Abstract
Background
Colonoscopy plays a vital role in assessing disease activity in ulcerative colitis (UC), and biopsy via colonoscopy helps to evaluate its histological activity. Endoscopists must report the endoscopic activity and rely on the biopsy results to predict the histological activity.
Methods
We aimed to develop a deep learning-based algorithm to evaluate the disease and histological activities of UC based on white-light endoscopic images obtained during the procedure in this research. A deep learning system for classifying the colonoscopic images for assessing the endoscopic and histological activities of UC patients was developed. Its performance was evaluated with an independent dataset. The system was utilized to analyze the captured video segments, and the results were compared with those of human endoscopists.
Results
A total of 375 video segments from 82 patients were utilized to develop the endoscopic and histological activity prediction assurance algorithm. Among the 375 video segments, 60%, 20%, and 20% were used for training, validation, and testing the proposed vision transformer (ViT) model, respectively. Moreover, four senior and six young endoscopists reviewed and scored the endoscopic and histological activities based on 77 testing video clips. The accuracies were 77.92%, 71.00%, and 83.12% for histological healing; and 74.35%, 72.51%, and 92.21% for complete mucosal healing (Mayo Endoscopic Score 0 vs 1–3), among senior endoscopists, junior endoscopists, and the ViT model, respectively.
Conclusions
Our novel deep learning-based model, based on endoscopic videos, was comparable to that of experienced endoscopists and surpassed that of young endoscopists in predicting histological remission and complete mucosal healing.
Introduction
Inflammatory bowel disease (IBD), including Crohn's disease and ulcerative colitis (UC), is a chronic condition characterized by persistent intestinal inflammation that requires continuous treatment and surveillance to monitor disease activity, thereby improving patients’ quality of life and preventing complications. 1 Although UC treatment has progressed from steroid, 5-aminosalicylic therapy to the recently introduced advanced therapy, the treatment goal has also evolved from clinical remission to endoscopic and even histologic remission. 2 Artificial intelligence (AI) is an emerging technology that can affect several aspects of healthcare. Most AI systems aim to provide diagnostic aid in decision-making or decrease the healthcare workers’ workload. In endoscopy, AI is now used to detect or characterize colorectal lesions during colonoscopy, 3 determine the peptic ulcer bleeding risk, 4 or as a quality assurance system.4–7
Endoscopic remission, a more accurate predictor of outcome, has been considered a long-term target in UC patients, and an accurate and reproducible assessment of endoscopic disease activity is central to effective disease management. 2 Traditional scoring systems, including the Mayo Endoscopic Score (MES) and UC Endoscopic Index of Severity (UCEIS), although widely adopted, are limited by subjectivity, interobserver variability, and a relatively coarse resolution of disease activity. 8 Even among expert reviewers, inconsistencies persist, particularly in distinguishing borderline remission cases from cases with mild activity, which are decisions that carry remarkable implications for upgrading or downgrading therapeutic strategies. 9
Beyond merely replicating human scoring, AI can identify subtle, previously unrecognized differences in mucosal appearance captured in endoscopic images and predict the corresponding histologic activity with high accuracy. Several groups have reported ML systems for UC activity assessment, most commonly using convolutional neural networks (CNNs). Maeda et al. 10 reported the use of a computer-aided diagnosis (CAD) system to predict histological inflammation by endocytoscopy. The system achieved a diagnostic sensitivity of 74%, specificity of 97%, and overall accuracy of 91%. Iacucci et al. 11 utilized a CNN using the Pentax system to assess endoscopic remission from white-light endoscopy (WLE) videos, achieving a sensitivity and specificity of 72% and 87%, respectively. Additionally, the model predicted histologic remission, defined as a Nancy Histological Index (NHI) of ≤1, with a sensitivity, specificity, and overall accuracy of 67%, 86%, and 84%, respectively.
Thus, integrating AI into the endoscopic assessment of UC holds considerable promise for improving the consistency and objectivity of disease activity scoring systems. Moreover, AI can potentially augment clinical decision-making by accurately predicting the underlying histologic inflammation. Unlike CNNs, vision transformers (ViTs) leverage self-attention to capture long-range dependencies across the entire endoscopic field, enabling integration of subtle, spatially distributed cues in mucosal texture and vascular patterning. This feature is well-suited to UC, where inflammation and healing often present as diffuse changes rather than focal. The present study aimed to develop and validate a ViT-based endoscopic model using standard, routinely available endoscopic equipment, compare its performance for endoscopic scoring against that of human endoscopists, and evaluate its utility in predicting histologic remission.
