Abstract
Extraction of DNA using magnetic bead-based techniques on automated DNA extraction instruments provides a fast, reliable, and reproducible method for DNA extraction from various matrices. Here, we have compared the yield and quality of DNA extracted from FTA cards using four automated extraction protocols on three different instruments. The extraction processes were repeated up to six times with the same pieces of FTA cards. The sample material on the FTA cards was either blood or buccal cells. With the QIAamp DNA Investigator and QIAsymphony DNA Investigator kits, it was possible to extract DNA from the FTA cards in all six rounds of extractions in sufficient amount and quality to obtain complete short tandem repeat (STR) profiles on a QIAcube and a QIAsymphony SP. With the PrepFiler Express kit, almost all the extractable DNA was extracted in the first two rounds of extractions. Furthermore, we demonstrated that it was possible to successfully extract sufficient DNA for STR profiling from previously processed FTA card pieces that had been stored at 4 °C for up to 1 year. This showed that rare or precious FTA card samples may be saved for future analyses even though some DNA was already extracted from the FTA cards.
Introduction
Biological material (blood or buccal cells) for forensic genetic analyses of reference samples may be collected on FTA cards. FTA cards are impregnated with chaotropic agents that inhibit infectious agents and reduce the potential biohazards of the sample, reducing if not eliminating potential hazards to the technical staff during sample processing.1,2 This enables storage of biological material on FTA cards at room temperature for extended periods. No refrigerators or freezers are required, which significantly reduces storage costs. Furthermore, shipment of samples is simplified as this may be done by traditional mail. 1
The use of short tandem repeats (STRs) is common to the majority of all forensic genetic analytical methods performed today.3–5 STR typing of biological material (e.g., blood, buccal cells) stored on FTA cards is performed by isolation of a small disk of the FTA card (e.g., by an automated puncher).6,7 Amplification of STR markers is subsequently performed on extracted DNA or directly on the FTA punch.8–10
Storage and extraction of DNA from FTA cards are fast and reliable. However, the usually used extraction methods fail to release the entire amount of DNA bound to the FTA card. Occasionally, the routine operation may fail to generate a full STR profile. 11 This may be due to too little biological material or a failure in the laboratory process (e.g., to extract sufficient amounts of DNA). Samples resulting in STR profiles with weak signals or no signal are usually reprocessed using 2 to 10 times the initial amount of the FTA card. Alternatively, DNA may be extracted from a larger part of the FTA card using automated protocols.11,12
We compared the efficiency of four automated protocols using two sample types for extraction of DNA from FTA cards. To enable comparisons between the different extraction protocols, all protocols and sample types were evaluated using eight different samples for each sample type and protocol. Following initial sample pretreatment and DNA extraction, the processed FTA card pieces were retained in the sample tubes and subjected to up to six additional DNA extractions ( Fig. 1 ) or stored for 1 year. STR typing was used to evaluate the quality of the extracted DNA.
