Abstract
Purpose
To address the reliance of task-specific deep learning models on large annotated datasets, this study investigates a Vector Quantized Variational Autoencoder (VQ-VAE) based few-shot learning framework for retinal vessel segmentation and disease classification.
Methods
A compact VQ-VAE was pretrained on unlabeled fundus photographs to learn transferable discrete representations. The pretrained encoder was used to initialize multiple downstream models, including segmentation networks (U-Net, SegNet, ERFNet) and three additional architectures (FR-UNet, Swin-Res-Net, RV-GAN), as well as classification networks (VGG-16, ResNet-50, EfficientNet-B0). Retinal vessel segmentation was evaluated on three public datasets (DRIVE, STructured Analysis of the Retina [STARE], and CHASE), while disease classification was assessed on the Retina and Ocular Disease Intelligent Recognition (ODIR) datasets. Segmentation performance was evaluated using Dice coefficient, Recall, Accuracy, Intersection over Union (IoU), mean IoU, and Average Offset Distance. Classification performance was assessed using accuracy, precision, recall, F1-score, and area under the receiver operating characteristic curve (AUC), with additional validation on the heterogeneous ODIR dataset.
Results
The VQ-VAE pretraining consistently improved performance under small-sample conditions. On the DRIVE dataset, Dice scores increased by approximately 2 percentage points across all segmentation backbones, with U-Net improving from 0.780 to 0.796 and SegNet from 0.670 to 0.692. Consistent performance improvements were also observed on the STARE and CHASE datasets. For disease classification on the Retina dataset, accuracy increased from 20% to 60% using only 70 labeled images. On the ODIR dataset, mean AUC improved across architectures, from 0.677 to 0.724 for VGG-16, 0.684 to 0.745 for ResNet-50, and 0.664 to 0.726 for EfficientNet-B0, indicating enhanced robustness across diverse disease categories and imaging conditions.
Conclusion
The proposed pretraining framework effectively reduces labeled data requirements while improving performance across multiple ophthalmic tasks, offering a scalable and resource-efficient solution for real-world clinical applications.
Introduction
Approximately 553 million people worldwide live with varying degrees of visual impairment, and timely detection is critical for preventing irreversible vision loss.1,2 Glaucoma, cataract, diabetic retinopathy, and age-related macular degeneration alone threaten the sight of hundreds of millions of people. Beyond these ocular conditions, retinal microvascular alterations also serve as sensitive indicators of systemic diseases such as hypertension, stroke, and coronary artery disease.
3
Systematic surveillance and prompt diagnosis are essential not only for preserving vision but also for guiding comprehensive patient management. Fundus photography is the only noninvasive imaging technique which provides direct,
In this study, motivated by these observations, we propose a label-efficient retinal analysis framework that integrates unsupervised Vector Quantized Variational Autoencoder (VQ-VAE) pretraining with lightweight transfer to both vessel segmentation and multidisease classification tasks. By leveraging unlabeled fundus photographs to learn transferable discrete representations, the framework reduces dependence on annotated data while maintaining strong task adaptability. This approach offers a scalable solution for retinal image analysis, with potential to support large-scale screening and improve accessibility in low-resource clinical settings.
