Abstract
Epithelial-mesenchymal transition (EMT) is essential for initiation of colorectal cancer (CRC) metastasis, but the diver proteins of EMT remain unclear. Special AT-rich sequence-binding protein 1 (SATB1) was found to be overexpressed in CRC cell lines, and its expression level was positively correlated with CRC progression. Strikingly, EMT process was regulated by SATB1, as SATB1 overexpression upregulated E-cadherin and SATB1 knockdown inhibited N-cadherin cell models. Mechanistically, SATB1 promoted EMT-mediated CRC metastasis via activation of Notch signaling pathway. Taken together, SATB1 plays a vital role in CRC metastasis and may act as a novel prognostic biomarker and a promising therapeutic target for CRC.
Introduction
Colorectal cancer (CRC) has been considered as the third most common type of cancer (1.8 million new cases) and second most frequent cause of cancer death (0.86 million) worldwide in 2018. 1 CRC is difficult to treat due to its tumor invasion ability and malignant metastasis. When CRC spread to distant organs (stage IV), the 5-year survival rate reduced to ~11%. 2 Therefore, it is urgent to investigate metastasis mechanisms of CRC with the aim to develop novel drugs or therapies to improve survival rate of CRC patients. Epithelial-mesenchymal transition (EMT) is defined as a process that epithelial cells trans differentiate into motile mesenchymal cells. 3 EMT plays a vital role in regulation of multiple signaling pathways in cancer metastasis.3,4 Wnt/β-catenin, TGF-β, and Notch signaling pathways have been report to mediate EMT progression. 5 However, the initiation progress of EMT in cancer remains unclear as well as requires further investigation.
Known as special AT-rich sequence-binding protein 1, SATB1 is a nuclear protein that acts as a transcription factor and chromatin organizer. 6 SATB1 couples with AT-rich motifs around heterochromatin and forms “cage-like” network to facilitate chromatin remodeling and gene regulation.6,7 SATB1 expression is closely related to tumor progression and prognosis, including increased tumor growth, metastasis, and reduced survival rate.8,9,10 In breast cancer cells, more than 1000 genes were reported to be regulated by SATB1. 8 SATB1 also promotes EMT and metastasis in CRC and prostate cancer.11,12 However, the exact molecular mechanisms of SATB1 in regulating cancer metastasis should be further clarified. In this study, SATB1 overexpression was found to correlate with aggressive invasion and migration of CRC cells. We also discovered the underlying mechanisms of SATB1-induced metastasis depended on Notch signaling pathway, indicating SATB1 might be a useful biomarker for CRC metastasis as well as a potential drug target for CRC further treatment.
Materials and methods
Cells and reagents
Human normal colorectal epithelial cell line FHC (CRL-1831) and CRC cell lines named SW480 (CCL-228), SW620 (CCL-227), LoVo (CCL-229), HCT116 (CCL-247), and HT29 (HTB-38) were acquired from the Chinese Academy of Sciences. Cells were incubated in Dulbecco’s modified Eagle medium (DMEM; Gibco, Carlsbad, USA) with 10% fetal bovine serum (FBS; Hyclone, Logan City, USA), penicillin (100 μg/ml; Gibco, Carlsbad, USA), and streptomycin (100 μg/ml; Gibco, Carlsbad, USA) at 37°C under 5% CO2. The 0.5 μg/ml recombinant Jagged 1 FC (R&D Systems, Minneapolis, USA) was used to stimulate the CRC cells. All cell experiments in this study were complied with the regulations of Biomedical Research Ethics Committee of Jinan University.
TCGA data analysis
Gene expression (RNA-seq) assay and clinical data of CRC patients were obtained from TCGA (The Cancer Genome Atlas) data portal (https://portal.gdc.cancer.gov/). UCSC Xena (https://xena.ucsc.edu/) was used to analyze the correlation between SATB1 expression and CRC overall survival (OS) rates. Specifically, survival rates of CRC patients stratified by quartiles of SATB1 expression. Cutoff values were set as follows: 1st < 8.572 (n = 107), 8.572 < 2nd < 9.316 (n = 100), 9.316 ⩽ 3rd < 9.817 (n = 108), 4th > 9.817 (n = 103), respectively.
