle classified as normal by human operators – suggesting good potential in post-diagnosis, to reduce human error.
Supervised ML has been used with EIM to diagnose motor neuron diseases and muscular dystrophies as well as to correlate with muscle mass,40,41,71 and to characterize EMG signals that would be helpful for diagnosis.23,38 In addition, it has potential in automating the analysis and reporting of routine NCS to save time, for common conditions such as diabetic polyneuropathy. 34
ML may be used to improve prognostication
There is potential for ML in the design of clinical decision support systems (CDSS), to assist clinicians with counselling and decision-making based on prior successful diagnoses, treatment, and prognostication. 4 ML has the advantage of providing results without variability from fatigue or environmental factors and having the ability to exhaustively review every part of the data every time. Such models may be used to stratify patients in clinical trials, predict individual patient disease trajectories and drug responses, and predict quality of life and caregiver burden. Rare diseases, however, are limited by already small patient databases and sparse longitudinal data may be further affected by selection and attrition bias, which makes it challenging to accurately analyze the disease course of heterogenous clinical populations.
In ALS and spinal muscular atrophy (SMA), disease progression may be complex and non linear.65,71–73 This poses challenges to clinicians, patients and caregivers when trying to make plans - preparing for disability, loss of income, expensive personalized disease-modifying treatment and care plans, and death. As these diseases have considerable clinical and biological heterogeneity, patient stratification is also necessary to enrol a homogenous patient population in clinical trials. Many models of ALS progression have used supervised ML to predict survival, weight change, and progression rate, with the most common predictors consistently being age, ALS Functional Rating Scale (ALSFRS) score, site of onset and disease duration.60,69,70 However, only the European Network for the Cure of ALS (ENCALS) model has been relatively reliable 74 and is available to registered clinicians as a personalized prediction tool, which suggests that future research needs to pay attention to methodological pitfalls and external validation in ML models. Clinical laboratory predictors, such as creatinine, creatine kinase, and phosphorus, have been identified as potential biomarkers of ALS progression and clinical outcome.69,72,75 Novel prognostic and monitoring biomarkers from genomics, transcriptomics, proteomics, lipidomics to serum metabolomics from patient-derived tissues have likewise been identified by supervised and unsupervised ML and may additionally shed light on pathological mechanisms relevant to both pre-symptomatic and symptomatic phenotypes of disease.52,54,56,59,62,64,132,134 Unsupervised ML approaches can cluster ALS patients in an unbiased manner to allow clinicians to draw insights from clinical progression patterns 73 or gene expression 26 to predict prognosis.
SMA treatment has been revolutionized by innovative survival motor neuron (SMN) protein repleting disease modifying drugs which are however extremely expensive. 76 Being able to predict individual trajectories can help shed light on clinical efficacies and durability of such drugs for particular patient subsets and create personalized treatment plans – when to start or stop a treatment, and whether to combine or switch therapies as children with SMA grow.65,71 In ALS, the clinical trials on edaravone used cohort enrichment to select for patients most likely to show progression to elucidate treatment effects. Interestingly, ML was used to show a statistically significant treatment effect in a cohort of participants with broader disease characteristics than the inclusion criteria, to mimic real-world clinical practice where edaravone is administered to all ALS patients. 77 Supervised and unsupervised ML have been used on multi-modal clinical, laboratory, electrophysiological and imaging data to predict response to edaravone treatment in ALS, 77 pulse corticosteroid treatment in chronic inflammatory demyelinating polyradiculoneuropathy (CIDP) 136 and intravenous immunoglobulin (IVIG) in inflammatory myopathies, 78 as well as to repurpose drugs for neuromuscular diseases. 50 Such research tools require validation in real-world settings to confirm their accuracies across racially heterogenous populations in different clinical centers, before they can be appropriately deployed to support clinicians to select treatments based on anticipated clinical response.