Related work
Supervised deep learning for retinal vessel segmentation and diagnosis
Within automated retinal image analysis, vessel segmentation and multidisease classification remain cornerstone tasks. Quantitative vascular biomarkers, including the arteriovenous ratio (the ratio between retinal arteriolar and venular calibers), branching angles, tortuosity, and vessel density, correlate closely with disease severity and therapeutic response. Deep-learning methods are advancing quickly and are reshaping clinical workflows.6–8
In ophthalmology, they support automated diagnosis, population-level screening and even risk prediction for systemic disease based on retinal images.9,10 Fu et al. reframed vessel segmentation as a boundary-detection problem and trained a convolutional neural network to generate pixel-wise vessel-probability maps. 11 They later combined a multilabel network with a polar transform to segment the optic disc and cup, enabling automated glaucoma assessment. 12 Yadav et al. developed a two-stage pipeline for retinal-detachment detection that first applies a best-basis stationary wavelet packet transform to enhance key features, then processes the result with a modified VGG-19 followed by a bidirectional long short-term memory layer. 13 Mehta et al. proposed a multimodal model for glaucoma that merges macular OCT volumes, color fundus photograph and patient demographic plus clinical data, yielding higher diagnostic accuracy than single-source approaches. 14
Despite these advances, leading retinal models still rely on large, precisely annotated training sets. Producing pixel-level labels is extremely time-consuming, labor-intensive and costly, also sufficient data is often unavailable for rare disorders. Imbalances in patient demographics or imaging devices can bias the training data, causing poorer performance in underrepresented groups. In addition, fully supervised models are designed for a single task, and adapting them to a new task requires slow fine-tuning or complete retraining, which restricts their use in real-world settings where tasks change frequently. 15 Hence, developing techniques that retain high accuracy while reducing the requirement for large, meticulously annotated datasets holds significant promise for the field.16,17
Few-shot, generative, and self-supervised learning in ophthalmic imaging
Few-shot learning has emerged as an effective approach to overcome the limitations associated with insufficient training data, incomplete annotations, and the lack of flexibility when adapting to new tasks. In ophthalmology, few-shot learning shows particular promise for diagnosing rare ocular diseases or reducing biases in artificial intelligence systems. Burlina et al. demonstrated that few-shot methods can effectively enhance retinal diagnostics, even with very limited labeled images. 16 Han et al. further constructed a few-shot eye disease screening framework on a mixed multidisease fundus dataset, combining metric-based few-shot learning with style-transfer data augmentation to improve generalization under scarce labels. 18
Recently, researchers have used deep generative models to augment data, aiming to build generalized models from small datasets.19,20 Two notable approaches within this category are generative adversarial networks (GANs) and variational autoencoders (VAEs).21,22 The GANs, despite their strong generative capabilities, frequently encounter significant training difficulties such as unstable training processes and mode collapse. In contrast, VAEs provide a more stable approach by effectively modeling data distributions and reliably generating synthetic data.23,24 The VAEs are particularly effective at extracting meaningful abstract features from raw inputs using neural network architectures, thereby overcoming common issues related to convergence and evaluation often observed in GANs.25,26 Furthermore, VAEs allow explicit control over the latent representation vector, facilitating integration with representation-learning methods to improve downstream task performance. By learning smooth latent representations of input data in an unsupervised manner, VAEs consistently generate realistic new samples, making them especially useful for augmenting small ophthalmic imaging datasets.27,28 Complementary to these reconstruction-based approaches, Kukačka et al. showed that contrastive self-supervised pretraining on large collections of unlabeled fundus photographs can improve retinal vessel and lesion segmentation across datasets, shorten training time, and enhance few-shot performance when fine-tuned with only a few annotated examples. 29 In keratoconus diagnosis, Agharezaei et al. used a VAE to synthesize additional corneal topography images and improve a single binary classifier. 28 Our work instead employs a discrete VQ-VAE to learn a shared representation from unlabeled color fundus photographs that can be transferred to both vessel segmentation and multidisease classification under strict label constraints.
Materials and methods
Overview
This research introduced a novel approach employing a VQ-VAE to enhance the few-shot learning capabilities for ophthalmic image segmentation and classification tasks (Figure 1). 30 Specifically, we first performed unsupervised pretraining of a VQ-VAE on unlabeled retinal images and then transfered the encoder parameters to downstream segmentation and classification networks to reduce the impact of limited labeled data. Unlike previous VAE-based augmentation strategies for keratoconus diagnosis, which focus on generating synthetic images to support a single classification model, our framework reused a compact VQ-VAE encoder as a common backbone for both retinal vessel segmentation and multidisease classification with very few labeled fundus images. 28

Proposed Vector Quantized Variational Autoencoder (VQ-VAE) pretraining and weight-transfer pipeline for label-efficient retinal analysis. Unlabeled fundus photographs are used to train a compact VQ-VAE whose encoder weights are subsequently transferred to downstream vessel segmentation (U-Net, SegNet, ERFNet) and multidisease classification (VGG-16, ResNet-50, EfficientNet-B0) models trained on limited labeled samples.
