Abstract
We have implemented and validated automated protocols for DNA extraction and PCR setup using a Tecan Freedom EVO liquid handler mounted with the Te-MagS magnetic separation device (Tecan, Mannedorf, Switzerland). The protocols were validated for accredited forensic genetic work according to ISO 17025 using the Qiagen MagAttract DNA Mini M48 kit (Qiagen GmbH, Hilden, Germany) from fresh whole blood and blood from deceased individuals. The workflow was simplified by returning the DNA extracts to the original tubes minimizing the risk of misplacing samples. The tubes that originally contained the samples were washed with MilliQ water before the return of the DNA extracts. The PCR was setup in 96-well microtiter plates. The methods were validated for the kits: AmpFSTR Identifier, SGM Plus and Yfiler (Applied Biosystems, Foster City, CA), GenePrint FFFL and PowerPlex Y (Promega, Madison, Wl). The automated protocols allowed for extraction and addition of PCR master mix of 96 samples within 3.5 h. In conclusion, we demonstrated that (1) DNA extraction with magnetic beads and (2) PCR setup for accredited, forensic genetic short tandem repeat typing can be implemented on a simple automated liquid handler leading to the reduction of manual work, and increased quality and throughput.
Keywords
Introduction
Automated extraction of DNA from blood or other types of biological material may be performed using a large variety of instruments and chemistries. Automated extraction of DNA for use in forensic genetic analysis of samples have been realized using either low-volume automated systems such as the Qiagen BioRobot EZl, 1 –4 the Promega Maxwell 16, 5 or medium- to high-throughput systems such as the Qiagen M48 BioRobot 6 or the QIAsymphony SP. 7 Customized solutions on commercially available automated liquid handlers such as the Tecan Genesis RSP 150/8 8,9 also have been described.
At the Section of Forensic Genetics, Department of Forensic Medicine at the University of Copenhagen, extraction of DNA from whole blood reference samples in criminal, paternity, and immigration cases was previously performed using a manual Chelex-based protocol. 10 To increase throughput and reduce the requirement for manual intervention, an automated solution for extraction of DNA from whole blood and subsequent automated PCR setup was sought.
We have developed and validated automated methods for a Tecan Freedom EVO 150/8 automated liquid handler (Tecan, Männedorf, Switzerland) according to the ISO 17025 standard for extraction and PCR amplification setup of 96 whole blood samples within 3.5 h. The methods are based on the M48 extraction kit (Qiagen GmbH, Hilden, Germany) and the AmpF
Experimental
Samples
Aliquots of blood were transferred from the original sample tubes to standardized sample tubes (1.5 mL, Eppendorf GmbH, Hamburg, Germany) at the day of arrival. For samples extracted using the manual Chelex protocol, 10,11 3 μL was used, whereas 6 μL was used for the automated DNA extraction method on the Tecan Freedom EVO.
Manual Chelex DNA Extraction Protocol
The manual Chelex DNA extraction protocol described previously 10,11 was used with modifications. In brief, one milliliter autoclaved MilliQ filtered water was added to each sample tube containing 3 μL blood, followed by mixing for 20 s on an IKA Vibrax VXR shaker (IKA Works, Inc., Wilmington, NC). After incubation for 30 min at room temperature, samples were mixed and centrifuged at 13,000 relative centrifugal force (RCF) for 5 min in a benchtop centrifuge (Eppendorf 5417 C). The supernatant was carefully decanted and discarded. To each sample, 200 μL of a 5% Chelex solution pH > 10.5 (Chelex 100 Resin, 100–200 mesh Na-form, BIO-RAD Laboratories, Richmond, CA) was added. The Chelex particles were prevented from sedimenting by continuous stirring on a magnetic stirrer (Variomag, Compact HP1, Daytona Beach, FL). After a brief mixing step on a IKA Vibrax VXR shaker, the samples were incubated for 1 h at 56 °C in a heating block (Techne Dri-Block, DB-3D, Techne, Cambridge, UK). The samples were vigorously mixed for 20 s on an IKA Vibrax VXR shaker and incubated for 8 min at 100 °C in a heating block, vigorously mixed for 20 s and centrifuged at 13,000 RCF for 5 min. The supernatant contained approximately 0.5-ng/μL extracted DNA and was used for the subsequent PCR. Figure 1 shows a flow sheet of the procedure.
