Abstract
We have implemented and validated automated protocols for a wide range of processes such as sample preparation, PCR setup, and capillary electrophoresis setup using small, simple, and inexpensive automated liquid handlers. The flexibility and ease of programming enable the Biomek 3000 to be used in many parts of the laboratory process in a modern forensic genetics laboratory with low to medium sample throughput. In conclusion, we demonstrated that sample processing for accredited forensic genetic DNA typing can be implemented on small automated liquid handlers, leading to the reduction of manual work as well as increased quality and throughput.
Keywords
Introduction
The application of forensic genetic analysis in crime case work has grown significantly the past two decades, increasing the workload on laboratories throughout the forensic community. The increase in workload has led to a shift from applying manual processes in the laboratories toward automated processes. With high workloads, automated processes are more attractive because of a reduction in process time, reduction in the requirement for manual intervention, reduction in the risk of sample misplacement, reduction in the per sample processing cost, reduction in musculoskeletal ailments of the technical staff, 1 standardized sample treatment, securing the chain of custody, and an increase in throughput. Implementation of laboratory automation should be scaled to the number of samples processed by the laboratory to be economically justifiable. 2 For high-throughput processes, application of automated solutions based on liquid handlers and Laboratory Information Management Systems (LIMS) is a necessity. However, these full to semi-automated solutions are often challenging and expensive to implement and validate in an accredited forensic laboratory as they require extensive testing. 3 Furthermore, as these automated production lines are dependent on many parts and complex interactions to function, they are vulnerable to breakdown, and local support and frequent maintenance may be required. For many processes in forensic laboratories, a fully automated production line is not appropriate, especially if experience with automated processes and liquid handlers is low. However, many small- and medium-sized laboratories may benefit by implementing small- to medium-sized automated liquid handlers (ALHs) to perform key processes.
We have previously reported that the Biomek 3000 may be used to perform complex laboratory processes. 4,5 Here, we present five simple methods developed for the Biomek 3000 liquid handler. The methods are easily applied in forensic laboratories, as well as validated and accredited according to the ISO 17025 standard. None of the methods requires LIMS integration or support and may be used as stand-alone applications.
Materials and Methods
Samples
Reference, paternity, and immigration samples were processed in the form of buccal samples on FTA cards (GE Healthcare, Piscataway, NJ), 6 anticoagulated blood or various other types of biological material extracted using a manual Chelex protocol, 7 the M48 extraction kit (Qiagen, Valencia, CA) on a Tecan Freedom EVO (Tecan, Männedorf, Switzerland), 8 a QIAsymphony SP with the QIAsymphony DNA Investigator Kit (Qiagen), or a Qiagen BioRobot EZ1 using the DNA Investigator Kit (Qiagen). For samples processed on the QIAsymphony SP, solubilization of the samples, addition of ATL buffer, and subsequent mixing were accomplished using a Biomek 3000 with a customized instrument deck and an in-house developed method. The samples were not quantified. The extraction protocols combined with selected elution volume were optimized to yield approximately 1 ng DNA in the volume used for the subsequent PCR.
Trace evidence samples were extracted using a manual Chelex protocol. 9 In brief, a 20% Chelex solution (Chelex 100 Resin; Bio-Rad Laboratories, Hercules, CA) was prepared by combining 100 g Chelex beads with 412.5 mL sterile RNase- and DNase-free water (product number W4502; Sigma-Aldrich, St. Louis, MO). Prior to autoclaving, the pH of the solution was adjusted with 10 N NaOH (Sigma-Aldrich) to pH 10.4–10.8. Prepared Chelex solutions were stored for up to 1 week at 4 °C. Following addition of 1 mL 20% Chelex solution to sample tubes containing trace items, sample tubes were incubated for 1 h at 56 °C in shaking incubators (VorTemp 56; Labnet International, Inc., Woodbridge, NJ), followed by 8 min at 100 °C in a heating block (SBH200D; Stuart, Bibby Scientific Limited, Staffordshire, UK) and centrifugation for 1 min at 16 000 relative centrifugal force (RCF) in a Heraeus Biofuge Pico bench top centrifuge (Heraeus, Hanau, Germany). The supernatant was transferred to Amicon Ultra 4 100K tubes (Millipore, Billerica, MA) containing 1.5 mL sterile RNase- and DNase-free water. Amicon tubes were centrifuged for 10 min at 4000 RCF in a centrifuge (Megafuge; Heraeus). Following addition of 1.5 mL sterile RNase- and DNase-free water to the Amicon tubes, the tubes were centrifuged again for 10 min at 3000 RCF. Tubes containing more than 50 µL liquid in the top reservoir were centrifuged for an additional 5 min at 2000 RCF. The DNA extract (30 µL) was transferred to a fresh sample 1.5-mL tube (Eppendorf, Hamburg, Germany). Samples containing less than 30 µL were adjusted to 30 µL by addition of sterile RNase- and DNase-free water. Samples were stored at −20 °C until quantification. Trace evidence samples were quantified using the Quantifiler Human DNA quantification kit (Applied Biosystems [AB], Foster City, CA). The quantifier reaction was automatically set up using the Tecan HID EVOlution qPCR/STR setup system as previously described. 10 Samples with amplifiable DNA concentrations less than 10 pg/µL were stopped and reported as too little DNA. Samples with amplifiable DNA concentrations between 10 and 20 pg/µL were concentrated using Amicon Ultra 4 100K tubes and requantified. Samples with amplifiable DNA concentrations greater than 20 pg/µL were normalized and, when required, diluted to enable 500 or 200 pg DNA to be added to each subsequent PCR depending on the concentration. PCRs were set up using 500 pg DNA whenever the DNA concentration allowed it.
