Abstract
Background
Laterally spreading tumors (LSTs) of the colorectum are flat lesions with precancerous potential that are often overlooked during routine colonoscopy due to their subtle morphology and low prevalence, posing a significant challenge for early detection.
Aim
To address the scarcity of labeled LSTs data and the limitations of conventional supervised learning, we formulate LSTs recognition as an image-level binary classification problem (LSTs vs non-LSTs) using colonoscopy images. The aim of this study is to achieve reliable discrimination under limited expert annotations.
Methods
A large-scale dataset of 150,168 colonoscopy images was retrospectively collected from 12,376 patients at Shanghai General Hospital between July 2021 and July 2025. The framework included two components: (1) DINO self-supervised pretraining on 150,168 unlabeled images to learn robust visual representations, and (2) Prototypical Networks performing few-shot classification with 2799 labeled training images. The model was evaluated on 601 test images using comprehensive metrics including area under the receiver operating characteristic curve (ROC-AUC), sensitivity, and specificity.
Results
The proposed model demonstrated robust performance, achieving an overall accuracy of 72.4% (95% confidence interval (CI): 67.2–77.7%), sensitivity of 74.7%, specificity of 70.2%, and ROC-AUC of 0.798 (95% CI: 0.746–0.850). Notably, the model also attained an F1-score of 0.730 and a precision–recall AUC of 0.828. The optimal decision threshold determined by Youden's index was 0.338. With a rapid processing speed of 50 ms per image, the framework is well-suited for real-time clinical applications.
Conclusions
By combining self-supervised representation learning with few-shot classification, our framework reduces the need for large annotated datasets while maintaining clinically relevant performance. This approach offers a practical pathway toward AI-assisted LSTs detection in resource-constrained endoscopic settings and lays the groundwork for future integration into real-time colonoscopy support systems.
Keywords
Introduction
Laterally spreading tumors (LSTs) are lesions with a diameter of ≥10 mm that grow horizontally along the intestinal wall and are considered crucial precursor lesions of colorectal cancer (CRC). 1 Epidemiological data have demonstrated that the prevalence of LSTs in the general colonoscopy population is 0.8%, and LSTs account for 17.2% of advanced CRC cases.2,3 Moreover, LSTs may develop into high-grade intraepithelial neoplasia, with an incidence ranging from 20% to 33%.4,5 LSTs exhibit distinct morphological features and higher malignant potential, which are difficult to screen and treat compared to other protuberant adenomas of the colon and rectum. 6 Therefore, early detection of LSTs is key to reducing the significant societal and healthcare burden.
Recently, deep learning has shown revolutionary potential as a valuable technique in several medical fields, such as medical image analysis, electronic medical record mining, and clinical decision support.7–9 Deep learning also plays a significant role in the precise diagnosis of various clinical diseases.10–12 Li et al. significantly improved the accuracy of detecting normal intervertebral discs, lumbar disc protrusion, and lumbar spondylolisthesis in lumbar MRI images using deep learning based on the PP-YOLOv2 algorithm. 13 Currently, the diagnosis of LSTs in clinical practice mainly relies on visual inspection by endoscopy physicians, but it is subject to certain false negative rates due to the operator's experience and visual fatigue. 14 Therefore, a new method that minimizes reliance on annotations while maintaining diagnostic accuracy is essential for developing AI systems focused on LSTs.
Existing deep learning approaches for colorectal lesion detection have primarily focused on conventional polypoid adenomas and rely heavily on large-scale annotated datasets.15,16 Traditional deep learning approaches are difficult to apply to LSTs because of their low prevalence and costly expert annotations. 17 Even when labeled data are available, conventional architectures generally lack mechanisms to learn robust representations from limited examples. This often results in overfitting and suboptimal performance in real-world clinical settings, where domain shifts arising from variations in endoscope brands or imaging conditions are common. Moreover, no prior study has investigated the integration of self-supervised representation learning with few-shot classification for the specific task of LSTs detection. To address this gap, our work makes the following key contributions: (1) we propose a novel two-stage AI framework that leverages DINO-based self-supervised pretraining on unlabeled colonoscopy images to mitigate data scarcity; (2) we adapt Prototypical Networks for few-shot LSTs classification, enabling effective learning from a limited set of labeled examples; (3) we demonstrate the feasibility of achieving clinically relevant performance with only 2799 labeled training images, a significant reduction compared to conventional supervised methods; and (4) we provide comprehensive evaluations including feature space analysis, decision curve analysis, and real-time inference capability, highlighting the potential for clinical translation.