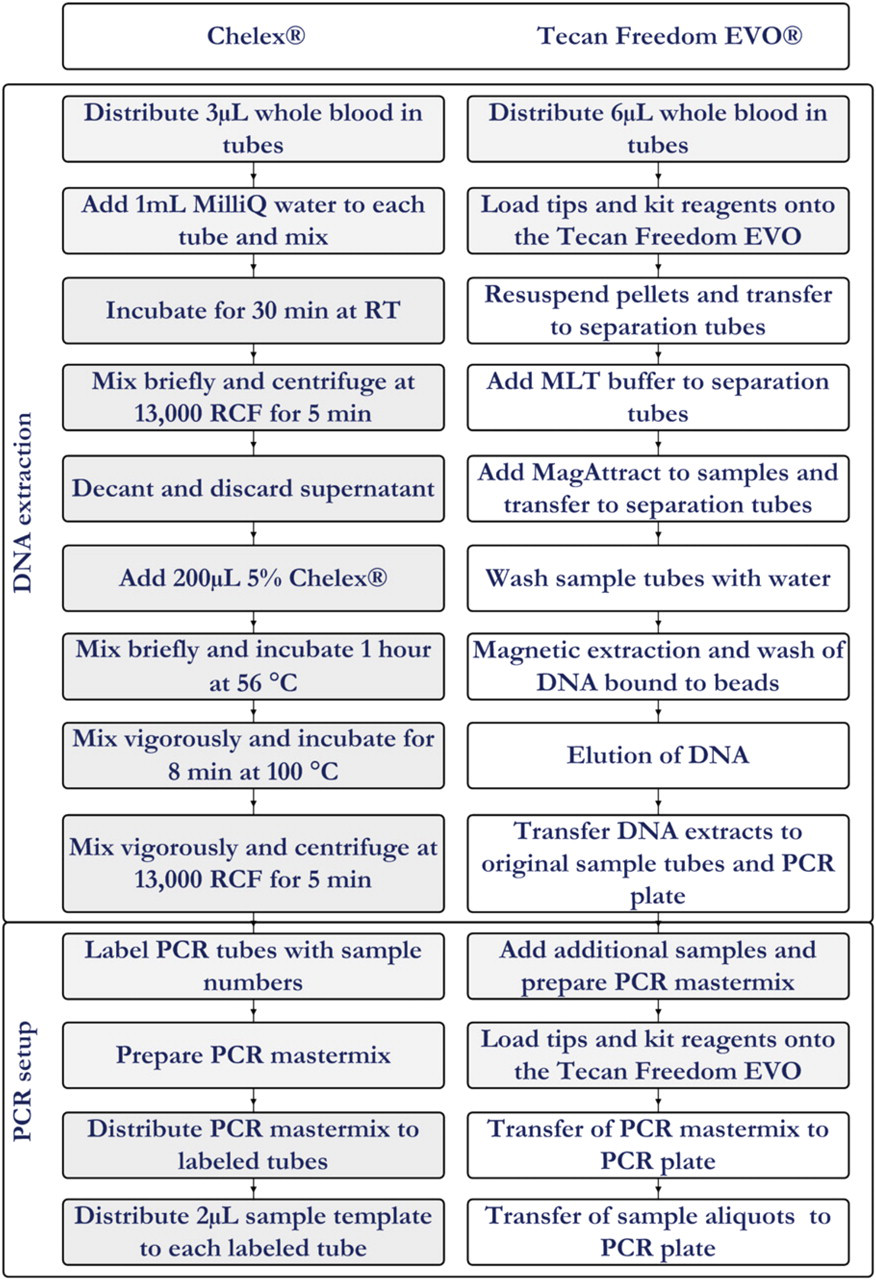
Flow diagrams for the manual Chelex procedure and the automated procedure on the Tecan Freedom EVO. Manual steps are indicated with a gray background.
