Abstract
Objectives:
To identify potential candidate proteins which are secretory in nature and present at a higher abundance in oral cancer patients with tobacco habits.
Methods:
Conditioned media of tobacco-treated and -untreated non-neoplastic oral keratinocytes were analyzed using iTRAQ-based mass spectrometry. Hypersecreted proteins; SPARC (osteonectin), cwcv and kazal like domains proteoglycan 1 (SPOCK1); prosaposin (PSAP); and protein tyrosine phosphatase, receptor type S (PTPRS) were validated by enzyme-linked immunosorbent assay (ELISA) using saliva samples from oral cancer patients who are tobacco users.
Results:
Proteomic analysis of tobacco-treated and -untreated cells led to the identification of 2873 proteins. Among these, 378 proteins showed high abundance and 253 proteins showed low abundance (2-fold cutoff) in conditioned-media of tobacco-treated cells. ELISA-based validation showed significantly higher levels of SPOCK1, PSAP, and PTPRS in oral cancer patients with tobacco chewing habits compared to healthy controls. However, PSAP showed low specificity compared to SPOCK1 and PTPRS.
Conclusions:
This study indicates significantly increased levels of SPOCK1, PSAP, and PTPRS in saliva of oral cancer patients with tobacco habits. These protein biomarkers might be useful to identify tobacco users with high risk of developing oral cancers.
Introduction
Oral cancer is the third most common cancer in India and this high prevalence can be attributed to the consumption of smokeless tobacco. 1,2 Despite advances in treatment modalities, the 5-year survival rate has been 40–50% which has not improved over the last three or four decades. 3 This poor survival rate is predominantly due to late diagnosis of the disease, which is usually presented at an advanced stage. The risk of development of oral cancer among tobacco users (chewers) is higher than nonusers. Also, significant differences of tumor morphology and behavior exist in oral cancer patients between tobacco users and nonusers. Tumors of oral cavity in tobacco users have been found to be poorly differentiated and more aggressive compared to nonusers of tobacco. 4,5 In addition, tobacco users show poor prognosis and poor survival with high rate of tumor recurrence. 6,7 Thus, there is a need to develop reliable and easily accessible biomarkers of clinical relevance to identify tobacco users who are at a higher risk of developing oral cancer. This is especially important in developing countries like India with a high population prevalence of smokeless tobacco users. 1
Saliva being the proximal fluid, several efforts have been undertaken to identify salivary biomarkers for the early diagnosis of oral cancer. In a recent study by Ishikawa et al., metabolites were found to be altered in saliva of oral cancer patients. 8 Profiling of microRNAs from oral cancer patients and healthy individuals revealed overexpression of miRNA-21 and miRNA-184. 9 Microarray analysis and polymerase chain reaction-based validation showed upregulation of several genes in saliva samples from oral cancer patients compared to healthy individuals. 10 In addition, several proteomic studies have been carried out to identify salivary biomarkers. More than 3400 proteins have been reported in saliva. 11 Hu et al. carried out the proteomic profiling of saliva samples from oral squamous cell carcinoma (OSCC) patients and healthy individuals and identified a panel of potential biomarkers which can be used in the diagnosis of oral cancer. 12 Wu et al., demonstrated the potential utility of salivary resistin (RETN) in the diagnosis of oral cancer. 13 Recently, Yu and group developed a panel of four saliva-based biomarkers to distinguish oral cancer samples from non-OSCC individuals. 14 Most oral cancer patients with tobacco chewing habits consume tobacco for several years before the onset of disease. Early detection biomarkers that could identify tobacco users with high risk of developing oral cancer will be extremely useful.
We developed an
Materials and methods
Adaptation of OKF6/TERT1 cells to chewing tobacco
Normal oral keratinocytes, OKF6/TERT1, were a gift from Dr James Rheinwald (Brigham and Women’s Hospital, Boston, Massachusetts, USA). Cells were authenticated by short tandem repeat analysis. Cells were cultured and treated with chewing tobacco extract as described previously. 15 In this article, OKF6/TERT1 cells not treated with tobacco (parental) are referred to as OKF6/TERT1 and tobacco-treated cells as OKF6/TERT1-tobacco.
