Abstract
Objective
We aimed to explore the role of structural maintenance of chromosomes 4 (SMC4) in malignant progression and immunology of colon adenocarcinoma (COAD).
Methods
The expression, genetic and protein features, and immune cell infiltration of SMC4 in pan-cancer were provided by public databases and websites. The protein expression of SMC4 in COAD tissues was screened by immunohistochemical assay. Si-RNA-mediated transfection was performed in COAD cells and the proliferation viability was measured using MTT, colony formation and EdU assays. Cell autophagy was detected by AO staining, western blots, and immunofluorescence staining. The migratory ability was determined using scratch and transwell assays. The expression of epithelial-to-mesenchymal transition (EMT) markers and transcriptional factors were detected using western blots.
Results
The expression of SMC4 was upregulated in pan-cancer and had relationships with prognosis, TMB, and MSI of cancer patients. Particularly, SMC4 protein was highly expressed in COAD tissues and correlated with poor prognosis of patients. Depletion of SMC4 inhibited cell proliferation, induced autophagy, and decreased migration through EMT progression in COAD cells. In addition, SMC4 was associated with infiltration of neutrophils, M2 macrophages, and CD4 + T cells in COAD, and had positive association with M2 macrophage markers and immune checkpoints.
Conclusion
SMC4 was correlated with patients’ poor prognosis, proliferation, metastasis, and immune cell infiltrates, and might function as a potential diagnosis and prognostic biomarker in COAD.
Keywords
Introduction
COAD is a relatively common malignancy of the digestive tract and one of the leading types of cancer that cause death worldwide. 1 In 2020, the global incidence rate was 10% and the mortality rate was 9.4%. 2 Due to the lack of effective biomarkers in the early diagnosis and treatment of COAD, patients with COAD are diagnosed in the advanced stage with poor prognosis. 3 Therefore, the further study of the abnormally expressed genes or proteins associated with COAD is of great significance for the early detection of COAD, the effective evaluation of prognosis, and the development of effective biomarkers.
Tumor microenvironment (TME) is a complex system composed of tumor cells, surrounding immune cells, inflammatory cells, fibroblasts, adjacent stromal tissue, and microvessels. 4 Inflammatory cells in TME play diverse roles in tumors, such as neutrophils, blocking the function of the immune cells of TME and promoting the growth of cancer cells. 5 Tumor-associated macrophages (TAMs) regulate immune escape and serve as an essential factor in tumor progression. 6 Studies have suggested that the pattern of immune cell infiltration could influence the prognosis and the clinical benefit of tumor immunotherapy.7,8 Immunotherapy, particularly immune checkpoint inhibitors (ICIs), has shown promising results in COAD and other tumor patients with mismatched repair defects or high levels of microsatellite instability (MSI-H).9,10 However, a considerable proportion of the patients had non-respond to ICIs. Therefore, the discovery of prognostic markers of immunotherapy efficacy and new immunotherapy targets can provide reliable directions for adjuvant therapy and improvement of COAD patient outcomes.
Structural maintenance of chromosomes 4 (SMC4) is one of the SMC family members, which is located in 3q25.33. This ATPase family maintains the stability of the chromosomal structure and participates in mitosis of eukaryotic cells. SMC4 is mainly involved in chromosome enrichment and separation, DNA recombination and repair.11–13 In recent years, studies have shown that SMC4 is a prognostic outlier gene with diagnostic value and therapeutic significance in cancers such as lung, liver, and breast cancer,14–16 which may be related to the dimers formed by SMC4 and SMC2, leads to genomic instability and abnormal gene regulation by interrupting the near chromatin binding.12,17 Ma et al. 16 found that SMC4 was upregulated in breast cancerous tissues and in T2-3N0 or ER/PR-positive patients, the higher mRNA expression level of SMC4 was associated with worse survival rates. SMC4 was reported to be corrected with glioma cell cycle and proliferation. 18 A similar function of SMC4 was found in liver cancer that SMC4 could affect cancer cell proliferation and migration through JAK2/STAT3 pathway. 19 Whereas, the role of SMC4 in COAD has rarely been reported. The main cause of death in patients with colorectal cancer is invasion and metastasis, 20 therefore, exploring and blocking the mechanism of SMC4 in COAD patients’ recurrence and metastasis may be the key to improving the survival of patients.