For diseases like ALS for which there is no cure and current therapies can only modestly reduce the rate of disease progression, the focus is on managing expectations and maintaining quality of life for patients and their caregivers. 79 ML models have been used to identify predictors of quality of life and caregiver burden in ALS1,80 in order to personalize support for patients and their caregivers. Using supervised ML, a CDSS was designed to notify clinicians when an individual with ALS is experiencing low quality of life so that support measures can be extended to them. 81 Supervised ML also suggested that implementing telehealth-based interventions can help with caregiver burden. 80
ML may be used to classify patients in new ways and shed light on underlying molecular mechanisms of neuromuscular diseases
The way clinicians classify patient subtypes based on clinical findings and diagnostic tests may not meaningfully identify patient subgroups and instead represent human constructs applied to the data based on empirical observations. ALS patients may be classified as fast progressors or slow progressors based on their clinical phenotype, however among the fast progressors there may still be a number of different biological processes underlying pathology, which brings biological heterogeneity to patients of the same clinical phenotype. 134 In an unbiased manner, patients could instead be grouped by affected biological pathways followed by clinical phenotyping of the biological groups to correlate molecular mechanisms with disease progression and identify patients most likely to benefit from drugs that target those affected biological pathways. 82 The ability to determine the correct number and nature of subgroups would aid in understanding the disease and support clinical care and clinical trial design.
Many newly designed therapeutic strategies undergoing clinical trials, such as antisense oligonucleotides (ASO), target pathology associated with specific genetic causes, such as SOD1 and C9orf72 ALS.83,84 In one study, unsupervised ML was used to identify ALS subtypes in deeply-phenotyped, population-based collections of patients from Italy, then supervised ML was used to build predictor models that could accurately classify individual patients. 24 Although the ML models were robust with usage of independent development and validation datasets, and the clinical parameters used were standard across the ALS field, all data originated from the Northern Italian population, and studies in other countries are required to test the models’ generalizability.
While identifying specific genetic-phenotype correlations identifies druggable targets for specific groups of patients and responsive genetic subtypes of patients from clinical trials, identification of common alterations across the ALS spectrum improves understanding of the mechanisms underlying common neurodegenerative pathways and opens new therapeutic scenarios for large portions of patients. In one study, ML was used to integrate layers of biological information from transcriptomics and deep-sequencing analysis. ML was used to analyze whole blood and spinal cord transcriptomes of ALS patients to identify top predictor gene sets, as well as data from whole genome sequencing of samples from the AnswerALS database to identify signatures that could discriminate between ALS and controls samples. 85 This study showed that ALS significantly correlates with ageing and DNA damage signatures and provides insights into how different genetic mutations and divergent molecular mechanisms can converge over time into a singular presentation of ALS.