Automated Extraction
Methods for extraction of DNA and PCR setup were developed using the software EVOware version 2.00.0018 (Tecan, Männedorf, Switzerland). Sample tubes containing 6 μL blood located in the bottom of the tube loaded in 16-tube carriers (Tecan) were inserted on the deck of the liquid handler. Extraction was accomplished using components of the MagAttract DNA Blood Mini M48 Kit (cat. no. 953336, Qiagen GmbH, Hilden, Germany). Liquid reagents other than MagAttract Suspension B were stored in 100-mL troughs on the deck of the liquid handler. All reagent volumes were added according to the number of samples being processed according to Table 1. The MagAttract Suspension B was manually distributed into the eight wells of the first column in a microtiter plate (twin.tec semi-skirted, Eppendorf, GmbH, Hamburg, Germany).
Reagent consumption for extraction of blood samples using the Tecan Freedom EVO
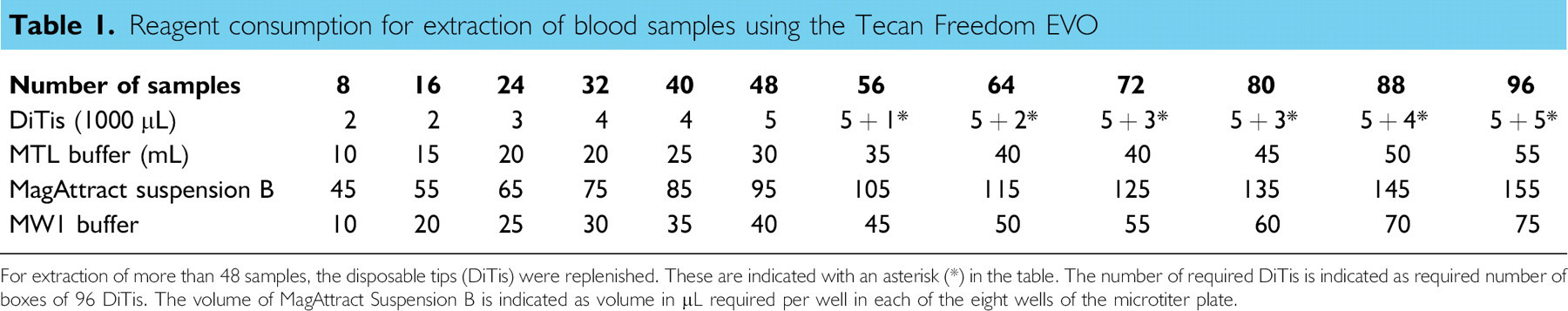
For extraction of more than 48 samples, the disposable tips (DiTis) were replenished. These are indicated with an asterisk (*) in the table. The number of required DiTis is indicated as required number of boxes of 96 DiTis. The volume of MagAttract Suspension B is indicated as volume in μL required per wellin each of the eight wells of the microtiter plate.
Before starting the extraction script, the magnetic particles in the MagAttract Suspension B were re-suspended by shaking the flask thoroughly for 1 min. The suspension was manually distributed in the first column (wells A1 through H1) of a 96-well microtiter plate (twin.tec, Eppendorf, GmbH, Hamburg, Germany) to reduce reagent consumption.