PCR
Reference, paternity, and immigration samples were amplified using the AmpFISTR Identifiler PCR Amplification Kit (AB) or the AmpFISTR Identifiler Direct PCR Amplification Kit. For samples requiring additional investigations, amplification using either the AmpFℓSTR Yfiler PCR Amplification Kit (AB) and/or SNP investigations as previously described 11 was also performed. Crime scene samples were amplified using the AmpFℓSTR SEfiler Plus PCR Amplification Kit (AB). PCR master mix was prepared according to the manufacturer’s instructions with the following modifications: For the AmpFISTR Identifiler PCR Amplification Kit and the AmpFℓSTR Yfiler PCR Amplification Kit, the total reaction volume was reduced to 10 µL. For the Identifiler Direct PCR Amplification Kit, the total reaction volume was reduced to 15 µL. PCR reactions were performed in 96-well semi-skirted twin.tec plates (Eppendorf) held in place on the ALHs by support bases (AB, part number N8010531). Following addition of PCR reagents, PCR plates were manually sealed using aluminum seal (PCR-AS-200; Axygen, Union City, CA) and amplified on gold-plated 96-well GeneAmp PCR systems 9700 (AB).
Capillary Electrophoresis
For analysis of the short tandem repeat (STR) amplicons, either 1.0 or 1.5 µL of the PCR products was combined with 15 µL formamide (AB) and size standards (AB) in a 96-well electrophoresis plate (Axygen or AB) held on 96-well plate bases (AB) on Biomek 3000 laboratory-automated workstations (Beckman Coulter, Fullerton, CA) using in-house developed methods. Formamide and size standards were manually distributed in 8-by-1 mini-tube strips (Axygen) prior to loading on the Biomek 3000 instrument. Capillary electrophoresis (CE) was performed on ABI 3130xL Genetic Analyzers (AB).
Data Analysis
Multicolor fluorescence CE results were analyzed using Genescan Analysis version 3.7 (AB), and allele calls were made using Genotype version 3.7 (AB) macros.
Method Development for the Biomek 3000 Instruments
All methods and associated pipetting templates and techniques for the Biomek 3000 instruments were developed using the Biomek software version 3.3 (build 14) (Beckman Coulter). All methods presented in this work were accompanied by methods for verifying the actual volume of liquid dispensed by the individual pipetting techniques and liquid types according to an in-house developed method described previously. 12
Results and Discussion
The number of forensic samples collected and analyzed has increased significantly over the previous years. Usually, increased workload also increases the risk of errors in manual operated processes. If more than one sample is processed manually, the risk of sample misplacement as well as variations in terms of pipetting performance between the laboratory staff exists. Furthermore, repetitive pipetting is associated with hand and shoulder muscular ailments. 1 Implementation of a small automated liquid handler such as the Biomek 3000 in key processing steps reduces the risk of sample misplacement and ensures a lower day-to-day variation in pipetting performance. The Biomek 3000 may be used in many parts of the sample processing. Currently, the Biomek 3000 instruments are applied in several processes in our laboratory ( Fig. 1 ).
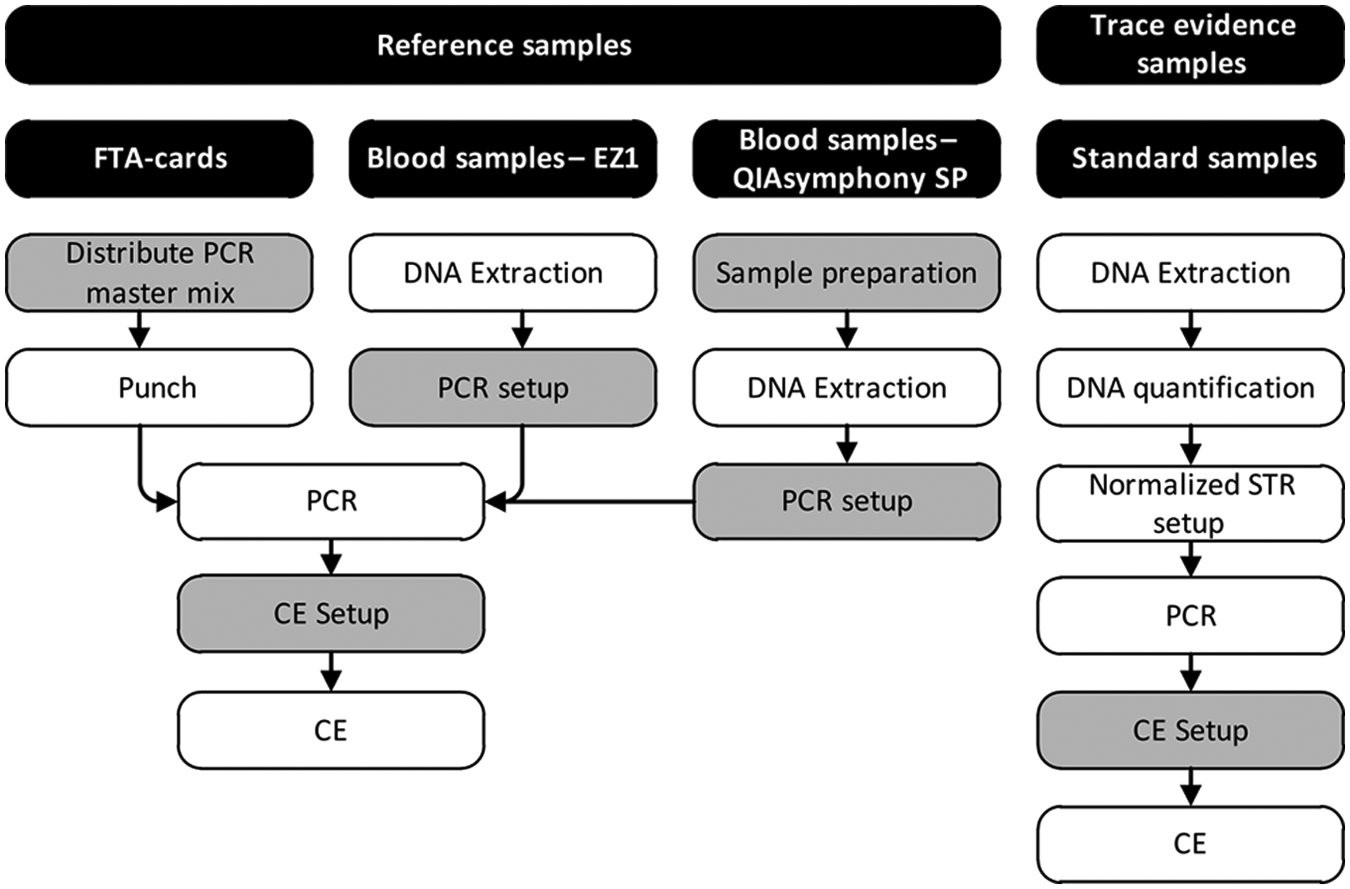
Flow gram of typical forensic genetic sample processing. Boxes with gray background represent steps performed by the Biomek 3000. Reference samples include samples for crime, paternity, and immigration cases. For FTA cards, only the protocol for direct short tandem repeat (STR) kits is presented. CE, capillary electrophoresis.