Machine learning may be used for drug discovery
ML may be used in combination with computational methods in high-throughput drug screening (HTDS) to test large libraries of compounds quickly and at low cost and aid lead compound optimisation.86,87 Using computational biology, millions of drugs can be tested for a predicted effect, such as binding a particular molecular target, and for other virtual parameters such as central nervous system penetrance, off-target effects, and pharmacokinetics/pharmacodynamics. One such study screened 1.5 million compounds for binding SOD1 in a way that can stabilize its dimer form, which is postulated to be neuroprotective in familial ALS, and found fifteen possible leads which also prevented SOD1 aggregation in lab-based assays. 88 To reduce off-target effects, the same group screened another 2.2 million compounds which could dock specific SOD1 sites with limited off-target binding. 89 ML approaches like Alphafold (AF), a 3D geometry modelling algorithm, can predict 3D protein confirmation at high accuracies and precisely estimate protein interactions. 90 Supervised and unsupervised ML has been used to combine -omics data analysis to predict drug properties, such as with PandaOmics, a cloud-based software platform that applies ML to multimodal -omics for therapeutic target and biomarker discovery. 91 PandaOmics has been applied on expression patterns of central nervous system samples and patient stem-cell derived motor neurons to discover novel therapeutic targets. 91 The acceleration of drug discovery and development by PandaOmics can be seen in the development of a new ALS drug which underwent AI-assisted drug discovery, FB1006. 92 It took less than two years from target identification to the completion of investigated-initiated clinical trial enrolment. Other AI-driven ways to improve drug discovery and development include incorporating ML in molecular simulations, de novo drug design, drug repurposing, prediction of drug-target interactions 93 and synthesis of new molecules (synthesis pathway generator). 94
Pitfalls of machine learning
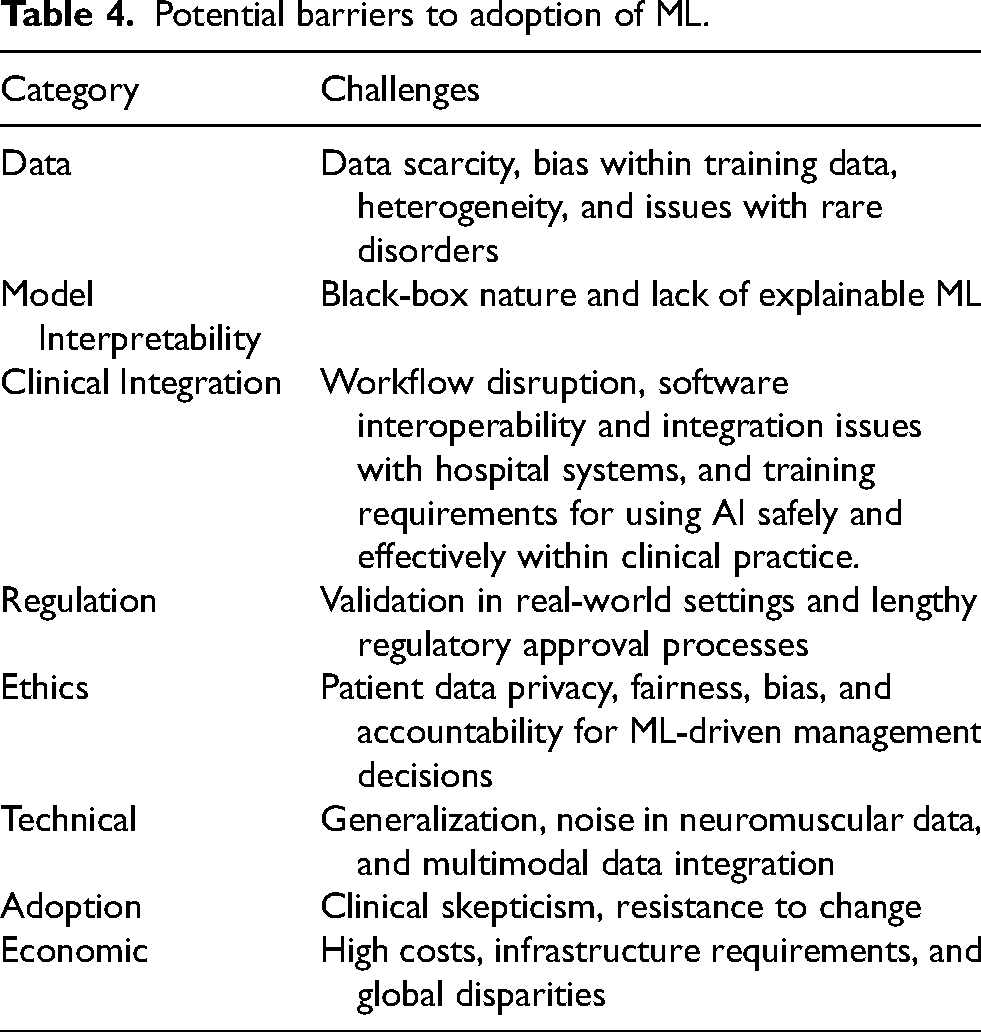
A summary of potential barriers to the adoption of machine learning is in Table 4. Of these, some points will be emphasized. To “first do no harm”, clinicians need to be aware of challenges which may limit their usage of ML, and implications on medical malpractice liability. Some ML models supply recommendations without directly explaining the underlying reasons for those results because of the “black box” of parameters which the model automatically derives and uses. 95 The opacity of “black box” medicine has implications on medical malpractice liability – how would liability apply to clinicians who are unable to understand the underlying mechanisms of the ML algorithms which recommend patient treatments? 96 Should they themselves understand how ML algorithms are developed and verified before using them, or is it sufficient to rely on the assurances of the technology developer? As it would not be practical to expect clinicians to be experts in ML, this primer serves to educate clinicians in the basics of ML so that they are equipped to evaluate the expertise of the developer.