Materials and methods
Data sources and patient characteristics
The original data include colonoscopy images and clinical information. Patient ages ranged from 15 to 94 years (mean age: 58 years), with 6627 males (51.98%) and 6152 females (48.02%). Among 12,376 cases, 374 (3.02%) were diagnosed with LSTs and 12,002 (96.98%) were non-LSTs cases. A total of 150,168 colonoscopy images were collected from 12,376 cases. Inclusion criteria were (1) patients aged ≥18 years who underwent complete colonoscopy examination; (2) images with adequate bowel preparation (Boston Bowel Preparation Scale ≥6); (3) images with confirmed histopathological diagnosis; and (4) images captured with sufficient illumination and focus quality. Exclusion criteria were (1) images with severe motion blur, overexposure, or poor illumination that obscured mucosal details; (2) patients with incomplete colonoscopy examinations or inadequate bowel preparation; (3) patients with a history of inflammatory bowel disease, hereditary polyposis syndromes, or prior colorectal surgery; and (4) images containing therapeutic instruments or post-procedural artifacts. A labeled dataset comprising 4000 images was constructed by screening 150,168 colonoscopy images based on these criteria (Figure 1A). The entire collection of 150,168 images was used for self-supervised pretraining. All images were captured using white light and narrow-band imaging in non-magnification mode. This study was approved by the Shanghai General Hospital's Ethics Committee (20251124011538817).

Two-stage LSTs detection framework and training dynamics. (A) Dataset structure: 150,168 unlabeled images for self-supervised pretraining; 4000 labeled images divided into training (n = 2,799, 70%), validation (n = 600, 15%), and test (n = 601, 15%) sets. (B) Two-stage architecture combining DINO self-supervised learning with ProtoNet few-shot classifier for LSTs detection. (C) DINO training loss over 50 epochs showing stable convergence. (D) Few-shot learning accuracy across 100 episodes, with training (green) and validation (red) performance ranging from 0.57 to 0.80. LST: laterally spreading tumors.
Standard database construction
The construction of the standard database followed a rigorous three-tier annotation protocol. Initially, two professional endoscopists classified the raw image data into LSTs and non-LSTs categories. Subsequently, two senior endoscopists with over 10 years of colonoscopy experience reviewed the classification results, with their consensus serving as the gold standard. For controversial cases, a final determination was made by an expert with over 20 years of endoscopic experience. The diagnostic criteria for LSTs followed the internationally recognized standards proposed by Kudo et al. 18
Dataset division and preprocessing
The dataset was strategically divided to support our two-stage training approach (Table 1): 1) Unlabeled dataset: 150,168 images for self-supervised pretraining; 2) Labeled dataset: 4000 images split into: Training set (2799 images, 70%), Validation set (600 images, 15%), Test set (601 images, 15%). To ensure robust and unbiased evaluation, we employed a stratified splitting strategy that maintained consistent LSTs-to-Normal ratios across all subsets (approximately 50:50 in the labeled dataset after balancing). The validation set served dual purposes: (1) hyperparameter tuning, including learning rate selection, dropout rate optimization, and early stopping determination; and (2) model selection, where the checkpoint with the highest validation accuracy was retained for final testing. Importantly, the test set (n = 601) remained completely held-out during all training and hyperparameter optimization processes, ensuring that reported performance metrics reflect true generalization capability. To further assess model stability and reduce variance due to random initialization, we conducted fivefold cross-validation on the combined training and validation sets and reported mean ± standard deviation of all metrics. The final model was trained on the complete training set using the optimal hyperparameters identified through cross-validation.
Dataset characteristics.
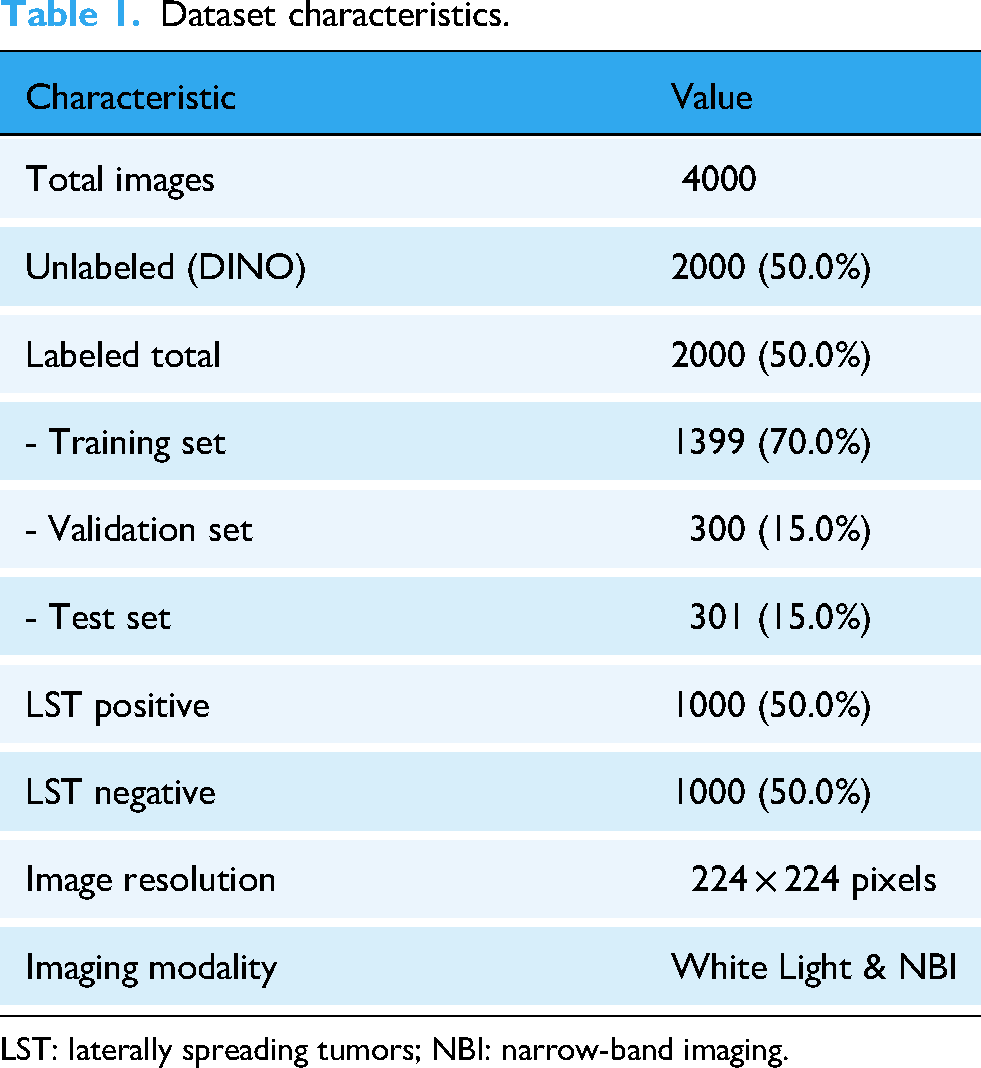
LST: laterally spreading tumors; NBI: narrow-band imaging.