Collection of secretome or conditioned media
OKF6/TERT1 and OKF6/TERT1-tobacco cells were grown to 80% confluence and starved overnight in media devoid of growth factors. Prior to starvation, cells were washed with 1X phosphate-buffered saline multiple times for complete removal of media with growth factors. Conditioned media from both cells was collected and centrifuged to remove cellular debris followed by filtration using 0.22 µm filter. Conditioned media was then concentrated using Amicon Ultra-15 centrifugal filter tubes with a 3 kDa membrane filter (Merck Millipore, Billerica, Massachusetts, USA) to remove salts and other small molecules. Protein estimation of the samples was carried out using bicinchoninic acid assay. 18
In-solution digestion and iTRAQ labeling
In-solution digestion and iTRAQ labeling of processed secretome samples was carried out as described previously. 17 Briefly, equal amounts of protein from secretome of OKF6/TERT1 and OKF6/TERT1-tobacco cells were reduced at 60°C with tris(2-carboxyethyl) phosphine for 1 h. Subsequently, the samples were alkylated using methyl methanethiosulfonate for 10 min at room temperature. Samples were then digested using trypsin (Promega, Madison, Wisconsin, USA) for 12 h at 37°C. Peptides from secretome of OKF6/TERT1 cells were labeled with iTRAQ reagent with reporter tags of m/z 114 and 115 and those from OKF6/TERT1-tobacco cells with reporter tags of m/z 116 and 117. Post labeling, the samples from both the conditions were pooled and subjected to fractionation.
Strong cation exchange chromatography
Strong cation exchange chromatography fractionation of iTRAQ labeled sample was carried out as described previously. 17 Pooled sample was diluted to 1 ml with solvent A (10 mM KH2PO4, 25% (v/v) ACN, pH 2.7). The pH of sample was adjusted to 2.7 using ortho-phosphoric acid. The peptides were loaded on PolySULFOETHYL A column (PolyLC, Columbia, Maryland, USA; 5 µm, 200 Å, 200 × 2.1 mm) using an Agilent 1260 Infinity series HPLC system (Agilent Technologies, Santa Clara, California, USA). Peptides were fractionated using a 50 min gradient from 0% to 40% solvent B (350 mM KCl in solvent A). A total of 96 fractions were collected and further pooled into 23 fractions based on peptide abundance reflected by chromatographic peaks. The pooled fractions were vacuum dried and desalted using C18 StageTips and stored at −20°C till further analysis.
LC-MS/MS analysis
LTQ-Orbitrap Velos mass spectrometer (Thermo Fisher Scientific, Bermen, Germany) interfaced with Proxeon Easy nLC system (Thermo Fisher Scientific, Bremen, Germany) was used for proteomic analysis. Peptides were enriched on a trap column (75 µm × 2 cm) packed in-house using C18 material (Magic C18AQ, 5 µm, 100 Å, Michrom Biosciences Inc., San Jose, CA, USA) with a flow rate of 3 µl/min using solvent A (0.1% formic acid) and resolved on an analytical column (75 µm × 10 cm, Magic C18AQ, 3 µm, 100 Å, Michrom Biosciences Inc.) at a flow rate of 350 nl/min using a linear gradient of 7–30% solvent B (95% acetonitrile, 0.1% formic acid) over 70 min. MS and MS/MS scans were acquired with a mass resolution of 60,000 and 15,000, respectively at 400 m/z using the Orbitrap mass analyzer. Precursor MS scan was set to m/z 350–1800. In each duty cycle, 20 most intense monoisotopic precursors were selected for MS/MS fragmentation using higher energy collision dissociation mode at 41% normalized collision energy. Isolation width was set to 1.9 m/z. Singly charged and unassigned charge precursor ions were rejected. Dynamic exclusion setting was enabled and acquired ions were excluded for 45 s. The automatic gain control for full MS and MS/MS were set to 1 × 106 and 5 × 104 ions, respectively. The maximum ion injection time was set to 100 ms for MS and 250 ms for MS/MS scans. The lock mass option was enabled using polydimethylcyclosiloxane ions (m/z, 445.120025) for internal calibration.
Data analysis
Sequest and Mascot (version 2.2.0, Matrix Science, London, UK) search algorithms through Proteome Discoverer (version 1.4.0.288, Thermo Scientific, Bremen, Germany) suite were used to search raw data against NCBI RefSeq human protein database (version 59 containing 36,208 protein sequences and known contaminants). Trypsin as proteolytic enzyme with one missed cleavage and oxidation of methionine as variable modification were included as search parameters. The static modifications included alkylation (methylthio) at cysteine and iTRAQ modification at N-terminus of the peptide and lysine. Mass tolerances of precursor and fragment ions were set to 20 ppm and 0.1 Da, respectively with a false discovery rate of 1% to report identification. The average of reporter ion intensities from technical replicates was used for iTRAQ quantitation. The data were normalized based on protein median. Subcellular localization and biological process associated with each protein was obtained from Human Protein Reference Database, a GO-Gene Ontology compliant database. 19
Data submission
Proteomics data generated in this study have been deposited to the ProteomeXchange Consortium via the PRIDE public data repository and can be accessed at http://www.proteomexchange.org with the dataset identifier PXD006523. Alternatively, the data can also be accessed through Human Proteinpedia at http://www.humanproteinpedia.org/data_display?exp_id=00857.