Methods
Patients and data preparation
This prospective, single-center study consecutively enrolled adult patients (aged ≥18 years) with a confirmed diagnosis of UC who underwent clinically indicated colonoscopy at the Endoscopy Center of Changhua Christian Hospital between March 2023 and February 2025. Eligible patients were identified through the institutional IBD registry and were invited to participate at the time of colonoscopy. All participants provided written informed consent before enrollment and video recording. Endoscopic images were reviewed and retrieved for subsequent analysis by two expert endoscopists, each with >15 years of experience. Scoring discrepancies were resolved through a discussion with a third independent endoscopist. MES (range: 0–3) was used to assess the endoscopic disease activity. For histological evaluation, biopsy samples obtained during routine clinical assessment were time-matched to the corresponding endoscopic images to ensure accurate labeling. Histological disease activity was assessed by experienced pathologists who are board-certified and have more than 10 years of experience interpreting colorectal biopsies from UC patients using the NHI (range: 0–4) based on a standardized evaluation form developed in Taiwan.
This study was conducted in accordance with the guidelines stipulated in the Declaration of Helsinki for research involving human subjects, including research on identifiable human materials and data. This study protocol was approved by the Institutional Review Board of Changhua Christian Hospital (Approval No. CCH IRB 230403).
Deep learning architecture of the ViT
In this study, we utilized a deep learning (DL) model known as the Transformer to classify UC severity. Although originally developed for natural language processing, transformers have been successfully adapted for image analysis tasks, demonstrating superior performance 12 over traditional neural networks, including CNNs, in various applications. Specifically, we opt for the original ViT 13 because of its simplicity and proven effectiveness across various image recognition tasks.
ViT processes images by partitioning them into nonoverlapping patches, which are subsequently transformed into linear embeddings through a learnable projection. The resulting sequence of patch embeddings is then fed into a standard Transformer encoder. 14 The ViT encoder leverages the self-attention mechanism, wherein the features from all patches are aggregated through weighted summation, with the weights determined by the pairwise similarity between the linearly projected representations of the patches. This mechanism enables the capture of long-range dependencies across all patches, thereby enhancing its capacity to model global contextual information effectively. At the end of the encoder, a fully connected layer maps the average pooled patch embedding to the output probabilities corresponding to each UC severity class.
The model was implemented using PyTorch 2.4.1 on an NVIDIA RTX 4090 GPU. We initialized a ViT-based architecture from the Timm library, 15 leveraging the model that was pretrained on the ImageNet dataset. 16 To address the ordinal nature of the classification problem, we utilized the Class Distance Weighted cross-entropy loss function, 17 with the power term set to 3. The model was trained for 50 epochs, with the first five epochs serving as the warm-up stages. A batch size of 64 was adopted, and the learning rate was initialized at 1e−6 and gradually adjusted using a cosine decay scheduler. The AdamW optimizer was used to enhance the training stability. All input images (512 × 512 pixels) were resized to 224 × 224 pixels during training and inference. Moreover, the original 30 frames per second of the video segment was reduced to three frames per second. Altogether, 225 (60%), 75 (20%), and 75 (20%) of 375 video segments were used to train, validate, and test the proposed ViT model.
Moreover, the ViT model was compared with the ResNeSt model, which is a deep residual network (ResNet) 18 variant, used in previous studies.6,7,19 We utilized 77 testing videos to compare the performance of the developed model with that of human endoscopists. Table 1 presents the 375 video segments used in this study according to the patients’ MES and NHI. Table 2 shows the distribution of the NHI and MES scores of the testing videos.
The distribution of 375 video segments from 82 patients used in the present study.
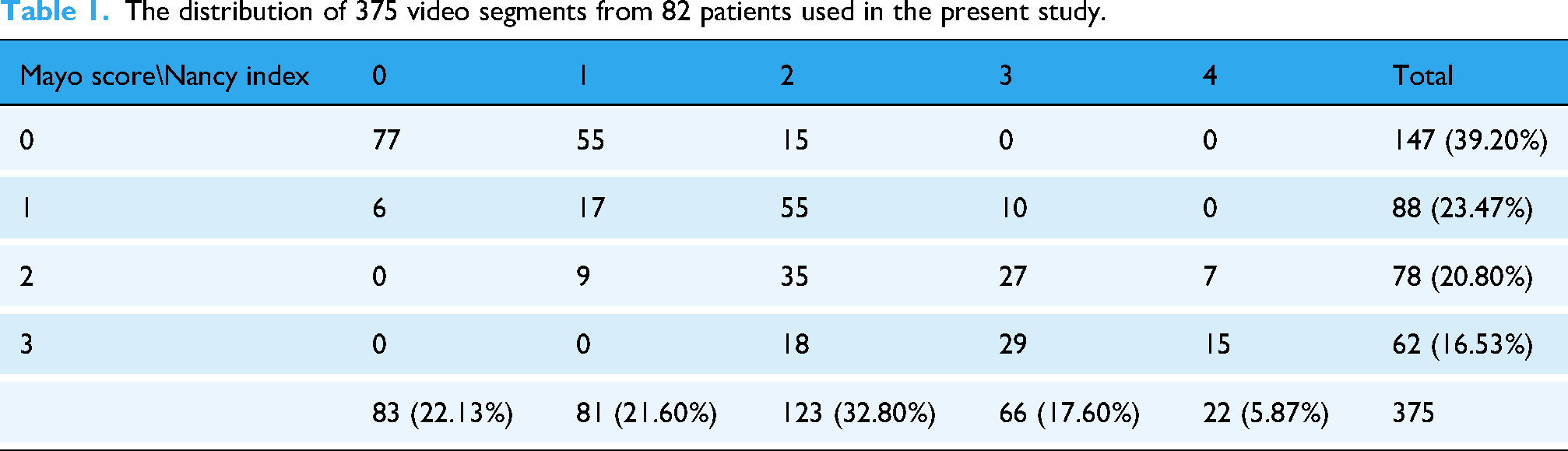
Distribution of the testing video segments used in the present study.