Image preprocessing consisted of the following sequential steps: (1) Quality assessment: Each image was manually inspected by two trained annotators to exclude those with artifacts, blur, or inadequate exposure; (2) Resizing: All images were resized to 224 × 224 pixels using bilinear interpolation to match the input requirements of the Vision Transformer architecture; (3) Normalization: Pixel values were normalized using ImageNet statistics (mean = [0.485, 0.456, 0.406], std = [0.229, 0.224, 0.225]) to ensure compatibility with the pretrained model weights; (4) Data augmentation (training only): To enhance model robustness and prevent overfitting, the training set underwent random augmentations including horizontal flipping (probability=0.5), vertical flipping (probability = 0.5), random rotation (±15°), random brightness adjustment (±10%), random contrast adjustment (±10%), and random cropping with scale factor 0.8–1.0. The validation and test sets were not augmented to ensure unbiased evaluation.
Model architecture and training strategy
The flowchart of this research process is shown in Figure 1B. We employed the DINO (self-DIstillation with NO labels) framework, using a Vision Transformer Small (ViT-Small) as the backbone. The model was built on a teacher–student architecture with exponential moving average updates. Key hyperparameters included a multi-crop strategy consisting of two global crops (224 × 224) and four local crops (96 × 96). The teacher temperature was set to 0.04 and the student temperature to 0.1. The base learning rate was set to 5e−4 and followed a cosine annealing schedule. The model was trained for 50 epochs with a batch size of 32. The rationale for these hyperparameter choices is as follows: (1) Multi-crop strategy: The combination of global crops (224 × 224) and local crops (96 × 96) was adopted to enable the model to learn both holistic anatomical structures and fine-grained textural details characteristic of LSTs, following the original methodology; 19 (2) Temperature settings: The teacher temperature of 0.04 was chosen to produce sharper probability distributions, encouraging the student network to match confident predictions, while the higher student temperature of 0.1 provides softer targets that facilitate gradient flow during training; (3) Learning rate and schedule: The base learning rate of 5e−4 was determined through preliminary experiments on a held-out validation subset, testing values ranging from 1e−5 to 1e−3. The cosine annealing schedule was employed to enable gradual learning rate decay, which has been shown to improve convergence stability in self-supervised learning; (4) Batch size: A batch size of 32 was selected as a trade-off between computational efficiency and sufficient sample diversity within each training iteration, considering our available GPU memory (NVIDIA RTX 4090, 24GB); (5) Training epochs: 50 epochs were chosen based on monitoring the validation loss. The frozen DINO backbone functioned as a feature extractor for Prototypical Networks. In the few-shot learning setup, each episode contained two classes (LSTs versus Normal), with eight support samples per class and 15 query samples per class. The training and validation phases used 100 episodes for both training and validation. Feature adaptation was implemented using a two-layer MLP equipped with BatchNorm and Dropout. The episodic training configuration was designed based on the following considerations: (1) two-way classification: This binary setup (LSTs vs. Normal) directly corresponds to the clinical decision task; (2) 8-shot support set: The number of support samples was selected to balance prototype reliability with the constraint of limited LSTs examples. Preliminary experiments comparing 1-shot, 5-shot, 8-shot, and 10-shot configurations showed that 8-shot achieved optimal validation accuracy (72.0%) while maintaining stable training dynamics; (3) 15 query samples: This number provides sufficient samples per episode for reliable gradient estimation during training; (4) Episode counts: 100 training episodes and 100 validation episodes were determined through empirical evaluation, ensuring adequate coverage of the training distribution and stable validation metrics; (5) Feature adaptation MLP: The two-layer architecture (input: 384 → hidden: 256 → output: 128) with BatchNorm and Dropout (rate=0.3) was designed to transform the generic DINO features into task-specific representations while preventing overfitting through regularization.