Validation of candidate proteins by ELISA
Unstimulated saliva samples from oral cancer patients were obtained from the Department of Oral Pathology and Microbiology, Institute of Dental Sciences, Odisha and Department of Biochemistry, Postgraduate Institute of Medical Education and Research, Chandigarh, and Dr. Ahmed Dental College and Hospital, Kolkata after informed consent from patients. Saliva samples from individuals with no habits of tobacco or alcohol use and no symptoms of any disease were considered as normal. The subjects were asked to refrain from eating at least 1 h before collection and asked to rinse his/her mouth with 10 ml sterile saline. Protease inhibitor cocktail (1 μl/ml whole saliva; Sigma, St. Louis, MO, USA) was added to saliva immediately after collection to prevent any protein degradation. The samples were then centrifuged to remove debris and the supernatant was collected and stored at −80°C until further use.
ELISA-based validation was carried out using saliva samples from healthy individuals and oral cancer patients who are tobacco users. Custom ELISA plates for prosaposin (PSAP), SPARC/osteonectin, cwcv, and kazal-like domains proteoglycan (testican) 1 (SPOCK1), and protein tyrosine phosphatase, receptor type S (PTPRS) were purchased from Cloud-Clone Corp., Houston, USA. ELISA-based validation of these proteins was carried out as per manufacturer’s instructions. Briefly, 100 µl of standard dilutions and 100 µl of diluted saliva samples were added to appropriate wells. The plates were sealed and incubated for 2 h at 37°C. Post-incubation liquid from each well was removed. Detection reagent A (100 µl) was added to each well, covered with plate sealers and incubated at 37°C for 1 h. Solution from each well was aspirated and wells were washed thrice with wash buffer. Detection reagent B (100 µl) was added and incubated at 37°C for 30 min. After incubation, the solution was aspirated and washed thrice with wash buffer. After washing, 90 µl of substrate solution was added and incubated at 37°C until color developed (for 15–25 min, not exceeding 30 min). The reaction was stopped by adding 50 µl of stop solution and absorbance was read at 450 nm. Statistical analysis of the data was carried out using MedCalc for Windows, version 17.5.3 (MedCalc Software, Ostend, Belgium, Europe).
Results
Secretome analysis of OKF6/TERT1 and OKF6/TERT1-tobacco cells
Chewing tobacco is one of the leading causes of oral cancer and yet there are no biomarkers either for early detection or to identify the high risk population prone to develop oral cancer, especially among tobacco chewers. Our group has demonstrated that chronic exposure to chewing tobacco results in cellular transformation and confers oncogenic potential to normal oral keratinocytes. 15 In this study, we profiled the secretome of oral keratinocytes chronically treated with chewing tobacco compared to the parental cells. Using iTRAQ-based LC-MS/MS approach, we identified a total of 2873 proteins. The experimental work flow followed is depicted in Figure 1. We observed 378 proteins that showed higher abundance in secretome of tobacco treated cells and 253 proteins that showed decreased abundance. Complete list of proteins and their peptides identified in the study are provided in Online Supplemental Tables S1 and S2, respectively. A partial list of proteins that were detected at a higher or lower abundance in response to chewing tobacco treatment is provided in Table 1. Among the hypersecreted proteins identified, we validated the levels of SPOCK1, PSAP, and PTPRS in saliva samples from OSCC patients compared to healthy individuals. These proteins were chosen if they possessed any of the following criteria: (i) prior evidence of association of these molecules with cancer; (ii) prior knowledge of presence of these molecules in saliva; and (iii) availability of commercially available antibodies or ELISA kits.
