Abstract
Background
Early diagnosis of pulmonary nodules is crucial for improving the survival rate of lung cancer patients. However, significant variability in nodule size, shape, and anatomical location presents ongoing challenges for automated detection systems, often resulting in high false-positive rates.
Objective
This study aims to develop a dual-stage pulmonary nodule detection framework based on cross-layer attention fusion, with the goal of improving sensitivity while reducing false positives in chest CT scans.
Methods
We propose a two-stage detection pipeline. In the candidate detection stage, we design an Attention-guided Spatial and Channel Residual Module that integrates multi-scale residual connections with cross-dimensional attention to enhance discriminative features while preserving spatial detail. For false positive reduction, we introduce a Multi-scale Progressive Perception Network, which processes candidates across three anatomical resolutions through parallel branches and integrates top-down semantic fusion with localized attention. The model is evaluated on the LUNA16 dataset.
Results
Experimental results demonstrate that the proposed method achieves a sensitivity of 90.0% at 0.55 false positives per scan on the LUNA16 dataset. Compared to state-of-the-art approaches, our framework provides a favorable balance between sensitivity and precision.
Conclusions
The proposed dual-stage detection framework effectively enhances the performance of pulmonary nodule detection by incorporating cross-layer attention mechanisms and multi-scale feature integration. These findings suggest its potential for clinical deployment in computer-aided lung cancer screening.
Keywords
Introduction
Lung cancer remains one of the most formidable threats to global health, ranking highest in both incidence and mortality among all cancer types. 1 Despite recent advances in therapeutic strategies, most patients are diagnosed at advanced stages due to the paucity of overt symptoms in early disease, resulting in limited treatment efficacy and poor prognosis. Clinical evidence suggests that overcoming this diagnostic bottleneck hinges on early detection, precise diagnosis, and timely intervention—factors that are decisive for improving five-year survival rates and long-term outcomes. 2 Central to early diagnosis is the accurate interpretation of medical imaging, particularly the effective identification and localization of pulmonary nodules, which constitute the critical control point in arresting disease progression. 3 Chest computed tomography (CT) stands as the most sensitive and widely adopted modality for nodule detection. However, conventional image reading relies heavily on the expertise and subjective judgment of radiologists, making it vulnerable to inter-observer variability and prone to missed or false diagnoses. The enormous volume and complex information content of CT datasets further exacerbate this challenge, as rapid yet accurate analysis of hundreds of slices places substantial cognitive demands on clinicians. 4 Consequently, there is a pressing need for computer-aided diagnosis (CAD) systems to assist radiologists by first flagging potential nodules with high sensitivity, and then suppressing false positives to streamline decision-making and enhance diagnostic accuracy. 5
With the rise of computer vision and artificial intelligence, deep learning-based CAD for automated pulmonary nodule detection has emerged as a major research focus. By training on large, annotated CT datasets, these systems learn to extract discriminative features associated with nodules, thereby achieving superior sensitivity and specificity compared to traditional, hand-crafted feature approaches.6–8 Numerous CAD solutions have been proposed to support radiological workflows; for example, Lu et al. 9 and Gu et al. 10 demonstrated conventional pipelines based on manually engineered features. Conventional methods often fall short in detecting pulmonary nodules with diverse shapes, sizes, textures, and anatomical locations, resulting in suboptimal detection performance. In contrast, deep learning approaches have demonstrated superior capability in capturing complex nodule characteristics compared to handcrafted features. 11 For instance, Setio et al., 12 Jiang et al., 13 Zuo et al., 14 and Xie et al. 15 proposed two-dimensional convolutional neural network (2D CNN)-based frameworks to extract nodule features. However, these 2D architectures inherently lack the ability to leverage the rich spatial context embedded in three-dimensional CT volumes. To address this limitation, Huang et al., 16 Wang et al., 17 Cao et al., 18 and Zhu et al. 19 have introduced 3D CNN-based detection models, achieving improved accuracy and robustness in pulmonary nodule identification.
Despite these advances, high false positive rates persist due to the extreme variability in nodule size, morphology, and anatomical context. To mitigate this issue, we propose a two-stage detection framework driven by cross-layer attention fusion, designed to enhance detection precision while effectively suppressing false positives. In the first stage, multi-layer feature fusion extracts local morphological details of candidate nodules, augmented by a dual-dimensional channel–spatial attention mechanism that strengthens global spatial awareness and positional encoding for precise coarse localization. The second stage employs an adaptive residual 3D CNN architecture, which captures hierarchical contextual features through multi-scale volumetric convolutions and further refines detection via an attention-guided fusion strategy to reduce false positives.
Extensive experiments on the LUNA16 dataset validate the effectiveness and competitiveness of our approach. The principal contributions of this work are as follows:
We design a multi-level feature fusion anchor prediction architecture that integrates intermediate semantic representations with upsampled high-resolution features, achieving precise characterization of small and morphologically complex nodules. We introduce the Attention-guided Spatial and Channel Residual Module (ASCRM), which simultaneously captures multi-scale features via parallel spatial-attention and channel-recalibration units, preserving spatial integrity while amplifying responses to subtle lesions. We develop an adaptive false-positive suppression network that utilizes cross-scale feature fusion to integrate hierarchical context, resulting in a reduction of false positives while preserving sensitivity.
Our framework demonstrates a sensitivity of 90.0% at just 0.55 false positives per scan on LUNA16, indicating its strong potential for clinical deployment.