Initially, the MagAttract Suspension B was re-suspended with a mixing step included in the customized liquid class by the robot before distribution of 10 μL into each tube on the Te-MagS separation device (Tecan). Figure 2 shows the work-table layout. This was required to achieve re-suspension of the magnetic beads. Secondly, 500 μL of MTL buffer was added to each tube on the Te-MagS. Transfer of samples to the tubes in the Te-MagS was facilitated by the addition of 80 μLautoclaved MilliQ water as carrier liquid to enable aspiration of the entire sample volume. Magnetic isolation of the DNA was performed on the Te-MagS in lid less 1.5 mL tubes (Sarstedt, Numbrecht, Germany) held in six by eight tube racks designed to fit into the Te-MagS separation device by allowing the magnetic beads to bind for 5 min. A magnetic separation step in which the magnetic unit was moved from one side of the tubes to the opposite side 15 times was performed. The magnetic unit Te-MagS was placed close to one side of the tubes to enable aspiration of the supernatant without aspiration of the magnetic beads to which the DNA was bound. After the removal of the supernatant, the beads were washed with 700 μL MW1 buffer. Each tube was subsequently washed twice with 500 μL 80% ethanol and once with 600 μL autoclaved MilliQ water using the above described procedure. Finally, 200 μL autoclaved MilliQ water was added and the samples were incubated at 56 °C for 5 min to elute the DNA from the magnetic beads. During the magnetic isolation of the DNA, the original sample containing tubes were washed with 700 μL autoclaved MilliQ water to dissolve and remove potential blood residues that might be inhibitory to the PCR. After aspiration and removal of the water used for washing of the tubes, the eluted DNA was transferred from the elution tubes in the Te-MagS unit to the original sample containing tubes in the tube carriers.
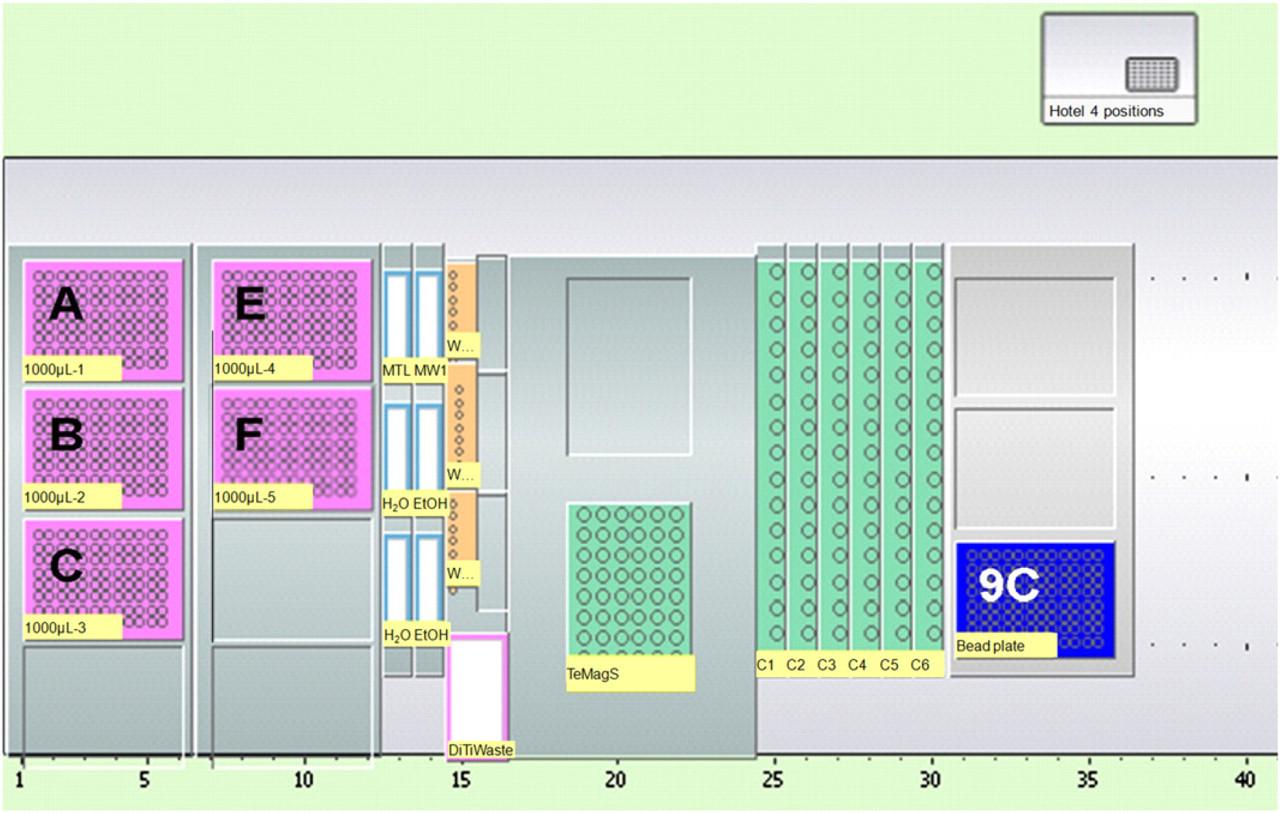
Worktable layout for the extraction method. Position A—F holds disposable conductive filter tips (1000 μL). Buffers and liquid reagents are held in 100 mL troughs. Position 9C holds a 96-well microtiter plate used to hold the magnetic beads before distribution.