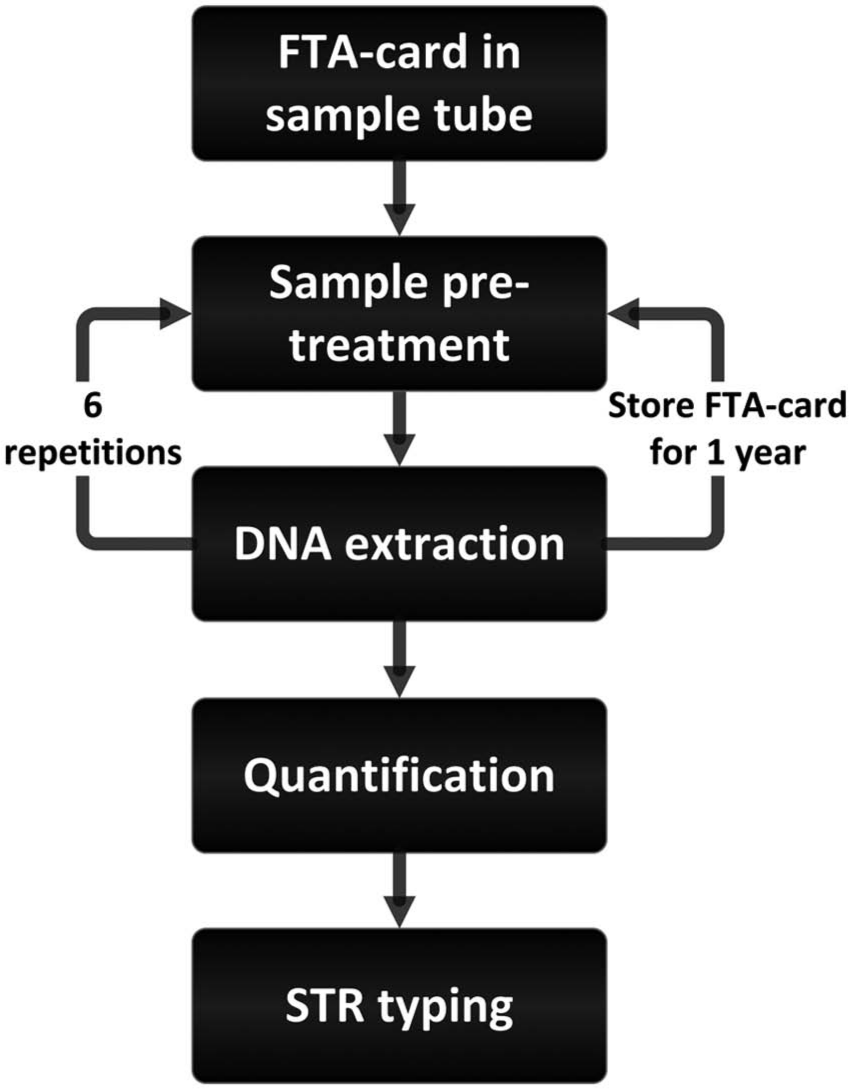
Extraction scheme. STR, short tandem repeat.
Materials and Methods
Samples
Buccal cells were isolated from the inside of the cheeks with a sterile foam-tipped applicator (GE Healthcare, Little Chalfont, England) and applied to indicating FTA cards (GE Healthcare) by pressing and rubbing each side of the foam onto the FTA membrane for 20 s. Blood samples were collected by venipuncture using tubes containing EDTA as an anticoagulant. From randomly selected samples, 200 µL blood was isolated with a manual pipette and transferred to FTA cards. Fingertip blood was isolated by puncturing a finger with a 1.75-mm Tenderlett finger incision device (ITC, Edison, NJ) and applying a couple of drops of blood onto an FTA card. The FTA cards were allowed to dry overnight at room temperature prior to further processing. For all extractions, a 1-cm2 piece of the FTA card containing biological material was manually isolated with a sterile scalpel (Swann-Morton Ltd, Sheffield, England).
DNA Extraction Using QIAamp DNA Investigator (QIAamp)
A total of 280 µL ATL buffer and 20 µL Proteinase K were added to each 1.5-mL sample tube (Eppendorf, Hamburg, Germany) containing an FTA card piece. Both reagents were part of the QIAamp DNA Investigator kit (Qiagen, Hilden, Germany). 13 Following gentle mixing, sample tubes were incubated for 1 h at 56 °C in a VorTemp 56 (Labnet International, Inc., Woodbridge, NJ) shaking incubator at 750 rpm. The supernatants were manually transferred to new sample tubes and DNA was extracted using a QIAcube (Qiagen) 14 and the DNA_QIAampDNAInvestigator_FTAAndGuthrieCards_BloodSpotLysisAndPurification_V2 protocol provided with the instrument. Extracted DNA was eluted in 50 µL water. Sample tubes containing the FTA card pieces were subjected to additional rounds of lyses by addition of ATL buffer and Proteinase K followed by DNA extraction as outlined in Figure 1 .
DNA Extraction with QIAsymphony SP with Qiagen Pretreatment (Q QSP)
FTA card pieces containing either buccal or blood material were transferred to 2-mL sample tubes (Eppendorf) and pretreated using the Qiagen-recommended pretreatment protocol. To each sample tube, 180 µL buffer ATL and 20 µL Proteinase K were added. Both reagents were part of the QIAsymphony DNA Investigator kit (Qiagen). 15 Following vortexing on an IKA Vibrax VXR shaker (IKA Works), the samples were incubated for 15 min at 56 °C in shaking incubators (VorTemp 56). Using a manual plunger operated pipette, the lysate was transferred to 2-mL sample tubes (Sarstedt, Numbrecht, Germany) and loaded onto the QIAsymphony SP (Qiagen). 16 DNA extraction was performed using the protocol ACS_REF_200 provided with the instrument. All samples were eluted in 100 µL MilliQ water in semi-skirted 96-well twin.tec plates (Eppendorf). Following the initial extraction, the sample tubes containing the FTA card pieces were subjected to a new round of the pretreatment and extraction procedure ( Fig. 1 ). The sample pretreatment and DNA extraction procedure was repeated up to a total of six times for each type of biological material. The elution plate was sealed with an aluminum seal (PCR-AS-200; Axygen Scientific, Inc., Union City, CA) to prevent evaporation of the eluted DNA and stored at −20°C until quantification and STR typing.