Methods
The proposed pulmonary nodule detection framework, shown in Figure 1, comprises two sequential stages: candidate generation and false-positive reduction. In the first stage, a U-shaped encoder–decoder network employs cascaded feature enhancement via the ASCRM module, fused with spatial coordinate priors, to preserve fine-grained CT details while leveraging multi-scale feature aggregation to boost candidate recall. To address the challenge of nodules adherent to the pleura or vasculature, as well as those exhibiting atypical morphology or diminutive size, we incorporate online hard-example mining during training to assign greater weight to these difficult cases—thereby enhancing sensitivity at the cost of an elevated false-positive rate. In the second stage, we introduce a three-branch progressive classifier in which each branch processes a specific scale of input; a cross-branch feature interaction mechanism enables bidirectional information flow and fusion across resolutions, enhancing the discrimination of true nodules from imaging artefacts.
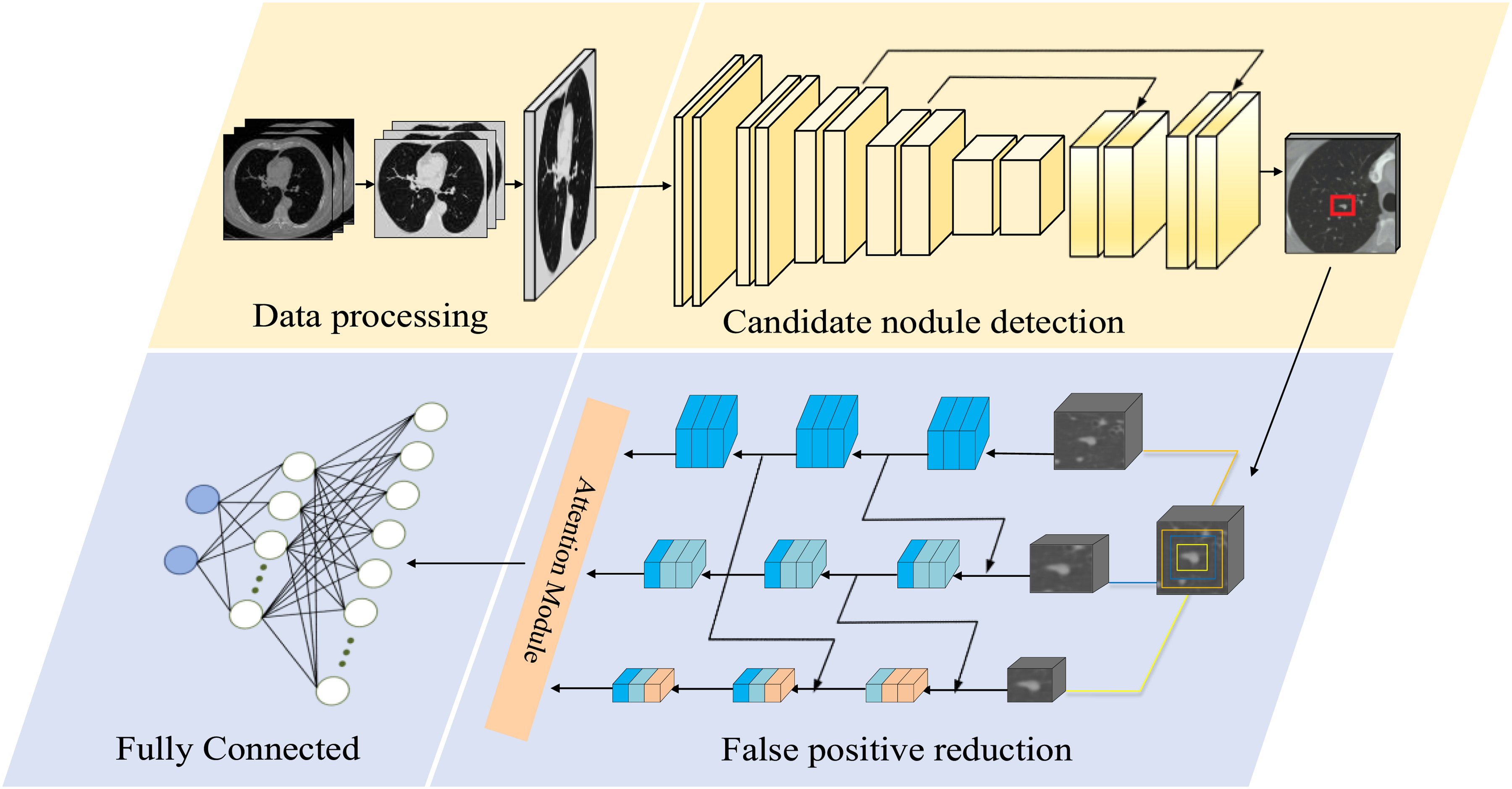
Workflow of the proposed pulmonary nodule detection framework.
To characterize the computational cost of the two-stage framework, Table 1 reports the training time, parameter count, average computation time, and floating-point operations (FLOPs) for each stage. The training time is measured per fold. Although the first-stage candidate detector has fewer parameters, GPU memory constraints require each CT scan to be processed in multiple 128 × 128 × 128 sub-volumes, followed by block-level reconstruction during training and validation, leading to longer runtime. The larger 3D input blocks in this stage also result in substantially higher FLOPs than those of the second stage. In contrast, the second-stage false positive reduction network has more parameters but operates only on 32 × 48 × 48 candidate nodule patches, which greatly reduces the computation scope and yields much lower FLOPs and faster training/inference.