PCR Setup
PCR master mix was prepared according to manufacturer's instructions with the following modifications: For the AmpF
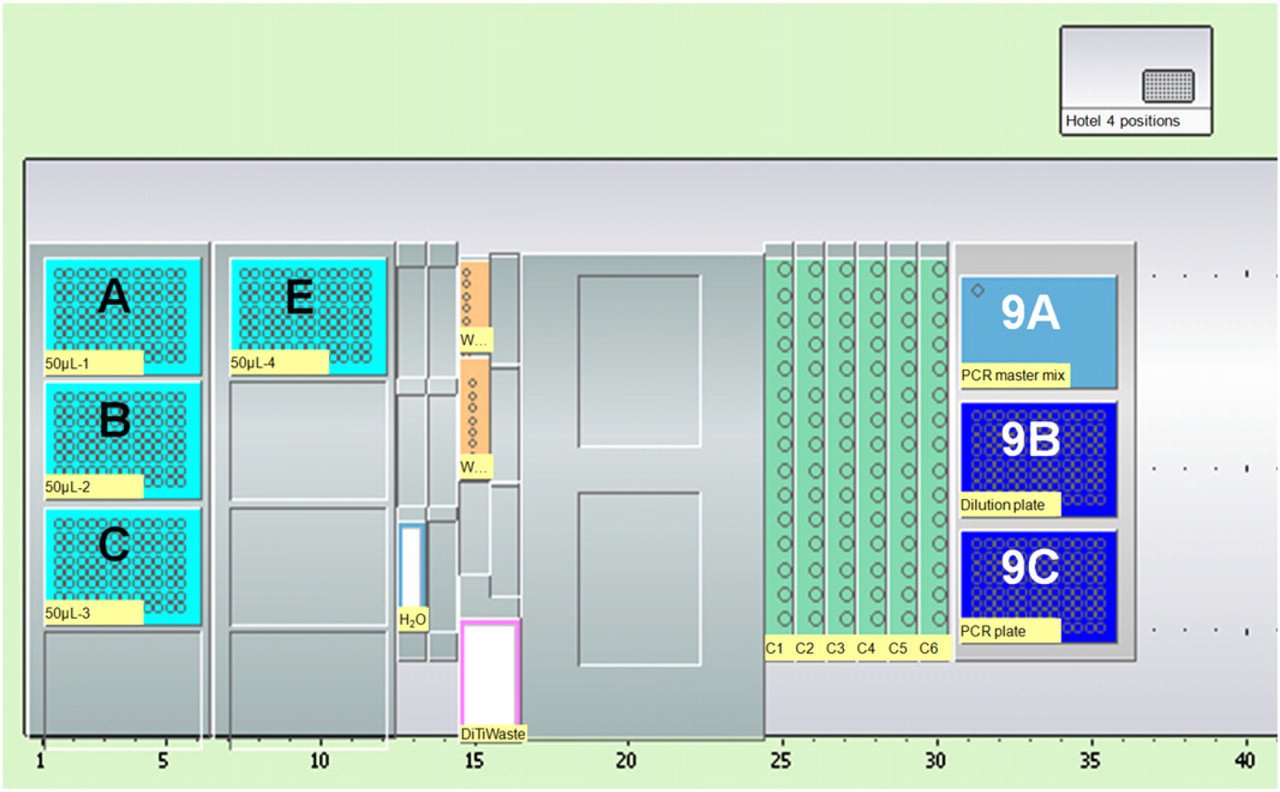
Worktable layout for the PCR setup method. Position A—E holds disposable conductive filter tips (50 μL). Position 9A holds the PCR master mix. Position 9B holds a sample dilution plate. Position 9C holds the PCR plate.