After six pretreatments and extractions with the QSP buccal protocol, 1.2-mm FTA punches from the eight processed FTA cards pieces were subjected to solid-phase PCR. Following manual transfer of the 1.2-mm FTA punches to 200-µL PCR tubes, 10 µL PCR master mix was added to each tube.
DNA Extraction with QIAsymphony SP and In-House Optimized Pretreatment (QSP)
The pretreatment protocol recommended by Qiagen was compared with an in-house optimized protocol. A total of 190 µL 0.5 × ATL buffer and 10 µL Proteinase K were added to the sample tubes containing FTA card pieces containing buccal, blood, or finger blood material and incubated for 15 min at 56 °C in a heating block (Techne Dri-Block; DB-3D, Duxford, England) interrupted by manual vortexing every 5 min followed by 5 min at 95 °C in another heating block. After vortexing, sample tubes were loaded onto the QIAsymphony SP, leaving the pieces of FTA cards in the tubes. Following the initial extraction, the sample tubes containing the FTA card piece were subjected to a new round of the pretreatment and extraction procedure ( Fig. 1 ) or stored for 1 year at 4 °C. Extraction and elution conditions were as described above.
DNA Extraction Using PrepFiler Express on AutoMate Express (PrepFiler)
FTA card pieces containing either buccal or blood material were placed in the LySep columns placed on top of a lysate collection tube included in the PrepFiler Express Forensic DNA Extraction Kit (Applied Biosystems [AB], Foster City, CA) 17 with sterile tweezers. To each sample, 500 µL freshly prepared lysis solution consisting of 500 µL PrepFiler Lysis Buffer included in the PrepFiler Express Forensic DNA Extraction Kit and 5 µL freshly prepared 1M dithiothreitol (Sigma-Aldrich, St. Louis, MO) were added. Following incubation of the LySep columns for 40 min at 70 °C in a shaking incubator (VorTemp 56) at 750 rpm, the columns were centrifuged for 2 min at 10,300 relative centrifugal force (RCF) in a Biofuge Pico bench-top centrifuge (Heraeus Instruments, Hanau, Germany). Following removal of the LySep column, the lysate collection tube was placed in the AutoMate Express instrument (AB) 18 and extracted immediately using the PF Express program provided with the instrument. The processed FTA card pieces were manually transferred using a pair of sterile tweezers to new LySep columns for the next rounds of extractions as outlined in Figure 1 . DNA extracts (50 µL) were stored at −20°C until quantification.
DNA Quantification
The extracted DNA was quantified using the Quantifiler Human DNA Quantification Kit (AB). 19 Quantification was performed on a 7900HT sequence detection system (AB) according to the manufacturer’s recommendations. All samples were quantified in triplicate.
PCR Amplification
PCR master mix was prepared according to the manufacturer’s instructions with modifications: For the AmpF
Direct amplification of 1.2-mm FTA card punches that were manually isolated using a hand-operated puncher (Harris; GE Healthcare) was performed on selected FTA card pieces that had been subjected to six rounds of extraction using the Q QSP protocol. Following isolation, the FTA card punches were transferred to 200-µL PCR tubes (AB) containing 10 µL PCR master mix and amplified. PCR conditions were as described above.
Capillary Electrophoresis and Data Analysis
For analysis of the STR amplicons, 1.5 µL of the PCR products was combined with 15 µL formamide (AB) and LIZ 500 size standards (AB) in a 96-well electrophoresis plate (Axygen Scientific) on a Biomek 3000 laboratory automated workstation (BC) using in-house developed methods. 21 Capillary electrophoresis was performed on ABI 3130xL Genetic Analyzers (AB).
Multicolor fluorescence capillary electrophoresis results were analyzed using Genescan Analysis version 3.7 (AB). Allele calls were done using Genotyper version 3.7 (AB) macros.