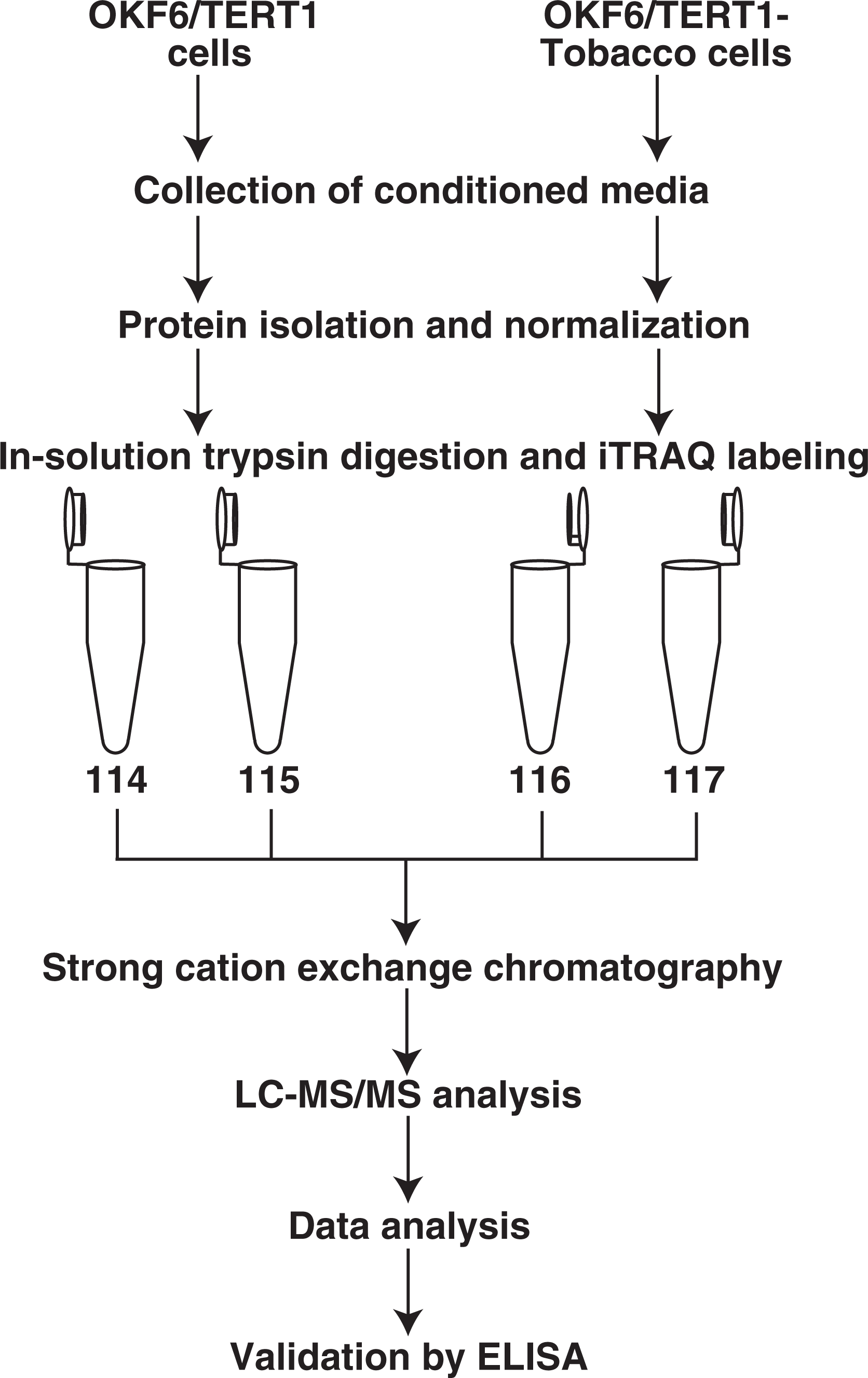
Workflow employed for discovery and validation of proteins secreted in response to chewing tobacco. Conditioned media from untreated and tobacco-treated OKF6/TERT1 cells was collected after overnight starvation in media devoid of growth factors. Proteins from secretome of OKF6/TERT1 and OKF6/TERT1-tobacco cells were isolated, normalized, and subjected to trypsin digestion. This was followed by iTRAQ labeling and strong cation exchange chromatography. The fractionated samples were then subjected to LC-MS/MS analysis. A subset of hypersecreted proteins were validated by ELISA.
Partial list of proteins found to be secreted at higher and lower levels in response to tobacco treatment.a.