Key performance metrics for two stages of pulmonary nodule detection.

Architecture of the cross-layer feature-fusion pulmonary nodule candidate detection network
In this section, we present a three-dimensional convolutional neural network (3D CNN) based on cross-layer feature fusion for the detection of pulmonary nodule candidates. The network takes a CT volume of size 128 × 128 × 128 voxels as input and adopts an encoder–decoder architecture as its backbone (Figure 2). To effectively capture the diverse morphological characteristics and complex spatial distribution of nodules, the model must possess strong feature representation capabilities, balancing global semantic understanding with fine-grained spatial detail preservation.

Architecture of the proposed candidate nodule detection network.
However, conventional convolutional modules exhibit limitations in modeling spatial and channel-wise attention. While they are proficient in extracting local spatial features, they lack explicit modeling of spatial positional information and offer limited interaction across channels. This hampers the network's ability to capture semantic dependencies and spatial regularities across scales—particularly problematic for nodules that are morphologically heterogeneous, diminutive, or ambiguously bounded. Furthermore, traditional convolutional operations are often deficient in integrating spatial and channel attention mechanisms, making it difficult to selectively enhance salient features, ultimately impairing detection sensitivity and accuracy.
To address these challenges, we introduce an ASCRM to enhance the network's representational and semantic modeling capacity. The encoder path incorporates five stacked ASCRM blocks as the core feature extraction units. Each ASCRM integrates spatial and channel attention to dynamically recalibrate feature responses, while employing multi-scale residual structures to facilitate cross-layer information exchange. This design enables the effective encoding of fine-grained semantic features critical to nodule detection. A detailed description of the ASCRM's architecture and fusion strategy is provided in the next section.
In parallel, the decoder path employs multi-level skip connections to fuse high-resolution structural cues (e.g. edges and textures) from shallow layers with upsampled deep features. This strategy mitigates spatial information loss during feature reconstruction, enhancing the delineation and fidelity of candidate nodule regions.
To further improve the network's spatial awareness, we introduce voxel-wise spatial coordinate maps during the downsampling stage. These coordinate tensors are concatenated as additional input channels, providing spatial priors that guide the model in learning anatomical layouts of the lung fields. This approach suppresses false positives outside lung regions and improves detection accuracy and generalization—especially in cases involving blurred boundaries or structurally complex nodules.
Finally, the network outputs a volumetric probability map of size 32 × 32 × 32, where each voxel denotes the likelihood of being the center of a pulmonary nodule. This map is subsequently used for candidate proposal generation and passed on to the false positive reduction stage.
Attention-guided spatial and channel residual module
To effectively enhance the network's capability of modeling fine-grained features during candidate nodule detection, we introduce the ASCRM as the core feature extraction unit. The architectural details of ASCRM are illustrated in Figure 3. Building upon conventional residual structures, ASCRM integrates dense skip connections and the Convolutional Block Attention Module (CBAM) 20 to strengthen semantic representation across multiple scales and improve the network's focus on critical spatial regions.

The proposed ASCRM module.
Specifically, the ASCRM module constructs a deep residual path by stacking multiple 3 × 3 × 3 convolution layers, while introducing long-range connections to progressively propagate high-resolution structural features from early shallow layers to deeper stages. This design effectively preserves fine-grained textures and boundary details of nodules. Within the residual path, each layer follows the formulation:
Let 
To further facilitate information exchange across different feature hierarchies, a cross-layer residual fusion mechanism is introduced into the ASCRM module. This mechanism integrates feature representations 
This design strengthens the expressive capacity of the fused features and ensures stable propagation of cross-layer information. By mitigating representational degradation that can occur during multi-scale feature integration, the residual fusion structure enhances the joint modeling of fine-grained structural cues and high-level semantics, thereby providing a more discriminative feature foundation for the subsequent attention modules.
To improve feature discriminability, the ASCRM module integrates the CBAM, constructing a dual attention pathway across spatial and channel dimensions. The channel attention branch utilizes both average and max pooling operations to capture global response features, generating a channel-wise attention map. The spatial attention branch aggregates pooled descriptors to produce a spatial response map that enhances critical regions. These attention mechanisms are defined as:


Where
Multi-scale progressive inference architecture
In the task of false positive reduction, the heterogeneous nature of pulmonary nodules—manifested in their diverse morphologies, variable densities, and particularly wide size range (Figure 4)—poses a significant challenge. Conventional fixed-scale detection strategies often struggle to concurrently capture the global structural patterns of large nodules and the subtle textural cues of smaller ones, leading to frequent misses of tiny nodules or misclassification of non-nodular tissues. To address these limitations, we propose a Multi-scale Progressive Perception Network (MPPN) that integrates a hierarchical context fusion scheme with attention-enhanced residual modules to enable precise classification of candidate nodules across scales (Figure 5).

Distribution of lung nodule sizes in the LUNA16 dataset.

The proposed false positive reduction model.