Capillary Electrophoresis
For analysis of the short tandem repeat (STR) amplicons, 1.5 μL of the PCR products were combined with 15 μL formamide and size standards (Applied Biosystems, Foster City, CA) in a 96-well electrophoresis plate (Axygen Scientific Inc., Union City, CA) on a Biomek 3000 laboratory automated workstation (Beckman Coulter, Fullerton, CA). Capillary electrophoresis (CE) was performed on ABI 3130xL Genetic Analyzers (Applied Biosystems, Foster City, CA).
Data Analysis
Multicolor fluorescence CE results were analyzed using Genescan Analysis version 3.7 (Applied Biosystems, Foster City, CA) and allele calls were made using Genotype version 3.7 (Applied Biosystems, Foster City, CA) macros.
Results and Discussion
DNA Extraction and Purification
Manual and automated DNA extraction processes have previously been compared. In general, these investigations reveal a slight advantage for manually extracted DNA regarding technical issues such as yield and purity of DNA even when identical chemistries were used. 2,12,13 The results 11 (Fig. 4) indicated that automated extraction method was comparable in sensitivity with the manual Chelex method. Reagent and instrument costs of the automated method supersedes that of the manual method by an estimated sevenfold. However, the automated method reduces the risk of sample misplacements, requires significantly less hands-on time, and minimizes injuries to the technical staff caused by intensive manual pipetting.
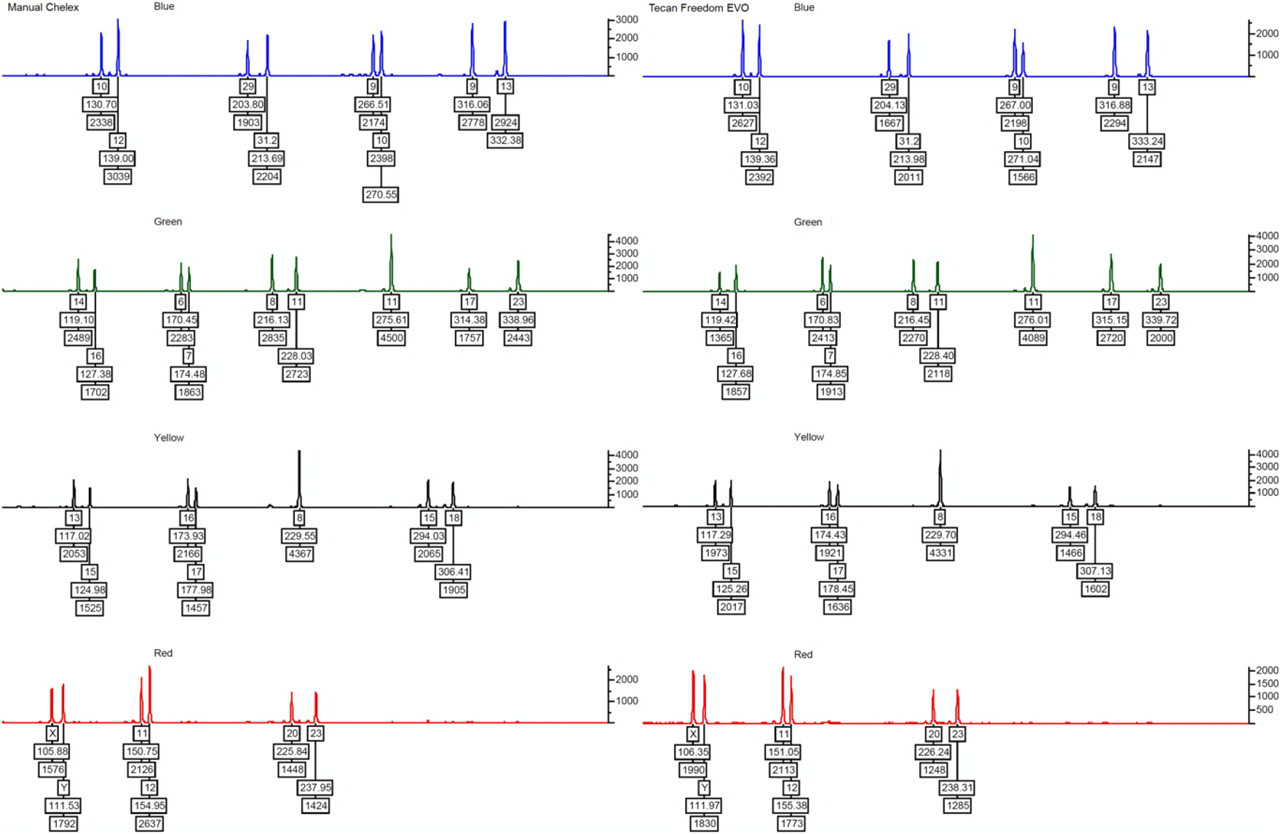
Comparison of short tandem repeat profiles obtained with automated (right) and manual (left) extraction of DNA from blood. Automated extraction was accomplished using 6 μL of blood, whereas Chelex extraction was accomplished using 3 μL blood.