Potential barriers to adoption of ML.
There are ways to make ML models more transparent and allow clinicians to understand how inputs influence predictions, such as by highlighting the region in a medical image the model uses to make predictions, so it is apparent what the predictions are based on. Such “interpretable AI” models can make predictions understandable and traceable and allow the clinician experts to manually modify mistaken concepts in the ML model’s decision-making pipeline. This can enhance clinical adoption, regulatory compliance, and informed decision-making. 97
With big data, there are risks of publishing false positive research findings, 98 particularly where studies conducted in a field are smaller - as might be the case in rare neuromuscular disorders, or when effect sizes are smaller – such as with heterogenous clinical populations in ALS, and when there is a greater number of tested relationships – for example using ML to test many parameters in a small number of patients. Clinicians need to be aware that while ML can accelerate the generation of hypotheses from multi-omics data, any single study merely provides a partial picture which can only be properly understood in the context of more testing outside this study and in the relevant field.
Overfitting (explained previously) and training on incomplete data may limit generalizability to the appropriate patient population. Data quality (explained previously) is critical for the performance, generalization, and trustworthiness of machine learning models, as poor-quality data can lead to inaccurate predictions and inefficiencies. Tools such as PROBAST can be used to assess the risk of bias and diagnostic or prognostic prediction model studies. 99
Risk mitigation
Clinicians need to know how to evaluate an ML model for clinical practice and research.
First, there should be independent comparison to an appropriate reference or gold standard. Gold standard is usually expert opinion on the diagnosis or prognosis of disease, or gold standard diagnostic tests.
Second, data should be from patients to whom the diagnostic test will be applied in clinical practice. If data is derived only from clinical trials, for example the ProACT database which amalgamates clinical data from ALS clinical trials,62,77,100,134 then this dataset may be biased and therefore should undergo validation in the general clinic patient population.24,56,59,81,133
Third, data from the training and validation sets should be independent from the testing set. As in all studies, the methods for obtaining the datasets and the procedure of analysis should be described in sufficient detail to be reproduced.
Fourth, results should be reported in the form of performance metrics as described in Table 1.
Fifth, the ML results should be repeatable and reproducible in the patient population of interest, and factors that may affect reproducibility in different institutions with different hardware and clinical practitioners should be considered.
Importantly, the clinician needs to be aware of institutional mechanisms and international guidelines on assessing validity of ML models and making direct comparisons of ML models.
Ethical use of ML models
ML models may amplify societal biases and exacerbate healthcare inequalities by underperforming in groups that are already disadvantaged by factors such as race, gender and socioeconomic background. 101 Patient privacy is also a concern, as recent public–private partnerships that collaborate on AI may have poorly protected privacy, and highly sophisticated algorithmic systems may cause data breaches even in anonymized datasets. 102
The international agreement on key principles for AI in healthcare currently include control of bias, explainability, transparency, systems of oversight and validation, and is reflected in European Union (EU) 103 and US regulatory frameworks. 104 Developers of AI technologies in healthcare should provide evidence of safety and validation which is approved by regulators.
Regulatory considerations
Regulators are rarely popular when there is a new technology that does not quite fit into existing categories. Some authors advocate regulatory guidelines on how to safely implement and assess AI, and understand the specific capabilities and limitations of its medical use. 105 However, it is more true to say that regulators such as the U.S. Food and Drug Administration (FDA) have the responsibility to ensure high standards of therapeutics and medical devices in the interest of projecting public health and safety 106 and while the FDA does produce guidelines, they are typically to explain their current thinking to guide those applying to them for marketing approval. Guidelines on how to implement approved products is not their responsibility.