As illustrated in Figure 5, MPPN stratifies candidates into three-scale categories based on nodule diameter, and accordingly extracts 3D voxel patches of size 32 × 48 × 48, 16 × 24 × 24, and 8 × 12 × 12, forming an adaptive multi-scale input representation. This design avoids spatial distortions and semantic degradation typically introduced by uniform resampling, and improves the network's ability to model the geometry and context of nodules at different scales. Each scale-specific input is processed by an independent 3D convolutional branch, constructing a coarse-to-fine multi-scale feature extraction pathway. To alleviate semantic isolation between scales, a top-down feature injection mechanism is introduced in the medium and small-scale branches, allowing high-level semantics from larger scales to guide the detection of small nodules. Additionally, each branch incorporates a CBAM to enhance the model's focus on discriminative regions. The high-level semantic features from all branches are concatenated along the channel dimension, followed by global average pooling to obtain a compact global representation. This representation is then recalibrated via attention and passed through a fully connected layer to output the probability of each candidate being a true nodule.
Experiment
Dataset and preprocessing
This study employs the LUNA16 21 benchmark dataset for the lung nodule detection task. The dataset is derived from the LIDC-IDRI database, a multi-center collaborative project initiated by the National Cancer Institute (NCI) and supported by the Foundation for the National Institutes of Health (FNIH), which includes chest CT scans annotated by multiple radiologists. 22
The LUNA16 dataset was rigorously selected from the LIDC-IDRI database, with inclusion criteria that primarily exclude cases with missing slices, inconsistent pixel spacing, or slice thicknesses greater than 3 mm. The final dataset consists of 888 high-quality CT scans, all provided in MHD/RAW format. In terms of nodule annotation, the LUNA16 dataset follows a strict expert consensus standard, including only nodules with a diameter of ≥3 mm, which were confirmed by at least three of the four radiologists. Based on this criterion, the dataset contains 1186 lung nodule samples, each accompanied by detailed center coordinates and scaling information. Additionally, segmentation images of the lung regions for all CT scans are included.
To enhance the model's generalizability and robustness, a binary segmentation mask was first applied to precisely extract the lung parenchyma region from each input image (Figure 6). Each CT slice underwent preliminary denoising via a Gaussian filter to mitigate errors caused by blurred tissue boundaries. Subsequently, a fixed intensity threshold of −400 HU was employed to binarize the image, enabling initial separation of lung tissue. Considering potential discontinuities and small holes at the lung parenchyma edges, morphological closing operations were further applied to repair the binary images and fill residual small cavities. After obtaining closed lung parenchyma candidate regions, connected component analysis was used to retain the two largest connected areas, thereby excluding non-pulmonary structures such as the trachea or external noise. Remaining internal holes were filled using a morphological binary fill operation, yielding complete masks for both left and right lungs.
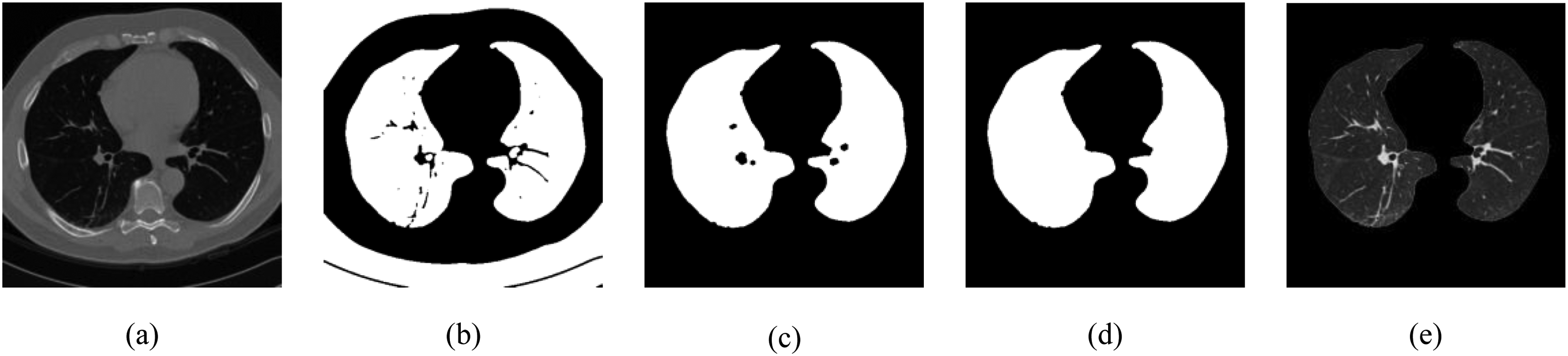
(a) Original image; (b) binary lung region mask obtained after thresholding; (c) initial extracted lung parenchyma mask; (d) lung parenchyma mask after morphological closing operation; (e) final segmented lung parenchyma.
To enhance contrast of pulmonary structures, CT values were linearly mapped within the lung window range of −1200 to 600 HU. Moreover, to address variations in spatial resolution arising from different manufacturers and scanner models, B-spline interpolation resampling was performed to achieve isotropic voxel spacing of 1 mm3, ensuring spatial consistency and comparability across all images.