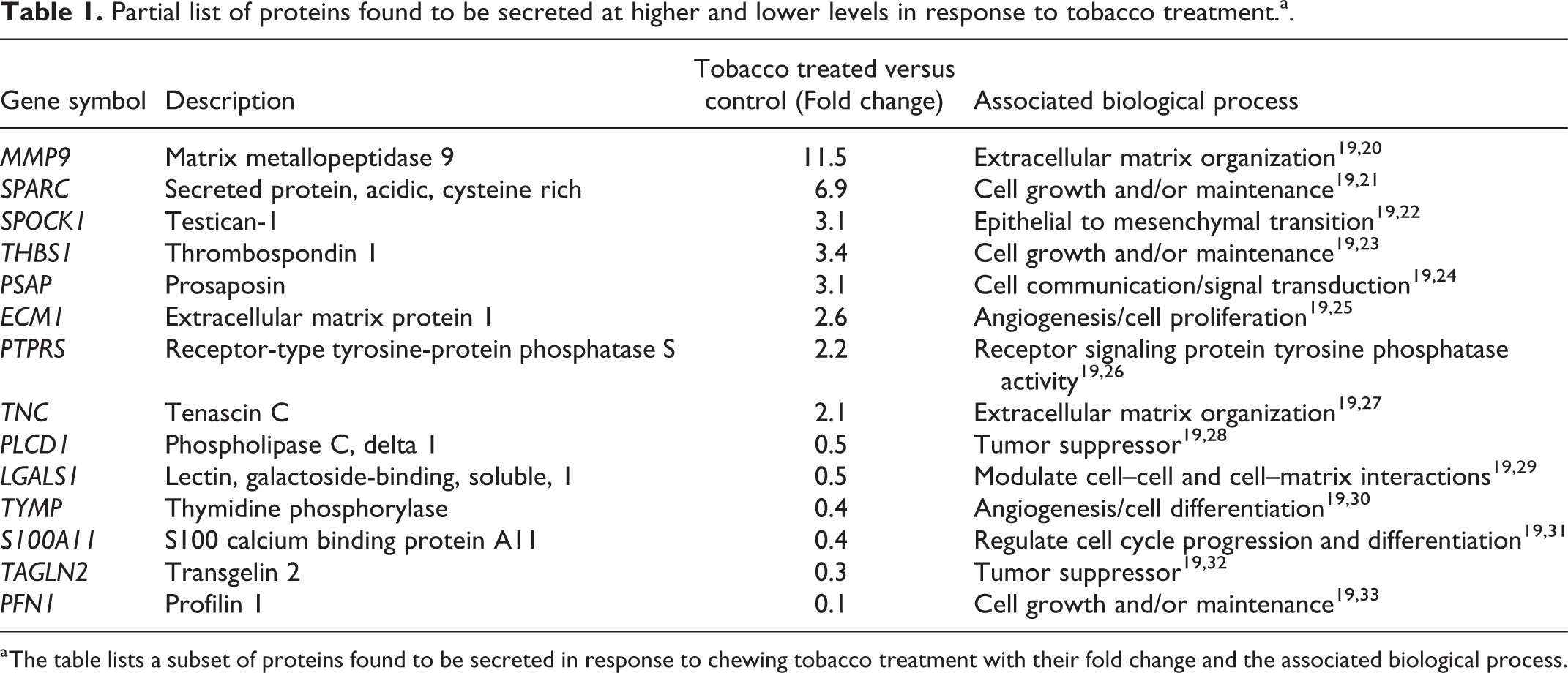
a The table lists a subset of proteins found to be secreted in response to chewing tobacco treatment with their fold change and the associated biological process.
SPOCK1 and PSAP are extracellular matrix proteins and are known to play a role in tumor growth and progression . 22,34 PTPRS is a receptor-type protein tyrosine phosphatase (PTP) and its epigenetic inactivation is shown to induce metastasis. 35 We evaluated the expression levels of these proteins in saliva samples of OSCC patients (who are tobacco users) compared to healthy individuals (who are nonusers of tobacco). This will enable us to determine whether the secreted form of these proteins could serve as early diagnostic markers to identify tobacco users who are at higher risk of developing oral cancer.
SPOCK1, PSAP, and PTPRS are hypersecreted in saliva of oral cancer patients
Mass spectrometry-based analysis of secretome from OKF6/TERT1-parental and tobacco-treated cells revealed over abundance of members of SPARC family proteins such as SPARC (6.9-fold), follistatin-like 1 (FSTL1, 4.7-fold) and SPARC (osteonectin), cwcv and kazal like domains proteoglycan 1 (SPOCK1) (3.1-fold). Members of this family are known to play a role in regulating cell proliferation, cell–cell adhesion, and migration. SPARC, SPOCK1, and FSTL1 are extracellular matrix proteins which are shown to be associated with tumor progression and metastasis.
22,36,37
SPOCK1 has been reported to be overexpressed in several cancers including colorectal cancer and head and neck squamous cell carcinoma (HNSCC).
38
SPOCK1 has also been reported to induce epithelial to mesenchymal transition (EMT) and drug resistance.
39
SPOCK1 has also shown to be overexpressed in HNSCC tissues and knock down of SPOCK1 inhibited cellular proliferation.