General Modification of the Deck Positions
The deck positions of the Biomek 3000 enable placement of many types of labware of the standard Society for Laboratory Automation and Screening (SLAS) format. We have modified key positions to ensure that misplacement and disorientation of labware rarely occur. The modifications consisted of placement of a 3-mm-thick stainless steel rod in either the lower right ( Figs. 2 – 3 ) or the lower left corner ( Fig. 3 ). For PCR plates, the modification also included manually removing the lower right corner of the support base ( Fig. 2 ). The steel rod also prevented nonmodified standard flat-bottomed SBS plates to be placed on the modified positions.
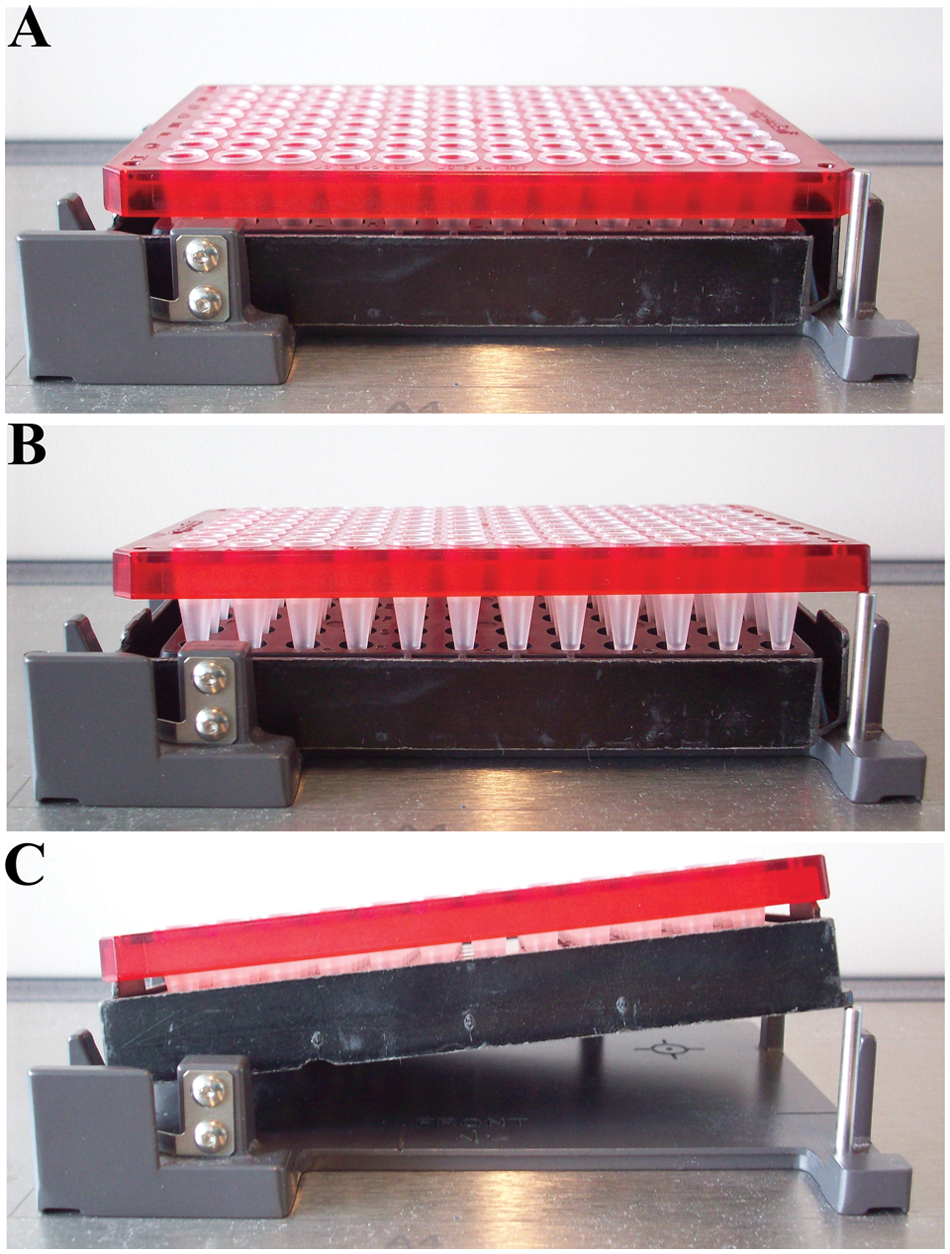

Modification to prevent misplacement and disorientation of microplates. (
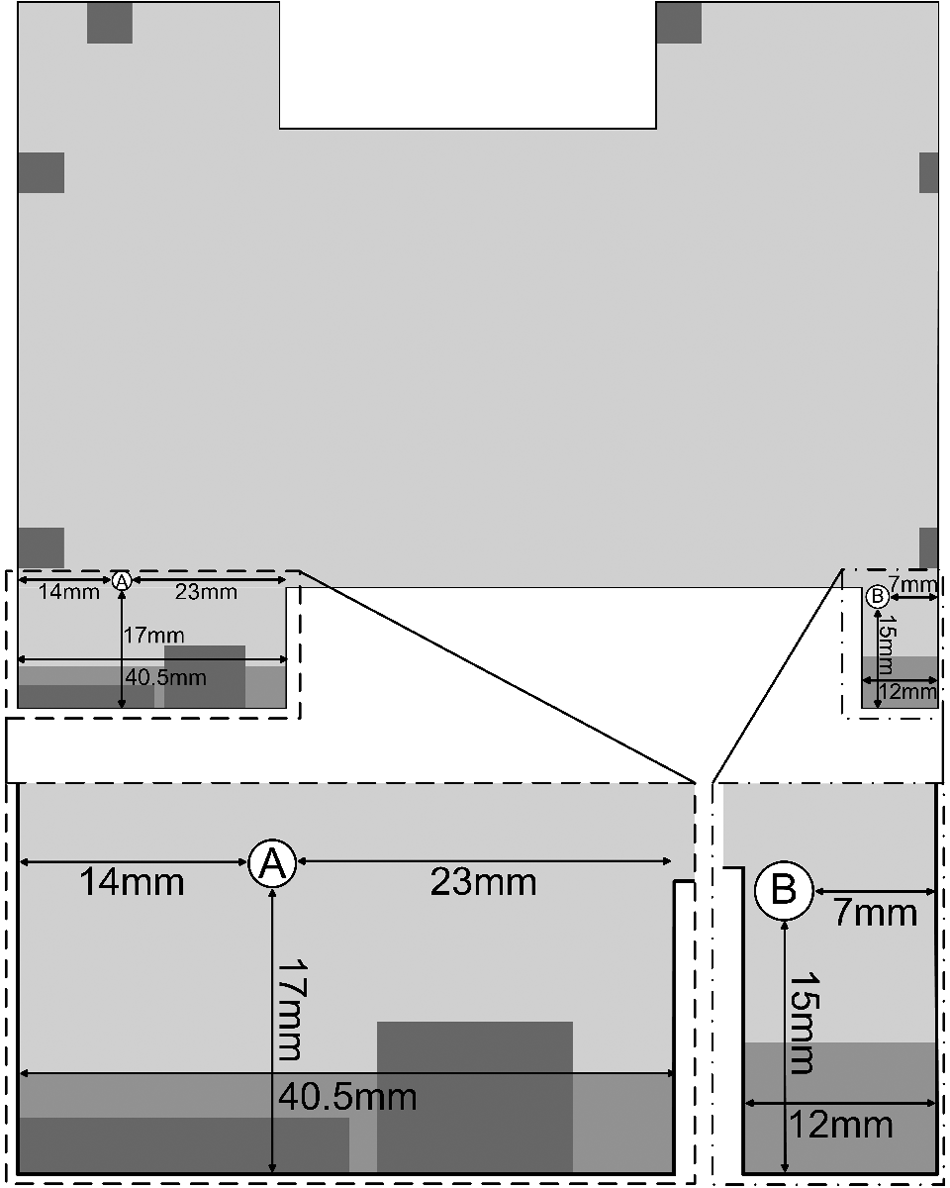
Line drawing of the modified Biomek 3000 deck positions. The darker shades of gray indicate various elevated prefabricated features of the positions. The lower left and right corners with the modifications are magnified below. The magnified areas are indicated with stippled rectangles. The position was modified in the lower left or lower right corner. The modification consisted of drilling a hole, enabling fixation of a 3-mm diameter stainless steel rod at a defined location. (
Sample Preparation for QIAsymphony SP Extraction of DNA from Blood Samples