Results and Discussion
Yield
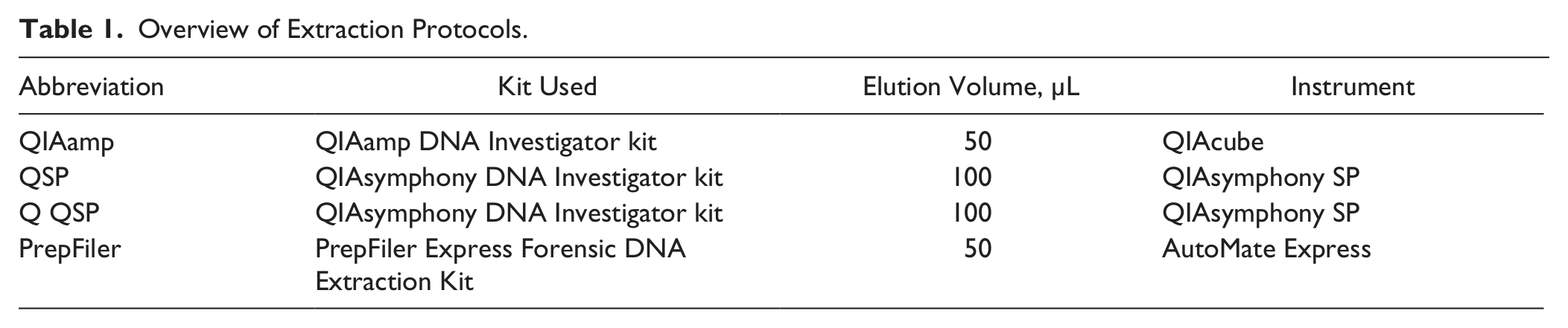
Four different automated DNA extraction protocols were evaluated using two different sample types applied onto FTA cards ( Table 1 ). Except for the Q QSP protocol, all protocols are provided with commercially available instruments and kits specially designed for use in forensic genetic laboratories. All instrument protocols are simple and fast to validate and implement in processing of samples for forensic genetic STR profiling. All instruments provide the laboratory with a flexible and standardized workflow that ensures reproducible and standardized processing of the samples. Following six extractions with each protocol, all DNA extracts (432 in total) were quantified in triplicates. The results from the eight different samples for each protocol were averaged ( Fig. 2 ). The results indicated that the entire amount of DNA bound to the FTA card was not released in the first extraction. This finding concerns both buccal and blood samples. Furthermore, there were significant differences between the extraction methods. The elution volume for both the QIAamp DNA Investigator kit and the PrepFiler Express kit was 50 µL, whereas the elution volume for extractions performed on the QIAsymphony was 100 µL. Hence, quantification