Dataset
Vector Quantized Variational Autoencoder architecture and pretraining
The VQ-VAE architecture is composed of three main components: an encoder, a discrete vector quantization module, and a decoder. The encoder–decoder structure closely matches the architecture of the subsequent segmentation networks (U-Net, SegNet, ERFNet) to facilitate direct and efficient parameter transfer.35–37 The vector quantization module employs a learnable codebook, where the embedding dimensionality is aligned with the channel dimension of the encoder feature maps, ensuring compatibility with downstream networks.
The training objective for the VQ-VAE combines reconstruction and quantization losses and is defined as: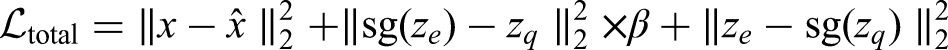
For downstream classification, we further address class imbalance by adopting a class-weighted cross-entropy loss, which reweights the contribution of each class proportionally to its inverse frequency. This adjustment increases the penalty for underrepresented classes, compelling the network to allocate greater learning capacity to minority categories without modifying the underlying loss structure. Formally, the weighted cross-entropy is written as:

We first pretrain the VQ-VAE using unlabeled retinal fundus images from the Retina dataset, which are resized to 448 × 448 pixels with three RGB channels. No data augmentation is applied at any stage in order to isolate the effect of VQ-VAE pretraining. After pretraining, the encoder weights are transferred to the downstream segmentation and classification networks, which are then trained on the corresponding small-sample labeled subsets. All models are optimized using Adam with a learning rate of 0.001 and a batch size of 8.
Parameter transfer strategy
Parameters obtained from the pretrained VQ-VAE encoder, covering convolutional and Batch Normalization layers, are directly transferred and shared with corresponding layers in the downstream segmentation and classification networks. All transferred parameters are initialized from the pretrained model and remain trainable during fine-tuning without any freezing steps.
Mathematically, this strategy can be interpreted through optimization lens: initializing with pretrained parameters
The adaptive fine-tuning without parameter freezing allows for smooth knowledge integration while maintaining the representational capacity to capture task-specific nuances. This approach has demonstrated superior performance across various downstream tasks, validating its efficacy in transferring pretrained representations to diverse application domains.
Downstream tasks
Retinal vessel segmentation
The retinal vessel segmentation task utilizes the standard DRIVE dataset, comprising 20 training images and 20 testing images. Segmentation models (U-Net, SegNet, ERFNet) receive weights transferred from the pretrained VQ-VAE encoder. To comprehensively assess segmentation quality, multiple evaluation metrics are employed, including Dice coefficient, Recall, Accuracy, Intersection over Union (IoU), mean IoU (mIoU) and the Average Offset Distance (AOD). These metrics collectively characterize overlap consistency, pixel-level correctness, sensitivity to thin vessels, and boundary-level discrepancies. The Dice coefficient measures the overlap between the predicted vessel region 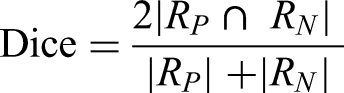
Recall evaluates the model's ability to correctly detect vessel pixels, particularly important for thin or low-contrast vessels, and is given by
Pixel Accuracy quantifies the proportion of correctly classified pixels across the entire image and is defined as
Intersection over Union measures the ratio between the intersection and union of predicted and ground truth vessel masks:
To account for the severe vessel–background imbalance, the mIoU is also computed by averaging the IoU of the vessel and background classes:
Finally, the AOD evaluates boundary-level consistency by quantifying misclassified pixels relative to the reference boundary length. Following prior definitions, AOD is computed as
To further evaluate the robustness and generalization of the proposed methodology, two complementary experimental settings were designed. First, the three baseline segmentation models (U-Net, SegNet, and ERFNet) were further validated on two additional retinal vessel segmentation benchmarks, namely the STARE and CHASE datasets, in order to assess their cross-dataset generalization performance beyond the DRIVE dataset.
Second, three state-of-the-art retinal vessel segmentation models (FR-UNet, 38 Swin-Res-Net, 39 and RV-GAN 40 ) were incorporated for comprehensive comparison. These models were evaluated across all three datasets (DRIVE, STARE, and CHASE) under a unified training and evaluation protocol. The VQ-VAE pretraining was used solely as an initialization strategy. After loading the encoder weights, each model was trained on the 20 labeled training images of the DRIVE dataset and evaluated on the remaining 20 DRIVE test images. Due to the limited number of annotated samples in the STARE and CHASE datasets, models trained on DRIVE were directly transferred and tested on STARE and CHASE without further fine-tuning, enabling an assessment of cross-dataset generalization under identical training conditions.