The FDA classifies AI software as “Clinical Decision Support (CDS) software. If a CDS software is classified as a medical device, it requires FDA marketing approval. However, a CDS software may be excluded from their definition of a medical device and may be classified as a Non-Device CDS if it meets the following criteria:
104
The software function does NOT acquire, process, or analyze medical images, signals, or patterns. The software function displays, analyzes, or prints medical information normally communicated between health care professionals (HCPs). The software function provides recommendations (information/options) to a HCP rather than provide a specific output or directive. The software function provides the basis of the recommendations so that the HCP does not rely primarily on any recommendations to make a decision.
Any software which provides a diagnosis or categorizes a patient's risk based on processing clinical information is likely to be classified as a medical device. Despite this stance, the FDA has approved over 950 medical devices with artificial intelligence features between 1995 and the end of 2024.
107
This however has been inadequate for many industry developers who feel it is inappropriate for AI software to be considered with the same rigid parameters as a traditional medical device. Additionally, contention has arisen around requirements that software be unchanged following approval – something which is reasonable for most hardware and software devices but would prevent an AI system from adjusting its underlying machine learning model weights to adapt to local conditions. Altering the ML model weights will result in inconsistent outputs, and it is not unreasonable to expect that a device produces the same outcome from the same inputs consistently.
In response to this, in June 2024, the FDA published some “guiding principles” for ML-enabled medical devices intended to support the development of “safe, effective and high-quality artificial intelligence/machine learning technologies that can learn from real-world use and, in some cases, improve device performance” 108 intended to address concerns that AI-enabled software did not fit well into the existing paradigm of medical device approval. This was followed by guidance in December 2024 which indicated that for software where modifications to the AI model are implemented automatically by software, also known as “continuous learning”, manufacturers can use a Predetermined Change Control Plan (PCCP) to prospectively specify and seek premarket authorisation for modifications to an AI-enabled device software function (AI-DSF). 109 This appears to address prior criticisms that AI software that continues to learn or adjust its ML model weights would require repeated FDA submissions and approvals. The use of a PCCP appears to allow modifications to be implemented to an AI-DSF without triggering the need for a new marketing submission. Premarket authorization for an AI-DSF with a PCCP must be established through the either the PMA pathway, 510(k) pathway, or De Novo pathway, as appropriate, as a PCCP must be reviewed and established as part of a marketing authorization for a device prior to a manufacturer implementing any modifications under that PCCP.
A draft guidance issued the next month on January 7, 2025 entitled “Artificial Intelligence-Enabled Device Software Functions: Lifecycle 3 Management and Marketing Submission Recommendations” 110 requests vendors provide detailed information about the technical characteristics of the underlying AI model(s) themselves and the algorithms and methods that were used in their development. This includes optimization methods, training paradigms (e.g., supervised, unsupervised or semi-supervised learning, federated learning, active learning); regularization techniques employed; training hyperparameters; and summary training performance.
The FDA has emphasized the importance of transparency in AI model training and operations, and the need to identify and reduce bias in AI models. Bias is defined as “a potential tendency to produce incorrect results in a systematic, but sometimes unforeseeable way, which can impact safety and effectiveness of the device within all or a subset of the intended use population (e.g., different healthcare settings, different input devices, sex, age, etc.,). 110 To guard against bias, the FDA recommends that companies “addressing representativeness in data collection for development, testing, and monitoring throughout the product lifecycle, as well as evaluating performance across subgroups of intended use.”. This will entail collecting evidence to evaluate whether a device benefits all relevant demographic groups similarly.
These publications issued in the final days of the FDA under the Biden Administration, and the discussions with industry that surround it, indicate a desire by the current FDA to develop guidance and resources for a total product life cycle approach to the oversight of AI-enabled devices. Stakeholders will naturally be observing carefully how this regulatory position develops under the new administration.