Extraction of DNA from blood using the QIAsymphony SP with the QIAsymphony DNA Investigator Kit and the Qiagen-provided ACS_REF_200 extraction protocol required that the blood was dissolved in a total volume of 200 µL ATL buffer included in the kit. Initial results indicated that extraction of DNA from 4 µL blood and subsequent elution in 100 µL ATE buffer provided with the DNA Investigator Kit were sufficient to enable robust STR amplification when 2 µL extract was combined with 8 µL Identifiler PCR master mix and 26 cycles of PCR amplification (data not shown). Automated addition of ATL buffer to samples aliquoted into 2-mL sample tubes held in a 24-tube carrier (Qiagen) was achieved using a Biomek 3000 and an in-house developed method. The method enabled a variable number of samples (1–24) to be processed in each run, enabling processing of only the relevant number of samples. The instrument deck layout was customized to hold the tube carrier by cutting a sheet of 4-mm-thick aluminum spanning four positions. When the piece of aluminum was placed over the positions, it enabled fixation of the Qiagen 24-tube carrier. A 39-mm diameter hole was drilled to the depth of 29 mm in a 127-by-86-mm piece of 30-mm thick Teflon to enable fixation and positioning of the ATL buffer flask on a standard, nonmodified deck position on the instrument deck ( Fig. 4 ). Addition of ATL buffer was achieved using in-house developed pipetting techniques and templates. Following addition of the buffer, the sample was mixed to ensure adequate mixing by aspirating and dispensing the buffer at different locations in each sample tube. The sample preprocessing combined with the continuous loading capabilities of the QIAsymphony enabled continuous processing of the samples throughout the working day.