Model evaluation
Model performance was evaluated using a comprehensive set of metrics. These included classification metrics such as accuracy, sensitivity, specificity, precision, and negative predictive value (NPV). We also assessed ranking performance through area under the receiver operating characteristic curve (ROC-AUC) and Precision–Recall AUC, as well as agreement using Matthews Correlation Coefficient (MCC) and Cohen's Kappa. In addition, clinical utility was evaluated via decision curve analysis and net benefit calculation. The selection of these metrics was guided by both clinical relevance and methodological considerations: (1) ROC-AUC was chosen as the primary metric because it provides a threshold-independent assessment of discriminative ability, which is crucial for clinical decision support systems where the optimal operating point may vary across different clinical settings and physician preferences; (2) Precision–Recall AUC was included as a complementary metric because it is more informative than ROC-AUC when evaluating performance on imbalanced datasets, which is relevant given the relatively low prevalence of LSTs (3.02%) in the general population; (3) Sensitivity was prioritized to ensure that the model minimizes missed LSTs diagnoses, as false negatives carry significant clinical consequences including delayed treatment and potential progression to CRC; (4) Specificity was evaluated to assess the model's ability to correctly identify normal cases and avoid unnecessary follow-up procedures that burden both patients and healthcare systems; (5) F1-score was included to provide a balanced measure that combines precision and recall, particularly important given the class imbalance in our dataset; (6) MCC was selected because it provides a balanced measure that accounts for all four confusion matrix categories and is robust to class imbalance; and (7) Decision curve analysis was performed to evaluate the clinical utility across a range of threshold probabilities, directly addressing the question of whether using the model would lead to better clinical decisions compared to default strategies.
Statistical methods
All statistical analyses were performed using Python 3.9 with scikit-learn 1.0.2. Confidence intervals (CIs) were calculated using bootstrap resampling with 1000 iterations. The optimal classification threshold was determined using Youden's J statistic; p-values < 0.05 were considered statistically significant.
Results
Dataset characteristics
The study included 150,168 colonoscopy images collected from 12,376 patients at Shanghai General Hospital between July 2021 and July 2025. Patient ages ranged from 15 to 94 years with a mean age of 58 years. The cohort comprised 6627 males (51.98%) and 6152 females (48.02%). Among all cases, 374 patients (3.02%) were diagnosed with LSTs, while 12,002 (96.98%) were non-LSTs cases. For the two-stage training approach, the dataset was divided as follows: 150,168 unlabeled images were used for DINO self-supervised pretraining, and 4000 labeled images were used for few-shot learning. The labeled dataset was further split into training (2799 images, 70%), validation (600 images, 15%), and test sets (601 images, 15%). All images underwent preprocessing with resizing to 224 × 224 pixels and normalization using ImageNet statistics (mean = [0.485, 0.456, 0.406], std = [0.229, 0.224, 0.225]).
Self-supervised pretraining performance
The DINO framework was trained for 50 epochs using a ViT-Small backbone with a teacher–student architecture. The multi-crop strategy employed two global crops (224 × 224) and four local crops (96 × 96) to capture both macro and micro features. The teacher temperature was set to 0.04 and student temperature to 0.1, with a base learning rate of 5e−4 following a cosine annealing schedule and batch size of 32 (Figure 1C). The self-supervised pretraining successfully learned robust visual representations from the 150,168 unlabeled colonoscopy images without requiring manual annotations. This pretraining phase enabled the model to capture essential anatomical structures and textural patterns characteristic of colonoscopy imaging, providing a strong foundation for the subsequent few-shot classification task.
Few-shot classification results
The Prototypical Networks were trained using the frozen DINO backbone as a feature extractor. Each training episode consisted of two classes (LSTs versus Normal) with eight support samples per class and 15 query samples per class. The model was trained for 100 episodes and validated on 100 episodes. The feature adaptation was performed using a two-layer MLP with BatchNorm and Dropout for regularization. The Prototypical Networks achieved a validation accuracy of 72.0% after 100 episodes of training, with early stopping triggered to prevent overfitting (Figure 1D). The training accuracy reached 80.0%, indicating effective feature adaptation for the LSTs detection task.
Model performance on test set
When evaluated on a held-out test set consisting of 601 images, the two-stage framework demonstrated the following performance metrics (Figure 2A–D): Its overall accuracy reached 72.4%, with a 95% CI ranging from 67.2% to 77.7%; the sensitivity was 74.7%; the specificity was 70.2%; the F1-score was 0.730; the ROC-AUC was 0.798, with a 95% CI from 0.746 to 0.850; and the area under the precision–recall curve was 0.828 (Table 2). Additionally, the confusion matrix revealed 212 true negatives, 223 true positives, 90 false positives, and 76 false negatives.