Training process
The training process is divided into two stages: candidate nodule detection and false positive reduction. During the candidate detection stage, considering the large volume of the entire CT scans, directly inputting the full volume into the network would cause GPU memory bottlenecks. Therefore, the images are partitioned into three-dimensional voxel blocks of size 128 × 128 × 128 to alleviate computational resource constraints. To enhance model robustness and generalization, data augmentation techniques such as random flipping and scaling transformations are applied during training. For generating training anchors, boxes with an Intersection over Union (IoU) greater than 0.5 are labeled as positive samples, while those with an IoU less than 0.02 are labeled as negative samples. Three anchor sizes of 5, 10, and 20 voxels are employed. The proposed loss function comprises two components: classification loss and regression loss, optimized jointly via a dynamic weighting mechanism. For classification, an improved binary focal loss (Binary Focal Loss) is adopted, whose core formulation is defined as follows:
Here,
For the regression task, the Smooth L1 Loss is employed, defined as: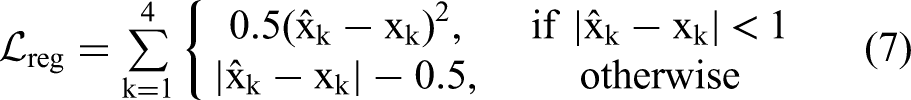
Here,
The total loss function balances the two objectives—classification and regression—via an adaptive weighting mechanism, and is expressed as: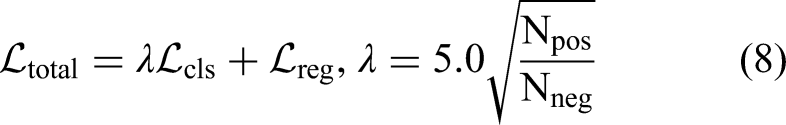
In the false positive reduction stage, the model is trained using image patches of three different sizes: 32 × 48 × 48, 16 × 24 × 24, and 8 × 12 × 12 voxels. To address the imbalance between true positive and false positive nodules, data augmentation techniques such as random flipping, translation, and rotation are applied. Additionally, the ratio of true positive to false positive samples is expanded to 1:3 to ensure a balanced training distribution. The model is optimized using the cross-entropy loss function, with an initial learning rate of 0.0001 and the Adam optimizer. All experiments are implemented using the PyTorch framework and trained on an Ubuntu server equipped with an NVIDIA GeForce RTX 3090 GPU.
Experimental results
To systematically evaluate the stability and generalizability of the proposed model under varying data splits, a six-fold cross-validation strategy was employed. The performance metrics across the six subsets are summarized in Table 2. On average, the model achieved a sensitivity of 94.9%, a Competition Performance Metric (CPM) score of 87.8, and generated only 4.2 candidate detections per scan. Notably, when the sensitivity was maintained at 90.0%, the average false positive rate was controlled within 0.55 per scan.
Results of six-fold cross-validation on the LUNA16 dataset.

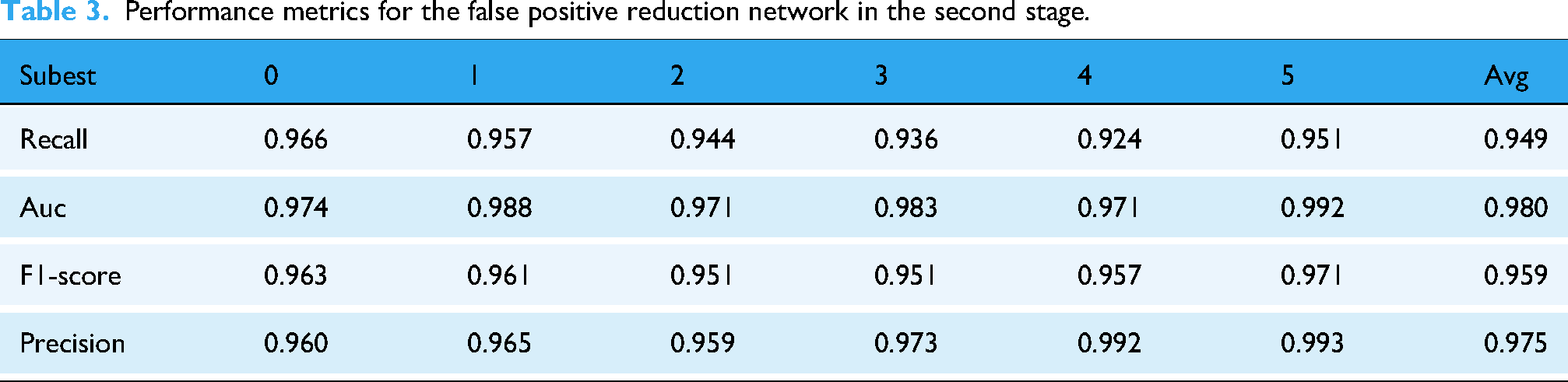
Building on the analysis of the system's overall performance, we further evaluated the independent classification capability of the second-stage false positive reduction network. As shown in Table 3, this network achieved an average AUC of 0.980, Precision of 0.975, and F1-score of 0.959 on the test set, while maintaining a recall of 0.949. These results indicate that the false positive reduction network demonstrates robust discriminative power in distinguishing true nodules from imaging artifacts, and its stable classification performance provides a reliable foundation for the system's low false positive rate.
Performance metrics for the false positive reduction network in the second stage.
To evaluate the generalizability of the proposed framework across different data sources, we conducted additional validation experiments on the LIDC-IDRI dataset. The six models obtained from the six-fold cross-validation on LUNA16 were applied to LIDC-IDRI for inference. Since LUNA16 is constructed from a subset of LIDC-IDRI, overlapping cases may introduce bias. Before running inference on LIDC-IDRI, we removed the CT scans that had been used to train each model, ensuring case-level independence between the training data and the LIDC-IDRI evaluation set. Performance metrics were computed for each of the six models and then averaged to obtain the final results. The LIDC-IDRI results of the first-stage candidate detection network and the second-stage false positive reduction network are summarized in Tables 4 and 5, respectively.
Validation results of the first-stage candidate detection network on the LIDC-IDRI dataset.