Herein, we exhibited the abnormal expression, genetic or proteic features of SMC4 in pan-cancer, and the role of SMC4 in malignant evolution and immune cell infiltration of COAD, the work partially was bioinformatics-based establishment combined with functional experimental validation. Our findings suggested that SMC4 could serve as a potential biomarker in diagnosis, predict prognosis and positive immunotherapeutic effects in COAD patients.
Materials and methods
Clinical specimens
The COAD tissue microarray was purchased from Shanghai Outdo Biotech Co. Ltd, included COAD tumor tissues and corresponding paracancerous tissues from 94 patients, who were selected at random and underwent routine treatment from January 2009 to October 2009, tissues were paraffin-embedded and met rigorous follow-up criteria. The follow-up period was 5.7-6.5 years. Clinical pathological parameters containing age, gender, tumor size, clinical stage, TNM stage, and 5-year survival, were seriously inspected and shown in Figure S1. The ages of the patients ranged from 27 to 90, with an average age of 58.5 years. 48:46 was the ratio of male to female. In conformity with the seventh edition of the American Joint Committee on Cancer, 21 COAD specimens were staged as early stage (I-II) and 36 as late stage (III-IV). The median survival time was 48 months. In addition, 78 cases from adjoining non-tumor tissues were obtained from the margin of the cancer resection in these patients. The present study was performed in accordance with the World Medical Association guidelines and approved by the Ethics Committee of Yanbian University College of Medicine and Shanghai Outdo Biotech Co. Ltd (Shanghai, China).
Cell culture and transfection
The COAD cell lines (SW620, SW480, HCT-116, HT-29, and Caco2) purchased from the ATCC were cultured in DMEM medium (Gibco, Gaithersburg, MD, USA) which supplemented with 10% FBS and appropriate antibiotics, and maintained in humidifies 5% CO2 at 37°C. Three kinds of SMC4 siRNAs (si-SMC4#1, si-SMC4#2, and si-SMC4#3) (RIBOBIO, China) were purchased. In the light of the knockdown effect, control siRNA, si-SMC4#1 and si-SMC4#3 were applied in this research. The sequence of siCon was 5′-CTTCCTGACAAACGAGTC-3′, si-SMC4#1 was 5′-CCACAAGAGTAGCATATCA-3′, si-SMC4#2 was 5′- GGATGATTATGAAGTCATT-3′, and si-SMC4#3 was 5′-GAAAGTCCTTGATGCAATA-3′. COAD cells (SW480 and HCT-116) were transfected with 30 nM of siRNA by adding Lipofectamine 3000 (Invitrogen, USA) as stated by the manufacturer’s protocol, and confirmed transfection efficiency by western blots.
Western blots
Western blots analysis was performed as previously described. 21 The primary antibodies against β-Actin (1:1000; cat. no. ab8226; Abcam), E-cadherin (1:500; cat. no. 3195S; CST), Vimentin (1:100; cat. no. 5741s; CST), Snail (1:50; cat. no. 3879S; CST), SMC4 (1:100; cat. no. 24758-1-AP; Proteintech), MMP-2 (1:50; cat. no. #A1035; Santa Cruz), MMP-9 (1:50; cat. no. #D2145; Santa Cruz), Beclin-1 (1:100; cat. no. 11306-1-AP; Proteintech), and LC3-II (1:1000; cat. no. 18725-1-AP; Proteintech) and horseradish peroxydase-coupled secondary antibody including anti-mouse (cat. no. ab97040) and anti-rat (cat. no. ab7097) (1:1000; Abcam) were used. Enhanced chemiluminescence (Millipore, China) was used to detect signals.