Future trends
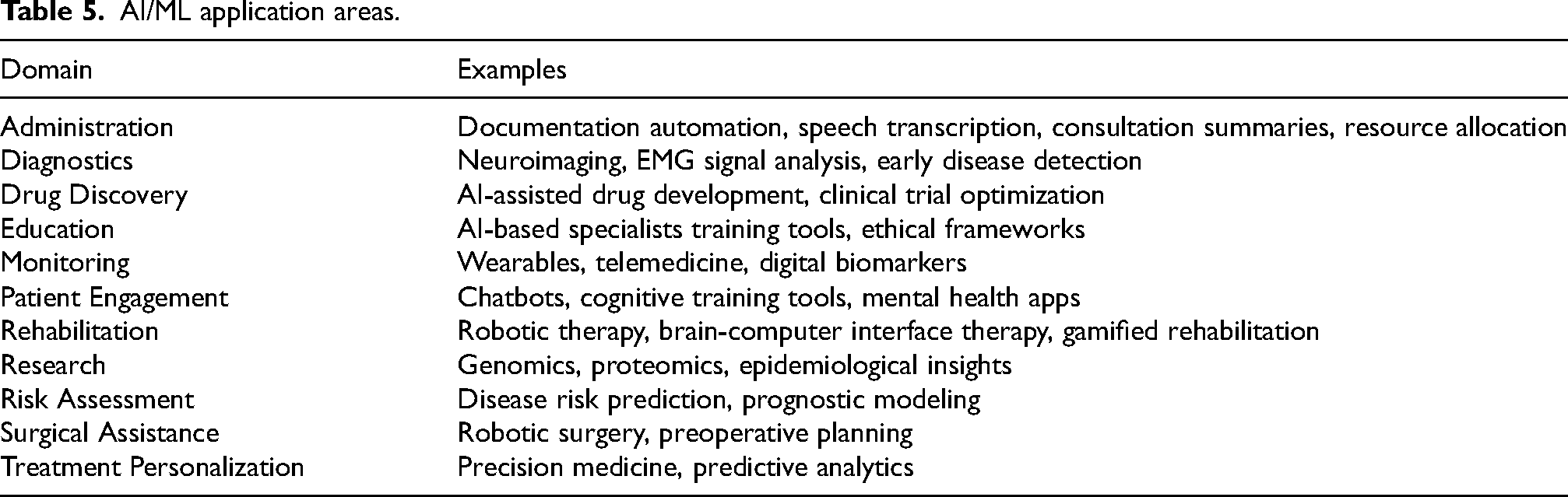
In an ideal future, ML will impact positively upon a neuromuscular clinician’s practice, enhancing diagnostics, improving treatment, and driving research among other application areas summarized in Table 5. What is more likely though is that change will be led by the progression of AI in consumer technology and neurology is at risk of lagging behind other specialties which have historically been faster to embrace technology and devices, such as radiology and surgery.
AI/ML application areas.
There are some obvious trends, such as the impact of generative AI. Large language models like ChatGPT and Foundation Models have already pervaded consumer technology. It is likely that agentic workflows, AI which operates mostly without supervision to execute a pre-defined role or function, will play a role in clinical workplaces of the future, for tasks which do not require FDA medical device approval such as document summarization, research topic monitoring and summarization, drug-drug interaction alerts based on literature mining, and patient education.
In the introduction, the known shortcomings of current generative AI, in particular its lack of true understanding of material and tendency to confabulate (“hallucinations”) were mentioned. AI-driven CDSS will benefit significantly from the introduction of logic and reasoning, and causality. 111 Discussing the research in automated reasoning112,113 is out of scope of this primer but is essential for AI to move to the next stage of evolution. Today, cloud-computing vendors such as Amazon are offering automated reasoning checks as a service to prevent factual errors from LLM hallucinations. 114 Error checking such as this may become mandatory for clinical software in the future.
With respect to early disease identification and tracking, perhaps ambient AI models, 115 which use information from contactless sensors in physical spaces such as the clinic or at home, can be used to analyze voice recordings, physical movement, and gait, and become approved for use in tracking clinical progression of motor neuron diseases such as ALS and SMA, or chronic muscular dystrophies, at the clinic or at home. Similarly, ambient AI models may be used to monitor and prevent impending respiratory failure of inpatients with myasthenia gravis or CIDP in the general wards and high dependency unit.