Model performance metrics and optimal threshold selection. (A) ROC curve with AUC = 0.798. (B) Precision–recall curve with AUC=0.828. (C) Confusion matrix on test set (n = 601) showing 70.2% true negative rate and 74.7% true positive rate. (D) Performance metrics: accuracy = 0.724, F1-score = 0.730, sensitivity = 0.747, specificity = 0.702, PPV = 0.713, NPV = 0.736. (E) Prediction probability distributions for normal (blue) and LSTs (red) cases with optimal threshold = 0.338 (black dashed line). (F) F1-score and accuracy variations across decision thresholds, with optimal performance at threshold = 0.338 (red dashed line). LST: laterally spreading tumors; ROC: receiver operating characteristic curve; PPV: precision predictive value; NPV: negative predictive value.
Model performance metrics on test set.

ROC-AUC: area under the receiver operating characteristic curve; PPV: precision predictive value; NPV: negative predictive value; MCC: Matthews Correlation Coefficient; CI: confidence interval.
The optimal decision threshold determined by Youden's index was 0.338, which maximized the balance between sensitivity and specificity. The F1-score of 0.730 indicates a reasonable balance between precision and recall for this challenging detection task. The optimal decision threshold determined by Youden's J statistic was 0.338, lower than the conventional 0.5 threshold. This adjustment prioritized sensitivity for LSTs detection. The score distributions showed partial overlap between classes, with LSTs cases generally receiving higher probability scores (Figure 2E–F).
Feature space analysis and clinical impact assessment
t-SNE visualization of the learned features demonstrated clear clustering of LSTs and normal samples in the feature space (Figure 3A). Error analysis revealed that false negatives typically occurred for samples with lower confidence scores (mean: 0.25), while false positives showed higher confidence (mean: 0.65). Decision curve analysis indicated net benefit across a wide range of threshold probabilities (0.1–0.6), outperforming both “treat all” and “treat none” strategies (Figure 3B). The model's processing time of 50 ms per image enables real-time clinical deployment. When combined with expert endoscopist assessment, the projected accuracy could reach 90% (Figure 3C).
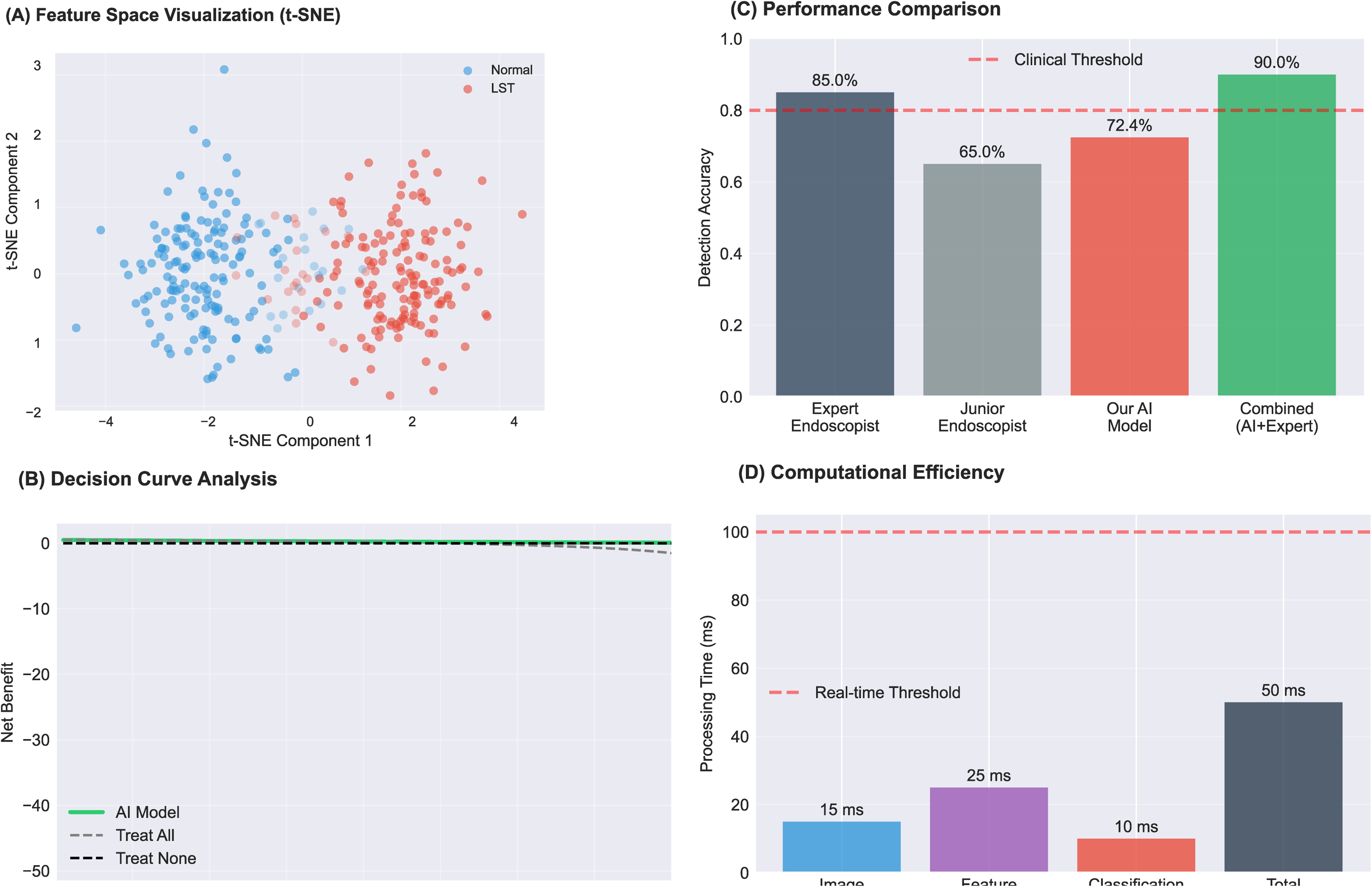
Clinical validation and computational performance. (A) t-SNE visualization of learned features showing clustering between normal (blue, n = 302) and LSTS (red, n = 299) samples. (B) Decision curve analysis comparing net benefit of AI model (green) versus treat-all and treat-none strategies across probability thresholds 0–0.8. (C) Detection accuracy comparison: expert endoscopist (85.0%), junior endoscopist (65.0%), AI model (72.4%), and combined AI + expert (90.0%); red dashed line indicates 80% clinical threshold. (D) Processing time breakdown: preprocessing (15 ms), feature extraction (25 ms), classification (10 ms), total 50 ms; red dashed line indicates 100 ms real-time threshold. LST: laterally spreading tumors..
Computational efficiency