results were presented in total yield to enable comparison between methods. The yields of DNA from the first extraction were compared with the combined yields from all six extractions ( Fig. 3 ). For blood samples, more DNA was extracted with the PrepFiler Express kit than with the QIAamp and QIAsymphony DNA Investigator kit, and most of the DNA was extracted in the first extraction. This may be due to a more effective lysis and/or release of bound DNA from the FTA card by either the lysis buffer in the PrepFiler Express kit or the use of LySep columns to isolate the lysate. For buccal samples, the kit resulting in the highest yield was the QIAamp DNA Investigator kit.
Overview of Extraction Protocols.

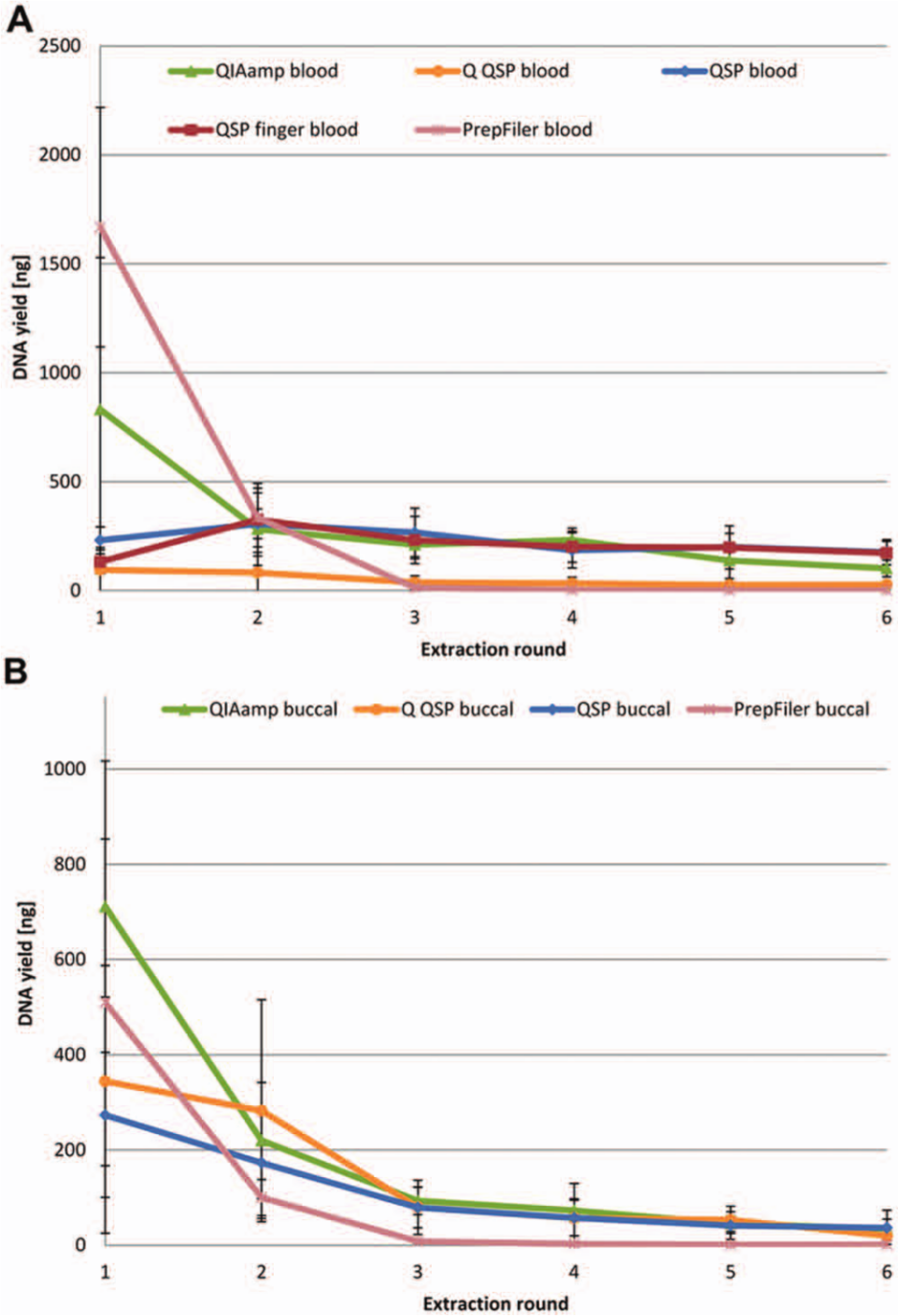
Quantification results. (
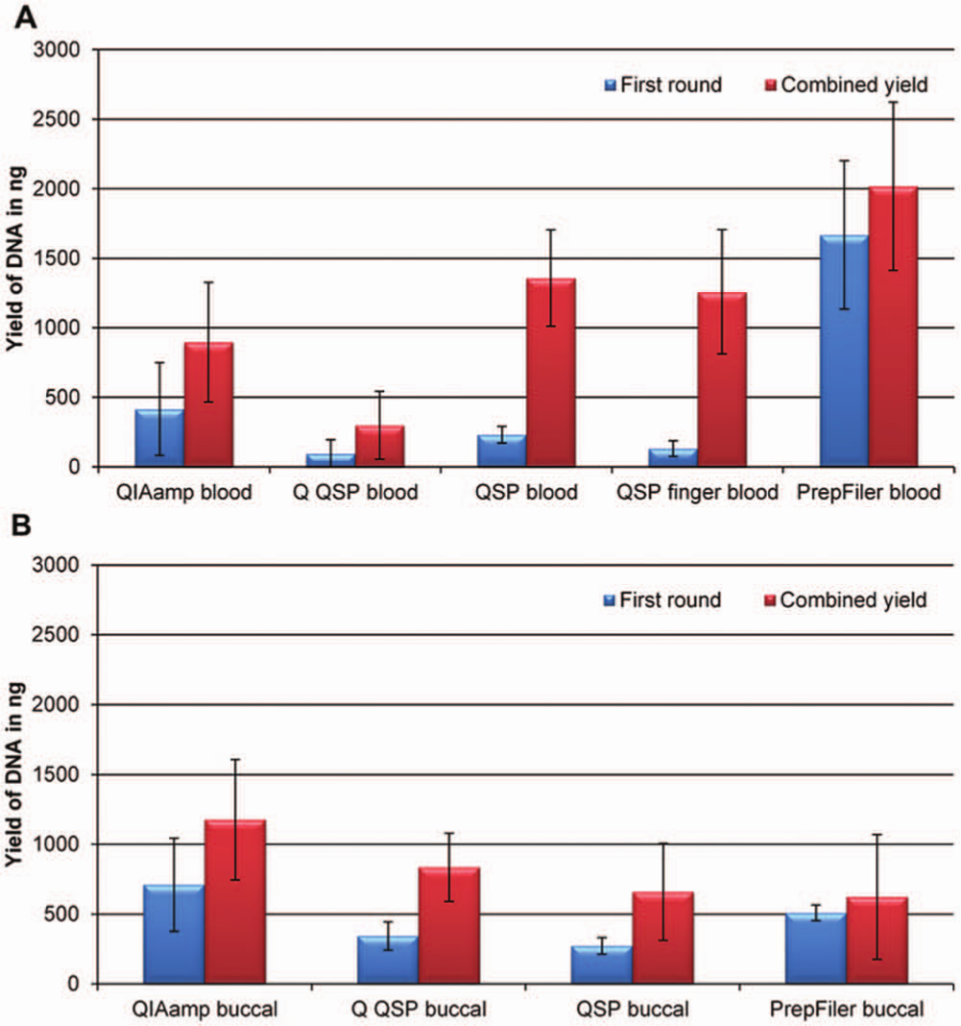
Comparison of the DNA yield from the first extraction with the overall yield of all six further extractions for the nine extraction protocols. Each extraction protocol was evaluated using eight samples for each round of extraction. All samples were quantified in triplicates. Error bars represent standard deviations. (