CDDS based on multi-modal ML models with neuromuscular ultrasound or MRI imaging and NCS/EMG data may become commonplace and automate routine diagnoses, screen for urgent conditions, distinguish between neuromuscular diseases and mimickers, anticipate progression, and predict response to therapies like corticosteroids or biologics.
Collaboration between clinicians and data scientists would be key to making ML useful for healthcare and medical research. Standardization and open sourcing of ML models by college boards and tertiary referral centers, and sharing of patient databases, could provide low-resource hospitals the means to improve the quality of their care.
Recent FDA guidance documents indicate an understanding of the significance of AI in clinical decision support but also a desire for transparency in how AI is trained and acute awareness of the risks of bias. It is likely there will be a strong preference in the clinic for ML models where output predictions are explainable and trustworthy. This may necessitate modifications to the training process or more detailed documentation in how training is conducted, and bias is detected and managed. Safety features to detect model performance drift and degradation are likely to also be necessary. A future where AI are also required to maintain continuing education certification is certainly plausible.
The use of cloud-based AI services is associated with data privacy and data transmission latency issues. The future will most likely see institutions use a mix of cloud-based AI, and AI developed to run using only local computing resources, so-called ‘inferencing at the edge’.83,84 Inferencing at the edge will allow hospitals and clinics to incorporate ML/AI tools into their workflow while still maintaining compliance with best practices in patient data privacy and data security.
Conclusion
ML underlies applications and devices which are assisting clinicians in the diagnosis, monitoring, formulation of prognosis, and treatment of patients with a spectrum of neuromuscular diseases. The prevalence of this will only grow further. However, how future practitioners of clinical medicine will look back upon this period of transition will be determined not by how much changed or by how fast clinicians embraced this change but by how much patient outcomes were improved and how safeguards are applied to prevent harm to patients when using these new technologies. These are key metrices which differentiate clinical practice from the other industries. For this to occur the same degree of rigor must be applied to AI as it is to any new clinical practice, and this must be done with understanding of the impact but also the underlying science. Clinicians should be able to assess the validity and impact of AI applications just as they do for other diagnostic or prognostic tools, and to do this requires some degree of understanding of machine learning processes. Understanding the relevance to clinical practice and research in neuromuscular and electrodiagnostic medicine alone may not be sufficient.
The history of medicine has a number of examples where new technologies, rules and tools actually have - looking back – have not improved patient outcomes but even caused harm. Concerns about AI in general, which have already manifest in some form include AI-driven automation resulting in loss of jobs, the spread of fake news, invasion of privacy, and AI-powered weaponry. In healthcare, failures in medical AI – breaches in patient confidentiality, erroneous medical evaluations, racial bias due to flawed training and limitations of current machine learning, could erode public trust in healthcare. Both technical and humanistic challenges regarding the social and ethical implications of AI must be discussed at each stage of clinical trial and implementation.
The tech industry grapples with ethical issues only when forced to and is unlikely to ever accept responsibility for the negative impacts of AI or the casualties or improperly-used AI. It therefore falls to the clinician to ensure their understanding of machine learning is of sufficient breadth and depth to be able to assess the specific challenges and ethics surrounding the medical use of these powerful technologies, in a fair and balanced manner.
Supplemental Material
sj-docx-1-jnd-10.1177_22143602251329240 - Supplemental material for A neuromuscular clinician’s primer on machine learning
Supplemental material, sj-docx-1-jnd-10.1177_22143602251329240 for A neuromuscular clinician’s primer on machine learning by Crystal Jing Jing Yeo, Savitha Ramasamy, F Joel Leong, Sonakshi Nag and Zachary Simmons in Journal of Neuromuscular Diseases
Footnotes
Funding
The authors report no financial disclosures and conflicts of interest. We confirm that we have read the Journal’s position on issues involved in ethical publication and affirm that this report is consistent with those guidelines.
Supplemental material
Supplemental material for this article is available online.
References
 . In:
. In: Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.