40
SPOCK1 was found to be hypersecreted (3.1-fold) in OKF6/TERT1-tobacco cells. A representative MS/MS spectrum of SPOCK1 is shown in Figure 2(a). ELISA-based validation of SPOCK1 was carried out using saliva samples from oral cancer patients who were tobacco users (

Representative MS/MS spectra of (a) SPOCK1, (b) PSAP, and (c) PTPRS. SPOCK1: SPARC (osteonectin), cwcv and kazal like domains proteoglycan 1 (SPOCK1); PSAP: prosaposin; PTPRS: protein tyrosine phosphatase, receptor type S.
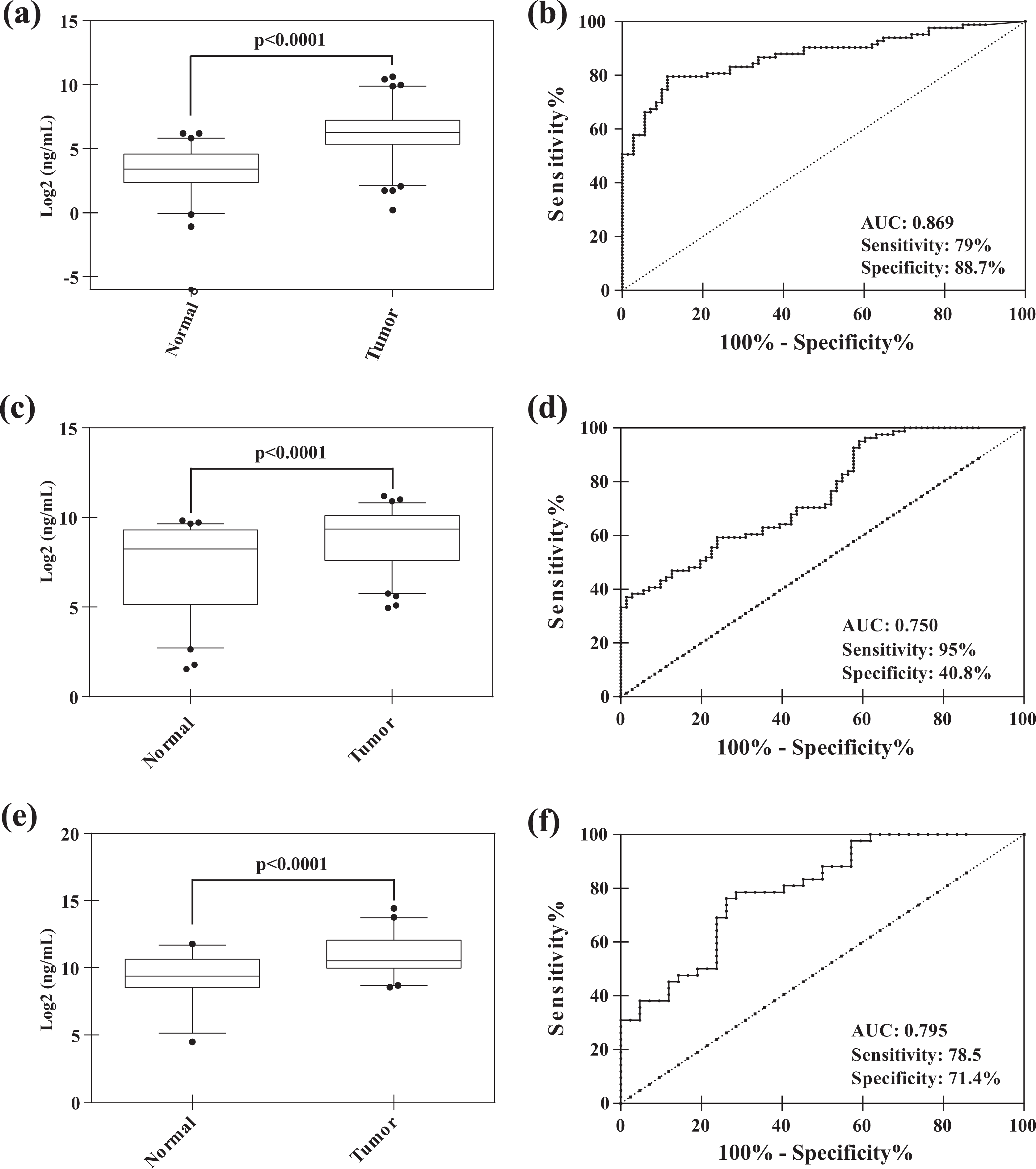
ELISA-based validation showed hypersecretion of (a and b) SPOCK1, (c and d) PSAP, and (e and f) PTPRS in saliva from oral cancer patients with tobacco habits compared to healthy individuals. SPOCK1: SPARC (osteonectin), cwcv and kazal like domains proteoglycan 1; PSAP: prosaposin; PTPRS: protein tyrosine phosphatase, receptor type S.