Blood samples extracted on the QIAsymphony using the QIAsymphony DNA Investigator kit displayed large differences between the yield of DNA in the first round of extraction and the combined yield ( Fig. 3 ). This indicated that the sample pretreatment or the extraction method may not be as efficient as the other methods. For blood samples, the pretreatment protocol recommended by Qiagen (Q QSP) seemed less efficient than the in-house optimized pretreatment method (QSP). For buccal samples, no significant difference between the pretreatment protocol recommended by Qiagen (Q QSP) and the in-house optimized protocol (QSP) was observed. Both anticoagulated blood (EDTA blood) and freshly drawn blood (finger blood) were included to investigate if EDTA treatment introduced positive or negative effects on the extraction yield and/or the quality of the extracted DNA. No difference was observed in terms of yield of DNA between the EDTA blood and freshly drawn finger blood ( Figs. 2 and 3 ).
Quality
All DNA extracts were amplified with the AmpF
Comparison of Number of Alleles in the Resulting STR Profiles following Multiple Rounds of Extraction and the Percentages of Full Profiles for Blood Samples.
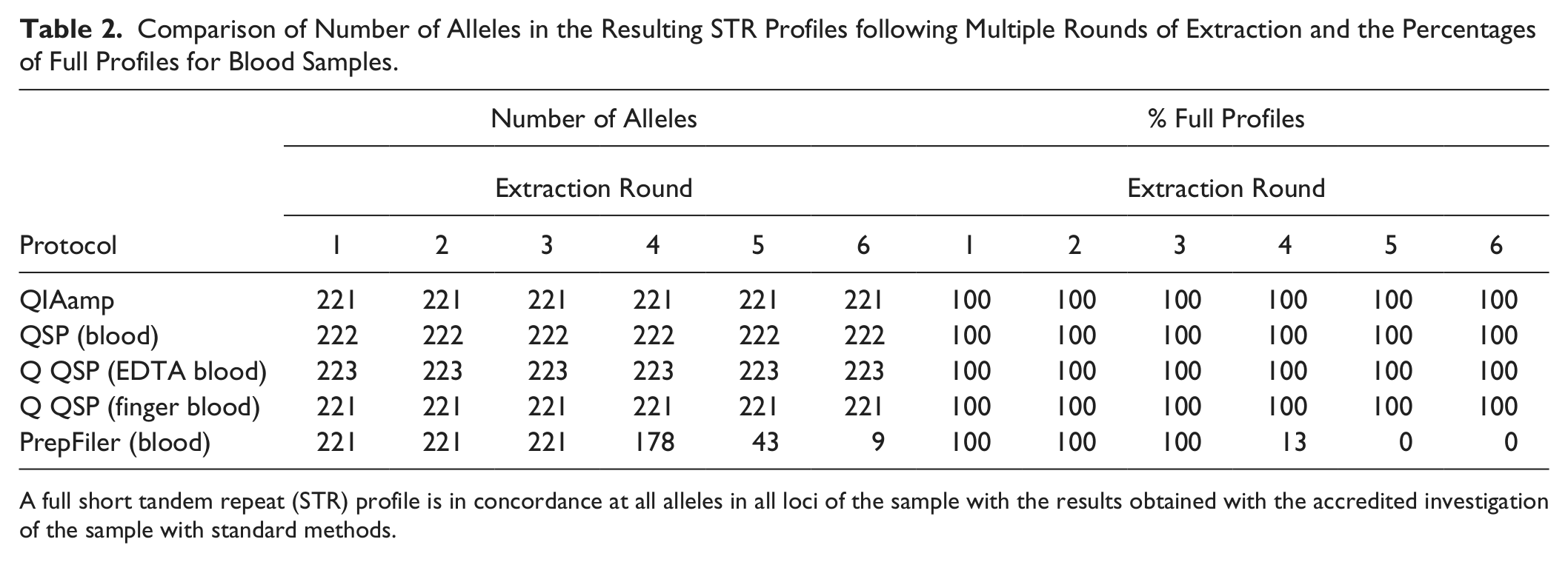
A full short tandem repeat (STR) profile is in concordance at all alleles in all loci of the sample with the results obtained with the accredited investigation of the sample with standard methods.