Retinal disease classification
The classification task uses the Retina dataset containing 601 images categorized into four distinct classes: normal and three common retinal diseases. A subset of 70 images serves as the training set, with another 30 as the testing set. The 70-image subset was constructed using stratified sampling, resulting in a relatively balanced distribution with 14 samples from each disease category and 28 samples from the normal class. Classification networks including VGG-16, EfficientNet-B0 and ResNet-50 are initialized with the pretrained VQ-VAE encoder weights. We measure classification performance using accuracy, precision, recall, and F1-score.
To further evaluate the robustness and generalizability of the learned representations, we conducted additional classification experiments on the ODIR dataset, a large and heterogeneous multicenter fundus image collection covering eight diagnostic categories. The dataset contains 7000 fundus photographs from patients with confirmed clinical labels. To quantitatively assess diagnostic performance across both settings, we computed receiver operating characteristic (ROC) curves and derived the corresponding area under the ROC curve (AUC) for each diagnostic category as well as the macro-averaged AUC. This evaluation provides a robust and comprehensive measure of model discriminative ability across heterogeneous imaging conditions, variations in acquisition devices and the broad disease spectrum represented in ODIR.
Implementation
All experiments were conducted on a high-performance workstation equipped with an NVIDIA RTX 4090 GPU (24 GB VRAM) and 64 GB RAM. Model development and training were implemented in PyTorch. All input fundus images were uniformly resized to 448 × 448 pixels with three RGB channels, and no additional normalization or data augmentation was applied in order to isolate the effect of VQ-VAE pretraining. For both segmentation and classification tasks, models were trained using the Adam optimizer with a fixed learning rate of 0.001 and a batch size of 8. For supervised fine-tuning, we employed an early stopping strategy based on loss stabilization to mitigate overfitting.
Statistical analysis
To assess the robustness and significance of performance differences between baseline models and those initialized with VQ-VAE pretraining, we conducted paired statistical analyses across segmentation baseline backbones (U-Net, SegNet and ERFNet). For each backbone, repeated training runs were performed under identical configurations, generating paired performance samples for Dice, Recall, Accuracy, IoU, mIoU, and AOD. Normality of paired differences was assessed using the Shapiro–Wilk test. Metrics exhibiting approximately normal distributions were evaluated using a paired
Experiments and results
Experimental design
In this study, two ophthalmic image analysis tasks including retinal vessel segmentation and multidisease classification were conducted under stringent label constraints to evaluate the efficacy of the proposed VQ-VAE-based pretraining strategy. The retinal vessel segmentation task utilized the DRIVE dataset, STARE and CHASE datasets, while the disease classification task employed the Retina dataset and ODIR dataset. Three network architectures (U-Net, SegNet, and ERFNet) and three state-of-the-art segmentation networks (FR-UNet, Swin-Res-Net, and RV-GAN) were also evaluated for comprehensive performance comparison, while VGG-16, ResNet-50, and EfficientNet-B0 were adopted for the classification task.
For the segmentation task, the comparison involved small-sample baseline (S-Base, 20 training images) versus small-sample with VQ-VAE (S-VQ, 20 training images with pretrained encoder). For the classification task, we evaluated three supervised training paradigms on the Retina dataset: large-sample baseline (L-Base, 570 training images), small-sample baseline (S-Base, 70 training images), and small-sample with VQ-VAE pretraining (S-VQ, 70 training images with pretrained encoder).