Immunohistochemistry (IHC)
IHC and the evaluation methods was performed as previously described in detail. 21 Tissues were deparaffinized using xylene, while rehydrated in graded alcohol washes. Antigen retrieval was performed by placing the slides in sodium citrate buffer, then endogenous peroxidase was blocked by 3% H2O2. The slides were subsequently incubated with the indicated antibody anti-SMC4 (1:200; cat. no. 24758-1-AP; Proteintech) followed by incubating with the secondary antibody, DAB was immunostained and hematoxylin was counterstained, the images were captured with Olympus BX51 light microscope (Olympus).
Colony formation assay
Cells (2000 cells) were seeded in 6-well plates in triplicate and incubated. After 2 weeks, cells were fixed with 4% paraformaldehyde and stained with hematoxylin followed by washing with PBS. The colonies (containing >50 cells) were counted directly and images were captured.
MTT assay
Approximately 2000 cells per well were seeded into 96-well plates, then added MTT (100 μL per well, Solarbio, China) at 0, 24, 48, and 72 h respectively. MTT was removed from the wells after 4 h and added 100 μL DMSO into each well. The OD value at 570 nm of absorbance was measured by Tecan Infinite 200 Pro microplate reader (Tecan, Switzerland). The relative number of viable cells was assessed by measuring the absorbance.
Scratch assay
Cells were seeded and cultured overnight. A sterile 200 μL pipette tip was used to create straight scratch through the monolayer, then removing cellular debris used PBS, and added a serum‐free medium. Cell migration was monitored at 0 h and 48 h, respectively, and images were captured using a light microscope (Eclipse 80i; Nikon Corporation).
Acridine orange (AO) staining
Cells were stained with 1 μg/mL−1 of AO (Macklin, Shanghai, China) for 15 min and washed with PBS at RT in the dark. The formation of acidic vesicular organelles (AVOs) was determined under a fluorescence microscope (AO, bright red fluorescence in acidic vesicles).
Immunofluorescence staining (IF)
Adherent cells were fixed with 4% paraformaldehyde, permeabilized with 0.5% Triton X-100 (CWBIO, China), and blocked with 3% BSA for 2 h. Cells were incubated with primary antibodies (Beclin-1 and LC3-II) at 4°C overnight, then incubated with Alexa Fluor 488-labeled goat anti-rabbit IgG secondary antibody (1:200, cat. no. A-11008; Invitrogen) for 1 h, nuclei were stained with DAPI solution and then captured by a fluorescence microscope (Leica SP5II).
Bioinformatics analysis
The abbreviations of tumor types used was according to OncoTree (https://oncotree.mskcc.org). The Human Protein Atlas (HPA, https://www.proteinatlas.org/), TIMER (https://cistrome.shinyapps.io/timer/), GEPIA (gepia.cancer-pku.cn), and UALCAN (https://uaLcan.path.uab.edu) databases were used to explore SMC4 expression in pan-cancer. Sangerbox (https://sangerbox.com/home.html) drew the relationship between SMC4 expression and patients’ survival, TMB, or MSI, and the receiver operating characteristic (ROC) curve, Sankey diagram in pan-caner and COAD. GSCA (https://bioinfo.life.hust.edu.cn/GSCA/#/) was used to show the relationship between SMC4 expression and patients’ stages. The cBioportal database (https://www.cbioportal.org/) contained large-scale cancer genomics data, the mutation frequency, copy number alteration, and mutation type of SMC4. PhosphoSitePlus (https://www.phosphosite.org/homeAction.action) provided altering mutations in SMC4 protein. Data of Go enrichment analysis was from LinkedOmics (https://linkedomics.org/admin.php) and STRING database (https://string-db.org/). Immune cell infiltration in cancer was searched on TISIDB website (https://cis.hku.hk/TISIDB/index.php), TIMER, Sangerbox, and Home for Researchers (https://www.home-for-researchers.com/static/index.html#/).