Validation results of the second-stage false positive reduction network on the LIDC-IDRI dataset.

As shown in Table 4, the first-stage candidate detector maintains a high sensitivity of 92.8% on LIDC-IDRI and generates 4.7 candidates per scan on average. When the operating point is adjusted to achieve 90% sensitivity, the false positives are controlled at 0.63 FPs/scan, indicating stable candidate generation under different operating thresholds. Table 5 shows that the second-stage false positive reduction network preserves strong discriminative capability on LIDC-IDRI, achieving an AUC of 0.968 with a balanced precision–recall profile (Precision = 0.942, Recall = 0.932). These results support the effectiveness of the overall pipeline in suppressing false positives on LIDC-IDRI.
Figure 7 provides a visual comparison of intermediate detection results at different stages of the pipeline. Column (a) shows the original chest CT images, column (b) presents the candidate nodule detection outputs, column (c) displays the refined results after false positive suppression, and column (d) shows the corresponding clinical ground-truth annotations. The proposed two-stage detection framework adopts a cascaded architecture: the first-stage candidate detection network emphasizes sensitivity through rich feature extraction (Figure 7(b)), but inevitably introduces a considerable number of false positives; the second-stage false positive reduction model leverages deep feature discrimination to suppress misclassified nodules effectively (Figure 7(c)). Experimental results demonstrate that this cascaded “detection-then-refinement” paradigm significantly enhances specificity while maintaining high sensitivity, thereby improving the overall robustness of pulmonary nodule detection.

Representative intermediate results of the proposed framework. Column (a) shows the original chest CT images; column (b) displays the detected candidate nodules from the first-stage detection model; column (c) presents the refined outputs after false positive reduction; and column (d) illustrates the corresponding ground-truth annotations.
Furthermore, to gain deeper insights into the decision-making mechanism of the first-stage candidate nodule detection model, Gradient-weighted Class Activation Mapping (Grad-CAM) was employed to visualize the spatial regions that contribute most to the model's predictions (Figure 8).The visualization results show that the model produces strong and spatially concentrated activations in regions corresponding to pulmonary nodules. These activations are consistently localized around the nodule areas across different samples, indicating that the model relies primarily on nodule-relevant spatial features when generating detection responses. The observed activation patterns suggest that the learned feature representations are closely associated with the morphological and intensity characteristics of nodules in CT images. These CAM visualizations provide an interpretable view of the detection process, offering qualitative evidence of how the model focuses on nodule-related regions during candidate generation.

Grad-CAM visualization of first-stage candidate nodule detection. Column (a) shows the original chest CT images; column (b) displays the Grad-CAM heatmaps highlighting the regions of interest identified by the model; column (c) presents the overlay of the Grad-CAM heatmaps on the original images, showing the model's attention in context.
Comparison with other methods
In this study, we propose a two-stage pulmonary nodule detection framework that takes three-dimensional chest CT scans as input. The first stage employs a U-shaped network to generate a nodule probability map and identify initial candidate regions. Subsequently, a MPPN, specifically designed for this task, is introduced to further discriminate these candidates, effectively suppressing false positives and improving detection accuracy. By integrating coarse candidate generation and refined classification, the two-stage architecture achieves a notable enhancement in overall detection performance.
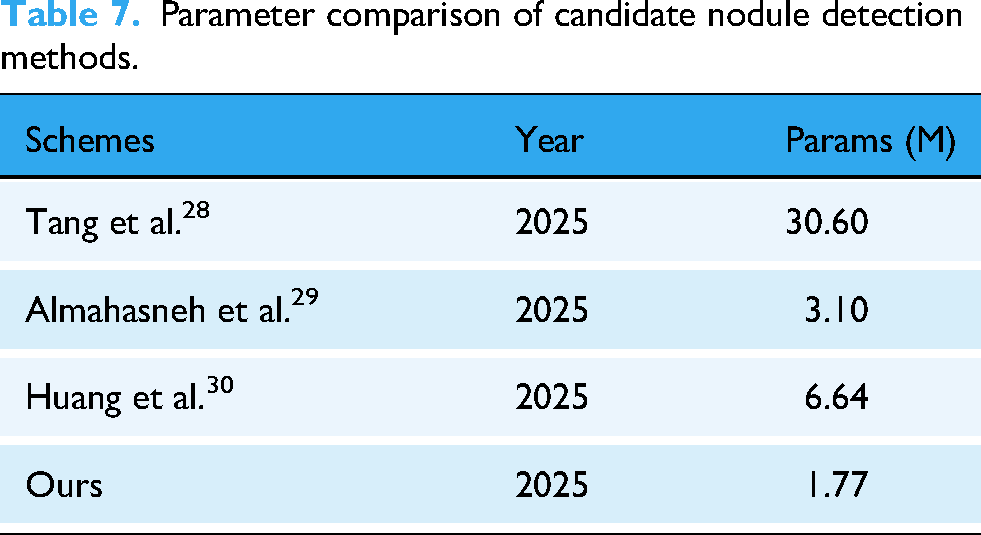
We conducted a comparative performance analysis of both the single-stage candidate detection model and the complete two-stage detection framework against several state-of-the-art methods. In the candidate detection stage, we benchmarked the sensitivity and the average number of candidates per scan (FPs/scan) of our method against those reported by Pereira et al. (2021), 23 Yuan et al. (2021), 24 Zhao et al. (2023), 25 Usman et al. (2024), 26 and Xiong et al. (2024). 27 As shown in Table 6, our approach achieved a sensitivity of 98.5%, while maintaining the lowest average number of false positives per scan, demonstrating its ability to retain high sensitivity with superior false positive control. Beyond detection performance, model size is an important consideration for computational cost and practical deployment. Accordingly, Table 7 compares the parameter count of our candidate detection network with recent representative methods. Our first-stage detector contains 1.77 M parameters, which is smaller than the compared networks, indicating a relatively compact model design.