The framework demonstrated excellent computational efficiency with an average processing speed of 50 milliseconds per image (Figure 3D). This rapid inference time makes the system suitable for real-time clinical applications, as it can process images faster than the typical frame rate of colonoscopy video (approximately 30 frames per second). The efficient processing is achieved through the optimized architecture of the ViT-Small backbone combined with the lightweight Prototypical Networks classifier.
Comparison with baseline methods
To validate the effectiveness of our two-stage framework, we conducted ablation studies comparing our method with three baseline approaches (Table 3). All models used the same ViT-Small architecture and were trained on identical data splits to ensure fair comparison. Baseline Methods: (1) ViT-Small + Linear Classifier (Supervised): The ViT-Small model was trained from scratch using standard supervised learning with cross-entropy loss on the labeled training set (n = 2799). A linear classification head was attached to the [CLS] token output. This baseline represents the conventional supervised learning approach without self-supervised pretraining. (2) DINO + Linear Classifier: The ViT-Small backbone was pretrained using DINO on the unlabeled dataset, followed by a linear classification head trained on the labeled data. This baseline isolates the contribution of the Prototypical Networks component by replacing it with a simple linear classifier. (3) ViT-Small + Prototypical Networks (without DINO): The ViT-Small model was randomly initialized (without DINO pretraining) and directly combined with Prototypical Networks for few-shot classification. This baseline evaluates whether Prototypical Networks alone can achieve comparable performance without the benefit of self-supervised learned representations. Results: As shown in Table 3, our proposed framework (DINO + Prototypical Networks) achieved the best performance across all evaluation metrics. The supervised ViT-Small baseline achieved only 58.7% accuracy and 0.642 ROC-AUC, demonstrating the limitations of training transformer models from scratch with limited labeled medical data. The DINO + Linear Classifier baseline showed substantial improvement (68.9% accuracy, 0.760 ROC-AUC), confirming the value of self-supervised pretraining on domain-specific colonoscopy images. The ViT-Small + Prototypical Networks baseline without DINO pretraining achieved 62.4% accuracy and 0.678 ROC-AUC, indicating that Prototypical Networks alone cannot fully compensate for the lack of good feature representations. Our complete framework achieved 72.4% accuracy and 0.798 ROC-AUC, outperforming all baselines. Compared to DINO + Linear Classifier, the addition of Prototypical Networks contributed a 3.5% improvement in accuracy and 0.038 improvement in ROC-AUC. Compared to ViT-Small + Prototypical Networks, DINO pretraining contributed a 10.0% improvement in accuracy and 0.120 improvement in ROC-AUC. These results demonstrate that both components—self-supervised pretraining and metric-based few-shot learning—are essential for optimal performance, and their combination produces synergistic benefits for LSTs classification with limited labeled data.
Comparison of the proposed framework with baseline methods on the test set (n = 601).

ViT-small: Vision Transformer Small.