For comparative experiments, retinal vessel segmentation performance was further evaluated on two additional datasets. Both three baseline models (U-Net, SegNet, and ERFNet) and three state-of-the-art models (FR-UNet, Swin-Res-Net, and RV-GAN) were compared under settings with and without VAE pretraining. Given the limited number of samples in the additional datasets, models trained on the DRIVE dataset were directly transferred and evaluated on these datasets without further fine-tuning, enabling an assessment of cross-dataset generalization performance. For the classification task on the ODIR dataset, two comparative experimental settings were designed to further assess the effectiveness of the proposed VQ-VAE pretraining strategy under heterogeneous imaging conditions. In the first setting, 5600 images were used for unsupervised VQ-VAE pretraining, and the remaining 1400 images were allocated for supervised classification, including 800 training images, 300 validation images, and 300 test images. Classification networks in this setting were initialized with the pretrained VQ-VAE encoder. In the second setting, the same supervised data split was used, but models were trained from random initialization without pretraining. This controlled design enabled a fair evaluation of the impact of VQ-VAE pretraining on classification performance using identical labeled data.
Retinal vessel segmentation
Table 1 and Figure 2 present the quantitative and statistical performance of the three segmentation backbones on the DRIVE dataset. Across all models, incorporating VQ-VAE pretraining leads to consistent and measurable improvements under the small-sample condition. For U-Net, the Dice coefficient increases from 0.780 ± 0.034 (S-Base) to 0.796 ± 0.034 (S-VQ), and Recall rises markedly from 0.721 ± 0.068 to 0.789 ± 0.073, representing one of the largest relative improvements among all backbones. The IoU and mIoU also show clear numerical gains (0.641 ± 0.045 vs. 0.663 ± 0.045 for IoU; 0.802 ± 0.024 vs. 0.812 ± 0.024 for mIoU), indicating more complete and structurally coherent vessel extraction. The SegNet and ERFNet display similar patterns. SegNet improves from 0.670 ± 0.053 to 0.692 ± 0.048 in Dice and from 0.544 ± 0.070 to 0.616 ± 0.074 in Recall, while ERFNet improves from 0.708 ± 0.045 to 0.721 ± 0.029 in Dice and from 0.633 ± 0.064 to 0.683 ± 0.057 in Recall. These increases across multiple metrics, including Dice gains of 2 percentage points and recall gains of 6 percentage points, consistently suggest enhanced sensitivity to small-caliber and low-contrast vessels. Accuracy remains largely stable across models, reflecting the dominance of background pixels and its limited discriminative power in retinal vessel segmentation.

Comparison of retinal vessel segmentation performance on the DRIVE dataset with and without Vector Quantized Variational Autoencoder (VQ-VAE) pretraining across three backbone networks (ERFNet, SegNet and U-Net).
Retinal vessel segmentation results on DRIVE (20 test images).

Figure 3 provides qualitative examples of the segmentation outputs, complemented by the errormaps shown in Figure S1. Models initialized with VQ-VAE produce more continuous vessel trees, fewer fragmented bifurcations, and more accurate delineation of narrow, low-contrast vessels compared with baseline models. The accompanying errormaps confirm these observations by showing reduced false-negative regions along thin vessels and fewer false-positive artifacts in background areas. Together, these qualitative results support the statistical findings and demonstrate that the discrete morphological priors learned by VQ-VAE are effectively transferred to downstream segmentation tasks.

Retinal vessel segmentation on DRIVE dataset. The top row represents results without Vector Quantized Variational Autoencoder (VQ-VAE) pretraining (S-Base), and the bottom row depicts results with VQ-VAE pretraining (S-VQ), demonstrating enhanced detail recovery and segmentation accuracy.
On the DRIVE dataset, the effectiveness of the proposed VQ-VAE pretraining strategy was further validated on three state-of-the-art segmentation models (FR-UNet, Swin-Res-Net, and RV-GAN). For all three models, VQ-VAE pretraining consistently improved segmentation performance across multiple metrics (Table S1). Specifically, for FR-UNet, VQ-VAE pretraining increased the Dice score from 0.7589 to 0.7706 and the IoU from 0.6126 to 0.6288, with a corresponding mIoU improvement from 0.7856 to 0.7949. For Swin-Res-Net, the Dice score improved from 0.7628 to 0.7809, while the IoU increased from 0.6174 to 0.6411, and the mIoU rose from 0.7881 to 0.8014. Similarly, RV-GAN achieved higher Dice and IoU scores after VQ-VAE pretraining. These results demonstrate that the proposed pretraining strategy remains effective even when applied to advanced segmentation architectures, confirming its compatibility with recent state-of-the-art models.