Performance comparison of candidate nodule detection methods.

Parameter comparison of candidate nodule detection methods.
We further compare our method with several representative approaches in the false positive reduction phase, as summarized in Table 8. Our method demonstrates stable performance across operating points and achieves a CPM of 0.878, which is the highest among the compared methods. At operating points of 0.5 and 1 FPs/scan, our method attains sensitivities of 0.891 and 0.939, respectively, indicating strong detection sensitivity under moderate false-positive constraints. Overall, the results in Table 8 suggest that the proposed framework provides a balanced trade-off across operating points and yields favorable CPM performance.
Comparative performance evaluation in the false positive reduction phase.

Bold text indicates the false positive rate per scanning round.
Figure 9 illustrates the FROC (Free-response Receiver Operating Characteristic) curve of our method on the LUNA16 dataset, which is used to assess detection performance under different false positive rates. As shown in the figure, even under a strict constraint of no more than 0.55 FPs/scan, our method still achieves a detection sensitivity of 0.9, clearly demonstrating its strong nodule recognition capability while maintaining a low false positive rate.
The superior performance of our approach is mainly attributed to the integration of multi-scale feature fusion and attention mechanisms, which enable the model to more accurately distinguish between nodules and non-nodules in the complex anatomical background of the lung. The combined results in Figure 9 and Table 8 indicate that our method achieves a favorable balance between high sensitivity and low false positives, showing solid potential for practical application—particularly in clinical lung nodule screening scenarios where strict control over the false positive rate is required.
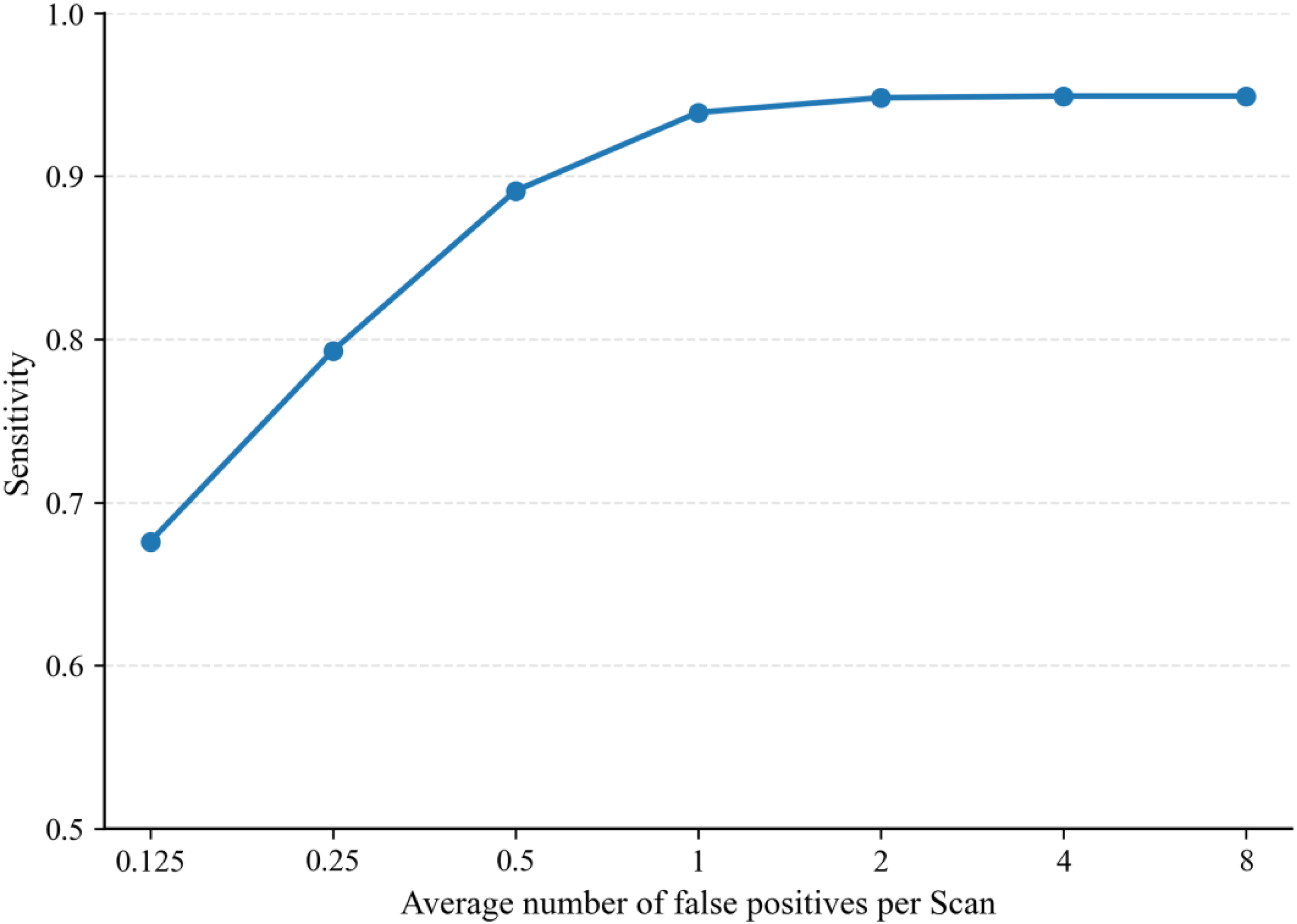
Free-response receiver operating characteristic (FROC) curve.