PCR Amplification
Validation of the pipetting capabilities of the Tecan Freedom EVO instrument using an in-house developed method 14 revealed that it was impossible to obtain acceptable accuracy and precision of pipetting for all eight pipettes for volumes below 5 μL (data not shown) with the 1000 μL diluters installed on the instrument. Hence, development of the methods required that all pipetting operations were of volumes greater than 5 μL. A 96-well microtiter plate serving as dilution plate was used enabling a 5 μL extract to be added to each PCR reaction without overloading the PCR with template DNA. The results indicated that this may be an efficient and cost-effective method rather than physically modifying the instrument by replacing some or all diluters with smaller versions, which may be a costly process and impractical as it precludes the use of the same instrument for both methods. The amplification setup method was adjusted to a final concentration of 0.2 ng/μL by adjusting the volume of the reagents used for the extraction in combination with the amount of sample material (data not shown). This allowed for PCR setup using 5 μL DNA extracts containing 1 ng of DNA. Samples with limited DNA were re-extracted using the double amount (12 μL) of blood on the next batch without any further modification. The number of amplification reruns could be reduced by quantifying the extraction before STR amplification and subsequent normalization of template DNA. However, the cost of quantifying all samples using real-time PCR 8 was estimated to exceed the cost of the additional amplification reruns. Furthermore, a quantification step would prolong the overall process time with approx. one half working day.
Comparing the STR types of samples extracted with the manual Chelex-based method with the automated protocol showed that all the STR profiles were concordant and that the number of full profiles obtained was similar (Fig. 5). A small fraction (1.1%) of the samples extracted with the manual Chelex method resulted in a partial profile lacking one or more STR loci. For samples extracted with the automated method, 0.9% resulted in a partial STR profile. The percentage of samples resulting in no STR profile was 0.4% and 1.7% for the manual and automated methods, respectively. The higher number of samples without any data obtained using the automated method is most likely caused by insufficient solubilization of blood and subsequent inhibition of the PCR.