Statistical analysis
The data analysis was performed by SPSS 25.0 software and GraphPad Prism 8.0 software. Paired T-test for independent means was used for group comparisons. Kaplan-Meier analysis calculated the survival curves. Univariate and multivariate hazard ratios of study variables were examined by Cox proportional hazards regression model. Group comparisons for continuous data were done by one-way ANOVA and followed by Tukey’s post hoc test. Biochemical experiments were performed in triplicate and at least three independent experiments were evaluated. The value of
Results
SMC4 was abnormally expressed in human pan-cancer
The HPA database revealed that SMC4 is highly expressed in most cancer tissues of patients, especially in colorectal and pancreatic cancers (Figure 1(A)). Analysis from the TIMER database also indicated that compared with normal tissues, SMC4 expression level was higher in a variety of tumor subsets, including BLCA (bladder urothelial carcinoma), BRCA (breast invasive carcinoma), CHOL (cholangiocarcinoma), COAD (colon adenocarcinoma), ESCA (esophageal carcinoma), HNSC (head and neck squamous cell carcinoma), KIRC (kidney renal clear cell carcinoma), LIHC (liver hepatocellular carcinoma), LUAD (lung adenocarcinoma), LUSC (lung squamous cell carcinoma), READ (rectum adenocarcinoma), SKCM (skin cutaneous melanoma), STAD (stomach adenocarcinoma) and UCEC (uterine corpus endometrial carcinoma) compared with the normal tissues (Figure 1(B), all The expression of SMC4 in human pan-cancer and the association between SMC4 expression level and survival of patients with cancers. (A) Expression of SMC4 in pan-cancers from HPA database. (B, C, D) Expression of SMC4 in pan-cancers compared with normal tissues from TIMER, GEPIA, and UALCAN databases. (E) Expression of SMC4 in different cancer cell lines from CCLE database (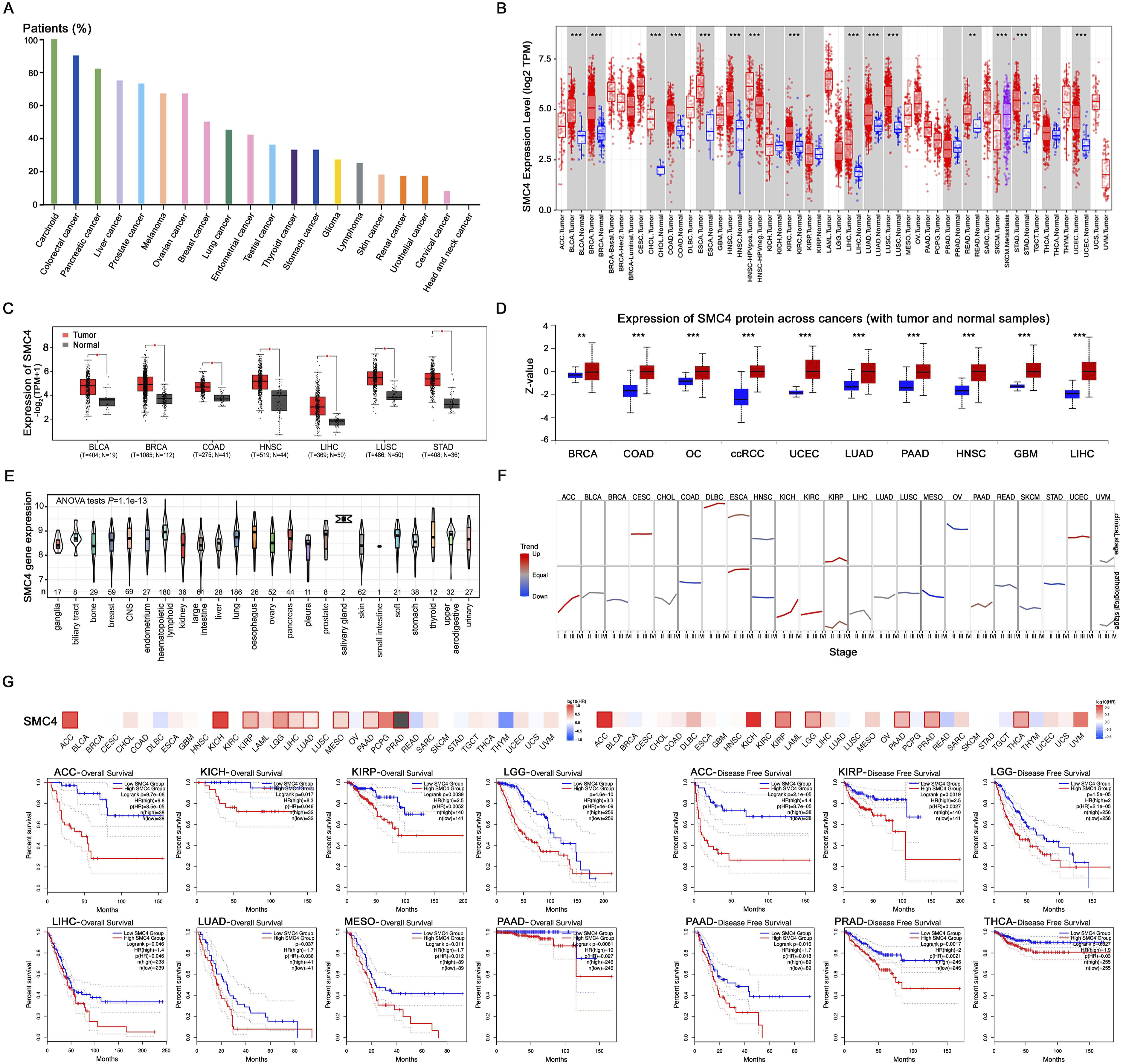
Genetic and protein features of SMC4 in pan-cancer
Genetic and protein alterations of SMC4 may cause its abnormal expression in cancers. According to the retrieval results of cBioportal website, we found that SMC4 has a high alteration frequency and mainly appears as amplification and mutation, especially in LUSC (23.41%) and ESCA (13.74%) (Figure 2(A)). The frequency of genetic alterations in all tumor types was shown as oncotree code in Figure 2(B). Genetic alteration was further detected among amino acids 0 and 1288aa of SMC4 gene, it was found that the mutation type missense was regarded as the dominant factor except for a variant of undetermined significance (VUS) mutation (Figure 2(C)). R1252 was the most frequent mutation site of SMC4, which was detected in UCEC ( Genetic and protein feature of SMC4 in pan-cancer. (A, B) Genetic alteration type and