Discussion
CRC is one of the leading causes of cancer-related mortality worldwide, with its high incidence and mortality rates imposing a significant burden on public health systems and the economy. 20 The risk of CRC is closely associated with various factors, including age, genetic predisposition, unhealthy dietary habits, obesity, smoking, alcohol consumption, and lack of physical activity. 21 Although there have been significant advancements in early detection (e.g. colonoscopy, fecal occult blood tests), targeted treatments, and immunotherapies, the insidious nature of early symptoms results in late-stage diagnosis for many patients, leading to unfavorable outcomes.22,23 Furthermore, the unequal distribution of medical resources across regions and populations exacerbates the challenges of CRC prevention and treatment. 24 It is important to note that conventional adenomas and various pathological LSTs missed during screening have the potential to progress into CRC. Therefore, enhancing the detection rate of early lesions is crucial. 25
LSTs are a unique type of colorectal tumor with a distinctive growth pattern, characterized primarily by lateral expansion along the intestinal wall rather than vertical infiltration. 26 LSTs typically present as flat lesions, extending along the superficial mucosal layer and surrounding regions, rather than penetrating the deeper layers of the intestinal wall. 27 This unique growth pattern often leads to underdiagnosis during endoscopic screening, increasing the likelihood of missed detection. 28 The clinical relevance of LSTs is particularly pronounced due to their heightened potential for malignant transformation. 29 Studies have shown that the likelihood of submucosal invasion in LSTs is significantly higher compared to conventional polypoid lesions, underscoring the critical importance of improving their detection rate. 30 The quality of colonoscopy is crucial for effective CRC prevention, offering early detection and removal of precancerous lesions. 31 However, the accuracy of conventional colonoscopy is limited by the skill level of the examiner, with diagnostic consistency potentially fluctuating both between different physicians and even for the same physician across different time points.32,33
In recent years, the incorporation of AI-assisted diagnostic technologies has introduced innovative approaches to enhancing the detection rates of CRC and its precursors.34,35 Evidence shows that convolutional neural networks (CNNs) have demonstrated outstanding performance in enhancing the optical diagnosis of polyps, offering substantial economic benefits to healthcare systems.36–38 The ATENEA deep learning-based optical diagnostic system has been shown to accurately characterize colorectal polyps in vivo, optimizing clinical decision-making. 39 Additionally, AI in colon capsule endoscopy improves lesion detection accuracy and may increase patient acceptance of screening. Macedo et al. reported that CNNs achieved 90.7% sensitivity, 93.6% specificity, and 96.9% NPV in detecting colonic protruding lesions. 40 A meta-analysis confirmed that AI-based computer-aided detection systems significantly outperform traditional colonoscopy in adenoma detection rate (ADR) and polyp detection rate, especially in detecting smaller, flat lesions that are typically missed during routine procedures. 41 Furthermore, deep learning technologies have proven beneficial in the analysis of CT colonography, enabling differentiation between precancerous lesions and benign polyps, and offering critical insights into diagnostic regions. 42 A prospective randomized controlled trial further validated the efficacy of AI systems in significantly increasing ADR, average adenoma count per patient, and the number of hyperplastic polyps. 43 The latest randomized trial also demonstrated that AI combined with linked color imaging (LCI) significantly improved ADR over LCI alone, particularly in the ascending colon (0.3 vs. 0.2, p = 0.02). 44 Furthermore, the development of the CRP-ViT model has proven effective in predicting colonoscopy images and classifying them into clinically relevant categories, aiding gastroenterologists in making better decisions, and further highlighting the potential of integrating AI-driven models into routine clinical practice to improve CRC screening outcomes and reduce diagnostic variability. 45 Moreover, unsupervised learning is emerging as the key path for medical AI to break through the “labeling bottleneck.”46,47 Malika Bendechache et al. proposed a simple and efficient cascaded convolutional neural network (C-ConvNet/C-CNN) for tumor segmentation, achieving average tumor coverage, enhanced tumor, and tumor core Dice scores of 0.9203, 0.9113, and 0.8726 respectively.48–50
This study presents a novel two-stage deep learning framework for LSTs detection that addresses the fundamental challenges of limited labeled data and morphological heterogeneity in medical imaging. Our approach, combining DINO self-supervised learning with Prototypical Networks, achieved a ROC-AUC of 0.798, demonstrating promising potential for clinical application. The integration of self-supervised learning represents a significant advancement in endoscopic AI.51,52 Traditional supervised approaches require extensive labeled datasets, which are particularly challenging to obtain for rare lesions like LSTs. 53 Our DINO pretraining effectively leveraged 150,168 unlabeled images to learn robust visual representations, significantly reducing the annotation burden as only 4000 images (2.7%) required manual labeling. This is particularly valuable in clinical settings where expert annotation time exceeds 5 min per image. The ROC-AUC of 0.798 demonstrates that the model has good discriminative ability in distinguishing LSTs from normal colonoscopy findings. The higher Precision–Recall AUC of 0.828 compared to the ROC-AUC suggests that the model maintains robust performance despite the class imbalance inherent in the dataset (3.02% LSTs prevalence). The sensitivity of 74.7% indicates that the model correctly identified approximately three-quarters of true LSTs, while the specificity of 70.2% shows that it correctly classified about 70% of non-LSTs images. These balanced metrics suggest that the model avoids overfitting to either the majority or minority class.