Discussions
The results of the comparative experiments demonstrate that the proposed two-stage detection framework performs competitively on the LUNA16 dataset, indicating that the framework design contributes to addressing the challenge of high false positive rates in pulmonary nodule detection. Traditional detection methods often face limitations when dealing with the wide variations in nodule size, shape, and anatomical location, particularly in cases where nodule morphology and anatomical background are complex. While methods based on 2D CNNs can effectively capture nodule features, they typically struggle to fully leverage the rich spatial context embedded in 3D CT data.
The proposed cross-layer attention fusion two-stage framework combines attention mechanisms with multi-scale fusion, effectively reducing false positive rates while maintaining high sensitivity. The ASCRM module enhances feature representation capabilities, improving the model's accuracy in nodule detection, while the MPPN network effectively suppresses false positives and enhances the fine discrimination of candidate nodules.
Ablation experiment
To validate the effectiveness of the ASCRM module and the false positive reduction network (MPPN), we conducted four ablation experiments: (i) baseline model only, (ii) baseline with CBAM-based attention mechanism, (iii) baseline with ASCRM module, and (iv) ASCRM module combined with the MPPN model. We validated the models on six subsets, and Table 9 provides the mean and standard deviation of the results from these six trials. The baseline model, without any enhancements, achieved a sensitivity of 91.63%. Incorporating the CBAM attention mechanism significantly improved sensitivity to 97.19%; however, it also dramatically increased the average number of detected candidates per scan from 3.81 to 14.26, and the false positive rate at 90.0% sensitivity rose from 1.20 to 2.92 per scan. This increase was primarily due to the detection of more true nodules at the cost of introducing excessive false positives under the same confidence threshold.
Results of four ablation experiments.

By integrating the ASCRM module, the model maintained high sensitivity while reducing the number of false positives compared to using CBAM alone. Furthermore, when the MPPN was added to the ASCRM-enhanced model, the false positive rate decreased to 4.18 per scan, with only a marginal drop in sensitivity. Overall, these results demonstrate that the ASCRM module and MPPN strategy effectively suppress false positives while preserving high detection sensitivity.
Additionally, we performed paired t-tests to calculate the significance of the differences between each model and the final two-stage model (Baseline + ASCRM + MPPN) for the FPs/Scan @ 90% Sensitivity metric. The results indicate that the p-values for all ablation models compared to the final two-stage model are less than 0.05, demonstrating statistically significant differences.
Conclusion
This paper proposes a two-stage pulmonary nodule detection method based on cross-layer attention fusion. In the candidate nodule detection stage, a multi-scale feature extraction and cross-layer attention fusion mechanism is introduced. The attention-enhanced module (ASCRM) effectively improves the model's sensitivity to tiny nodules and enhances the recall rate of candidate regions. In the false positive reduction stage, a MPPN is constructed to integrate semantic information across different spatial scales, thereby improving the model's ability to distinguish true nodules from artifacts and significantly reducing the false positive rate.
We conducted comprehensive experimental evaluations on the LUNA16 dataset, and the results demonstrate that the proposed method achieves high recall while significantly improving detection accuracy. Compared with several state-of-the-art detection models, our method shows superior performance, validating its effectiveness and robustness. Future work will focus on enhancing the clinical interpretability and lightweight deployment of the model to promote its application in real-world CAD of lung cancer.
Nevertheless, despite the proposed method achieving certain results in experimental performance, several limitations deserve further attention. The training and validation of the model rely primarily on the LUNA16 dataset, whose homogeneous imaging conditions and single-source origin may restrict the model's generalizability in real-world clinical environments. The lack of independent validation on real clinical data remains one of the major limitations of the current work. Moreover, the detection of nodules smaller than 3 mm or those closely attached to the pleura or vasculature remains challenging, as such nodules typically exhibit low contrast and irregular morphology, placing higher demands on feature representation. In terms of computational efficiency, although the two-stage architecture performs well overall, the first stage still requires dense feature extraction across the full 3D CT volume, resulting in considerable computational overhead that may hinder deployment on resource-constrained systems. In addition, although Grad-CAM visualizations provide some insight into the model's behavior, the decision-making process retains a degree of “black-box” characteristics, indicating that further efforts are needed to enhance interpretability.
Future work will focus on improving the model's generalizability to real clinical data, optimizing computational efficiency, strengthening interpretability, and validating the approach on larger and multi-center datasets.
Footnotes
Acknowledgements
The authors would like to thank the project team for providing access to computing resources.
Author contributions
Lixin Wang contributed to methodology, investigation, formal analysis, and writing the original draft. Xiaowen Lan was responsible for conceptualization and writing—review & editing. Kaikai Zhang contributed to conceptualization and visualization, while Yanhui Wang was involved in formal analysis and visualization. Shaofeng Wang and Wenjing Liu managed project administration and participated in writing—review & editing. All authors have read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Inner Mongolia Autonomous Region (grant number 2025LHMS06016).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