frequency of SMC4 in pan-cancer of TCGA by cBioportal website. (C) Mutation sites (up) of SMC4, which contained 212 missense, 36 truncating, 1 inframe, and 5 splice mutations, and PTM (down) of SMC4, which contained phosphorylation, acetylation, ubiquitination, and methylation in cBioportal website. (D) 3D structure showed the R1252 mutation site of SMC4 in cBioportal website. (E) Proportion of tumors with altering mutations of SMC4 protein in PhosphoSitePlus database. (F) The correlation analysis between SMC4 expression in pan-cancer and TMB was described in Sanger Box. (G) The correlation analysis between SMC4 expression in pan-cancer and MSI was described in Sanger Box.
High expression of SMC4 was associated with poor prognosis in COAD patients
Based on the genetic alteration of SMC4 and the significant association with TMB and MSI in COAD, we further explored the clinical value of SMC4 in COAD. It was discovered that the mRNA expression of SMC4 was markedly increased in COAD tissues from GEO DataSet (GDS4515/201664_at, The expression of SMC4 in COAD and the association between SMC4 expression level and clinicopathological features or survival of patients with COAD. (A) Box plots derived from gene expression data in GEO DataSet (GDS4515/201664_at) and TCGA database comparing the mRNA expression of SMC4 in normal and COAD tissues (both 
Depletion of SMC4 reduced the proliferation, colonization, and induced autophagy in COAD cells
Then we analyzed the co-expression gene and functions of SMC4 in COAD using the LinkedOmics website. As the volcano plot shown in Figure S2(C), 3293 genes had significant positive correlation with SMC4, whereas 3129 genes had negative correlation with SMC4 in COAD (false discovery rate, FDR <0.01). The top 20 genes which correlated with SMC4 were shown in the sequence map, including positively connected with cellular proliferation-related gene MKI67, telomerase RNA component (TERC), and negatively connected with epithelial cell migration and tissue repair-related gene LUM (Figure S2(D)). Followed by performing biological process, molecular function, protein domains, and pathway enrichment analysis, it was explored that SMC4 related to kinetochore and membrane organization process, histone, threonine and serine kinase activity, and cell cycle pathway in COAD (Figure 4(A)). These findings provided the basis for the follow-up studies which revealed the biological function of SMC4 in COAD. SMC4 was related to proliferation, colonization and autophagy of COAD cells. (A) Classification for SMC4 related process and functions in COAD. A: biological process; B: molecular function; C: protein domains; D: pathways. (B) Western blots confirmed the protein expression of SMC4 in COAD cells. β-Actin was used as