Performance of the DL model in comparison with human endoscopists
Altogether, 77 distinct video clips (each up to 10 s) were prepared for subsequent comparison between the DL model and human endoscopists. Ten endoscopists from our institution participated and were stratified into the following two groups: (A) senior endoscopists with at least five years of experience in managing IBD and performing endoscopic assessments; and (B) young endoscopists, including fellows and early-career staff, who received their training at our unit and were familiar with the MES system application. Each endoscopist independently reviewed the video clips and provided their assessment of the MES for endoscopic remission (MES 0–1 vs 2–3) or complete mucosal healing (MES 0 vs 1–3) and their binary prediction of the histologic remission status based on the corresponding biopsy findings using the NHI, with NHI of ≤1 and ≥2 indicating histological remission or nonremission, respectively. 20
Statistical analysis
Descriptive statistics were used to summarize the classification performance across the following three groups: ViT model, experienced human raters, and young raters. The performance evaluation encompassed four metrics, including accuracy, sensitivity, specificity, and F1 score. To assess the group-level differences across these, a one-way analysis of variance (ANOVA) was conducted. Upon identifying significant effects (p < .05), Tukey's Honest Significant Difference (HSD) test was utilized for post-hoc pairwise comparisons to determine group disparities.
As the ViT model produced a single observation per metric, bootstrapping with 1000 iterations was implemented to simulate plausible performance distributions, assuming a standard deviation of 0.5%. These synthetic distributions facilitated variability approximation and enabled the calculation of Cohen's d effect sizes for pairwise comparisons between experienced and young raters. To compare the Area Under the Receiver Operating Characteristic (AUROC) curves between the ViT model and experienced and young endoscopists, DeLong's test was applied.
All statistical analyses were performed using Python, leveraging libraries such as NumPy, SciPy, Statsmodels, and Matplotlib. A significance threshold of α = 0.05 was maintained for all procedures.
Furthermore, gradient-weighted class activation mapping (Grad-CAM) 21 was employed to visually and interpretably assess the model's prediction.
Results
Trained ViT and ResNeSt model performances
To establish the model, the images were reviewed and divided into the training, validation, and testing subsets. Figure 1 shows the training and validation accuracies and losses for NHI (binary classification) and MES (four classifications). The ViT model MES (4-class) accuracies for training, validating, and testing are 92.28%, 58.80%, and 71.43%. The ResNeSt model MES (4-class) accuracies for training, validating, and testing are 87.82%, 58.21%, and 64.94%. The ViT model NHI (2-class) accuracies for training, validating, and testing are 98.81%, 82.48%, and 83.12%. The ResNeSt model NHI (2-class) accuracies for training, validating, and testing are 97.46%, 79.02%, and 79.22%. Table 3 presents the improved performance of the ViT model in comparison with the ResNeSt model for NHI, MES 0 versus 1–3, and MES 0–1 versus 2–3. Figure 2 illustrates representative Grad-CAM visualizations, highlighting the regions within the endoscopic images that the AI model considered most influential for predicting disease activity.

The training and validation accuracy and loss learning curves. Nancy learning curves: (a) accuracy and (b) loss. Mayo learning curves: (a) accuracy and (b) loss.

The figure illustrates representative Grad-CAM of endoscopy figure (left) and model prediction (right). (a) Endoscopic Mayo score of 3 (b) Endoscopic Mayo score of 0 (c) Nancy histological score of 0 (d) Nancy histological score of 4.
Performance of the ViT model and ResNeSt.

Note: MES: Mayo Endoscopic Score; ViT: vision transformer.