Comparison of Number of Alleles in the Resulting STR Profiles following Multiple Rounds of Extraction and the Percentages of Full Profiles for Buccal Samples.
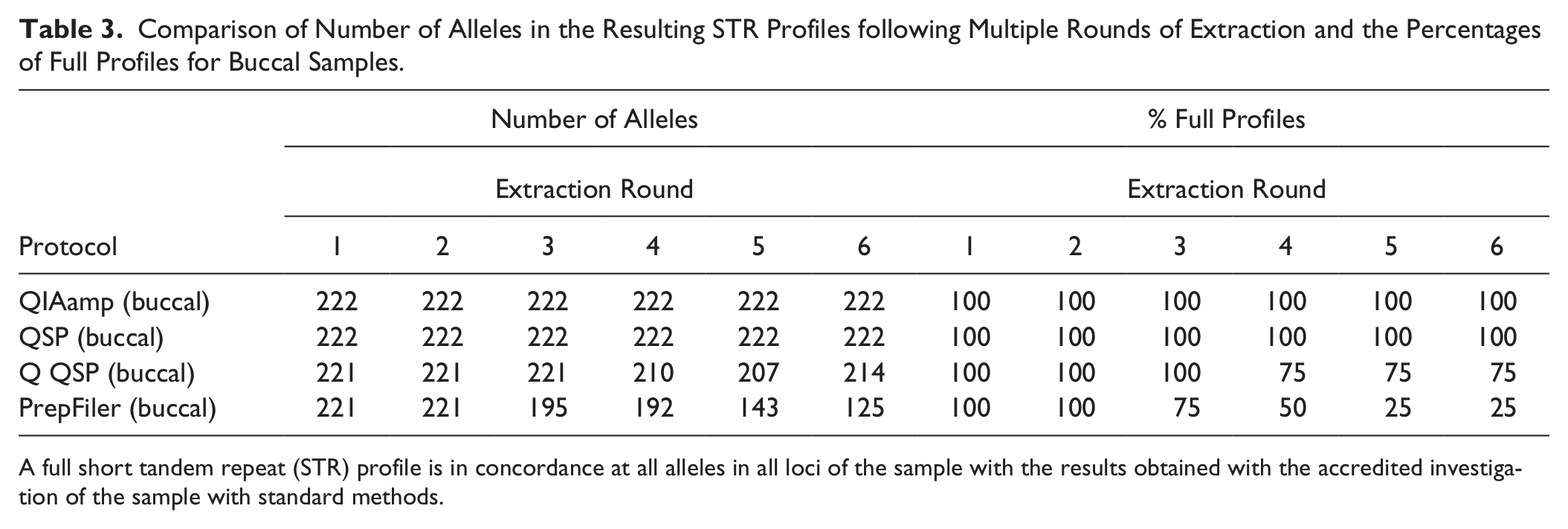
A full short tandem repeat (STR) profile is in concordance at all alleles in all loci of the sample with the results obtained with the accredited investigation of the sample with standard methods.
For samples subjected to solid-phase PCR, full STR profiles were obtained from out of the eight samples. One sample did not result in any signal. The remaining four samples gave partial profiles where 60% to 97% of the alleles were amplified. In total, 163 alleles out of the expected total of 221 alleles (74%) were obtained (data not shown).