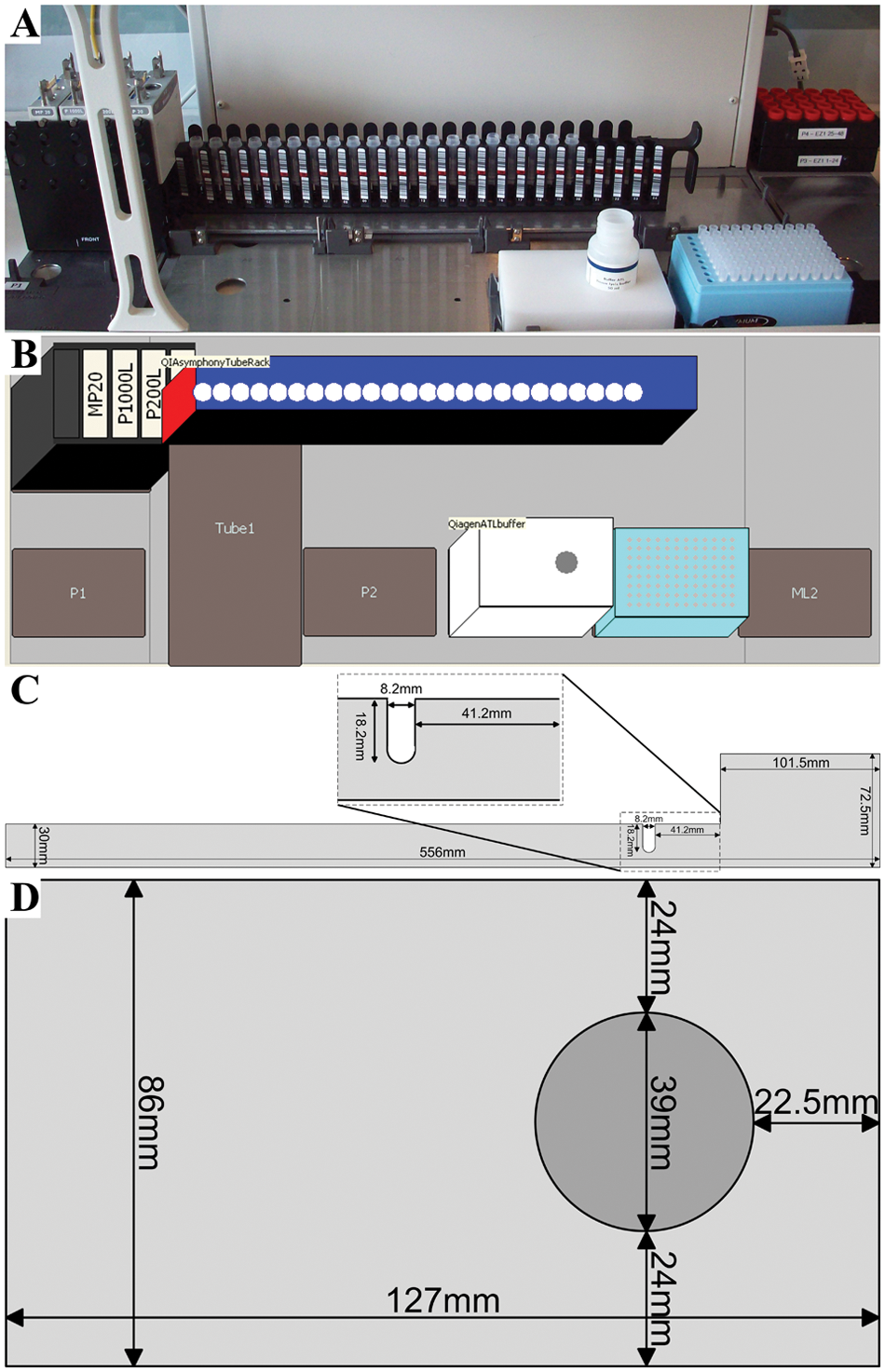
Deck layout of the instrument. (
PCR Setup
Three slightly different methods were developed for setting up PCR reactions. The first was applied when the extracted DNA was in the microplates. Following extraction of DNA, PCR reactions were set up using either 8 or 9 µL manually prepared PCR master mix and 2 or 1 µL extracted DNA, resulting in a total volume of 10 µL. PCR setup was achieved on a Biomek 3000 using an in-house developed method distributing PCR master mix and subsequently DNA extract in the required wells, omitting pipetting in predefined selected wells used for the ladder in the subsequent CE. The method enabled a variable number from 1 to 90 samples to be processed. The deck layout is presented in Figure 5 .
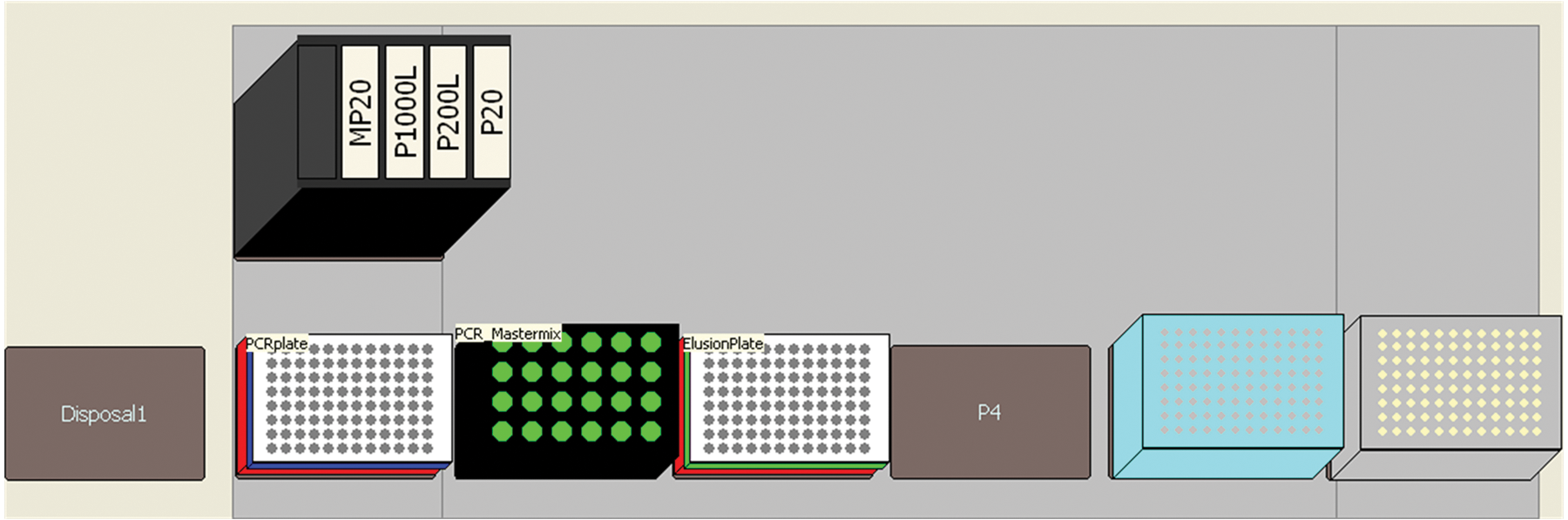
Deck layout for PCR setup from plate to plate.