Comparison of the performance of the DL model and human endoscopists
Prediction of endoscopic remission (MES 0–1 vs 2–3)
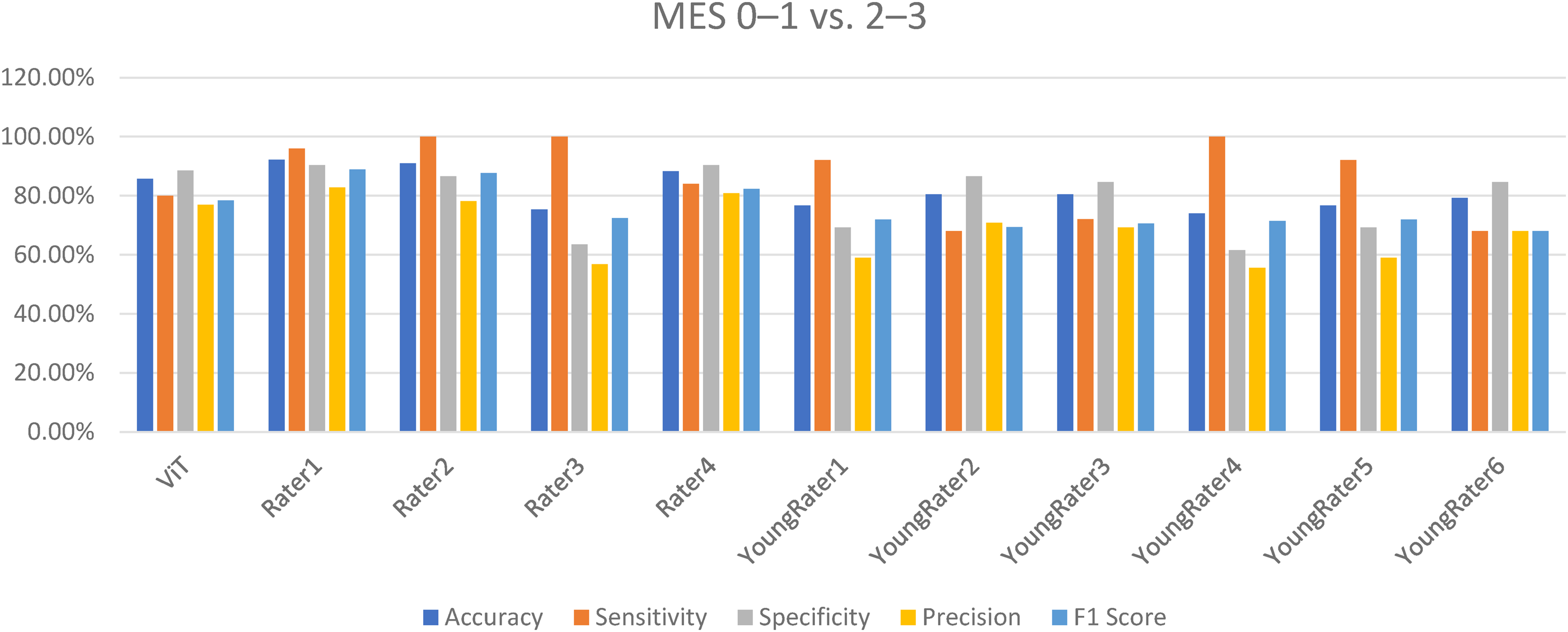
Performance of the ViT model and raters for predicting MES 0–1 versus 2–3.

The ROC of the ViT model and raters for predicting MES 0–1 and 2–3.
Prediction of complete mucosal healing (MES 0 vs 1–3)
Figure 5 demonstrates the classification performance metrics of the ViT model, experienced human raters, and young raters. The ViT model exhibited robust and consistent performance, with accuracy, sensitivity, specificity, and F1 score of 92.21%, 93.02%, 91.18%, and 93.02%, respectively. Experienced raters demonstrated lower and more variable results, with accuracy of 66.23%–80.52% and specificity of 23.53%–55.88%. Young raters showed similar variability, particularly in specificity and F1 score. ANOVA revealed significant group differences in sensitivity (p < .001), specificity (p < .001), F1 score (p = .004), and accuracy (p = .010). Post-hoc Tukey's HSD tests indicated that the ViT model significantly outperformed both rater groups across multiple metrics, especially in specificity and F1 score. Comparing the AUROC curves (Figure 6) demonstrated that the ViT model achieved a significantly higher AUROC than the young (p < .001) and experienced (p = .004) raters.