Removal of the area containing the biological material from FTA cards effectively prevents reprocessing of the sample. However, as shown above, the FTA cards contained sufficient DNA for future investigations even though DNA was extracted from the card one to three times. DNA from eight FTA cards with blood samples was extracted once and the FTA pieces were left in the sample tube and stored for 1 year. A second extraction was performed and the DNA was amplified with the AmpF
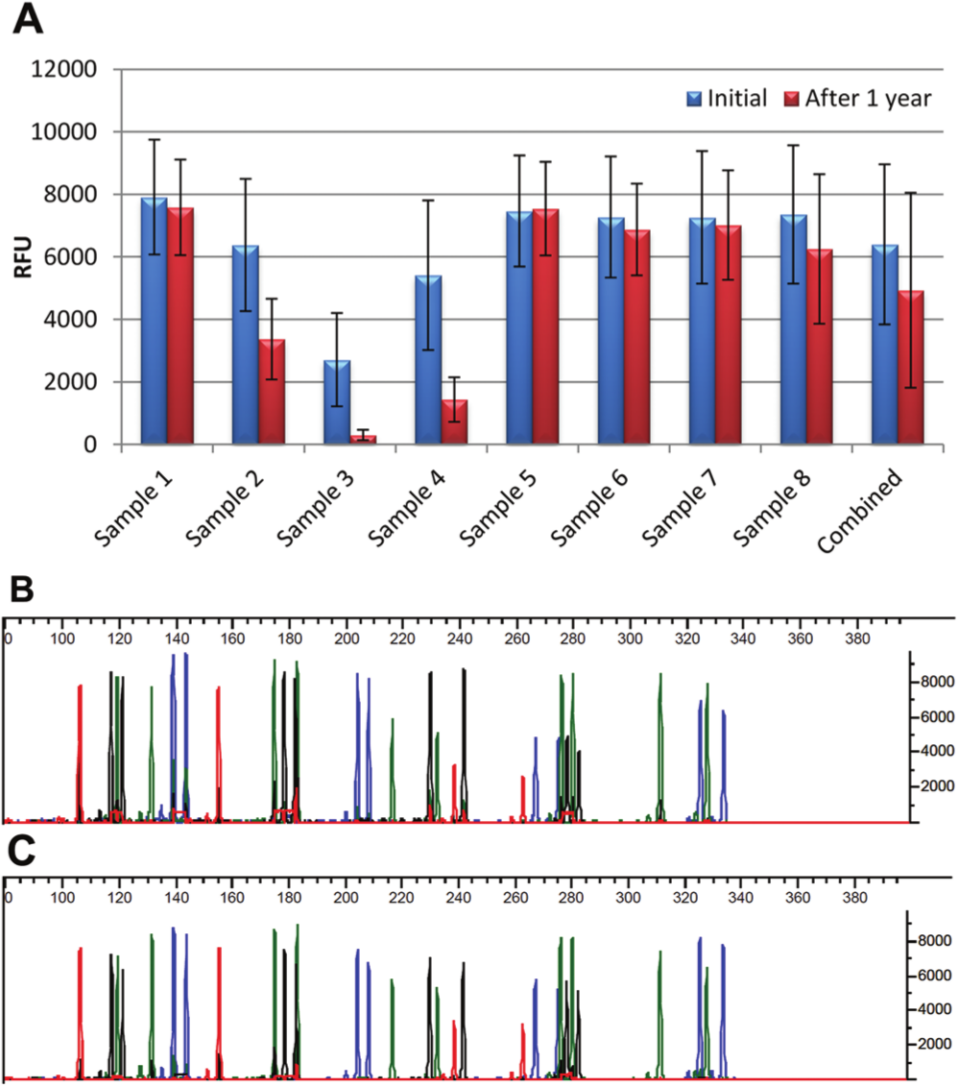
Comparison of peak heights from short tandem repeat (STR) profiles following extraction immediately after sample collection and after extraction of the same pieces of the FTA card containing EDTA blood after 1 year of storage. (
All three instruments provide an alternative to manual processing of samples for forensic genetic extraction of DNA for reference samples. The QIAsymphony provides the highest and most flexible throughput of the instruments tested, but the required manual sample preprocessing hampers in practice this throughput. The AutoMate Express with the PrepFiler Express kit extracts the largest proportion of the bound DNA in the first extraction. However, all protocols tested fail to release the entire amount of bound DNA in the first extraction. Optimization of the sample pretreatment protocols may be relevant if the amount of DNA released in the first extraction is to be increased.
Footnotes
Acknowledgements
We thank Anja L. Jørgensen, Signe A. Hansen, and Marianne Olesen for technical assistance.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