Western blot
Cells were lysed in RIPA lysis buffer (Beyotime, Shanghai, China), and the concentration of protein extracts was detected by BCA Protein Assay Kit (Thermo Fisher Scientific, Waltham, USA). Equivalent amounts of protein were separated by sodium dodecyl sulfate–polyacrylamide gel (SDS-PAGE), and then transferred onto PVDF (polyvinylidene difluoride) membranes (Millipore, Billerica, USA). After blocking in 5% skim milk for 2 h, membranes were incubated with primary antibodies (SATB1, E-cadherin, N-cadherin, Vimentin, Snail, Slug, NICD, Hes1, β-actin; Abcam, Cambridge, UK) overnight at 4°C. Washing 3 times with tris-buffered saline containing 0.1% polysorbate 20, membranes were then incubated with secondary antibodies (Abcam, Cambridge, UK) for 1 h. Proteins were visualized by enhanced chemiluminescence (Thermo Fisher Scientific, Waltham, USA) and Image Quant LAS 4000 Mini (GE Healthcare Life Sciences, Boston, USA). Proteins bands intensities were analyzed with Image J.
Establishment of SATB1-knockdown cell line
Knockdown vectors of SATB1 (siRNA-SATB1) and control vectors (siNC) were purchased from Sangon Biotech (Shanghai, China). Lipo2000 (Invitrogen, Carlsbad, USA) was used to transfect these vectors into LoVo cells. Stable SATB1-Knockdown LoVo cells were screened using 1 mg/ml G418 (Selleck, Houston, USA).
Migration and invasion assay
Cell migration and invasion assay was performed with transwell chambers (8 μm pore size; Corning, Corning City, USA). For cell invasion assay, Matrigel (Corning, Corning City, USA) was pre-coated on chambers. The upper chamber seeded with CRC cells, and the lower chamber filled DMEM with 10% FBS. Chambers were incubated for 12 h, and non-migratory cells were wiped from upper chambers. 4% paraformaldehyde and Crystal Violet Solution (Beyotime, Shanghai, China) were used to fix and stain the cells in lower chamber. Subsequently, five random fields were selected to count the migrated or invaded cells under microscope (Olympus, Tokyo, Japan).
Quantitative real-time PCR
Total RNAs in LoVo cells were isolated with TRIzol reagent (Invitrogen, Carlsbad, USA). PrimeScriptTM RT reagent Kit (Takara, Shiga, Japan) was used to accomplish the cDNA synthesis according to manufacturer’s protocol. Quantitative real-time PCR was carried out with ABI 7500 Real-Time PCR system (Applied Biosystems, Foster City, USA). –2∆∆Ct method was used to determine relative mRNA expression. Primer sequences used in this study were listed in Supplementary Table S1.
Statistical analysis
Experiments in this study were performed in triplicate, and results were presented as mean ± SD (standard deviation). Statistical significance between two groups was determined with unpaired Student’s t-test using SPSS 16.0 software. OS was compared with Kaplan–Meier survival curve, and statistical significance was evaluated with log-rank test. A value of
Results
SATB1 overexpression was associated with poor outcome of CRC
Normal colorectal epithelial cell line (FHC) and CRC cell lines (SW480, SW620, LoVo, HCT116, HT29) were used to measure the expression levels of SATB1. FHC cells expressed relatively low level of SATB1 protein, while CRC cells expressed higher SATB1 (Figure 1(a)). LoVo cells, derived from distant metastatic CRC, showed the highest expression level of SATB1 (Figure 1(a)). As a contrast, SW480 cells, derived from a primary CRC patient, expressed modest level of SATB1 (Figure 1(a)). Subsequently, effect of SATB1 on OS of CRC was determined using clinical data in TCGA. In 210 CRC cases, the expression of SATB1 showed notable correlation with OS of CRC patients (Figure 1(b)), suggesting that SATB1 expression might play an important role in CRC progression.

SATB1 overexpression in CRC cells and correlated with poor outcome of CRC. (a) Protein expression of SATB1 in control colorectal epithelial cell line (FHC) and CRC cell lines (SW480, SW620, LoVo, and HCT116HT29). Expression level was normalized to β-actin. (b) OS time of CRC patients as well as SATB1 expression.
SATB1 correlates with aggressive CRC migration and invasion
To investigate the role of SATB1 expression in CRC progression, we established a stable SATB1-knockdown LoVo cell line. Compared with that in negative control expressing siRNA (siNC), SATB1 protein expression was effectively downregulated in LoVo cells upon SATB1 siRNA (Figure 2(a)). Subsequently, the migration and invasion abilities of these two LoVo cell lines were examined. In migration assay, cell motility was significantly decreased in SATB1 knockdown cells (***

SATB1-knockdown inhibited CRC migration and invasion in vitro. (a) Immunoblot of SATB1 expression in LoVo cells upon siNC or siRNA transfection. Protein expression level was normalized to β-actin. (b) Migration analysis of LOVO cells expressing siNC or siRNA. (c) Invasion analysis of LOVO cells expressing siNC or siRNA.