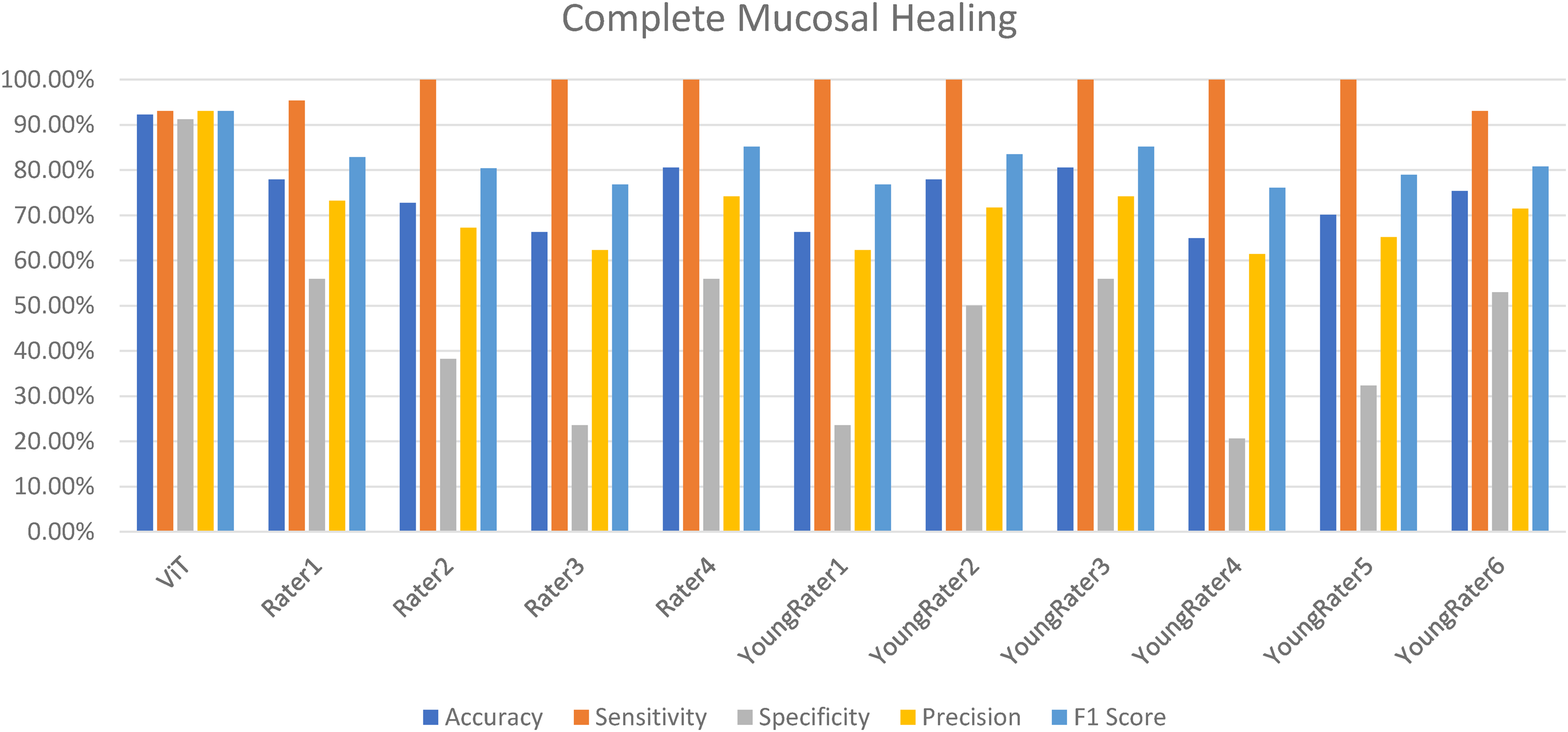
Performance of the ViT model and raters for predicting complete mucosal healing (MES 0 vs 1–3).
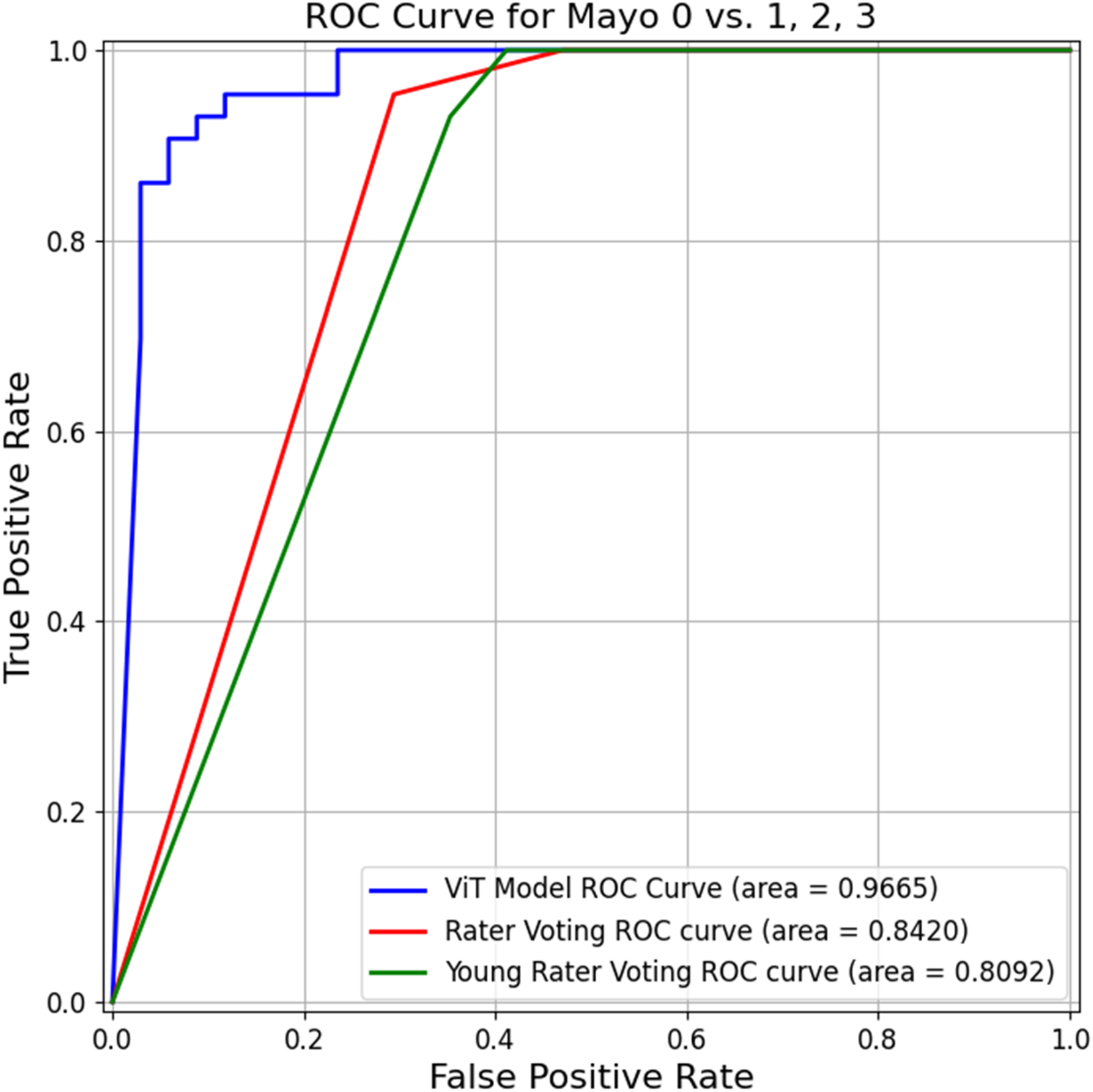
The ROC of the ViT model and raters for predicting complete mucosal healing (MES 0 vs 1–3).
Prediction of histological healing (NHI ≤ 1) or nonremission (NHI ≥ 2)
Figure 7 demonstrates the classification performance metrics of the ViT model, experienced human raters, and young raters. The ViT model exhibited a robust and balanced performance, achieving accuracy, sensitivity, specificity, and F1 score of 83.12%, 80.95%, 85.71%, and 82.02%, respectively. Contrarily, experienced raters demonstrated moderate variability, with accuracy and F1 scores of 76.62%–79.22% and 75.68%–81.40%, respectively. Young raters generally showed lower and more inconsistent performance, particularly in terms of sensitivity and specificity.

Performance of the ViT model and human endoscopists in predicting histological healing.
Statistical analysis revealed significant group differences in sensitivity (p = .027) and specificity (p = .038), with marginal significance observed for the F1 score (p = .065) and accuracy (p = .109). Post-hoc Tukey's HSD tests indicated that the ViT model significantly outperformed young raters in sensitivity and specificity; no significant differences were found between the ViT model and experienced raters. Furthermore, comparisons of the AUROC curves (Figure 8) demonstrated that the ViT model achieved a significantly higher AUROC than the young raters (p < .01); no significant difference was observed when compared to the experienced raters (p = .34).

The ROC of the ViT model and raters for predicting histological healing.
Discussion
The current study developed a DL-based approach to evaluate the performance of an AI system based on prospectively collected colonoscopy videos. Our study is the first to report the use of a ViT as a based architecture for model development based on prospectively collected endoscopic videos. 22 Instead of utilizing retrospectively stored still images as training material, which might raise concerns about selection bias, our findings are consistent with the findings of a previous report demonstrating the high accuracy of AI models in predicting endoscopic remission with MES and histological remission with the NHI via video-based evaluations.11,23 Our ViT model demonstrated improved performance in comparison to previous utilized ResNeSt CNN model. 