To evaluate cross-dataset transferability, models trained on the DRIVE dataset were directly tested on the STARE and CHASE datasets without further fine-tuning. Figure 4 showed Qualitative retinal vessel segmentation results on the STARE and CHASE datasets. As shown in Table S2 and Table S3, VQ-VAE pretraining led to substantial and consistent performance improvements across all three SOTA models. On the STARE dataset, VQ-VAE pretraining improved the Dice score of FR-UNet from 0.5330 to 0.6112, and increased its IoU from 0.4003 to 0.4644. For Swin-Res-Net, Dice increased from 0.5753 to 0.6069, while IoU improved from 0.4245 to 0.4544. RV-GAN showed a Dice improvement from 0.5640 to 0.6036, accompanied by an IoU increase from 0.4185 to 0.4556. On the CHASE dataset, Dice scores increased from approximately 0.27–0.51 to 0.48–0.56, with corresponding Accuracy values increasing from approximately 0.94 to 0.95 after VQ-VAE pretraining. Notably, the performance gains on STARE and CHASE are substantially larger than those observed on DRIVE, indicating that VQ-VAE pretraining significantly enhances model robustness under domain shift and limited-data conditions.

Qualitative retinal vessel segmentation results on the (A) STructured Analysis of the Retina (STARE) and (B) CHASE datasets. For each dataset, the original retinal image and corresponding ground truth are shown, followed by segmentation outputs generated by FR-UNet, Swin-Res-Net, and RV-GAN under baseline training (without Vector Quantized Variational Autoencoder [VQ-VAE] pretraining) and VQ-VAE–pretrained settings.
Retinal disease classification
Table 2 and Figure 5 detail the classification performance across the three tested architectures and training regimes. Significant performance gains are observed when using the VQ-VAE pretrained encoder, particularly in conditions of limited labeled data. For example, the VGG-16 network improves by 40 percentage points, from an accuracy of 0.200 in the small-sample baseline (S-Base) to 0.600 with VQ-VAE pretraining (S-VQ), which represents a threefold relative increase and even exceeds the performance of the large-sample baseline (0.567). Figure 6 visually presents the confusion matrices for each model and training strategy. The matrices clearly indicate that VQ-VAE pretraining considerably reduces misclassification rates, especially in categories that are prone to confusion in small datasets, such as cataract and glaucoma, where both conditions can present with diffuse changes in overall image brightness and optic-disc appearance rather than highly localized lesions. The confusion matrices of the S-VQ models (bottom row) closely approximate the diagonal-dominant structure typical of well-performing classifiers, which is comparable to those of the L-Base models (top row).

Comparative quantitative performance for retinal disease classification on Retina dataset. Blue bars represent S-Base, red bars S-VQ, and violet bars represent L-Base (classification only) on Retina dataset.

Confusion matrices illustrating classification performance improvements achieved by Vector Quantized Variational Autoencoder (VQ-VAE) pretraining (bottom row) compared to small-sample baseline (middle row) and large-sample baseline (top row) on Retina dataset.
Four-class disease classification results on the Retina dataset (30 test images).

Figures 7–8 report the classification performance on the ODIR dataset in terms of per-disease and overall AUC for three representative backbone networks. Overall, models initialized with pretrained VQ-VAE encoders achieved consistent improvements in average AUC across all three architectures. Specifically, the mean AUC increased from 0.677 to 0.724 for VGG-16, from 0.684 to 0.745 for ResNet-50, and from 0.664 to 0.726 for EfficientNet-B0, indicating that VQ-VAE pretraining enhances discriminative representation learning under heterogeneous and multidisease settings. At the individual disease level, VQ-VAE pretraining resulted in higher AUC values for most categories across different backbones. For cataract, AUC increased from 0.806 to 0.933 (VGG-16), 0.891 to 0.935 (ResNet-50), and 0.860 to 0.968 (EfficientNet-B0). Similarly, pathological myopia showed consistent improvements, with AUC rising from 0.893 to 0.938, 0.867 to 0.980, and 0.818 to 0.992 for VGG-16, ResNet-50, and EfficientNet-B0, respectively. Moderate AUC gains were also observed for normal, diabetes, and hypertension classes across most architecture. For other categories, including glaucoma, age-related macular degeneration, and other abnormalities, performance remained comparable across settings, with improvements observed in some backbones and marginal decreases in others.