Comparisons between samples prepared manually for the QIAsymphony and subsequently set up for PCR manually with samples on a Biomek 3000 indicated that the results were similar ( Fig. 6 ).
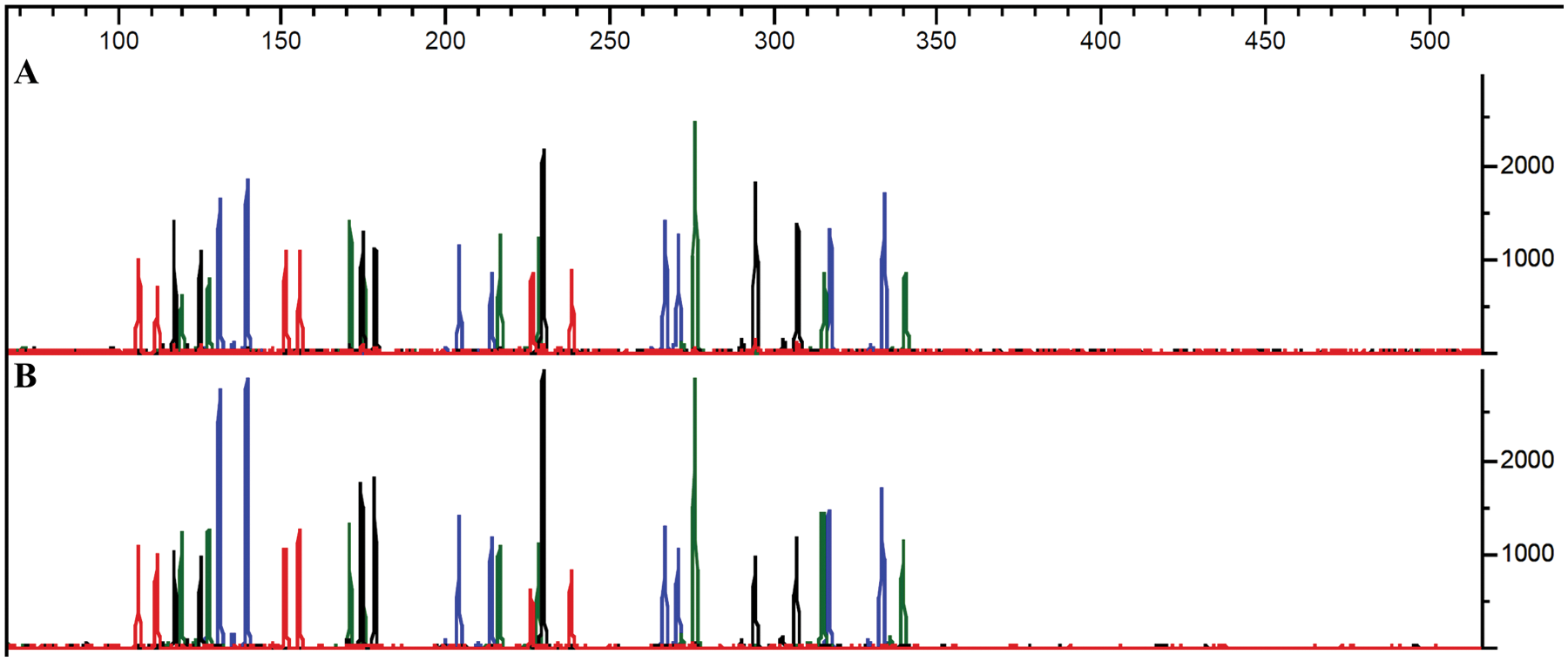
Comparison of manual and automated sample preparation and PCR setup. (
Recently, several STR kits that enable direct amplification have become commercially available. In theory, these kits enable STR amplification directly on FTA punches containing buccal or blood samples 14,15 without a requirement of washing the FTA punch prior to PCR setup. A recent study on cotton swabs indicated that direct PCR kits resulted in better STR profiles than conventional extraction followed by nondirect STR kits. 16 Hence, a second PCR setup method for the Biomek 3000 based on the PCR method described above was developed simply by removing the section of the method adding the DNA ( Fig. 7 ).