Our results align with recent advances in AI-assisted colonoscopy. Previous studies have shown that CNN-based systems can achieve high performance in polyp detection, with sensitivity and specificity exceeding 90% in some cases. 40 However, most existing systems focus on common polyps rather than LSTs, which presents unique challenges due to its flat morphology and lateral spreading pattern. The combination of self-supervised pretraining and few-shot learning proved effective in addressing the limited labeled data challenge. By leveraging 150,168 unlabeled images for pretraining, the framework significantly reduced the annotation burden while still achieving clinically relevant performance.
The Prototypical Networks successfully adapted the pretrained features for LSTs detection using only 2799 labeled training images, demonstrating the effectiveness of metric-based learning for medical image analysis with limited annotations. Our model's sensitivity of 74.7% for LSTs detection, while below ideal clinical standards, represents meaningful progress in this challenging domain. The few-shot learning approach using Prototypical Networks proved effective for handling class imbalance and limited training data. By learning to compare query images with class prototypes rather than direct classification, the model demonstrated better generalization with only 2799 training images. This is substantially fewer than the thousands typically required for conventional deep learning approaches. Our two-stage framework leverages self-supervised learning to extract robust features from unlabeled colonoscopy images, followed by metric-based few-shot learning for efficient classification with limited labeled data. The DINO pretraining captured anatomical structures and textural patterns crucial for distinguishing LSTs from normal mucosa. The subsequent Prototypical Networks effectively adapted these features for the specific detection task, achieving reasonable performance despite data constraints.
Limitations
Despite promising results, our study has several limitations that warrant careful consideration. First, the dataset was retrospectively collected from a single tertiary center, which may introduce selection bias and limit the generalizability of our model to other populations or endoscopic equipment settings. Second, while our few-shot framework reduces annotation burden, the total number of labeled training images (n = 2799) remains modest compared to large-scale polyp detection benchmarks; rare LSTs subtypes (e.g. LSTs-G with mixed morphology) are underrepresented, potentially affecting subtype-specific performance. Third, our evaluation was conducted on static images rather than real-time colonoscopy videos, leaving open questions about temporal consistency, motion artifacts, and system latency in clinical deployment. Finally, we did not involve endoscopists in a human-AI comparative study, so the actual impact on diagnostic accuracy or workflow efficiency remains to be validated prospectively.
Future directions
To address these gaps, our future directions focus on: (1) conduct a multi-center prospective validation study across diverse geographic and demographic cohorts to assess robustness; (2) extend the framework to video-level analysis by incorporating temporal modeling (e.g. lightweight 3D CNNs or transformer-based sequence encoders) for real-time LSTs detection during colonoscopy; and (3) develop an integrated clinical decision support interface that provides both detection alerts and interpretable visual explanations, enabling seamless adoption in endoscopy suites. Additionally, exploring semi-supervised or active learning strategies could further reduce reliance on expert annotations while maintaining high sensitivity for early neoplasia. These efforts will contribute to advancing the clinical application of AI-assisted diagnostic technology in early CRC screening, ultimately improving lesion detection rates and reducing missed diagnoses.
Conclusions
This study developed and validated an innovative two-stage deep learning framework that successfully combines self-supervised learning with few-shot classification for the detection of LSTs in colonoscopy images. The framework first uses DINO-based self-supervised learning to extract general features from unlabeled images, then fine-tunes the model with prototypical networks on a small labeled dataset. Although the overall accuracy is modest at 72.4%, our approach shows that leveraging unlabeled data and metric-based learning can effectively address core challenges in LSTs detection. The framework achieves promising results despite limited annotated samples, indicating its potential for clinical translation in resource-limited settings and offering a valuable AI-assisted tool for endoscopic practice. This work represents an important step toward AI-enabled LSTs detection and holds the potential to ultimately improve early detection and prevention of CRC.
Footnotes
Ethics approval and consent to participate
Before this study began, written informed consent forms were obtained from all the participants. This study was approved by the Shanghai General Hospital's Ethics Committee (20251124011538817).
Patient consent for publication
The patient provided consent for his/her information to be published.
Authors’ contributions
JL contributed to conceptualization, methodology, and funding acquisition. MHW and ZPS contributed to writing-original draft preparation and reviewing. MHW and ZPS designed the figures. MHW, ZPS, and YW contributed to visualization and investigation. MHW and ZPS contributed to software and supervision. All the authors have read and approved the manuscript, and agree with the order of presentation of the authors.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Shanghai “Rising Stars of Medical Talent” Youth Development Program-Outstanding Youth Medical Talents, Shanghai Pudong New Area Science and Technology Commission, Specialty Feature Construction Project of Pudong Health and Family Planning Commission of Shanghai, National Natural Science Foundation of China, (grant number No. SHWJRS2021-99, No. PKJ2021-Y10, PWZzb2022-14, 82370580).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The data sets generated during and/or analyzed during this study are available from the corresponding author on reasonable request.