a loading control. (C). Different si-SMC4 sequence transfection efficiency was detected by Western blots. β-Actin was used as a loading control. (D-F) Ability of cell proliferation was examined by MTT (D), colony formation (E) and EdU (×100) (F) in the constructed cells. (G) The formation of autophagosomes was evaluated by AO staining. Magnification: 200x. (H) Expression of autophagic process markers LC3-II and Beclin-1 was confirmed by Western blots in the constructed COAD cells. (I) IF staining of LC3-II and Beclin-1 in the constructed COAD cells. Magnification: 400x. Data were represented as mean ± standard error of at least three independent experiments, *
To elucidate the regulation of SMC4 on the biological function of COAD, we first detected the protein expression of SMC4 in COAD cell lines (SW480, SW620, HCT-116, HT-29, and Caco2) and found that SMC4 was upregulated in SW480 and HCT-116 cells (Figure 4(B)). Then, SW480 and HCT-116 were transfected with SMC4-specific si-RNAs (Figure 4(C) and S2(E)) and confirmed that silencing of SMC4 expression significantly inhibited the growth of COAD cells by MTT assay (Figure 4(D)). In the colony forming experiment, SMC4 depletion decreased the colonization of COAD cells (Figure 4(E)). In addition, EDU assay showed that SMC4 depletion inhibited the ability of DNA replication in COAD cells (Figure 4(F)). Therefore, these studies showed that SMC4 depletion impaired the proliferation and colonization of COAD cells. Under different conditions, autophagy could enhance cell survival or increase cell death. Then, AO staining was performed and showed orange-colored autophagosomes were observed following the depletion of SMC4 (Figure 4(G)), indicating that SMC4 could induce COAD cells to generate the autophagic response. The expression of autophagy markers (LC3-II and Beclin-1) was activated and upregulated in SMC4 depleted cells (Figure 4(H) and S2(F)). IF staining further verified our findings (Figure 4(I)). These results revealed that SMC4 could regulate proliferation, colonization, and autophagy in COAD cells.
Suppression of SMC4 inhibited the migration of COAD through EMT
High SMC4 expression was associated with epithelial cell migration and LN metastasis in COAD patients, according to the results of IHC staining and public database analysis. Hence, we further explored the function of SMC4 in COAD cell migration. Through the scratch and transwell assays, we found that the downregulation of SMC4 expression led to the inhibition of COAD cell migratory ability (Figure 5(A) and (B)). Studies have shown that EMT promotes the metastatic potential of cancer cells. Therefore, we examined the effect of SMC4 on EMT related markers. Western blots showed that the expression of E-cadherin