Area under the receiver operating characteristic curve (AUC) comparison on the Ocular Disease Intelligent Recognition (ODIR) dataset. Bar plots showing per-disease AUC values for multidisease classification on the ODIR dataset using VGG-16, ResNet-50, and EfficientNet-B0, trained with and without Vector Quantized Variational Autoencoder (VQ-VAE) pretraining.

Receiver operating characteristic (ROC) curves on the Ocular Disease Intelligent Recognition (ODIR) dataset. The ROC curves for multidisease classification on the ODIR dataset using VGG-16, ResNet-50 and EfficientNet-B0, trained with and without Vector Quantized Variational Autoencoder (VQ-VAE) pretraining. The false positive rate (FPR) is shown on the
Discussion
In this study, we introduced a VQ-VAE-driven framework for small-sample retinal vessel segmentation and multi-disease fundus classification. To the best of our knowledge, this is the first empirical exploration of VAE-based few-shot deep-learning models simultaneously tackling these two cornerstone ophthalmic tasks. Training high-performing deep networks typically requires large, meticulously annotated datasets, a prerequisite that is often impractical in ophthalmology, where expert graders are scarce and disease prevalence is highly imbalanced. Insufficient or biased data routinely cause degraded performance in underrepresented sub-populations. We pretrained a compact VQ-VAE on unlabeled photographs and transferred the encoder to six downstream backbones. This strategy yielded consistent gains in Dice, IoU, accuracy, precision, recall, and F1-score while using only 20–70 labeled images. The generative pretraining strategy therefore alleviated annotation burden without sacrificing diagnostic fidelity, offering a scalable solution for real-world screening settings.
While several state-of-the-art retinal vessel segmentation models, such as FR-UNet, Swin-Res-Net, and RV-GAN, have reported strong performance on the DRIVE dataset under fully supervised settings,38,40,41 their evaluations assume access to sufficient annotated data and do not explicitly address performance under label-scarce conditions or cross-dataset generalization. In contrast, our study focused on improving segmentation behavior when only a very limited number of labeled samples were available, by introducing VQ-VAE pretraining as a model-agnostic initialization strategy. Importantly, we directly evaluated FR-UNet, Swin-Res-Net, and RV-GAN with and without VQ-VAE pretraining under the same training protocol. The results show that VQ-VAE pretraining not only improves segmentation performance on the DRIVE dataset but also leads to consistently higher performance when models trained on DRIVE are directly tested on the STARE and CHASE datasets without additional fine-tuning. These findings indicate that VQ-VAE pretraining enhances the transferability and robustness of state-of-the-art segmentation models under domain shift and limited supervision, rather than merely improving performance on a single benchmark. Moreover, compared with other self-supervised learning methods such as SimCLR and BYOL, our approach is more suitable for small data regimes where contrastive learning or adversarial training often becomes unstable or ineffective.42,43 The VQ-VAE provides a lightweight and data-efficient alternative for learning transferable morphological priors from a modest number of unlabeled images. Thus, the contribution of this study lies in improving learning stability and segmentation robustness when annotation is limited, offering a practical solution for real-world ophthalmic applications.