SATB1 promotes EMT in CRC
EMT process played an essential role in CRC metastasis; therefore, effect of SATB1 disruption on EMT-associated gene expression was further analyzed. E-cadherin, a marker of epithelial cells, was upregulated. However, N-cadherin and Vimentin, two markers of mesenchymal cells, were downregulated in SATB1-knockdown LoVo cells (Figure 3(a) and (b)). Slug and Snail, two EMT-associated transcription factors, were also inhibited in SATB1-knockdown LoVo cells at mRNA and protein levels (Figure 3(a) and (b)), suggesting SATB1-promoted CRC migration and invasion by regulating EMT in CRC.
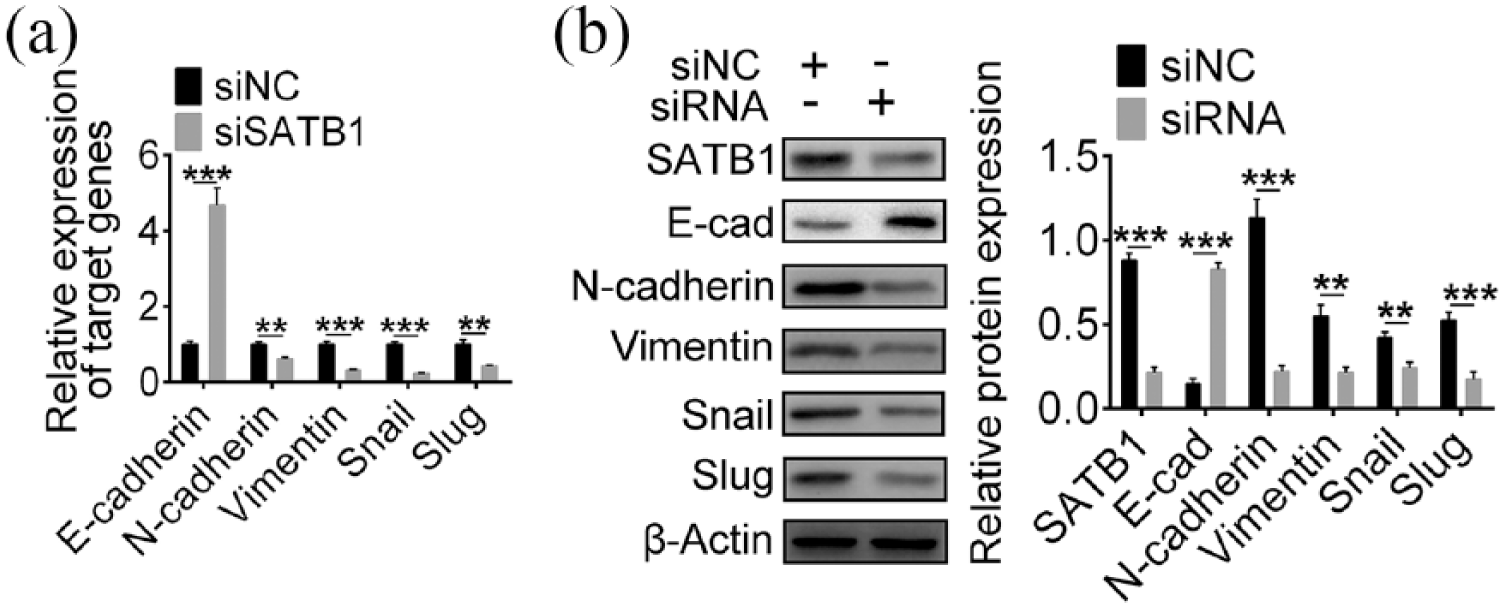
SATB1-knockdown suppressed EMT process in CRC cells. (a) qPCR analysis of mRNA level of E-cadherin, N-cadherin, Vimentin, Snail and Slug in LOVO cells upon siNC or siRNA transfection. (b) Immunoblot analysis of E-cadherin, N-cadherin, Vimentin, Snail and Slug in LOVO cells expressing siNC or siRNA transfection. Protein expression level was normalized to β-actin.