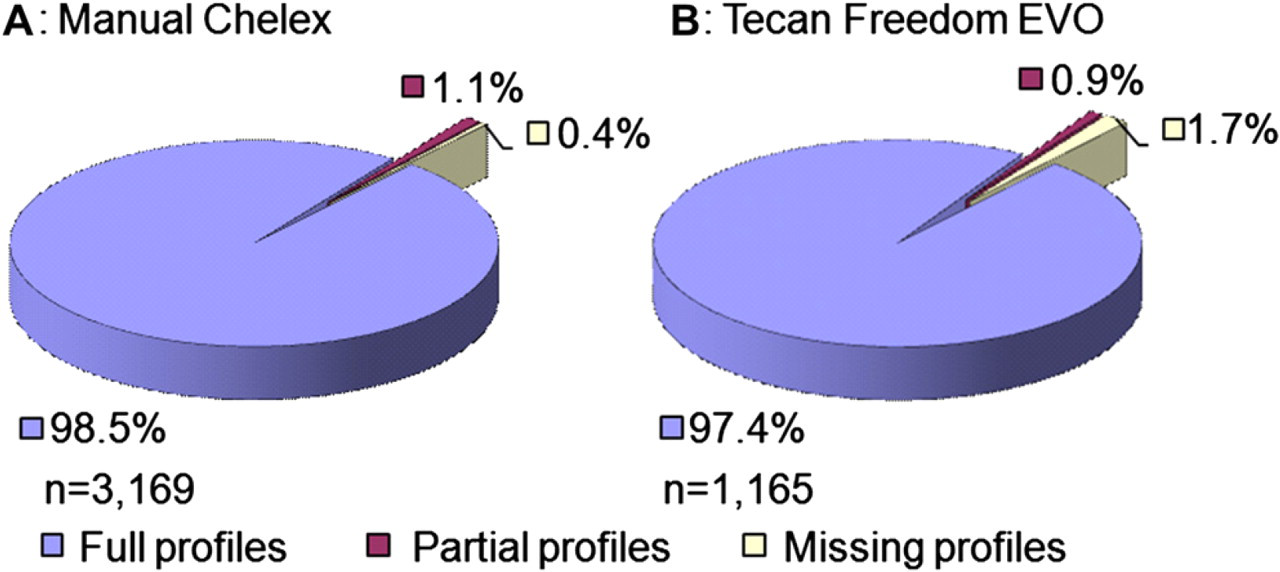
Approval rates of short tandem repeat typing results using the AmpF
Workload
Manual Chelex extraction enabled extraction of DNA from a total of 60 samples including controls per run. The time to completion of the extraction was around 5 h including setting up the PCRs. Combined, this allowed for only one extraction run within one working day per technician. Implementation of the automated extraction method enabled extraction of up to 96 samples including controls within 4 h. This allowed for processing of two runs, each of up to 96 samples including controls within one 8-h working day.
Comparison of the Manual and Automated Extraction Method
The advantages of the automated extraction method include (1) a more than threefold increase in throughput without increasing the workload on the technical staff, (2) a significant reduction of the risk of sample misplacement during processing, (3) simplified downstream processing because of the handling of samples in plates rather than individual tubes, (4) a simplified extraction process requiring minimal manual intervention, and (5) standardized technician-independent processing of samples. The most significant disadvantage is more than sevenfold larger reagent cost per sample ignoring instrumentation costs (data not shown). If wages and the requirement for technical competence of the technical staff are low, there is no economic argument for the automated method. Training of technical staff to the required level of competence for working in forensic genetic laboratories accredited under ISO 17025 is time consuming and costly. Occupational injuries such as musculoskeletal ailments are associated with long-term exposure to extensive manual repetitive pipetting. 15 Implementation of laboratory automation reduces the amount of manual repetitive pipetting operations. Reducing the risk of sample misplacement increases the credibility of the obtained results and, hence decreases the economic disparity between the automated and the manual method. The final choice between an automated or a manual solution depends on local laboratory specific factors, such as sample throughput and certification degree, and regional factors, such as wages and staff availability.
Conclusion
A manual DNA extraction protocol for STR typing in a forensic genetic laboratory was replaced with an automated protocol without affecting the quality of the STR typing results judged by the proportion of partial and missing STR profiles obtained. The automated protocol was implemented on an open robotic system making it flexible in terms of kit supplier. The automated protocol increased the daily throughput more than threefold without the requirement for additional staff, significantly reduced the probability of sample misplacement during processing, simplified extraction process requiring minimal manual intervention, simplified downstream processing because of handling of samples in plates rather than individual tubes, and provide a standardized technician-independent processing of samples. However, the automated extraction method increased the extraction reagent costs more than sevenfold per sample. The developed methods for the Tecan Freedom EVO automated liquid handler are available as Supplementary Material.
ACKNOWLEDGMENTS
Competing Interests Statement: The authors certify that they have no relevant financial interests in this article.
SUPPLEMENTARY MATERIAL
Supplementary material associated with this article can be found, in the online version, at doi:10.1016/j.jala.2010.11.003.