11 The ViT architecture captures long-range dependencies across the entire endoscopic field through its self-attention mechanism. This enables the model to integrate global mucosal and vascular patterns rather than focusing on small image patches, thereby offering a more comprehensive assessment of disease extent and subtle inflammatory changes. To our knowledge, this study represents the first prospective evaluation of a ViT-based model using colonoscopy videos to predict both endoscopic and histologic activity in UC. Additionally, its classification performance was comparable to that of experienced endoscopists and surpassed that of young endoscopists in predicting histological remission and complete mucosal healing. These outcomes are clinically significant, as histological remission and mucosal healing serve as important surrogate markers for favorable patient outcomes in UC.4,23,24 Investigating the use of AI technologies is crucial, particularly in regions with a low IBD incidence, where experienced endoscopists who could manage UC are lacking.1,25
To date, several endoscopic scoring systems are utilized for assessing the UC severity, including MES and UCEIS. 26 The former is simple to classify the disease into four grades, and the latter further classifies the endoscopic findings into three components, including bleeding, erosion/ulceration, and vascular pattern. These scoring systems may standardize and objectify the evaluation of inflammation during colonoscopy, enabling consistent communication among specialists in daily clinical practice and clinical trial assessments. The former method is straightforward and easier to implement; however, it has several limitations, including the suboptimal interobserver agreement for intermediate scores (1–2), subjectivity in interpretation, poor characterization of severe disease due to the inability to distinguish between deep and superficial ulcers and differing prognostic implications between scores 0 and 1. The latter method was developed through a more rigorous process and incorporates objective parameters and demonstrates a strong prognostic value; however, it is more complex and thus less commonly used. 27 A recent meta-analysis 8 found pooled agreement rates of 0.58 and 0.66 for MES and UCEIS, respectively. Experts show less interobserver variability than nonexperts, and training programs can help reduce this variability. Accurate endoscopic assessment is crucial for evaluating disease activity, but the training process is resource-intensive, requiring considerable time, money, and effort to turn a young into an expert gastroenterologist. One approach to addressing this issue is to utilize an AI system that assists endoscopists in decision-making and provides consistent ratings.