Demographic characteristics of patients.

PSAP is a glycoprotein known to exist as lysosomal and secretory protein. As a lysosomal protein, it brings about metabolism of glycospingolipids. Secreted PSAP has been identified as a neurotropic factor.
41
-43
In addition to its role as neurotropic factor, the secretory form of PSAP has been shown to be overexpressed in various cancers including breast, gallbladder, and esophageal cancer.
44
-47
Studies have shown the role of PSAP in the pathogenesis of cancer and PSAP is considered as a tumor promoting factor.
47
Secreted levels of PSAP were found to be increased in serum from advanced prostate cancer patients compared to primary prostate cancer.
48
In this study, PSAP was found to be 3.1-fold more abundant in the secretome of the OKF6/TERT1-tobacco cells (Figure 2(b)). Further, ELISA-based validation confirmed hypersecretion of PSAP in the saliva of oral cancer patients (
Protein tyrosine kinases and phosphatases play a key role as molecular drivers for tumor onset and progression. PTPs regulate the activity of protein tyrosine kinases that are involved in protein phosphorylation and signaling. Apart from behaving as tumor suppressors, PTPs also exert oncogenic functions. 26,49 In this study, chronic tobacco exposure induced hypersecretion of several PTPs including PTPR type U (PTPRU, 4.7-fold), PTPR type K (PTPRK, 4.2-fold), PTPR type F (PTPRF, 2.2-fold), and PTPR type S (PTPRS, 2.2-fold). Among these, PTPRS and PTPRF have been reported to be secreted in saliva. 11 We observed a 2.2-fold increase in the abundance of PTPRS in the secretome of tobacco treated cells (Figure 2(c)). PTPRS is a receptor-type PTP and one of the PTPs known to function as tumor suppressor. 35 A recent study by Wang et al. showed that epigenetic inactivation of PTPRS induces EMT and metastasis in hepatocellular carcinoma by activation of epidermal growth factor receptor (EGFR) signaling. 50 PTPRS is found to be deleted in HNSCC. 50 This deletion induces activation of EGFR pathway and modulates sensitivity to EGFR inhibitors. 50 Though PTPRS is shown to exert its functions by regulating EGFR signaling, its role in secreted form is not known. To test the utility of PTPRS in early diagnosis of oral cancer, levels of PTPRS were validated in saliva samples from 42 oral cancer patients and 42 healthy individuals. ELISA-based validation confirmed hypersecretion of PTPRS in oral cancer patients compared to healthy individuals and AUC was found to be 0.795 with 78.5% sensitivity and 71.4% specificity (Figure 3(e) and (f)).
Discussion
India accounts for a high number of chewing tobacco users in the world. About 50% of oral cancer cases can be attributed to tobacco use. 1 Although such high incidences of oral cancer cases are reported in India, early diagnostic markers approved for clinical use are not available till date. Thus, oral cancer is often diagnosed at an advanced stage and is associated with high morbidity, mortality, and poor survival rates. Secreted proteins are important class of molecules known to be involved in various biological processes such as cell–cell signaling, cell communication, migration, and growth. 51,52 Secreted proteins thus reflect various stages of pathological conditions and serve as an incredible source for novel biomarker discovery. A practical approach to identify early diagnostic markers for oral cancer is to explore secreted proteins in saliva as oral cancer cells bathe in salivary milieu. Once identified, simple non-invasive diagnostic assays can be developed to screen individuals who are at a higher risk of developing the disease.
In this study, we analyzed secretome of non-neoplastic oral keratinocytes chronically treated with chewing tobacco and compared it with the parental cell line that was untreated to identify proteins that are secreted due to tobacco treatment. We evaluated the possibility of detecting these differentially secreted proteins in saliva from oral cancer patients with tobacco chewing history. Proteomic analysis of spent media of tobacco-treated and normal oral keratinocytes led to the identification of wide range of proteins including certain growth factors, cell adhesion proteins, extracellular matrix proteins, metalloproteinases, cell surface receptors, and cytokines. Matrix metalloproteinases such as matrix metallopeptidase 3 (MMP3) and matrix metallopeptidase 9 (MMP9) and cadherins family of proteins including N-cadherin implicated in EMT were found to be hypersecreted in response to chewing tobacco. In addition, LC-MS/MS analysis of secretome revealed secretion of several molecules which are regulators of cell proliferation. These include syndecan-1 (SDC1, 5.1-fold), growth arrest specific 6 (GAS6, 3.9-fold), and midkine (MDK, 4.0-fold) were secreted at a higher level in the secretome of OKF6/TERT1-tobacco cells. These proteins are known to be positive regulators of cellular proliferation. 53 -55 In addition, certain proteins such as phospholipase C, delta 1 (PLCD1, 0.5-fold), and promyelocytic leukemia (PML, 0.3-fold) were downregulated. Previous studies have shown that downregulation of these molecules promotes tumor progression. 56,57 This further implicates the cellular transformation of normal oral keratinocytes in response to chewing tobacco.