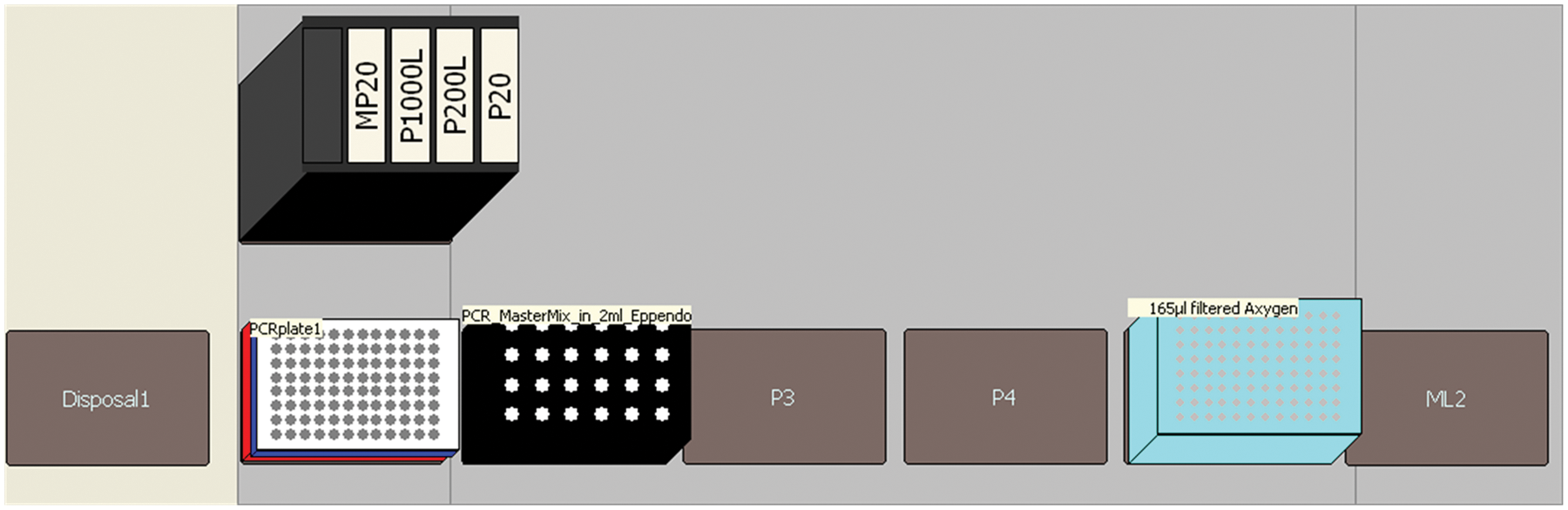
Deck layout for Identifiler direct setup.
Small DNA extractors such as the BioRobot EZ1 from Qiagen, the AutoMate Express from Applied Biosystems, or the Maxwell 16 from Promega (Madison, WI) are often convenient when either small batches of samples are being processed or multiple sample types are required to be processed differently. However, they elute the extracted DNA in either 1.5- or 2-mL sample tubes, and it may be cumbersome to manually prepare many different PCRs in individual tubes. Combining several extraction runs into one PCR plate not only saves time but also reduces the requirement for multiple runs with few injections on the electrophoresis systems. Hence, a third PCR setup method was developed, enabling PCR setup from up to forty-eight 2-mL sample tubes held in 6-by-8 tube holders. By storing the response of initial user queries on how many samples were in each tube holder in variables, the method enabled processing of a variable number of samples in each tube holder. The deck layout is presented in Figure 8 . Comparison of the STR profiles indicated that the results obtained with the Biomek 3000 did not differ from those obtained by manual setup (data not shown).
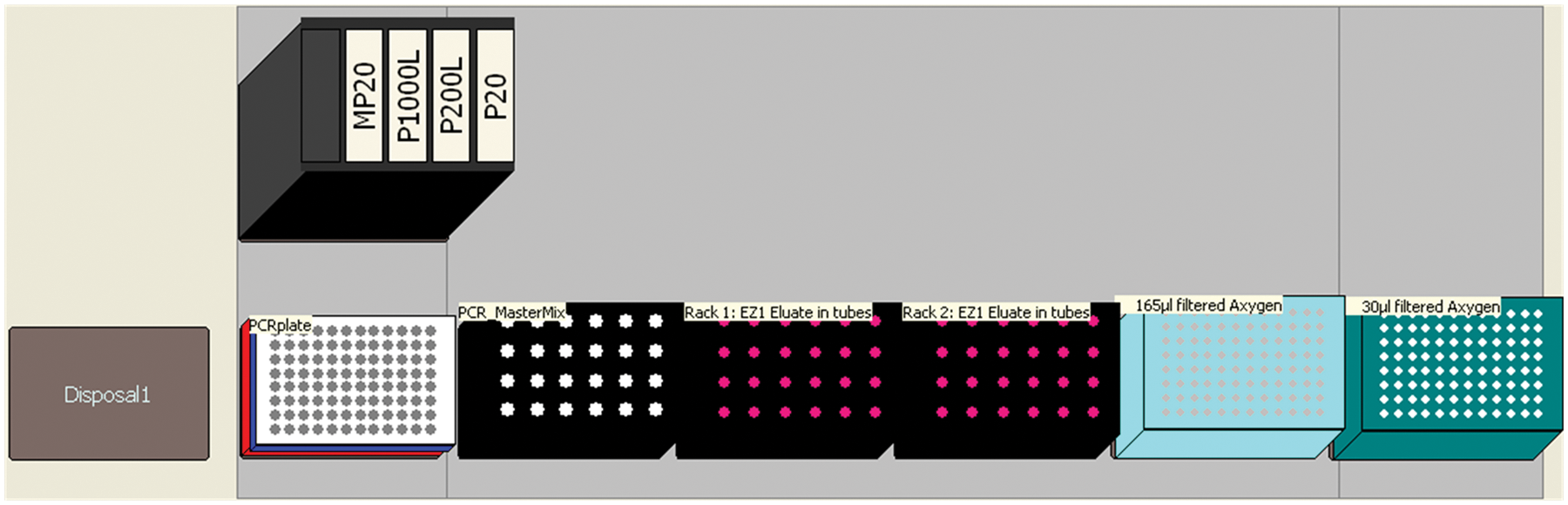
Deck layout for PCR setup from samples held in tubes.