AI reportedly can evaluate the endoscopic disease activity of IBD patients. Ozawa et al. 28 reported the use of a CNN-based CAD system with a high performance level with AUROCs of 0.86 and 0.98 to identify MES 0 and 0–1 cases, respectively, using large colonoscopy imaging datasets from UC patients. Stidham et al. 9 demonstrated the excellent performance of a CNN model using 16,514 images from 3082 unique patients in distinguishing endoscopic remission from moderate-to-severe disease, with a sensitivity of 83.0% and specificity of 96.0%. The agreement between CNN and experienced raters was also good for identifying the exact MES.
A subsequent study, 24 including our study, showed that AI models perform well in assessing the endoscopic severity of UC patients, which may aid in decision-making and standardizing clinical practice. Although most research comes from high-incidence IBD countries, 29 such as Western countries, Japan, and Korea, few studies have emerged from low-incidence areas, such as Taiwan.3,30 The present study contributes to the growing body of evidence supporting the potential of AI models in assessing the disease activity of UC. Most previous studies have demonstrated that the AI systems perform comparably to expert endoscopists, but only a few have investigated their performance in comparison with that of less experienced endoscopists.23,31,32 Kim et al. 31 constructed a DL model for distinguishing MES 0 and 1 UC cases. The model achieved an F1 score of 0.92, outperforming the consensus of seven young endoscopists, who tended to overestimate the disease activity. Qi et al. 33 reported a significant difference in the accuracy rates among the AI model, senior endoscopists, and best-performing young endoscopists, with values of 0.908, 0.849, and 0.773, respectively. Lo et al. 23 trained a CNN using MESs from 2561 images and 53 videos obtained from 645 patients. The model achieved an overall accuracy of 82%, with no significant difference in performance in comparison to that of the expert endoscopists. When used as a decision-support tool, the AI system improved the diagnostic accuracy of non-IBD specialists by 12%. We found that the AI model outperformed young endoscopists across three key outcomes, including estimation of histological healing, endoscopic remission, and complete endoscopic healing. Notably, the model demonstrated superior accuracy in distinguishing complete endoscopic healing (MES 0 vs 1–3) as compared to human endoscopists of different skill levels. These findings help bridge a critical gap in the literature by providing data that are particularly relevant to clinical settings with limited access to expert endoscopists, especially in low-incidence IBD regions.
One strength of our study is that the data were prospectively collected to evaluate the utility of using WLE to predict the histological activity of UC. The training videos were collected before the endoscopic biopsy to avoid the interference of postbiopsy bleeding for model training. Some studies utilized a special endoscopy system to develop an AI system for predicting histological activity. Maeda et al. 10 reported a 91% accuracy of a CAD system using endocystoscopy with a 520-fold ultramagnifying endoscope in predicting persistent histological inflammation with perfect reproducibility. Pieter et al. 34 reported diagnostic accuracy rates of 83.3% and 67.5% (p < .005), respectively, in predicting histological remission (Geboes score ≤ 2 B.0) for nonmagnifying single-wave endoscopy and WLE (Prototype, FUJIFILM, Tokyo, Japan), respectively. The diagnostic accuracy improved to 95.2%, whereas the case number increased from 42 patients to 112 patients. Despite these promising results, these utilized endoscope systems were not available for daily use in our clinical practice. Although white-light imaging only assesses the mucosal surface structure and vessel pattern and is not able to evaluate the inflammatory infiltrate in the lamina propria, the use of virtual chromoendoscopy (VCE) enhances the contrast of the mucosal and vascular architectures of UC patients, and the Paddington International virtual ChromoendoScopy ScOre (PICaSSO) showed a good correlation with histological activity. 35 Thus, an AI system might be able to detect the subtle changes using WLE without performing chromoendoscopy. Given that WLE systems are more widely available in clinical practice, developing such an AI system could have a considerable clinical value. Although most previous studies on AI for classifying endoscopic severity in UC have focused on endoscopic scores, only a few have used histological remission as the target outcome. The DNN developed by Takenaka et al., 36 which was trained on retrospectively collected still endoscopic images, reported a 92.9% accuracy in predicting histologic remission (Geboes score ≤ 3). Their subsequent study 37 utilized prospectively collected still endoscopic images, and the endoscopy videos showed an accuracy of 88.8% in predicting histological remission. They found a small discrepancy in the data between the AI model and pathologist assessment, which was due to poor bowel preparation. Iacucci et al. 11 developed an AI model using prospectively collected endoscopic videos to predict histological remission, as defined by NHI of ≤ 1), Robarts Histopathology Index (RHI) of ≤ 3, and PICaSSO Histologic Remission Index (PHRI) of 0. When trained with chromoendoscopy data, the