increased, vimentin, Snail, and MMP2, MMP9 decreased in SMC4 silenced cells inversely (Figure 5(C)). It was indicated that SMC4 induces the metastatic process of COAD by activating EMT. Depletion of SMC4 promoted the COAD cells migration via EMT. (A) The rate of migration into scratched area in the constructed cells was detected by scratch assay. (B) Transwell migration assay detected the effect of SMC4 expression on migratory ability of COAD cells. (C) The effect of SMC4 differential expression on EMT markers (E-cadherin, Vimentin, Snail, MMP-2, MMP9) was tested by Western blots in COAD cells. β-Actin was used as a loading control. **
SMC4 modulated immune infiltration in COAD
TISIDB web portal showed that the expression of SMC4 was significantly associated with different immune subtypes of COAD, especially in lymphocyte-depleted and immunologically quiet types (Figure 6(A)), which may be related to the involvement of SMC4 in immune cell infiltration. TIMER website displayed that SMC4 expression was related to the infiltration of immune cells in colorectal cancer, including dendritic, CD8+T, and B cells (Figure 6(B)). It was also explored that the expression of SMC4 was associated with infiltration of neutrophils, M2 macrophages, and CD4+T cells in COAD using QuanTIseq computational pipeline in Sanger Box (Figure 6(C)). Kaplan-Meier plots in TIMER website showed that low infiltration of macrophages indicated a favorable prognosis in colorectal cancer patients, however, infiltration of other immune cells is not associated with survival of patients (Figure 6(D) and S2(G)). Importantly, the correlation analysis of SMC4 and immune cell markers was assessed, and displayed that SMC4 has a positive association with M2 macrophage markers (ARG2, MRC1, and CD163, The relationship between SMC4 and immune cell infiltration in COAD. (A) Expression of SMC4 in different immune subtypes of COAD in TISIDB web portal (n = C1: 332, C2: 85, C3: 9; C4: 12. C5: 0, C6: 3; C1: wound healing, C2: IFN-gamma dominant, C3: inflammatory, C4: lymphocyte depleted, C5: immunologically quiet, C6: TGF-b dominant). (B) Relationship between SMC4 expression and infiltration of immune cells in colorectal cancer in TIMER database. (C) Score of immune cell infiltration detected by scRNA-seq methods in Singer Box. (D) Kaplan-Meier analysis was used to compare the high and low infiltration groups of macrophages and neutrophils using the TIMER algorithm. (E) Correlation analysis of SMC4 expression and immune cell markers in Home for Researchers. (F) Relationship between SMC4 expression and immune checkpoints was analyzed in Home for Researchers. Red represented COAD patients with high expression of SMC4 and blue represented COAD patients with low expression of SMC4.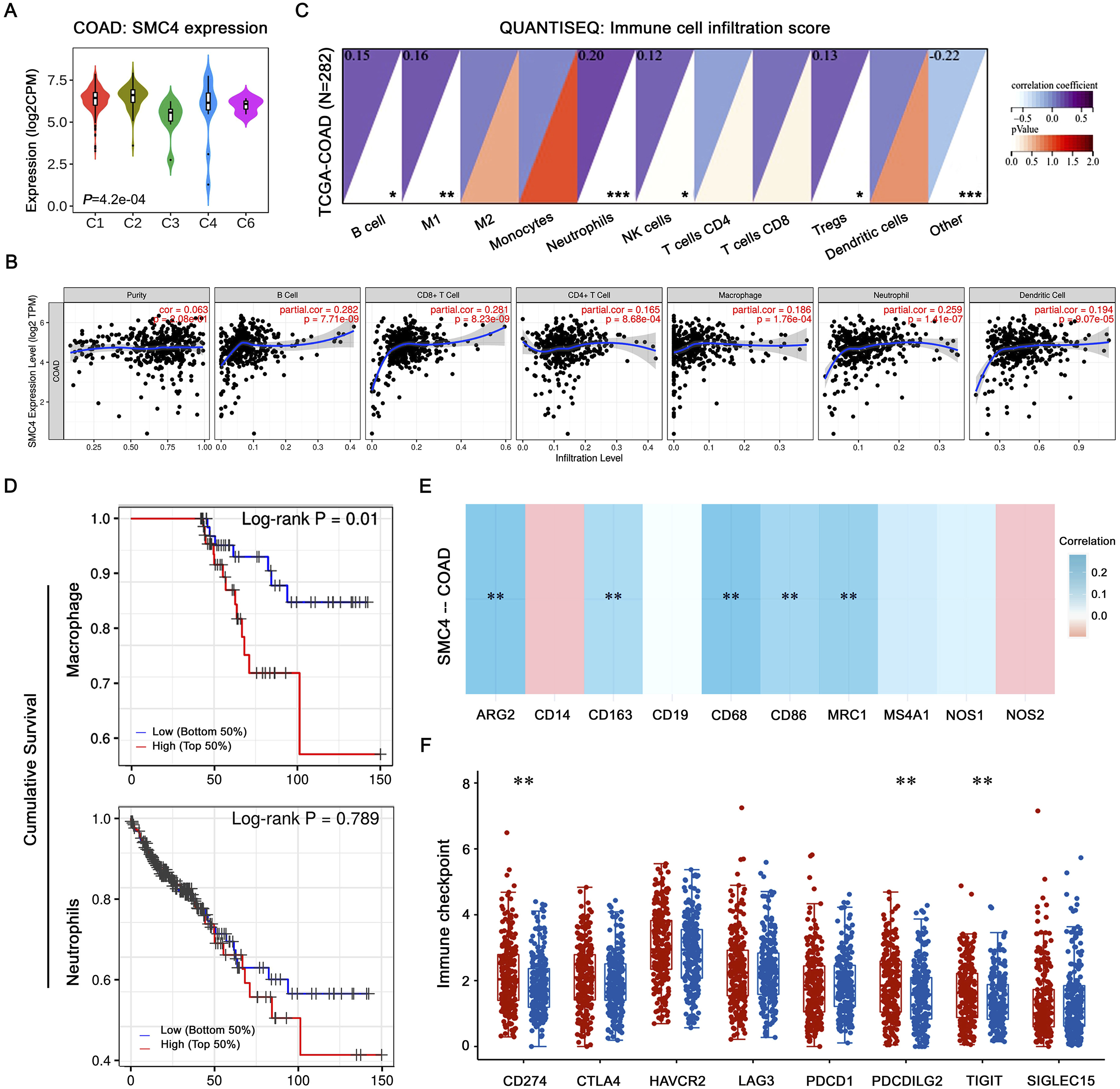