Proteins identified in secretome of tobacco-treated cells have been compared to proteins secreted by established HNSCC cell lines. 17 Of the 2873 proteins cataloged in this study, 1894 proteins were common to proteins identified from secretome of HNSCC cells. 17 Some of the proteins included insulin-like growth factor binding protein 7 (IGFBP7, 4.5-fold), thrombospondin 1 (THBS1; 3.4-fold), transforming growth factor, beta-induced (TGFBI, 3.5-fold), lectin, galactoside-binding, soluble, 3 binding protein (LGALS3BP, 3.0-fold) and plasminogen activator, urokinase (PLAU, 2.6-fold). Further, to determine whether the identified proteins have also been reported in body fluids, the data were compared to salivary and plasma proteome. 11,58 Of 2873 proteins, 1547 and 2293 were reported in saliva and plasma, respectively. In addition, proteins such as thrombospondin 2 (THBS2), S100 calcium binding protein A9 (S100A9), serpin family A member 1 (SERPINA1), MMP9, and SPARC have been reported to be hypersecreted in saliva samples from oral cancer patients and were found to be abundant in secretome of tobacco-treated cells. 12,16,59,60 Although, we identified a number of proteins from the spent media (secretome) of tobacco-treated cells, we cannot conclude that all of the secreted proteins to be associated with oral cancer or tobacco use. To corelate them, an additional validation has to be carried out in samples from (i) oral cancer patients with no habits of tobacco and (ii) individuals with tobacco habits but no symptoms of disease. Since, we did not have access to such samples, in this study, we could not corelate all proteins identified from proteomics study to be associated to oral cancer or tobacco use.
To define the diagnostic utility, a subset of secreted proteins was validated by ELISA using saliva from oral cancer patients who are users of tobacco compared to healthy individuals who are nonusers of tobacco and have no symptoms of the disease. SPOCK1, PSAP, and PTPRS were found to be hypersecreted in saliva from oral cancer patients with tobacco using habits compared to healthy individuals with no history of tobacco use. SPOCK1 and PTPRS showed high specificity and sensitivity, however, the specificity for PSAP was found to be lower than the other two proteins, implying that SPOCK1 and PTPRS can serve as early biomarkers to identify individuals who are at high risk of developing oral cancer due to tobacco use. However, to be used in a clinical setting for surveillance in a high risk population, the expression levels of these proteins need to be studied in follow-up studies using larger cohorts.
Conclusions
In summary, our results suggest SPOCK1 and PTPRS are promising biomarkers for early detection of oral cancer that should be studied in larger cohorts. These oral cancer biomarkers will be useful not only in oral cancer prevention and early detection but also aid in clinical management studies that will identify high risk patients, both who are chewers of tobacco and patients with precancerous lesions like leukoplakia, to predict their chances of progressing to carcinoma. Resources can then be utilized on the validation of the initial findings in larger number of samples from independent cohorts to finally develop a point-of-care device. This is of utmost importance in developing countries like India where majority of the patients do not have adequate access to health care facilities.
Footnotes
Acknowledgments
We thank Dr Rupesh Dash from Institute of Life Sciences, Bhubaneswar, Odisha for his inputs on the manuscript. We thank the Department of Biotechnology (DBT), Government of India for research support to the Institute of Bioinformatics (IOB), Bangalore. We thank the “Infosys Foundation” for research support to the Institute of Bioinformatics.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of following financial support for the research, authorship, and/or publication of this article: This work was supported by Department of Science and Technology (DST) grants (SERC/LS-439/2011 and SR/SO/HS/0208/2013) and EMR/2016/003253 grant of Dr Arnab Pal. This work was partially supported by grant from the Vision Group of Science and Technology (VGST), Government of Karnataka. IOB is supported by DBT Program Support on Neuroproteomics and infrastructure for proteomic data analysis (BT/01/COE/08/05). This work was supported by NCI’s Clinical Proteomic Tumor Analysis Consortium initiative (U24CA160036) and FAMRI-funded 072017_YCSA.