model achieved accuracies of 83%, 81%, and 83% for NHI, RHI, and PHRI, respectively, with AUROCs of 0.83, 0.81, and 0.81, respectively. When applied to videos obtained via high-definition WLE, the model still performed well, achieving accuracies of 80%for RHI, 81% for NHI, and 80% for PHRI, with AUROCs of 0.80, 0.81, and 0.79, respectively. Notably, the use of VCE and high-resolution imaging was associated with an improvement in accuracy of up to 5%. Jiang et al. 33 reported a high accuracy of 91.28% for a CNN model in predicting histological remission (Geboes score ≤ 3 points), as compared to human endoscopists (87.46%). Our model achieved an 83.12% accuracy in predicting histological healing (NHI ≤ 1), which was comparable to the rate reported in the previous study, illustrating the potential utility of AI in this field. Moreover, our study was novel, as it was the first to compare the performance of the ViT model with those of human endoscopists of different skill levels. The AI model could achieve a better performance with more balanced sensitivity and specificity in comparison with human endoscopists, especially young endoscopists. Incorporating such a system into daily practice could help endoscopists more precisely perform biopsies from areas of interest (i.e. to selecting areas with endoscopic remission but without histological remission) and could reduce the need for unnecessary biopsies.
The current study has several limitations. First, it was conducted using prospectively collected, high-quality colonoscopy videos from a single endoscopy unit. Thus, the model's performance may not be generalizable to other clinical settings, particularly those using different endoscopic systems or those with suboptimal video quality. One important limitation of our study is the lack of external validation. All data used for model development and testing were obtained from a single center, using videos generated from the same endoscopic system and interpreted by local expert endoscopists with similar training backgrounds. This raises concerns regarding potential overfitting and limits the generalizability of the model to other clinical environments, particularly those with different equipment, image quality standards, or population characteristics. Future studies are needed to externally validate the model's robustness, assess its adaptability across diverse real-world settings, and refine the algorithm to support broader clinical integration. Nevertheless, this study represents the first of its kind, as it was conducted at a region with low IBD incidence, whereas most prior studies originated from high-incidence areas, such as Western countries, Japan, and Korea. These findings support the feasibility and potential utility of developing AI-assisted tools to aid endoscopists in assessing UC, particularly in settings with limited access to expert interpretation. Moreover, the ground truth used for model training was provided by expert endoscopists from the same institution, who all shared similar training backgrounds. Future work should focus on external multicenter validation across diverse endoscopic systems, patient populations, and disease severities to confirm model robustness. Second, the current model was trained exclusively on WLE videos. The model was not designed to detect neoplastic lesions or to integrate image-enhanced endoscopy modalities—such as narrow-band imaging, blue-laser imaging, and linked-color imaging—which represent important directions for future development aimed at improving its clinical applicability.1,22 Another limitation is that although variability between experienced and young endoscopists was discussed, interrater agreement was not formally quantified. While the present study focused on comparing the overall classification performance of the ViT model with aggregated human scores, future analyses should incorporate statistical measures such as Cohen's kappa or Fleiss’ kappa coefficients to objectively assess agreement between raters. This would provide a clearer understanding of the degree of variability in human interpretation and further contextualize the performance of the AI model.
Conclusion
Our novel deep learning-based model, trained on endoscopic videos, demonstrated performance comparable to that of experienced endoscopists and outperformed young endoscopists in predicting histological remission and complete mucosal healing in patients with UC. Such AI-based algorithms hold significant potential and clinical value, particularly in regions with a low incidence of IBD and limited endoscopic expertise.
Footnotes
Acknowledgments
Ethics statement
The study was approved by Institutional Review Board of Changhua Christian Hospital (Approval No. CCH IRB 230403).
Author contributions
Conceptualization: Hsu-Heng Yen and Yuan-Yen Chang; methodology: Yuan-Yen Chang and Han-Po Yang; formal analysis: Hsu-Heng Yen and Yuan-Yen Chang; writing—original draft preparation: Hsu-Heng Yen, Yuan Yen Chang, and Han-Po Yang; writing—review and editing: Yuan-Yen Chang and Hsu-Heng Yen. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Changhua Christian Hospital and Ministry of Science and Technology, Taiwan (Grant Nos. 110-CCH-IRP-020, 114-CCH-IRP-007, and 113-2221-E-025-007).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and material
The data that support the findings of this study are available from the corresponding author upon reasonable request.