Few-shot learning has become a prominent remedy for data scarcity. Meta-learning approaches such as the difficulty-aware, task-augmentation model (DaTa-ML) improve diabetic-retinopathy grading by explicitly modeling task complexity. 44 Generative adversarial networks have also been leveraged: DeepDrRVO augments color-fundus photographs to boost early recognition of retinal vascular occlusion, while GAN-based augmentation in OCT images significantly improves the diagnosis of rare retinal diseases under few-shot constraints.45,46 Gradient-guided Retina-TransNet advances retinal-vessel segmentation by combining transformers with episodic learning. 47 These transformer-based segmentation networks report improved delineation of thin and low-contrast vessels compared with purely convolutional baselines, which indicates that modeling long-range spatial dependencies and global context is beneficial for fundus images. Despite these advances, these methods still exhibit notable drawbacks, such as unstable training, hyperparameter sensitivity, and limited adaptability across diverse ophthalmic tasks. Recent ophthalmology-specific applications demonstrate the utility of VQ-VAE in medical imaging. For example, Li et al. integrated VQ-VAE into a generative video modeling pipeline to compress and represent complex ophthalmic image sequences efficiently, which may indirectly benefit downstream tasks like classification. 48 Meanwhile, Jebril et al. used VQ-VAE for unsupervised anomaly detection in OCTA by learning normal perfusion patterns from healthy scans and localizing deviations as anomalous regions without dense pixel-level labels. 49 These works primarily treat VQ-VAE as a task-specific generative or anomaly detection model. By contrast, we pretrain a compact VQ-VAE on unlabeled color fundus images and reuse its encoder as a shared initialization for several supervised vessel segmentation and multidisease classification networks in a few-shot regime. This design allows a single discrete representation to directly support multiple downstream retinal tasks while still benefiting from limited annotated data.
The VQ-VAEs replace the continuous latent space of traditional VAEs with a discrete codebook of embeddings, offering three key advantages for few-shot learning. First, discretization prevents posterior collapse and enhances latent representation diversity. Second, the shared codebook acts as a regularizer, stabilizing training in label-scarce settings. Third, the learned tokens can be reused across tasks with minimal retraining cost, enabling effective transfer learning. These benefits have been validated across a range of medical imaging applications in the past three years. Majoral and Domnich introduced
This study has several limitations. Experiments were conducted on moderate-sized public datasets, and external multicenter validation was not performed. Consequently, important sources of dataset bias, including demographic underrepresentation, uneven disease prevalence and variation in imaging devices, were only partially evaluated. Potential domain shifts related to differences in camera types, patient populations, or clinical environments may therefore influence generalizability in real-world applications. Additionally, we did not investigate semisupervised fine-tuning strategies, more aggressive data augmentation techniques or fairness-oriented evaluation frameworks, all of which may help reduce bias and improve robustness. Practical challenges associated with clinical deployment also remain, such as variability in image quality, atypical patterns of disease presentation, and the need for prospective and multi-institutional validation. Future work will extend evaluation to larger and more diverse cohorts, incorporate data from multiple centers and ethnic groups, and explore hybrid VQ-VAE and transformer architectures within semisupervised or continual-learning settings. Transformer-based models may be particularly advantageous for modeling long-range spatial relationships in fundus photographs, including vascular topology and the structural relationship between the optic disc and macula, which may not be fully captured by conventional convolutional networks. This motivation is consistent with recent retinal vessel segmentation and diabetic retinopathy studies that employ transformer or hybrid CNN–transformer backbones to combine local detail with global context.
Conclusion
We presented a label-efficient retinal analysis pipeline that couples unsupervised VQ-VAE pretraining with lightweight transfer to segmentation and multidisease classification networks. Using only 20–70 annotated fundus photographs, our method consistently outperformed task-specific baselines and even rivaled large-sample regimes on DRIVE and Retina datasets, underscoring the strength of discrete generative representations as a universal scaffold for ophthalmic AI. By markedly lowering annotation requirements, the framework can accelerate population-level screening programs and expand access in resource-constrained settings. Future work will validate the approach in multicenter cohorts, explore semisupervised fine-tuning, and integrate transformer backbones to further enhance generalizability by explicitly capturing long-range dependencies and global retinal context that are difficult to model with convolutional networks alone.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076261433086 - Supplemental material for A unified VQ-VAE framework for few-shot retinal vessel segmentation and multidisease classification
Supplemental material, sj-docx-1-dhj-10.1177_20552076261433086 for A unified VQ-VAE framework for few-shot retinal vessel segmentation and multidisease classification by Haojun Yu, Zongcai Tan, Huazhen Liu and Xinyu Xu in DIGITAL HEALTH
Footnotes
Acknowledgments
The authors thank all the authors for their research contributions.
Contributorship
Haojun Yu contributed to conceptualization, data curation, formal analysis, investigation, writing – original draft. Zongcai Tan contributed to conceptualization, methodology, validation, visualization, writing – original draft. Huazhen Liu contributed to data curation, formal analysis, methodology, supervision, writing – review & editing. All authors read and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