Setup of STR Products for Capillary Electrophoresis
Determination of STR fragment length was performed by CE. Prior to CE, PCR products were combined with formamide and sizing standards to ensure denaturing conditions during CE and correct size determination. The electrophoresis was to be performed on ABI 3130xL instruments with 16-capillary arrays. Hence, a 3130xL will divide a full 96-well plate into six injections. To enable processing of partial plates, all methods were designed by separating the plate into six injections. A method for Biomek 3000s was developed enabling filling of a 96-well CE plate held in place in a 96-well base plate (AB part number 4317237) with formamide containing size standards and subsequent addition of PCR products based on the number of injections to the ABI 3130xl instruments. Formamide and sizing standard were manually combined in strip-tubes prior to starting the method. The addition of PCR products was hampered by the aluminum seal on the PCR plate. A special pipetting template and technique were developed using an initial aspiration of a 10-µL air gap followed by a slow puncture of the aluminum plate seal and subsequent lateral movements of the tips, enlarging the hole to prevent the plate from being moved along with the pipette tips. Prior to moving the tips down into the PCR products, the air gap was rapidly dispensed, removing any sealing film potentially blocking one or more tips. Highly deionized (Hi-Di) formamide was used to ensure prolonged denaturing conditions of the DNA. Because formamide penetrates plastic, the commercially available disposal bag system for the Biomek 3000 was not practical to use. Returning the used tips to the tip-box could result in accidental failure of the technical staff to replenish the tips following a run and hence an accidental reuse of the tips. To prevent this, a disposal unit was added to the instrument decks in the software. However, instead of physically adding the original disposal option to the deck of the instruments, a conventional 1.5L Pyrex beaker glass was used as a waste container. Formamide fumes were removed from the laboratory by placing the Biomek 3000 in an in-house designed aluminum framed container with acrylic sides and top-mounted point suction ( Fig. 9A ). Used tips were left in the beaker glass and emptied directly in single-use chemical waste containers. The deck layout is presented in Figure 9B .
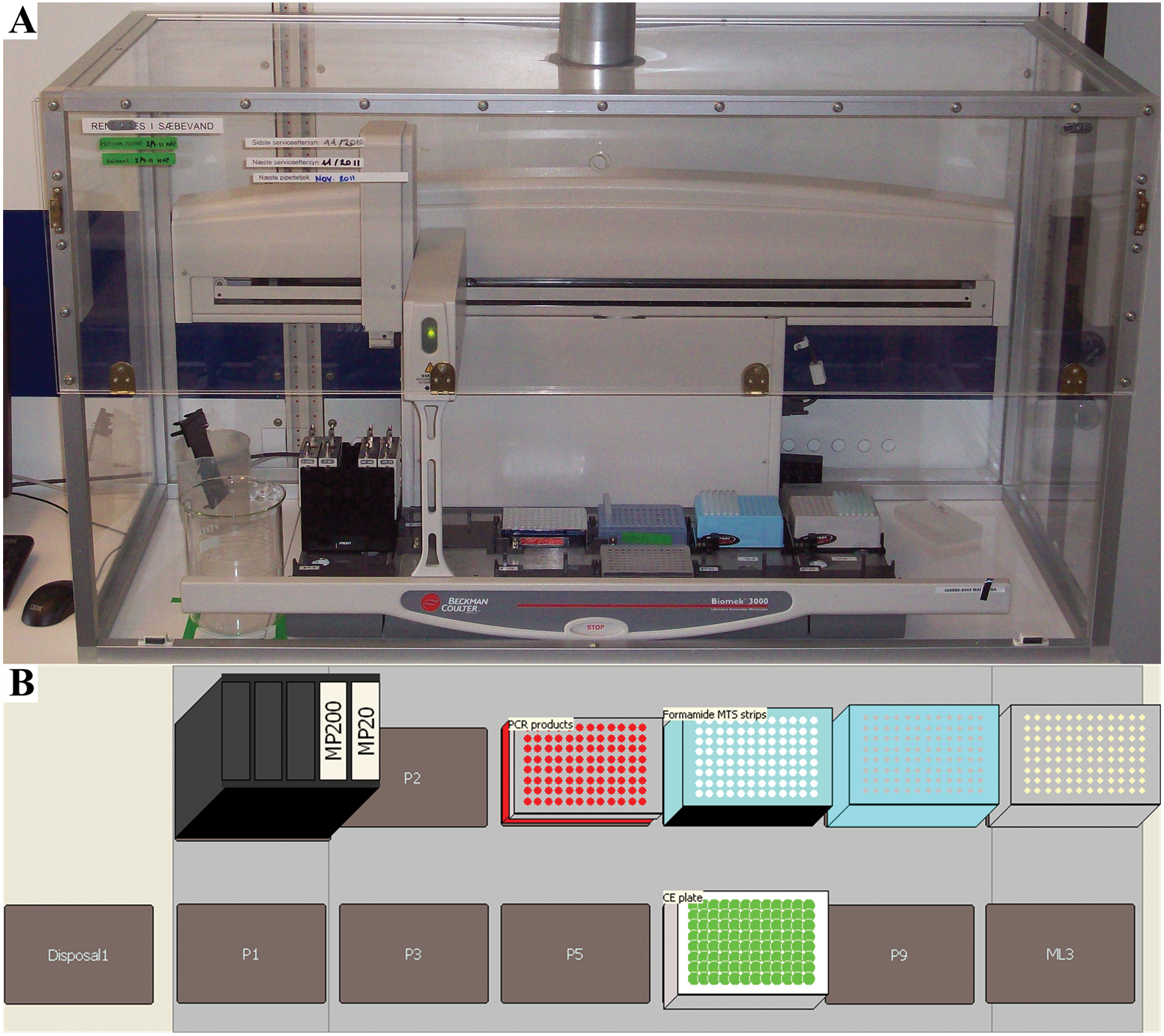

Deck layout for capillary electrophoresis (CE) setup. (
A small and flexible automated liquid handler such as the Biomek 3000 may be an invaluable tool in a modern forensic genetic laboratory. Automation of key processes may not only increase throughput and reduce the risk of sample misplacement but also reduce occupational injuries to the technical staff. We have automated multiple simple processes from sample preparation, over PCR setup to CE setup, as well as complex cumbersome processes such as the GenPlex, 4,5 using Biomek 3000 instruments as illustrated in Figure 1 . In conclusion, the Biomek 3000 is a flexible and robust liquid handler capable of processing samples in forensic genetic laboratories with small or medium sample throughput. All developed methods are available as supplementary information.
Footnotes
Acknowledgements
We thank Anja L. Jørgensen and Søren Lorentzen for technical assistance.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