SATB1 interacts with Notch signaling pathway to regulate EMT
Abnormal activation of Notch signaling is related to tumorigenesis and EMT process in a variety of tumors.13,14 Therefore, the role of SATB1 expression in Notch signaling was investigated. NICD (intracellular domain of Notch) and Hes1 (hairy and enhancer of split-1), two important components in Notch signaling, were inhibited in SATB1-knockdown LoVo cells (Figure 4(a)). Jagged-1, a Notch ligand that could activate Notch signaling pathway, was used to evaluate the interaction between SATB1 and Notch signaling. In SATB1-knockdown LoVo cells, EMT process and Notch signaling were inhibited (Figure 4(b)). However, the stimulation of Jagged-1 could reverse the effect of SATB1 downregulation, accompanied with decreased E-cadherin expression and increased NICD, N-cadherin, Vimentin, Snail and Slug expression levels (Figure 4(b)). These results demonstrated SATB1-facilitated CRC metastasis via Notch signaling pathway.

SATB1 interacts with Notch signaling pathway to promote EMT. (a) Western blot analysis of NICD, Hes1, SATB1 in LOVO cells upon siNC or siRNA transfection. (b) Immunoblot analysis of E-cadherin, upregulation of NICD, N-cadherin, Vimentin, Snail as well as Slug in SATB1-knockdown LOVO cells upon Jagged-1 treatment. Protein expression level was normalized to β-actin.
Discussion
In present study, SATB1 overexpression was found in several CRC cell models, and SATB1 expression was closely associated with poor prognosis of CRC. EMT process, the initiation of cancer metastasis, could be induced by SATB1 overexpression through activating Notch signaling. It is reported SATB1 was overexpressed in CRC in vitro and in vivo.15,16 Furthermore, SATB1 has been reported to induce aggressive CRC symptoms, including advanced TNM stage, distant metastasis, and poorer prognosis. 16 Taken together with previous studies, our study demonstrated that SATB1 played a critical role in CRC progression.
EMT is considered as an essential process inducing CRC metastasis. 17 The typical characteristics of EMT are the decrement of E-cadherin (epithelial marker) and the increment of N-cadherin and Vimentin (mesenchymal marker). 18 In this study, SATB1 knockdown upregulated E-cadherin expression while downregulated N-cadherin and Vimentin expression, suggesting SATB1 was involved in the initiation of CRC metastasis. Transcription factors Snail and Slug are the key EMT regulators that determining the EMT progression. 19 Downregulation of Snail and Slug in SATB1-knockdown CRC cells indicated SATB1 could promote EMT progression as well.
As an important pathway during development, Notch signaling contributes to tumor angiogenesis and metastasis.14,20 In SATB1-knockdown cells, Notch signaling was inhibited, based on the expression of intracellular domain of Notch receptor NICD and the downstream target Hes1, illustrating that the upstream mediators of Notch signaling were regulated by SATB1. Interestingly, it has been reported SATB1 could bind to the promoter region of Notch 1 and thus activating the Notch signaling in dendritic cells. 21 In hypoxic environment, Notch signaling promoted EMT and tumor metastasis by upregulating Snail and Slug.22,23 Utilization of Notch ligand Jagged-1 could reverse the inhibitory effect on EMT process in SATB1-knockdown CRC cells, indicating that cancer metastasis induced by SATB1 depends on Notch signaling pathway. Taken together, SATB1 could enhance the malignant degree of CRC via Notch signaling pathway. However, the exact molecular mechanisms of the interaction between SATB1 and Notch signaling remain to be further explored. In conclusion, SATB1 is a critical regulator in migration and invasion of CRC through activating Notch signaling to promote EMT process, highlighting that SATB1 may act as a novel prognostic biomarker and a promising drug target for CRC therapeutics from bench to clinic.
Supplemental Material
Table_S1 – Supplemental material for SATB1 promotes epithelial-mesenchymal transition via Notch signaling pathway in colorectal cancer
Supplemental material, Table_S1 for SATB1 promotes epithelial-mesenchymal transition via Notch signaling pathway in colorectal cancer by Jun Tang and Jingfang Yang in European Journal of Inflammation
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/ or publication of this article: This work was supported by Open Subject of Key Laboratory of High Incidence Prevention and Treatment in Guangxi West Region: Effect of Folfox Neoadjuvant Chemotherapy on COX-2 Expression in Rectal Cancer. (Grant No. Kfkt2016002).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