Discussion
In the current study, we summarized specific results through bioinformatics analysis that SMC4 predicted the poor prognosis of patients with ACC, KIRP, LGG, and PAAD, and combined with previous studies,22,23 hinted that SMC4 could be a prognostic indicator in cancers.
We also found SMC4 has high mutation and amplification rates in tumors, the most frequent mutation site was R1252 VUS mutation and occurred in UCEC and COAD. The amplification had high alteration frequency in LUSC, ESCA and OV. In Zhu
Similar to the results of database and Feng
ICIs have demonstrated significant clinical effects on patients with MSI-H colorectal cancer,
9
whereas, a low treatment response rate has limited their practical application. Tumor progression and the efficacy of immunotherapy are strongly influenced by the composition and abundance of immune cells in the TME.
36
Wang
Some limitations of this study should be noted. First, we found that SMC4 was modulated immune infiltration, especially M2 macrophages in COAD, however, it’s still need evidence from the results of experiments in vitro and
Conclusion
SMC4 overexpression is significantly related to the poor prognosis of patients and the immune microenvironment of COAD. Depletion of SMC4 inhibited proliferation, migration, and EMT of COAD cells. The clinical implications of these findings lie in providing a potential diagnostic and prognostic marker and a novel antitumor strategy for COAD.
Supplemental Material
Supplemental Material - SMC4 serves as a potential marker for the diagnosis and prognosis of colon adenocarcinoma
Supplemental Material for SMC4 serves as a potential marker for the diagnosis and prognosis of colon adenocarcinoma by Xinyue Wang, Wenxin Cheng, Ying Liu, Liang Ma, Yao Sun, Hongzhen Wang, Haifeng Liu, Li Nan, Yang Yang and Dawei Yang in International Journal of Immunopathology and Pharmacology.
Footnotes
Author contributions
D.-W.Y. and Y.Y. were involved in the conception and design of the study, X.-Y.W., W.-X.C. and Y.L. performed in vitro experiments, bioinformatics analysis and drafting of the manuscript; L.M., N.L. and Y.S. performed data analysis; H.Z.-W., H.-F.L., and Y.Y. were manuscript revisions. All authors listed approved the final version of the manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (No. 82160552), the Project of Science and Technology Department of Jilin Province (No. YDZJ20221ZYTS245), the Startup Foundation for Doctors (Science and Technology Administration Contract in 2019, No.20) and Application Basic of Yanbian University and the Science and Technology Project of Wenzhou, China (No.Y20210080).
Ethical statement
Data Availability statement
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
