Abstract
Background
Thalassemia is a common hereditary anemia that poses diagnostic challenges during pregnancy due to physiological changes in hematological parameters. Misdiagnosis or delayed diagnosis can result in serious maternal and fetal complications. Although genetic testing is the gold standard for thalassemia diagnosis, its high cost and limited availability restrict its use in primary care settings. This study aimed to develop and validate a clinically interpretable machine learning model to identify thalassemia in pregnant women using routine complete blood count indicators.
Methods
A total of 523 pregnant patients were retrospectively enrolled in this study. Eight predictive features were selected using the Boruta algorithm. Multiple machine learning algorithms were trained and evaluated using ten-fold cross-validation. An ensemble model was constructed through bagging and weighted integration of top-performing models. Model performance was assessed using a series of metrics such as AUROC, AUPRC, accuracy, F1-score, and MCC. Comparative evaluation was conducted against traditional diagnostic indices and a recently proposed model. Model interpretability was analyzed using permutation importance and SHAP methods.
Results
The Thal-classifier, constructed by integrating four bagged machine learning models, was developed for accurate identification of thalassemia. It exhibited excellent diagnostic performance, with an AUROC of 0.945 on the cross-validation and 0.899 on the independent testing dataset. Compared with the existing indices and models, it achieved superior performance across multiple evaluation metrics. Feature importance analyses consistently identified RDW-SD and MCHC as the primary predictive features.
Conclusion
Thal-classifier is a practical and reliable tool for early detection of thalassemia in pregnant populations, offering valuable support for prenatal screening and individualized clinical decision-making.
Introduction
Thalassemia is an autosomal recessive genetic disorder resulting from mutations or deletions in the alpha- or beta-globin genes, characterized by impaired hemoglobin synthesis and ineffective erythropoiesis. 1 In China, the disease exhibits a distinct regional distribution pattern, with carrier rates ranging from 6% to 15% in high-prevalence southern provinces such as Guangxi, Guangdong, and Hainan. 2 In contrast, the carrier rate in the Yangtze River Basin and northern regions is approximately 1% to 3%.3,4 However, due to increased population mobility and inter-regional marriage, thalassemia cases are increasingly being reported in non-high-incidence areas, warranting attention.
Anemia is a common complication during pregnancy, with multiple potential etiologies including iron deficiency and folate deficiency. Thalassemia, as a congenital microcytic anemia, constitutes a significant proportion of anemia cases encountered during pregnancy. 5 Given the substantial differences in pathogenesis, treatment approaches, and prognosis between thalassemia and other microcytic hypochromic anemias such as iron deficiency anemia, it is crucial to distinguish between these conditions. For instance, thalassemia may necessitate regular blood transfusions and concomitant iron chelation therapy to manage both anemia and secondary iron overload, 6 whereas iron deficiency anemia during pregnancy can typically be effectively managed through iron supplementation and dietary interventions, 7 with a more favorable prognosis. Failure to accurately diagnose and appropriately manage thalassemia during pregnancy may result in severe maternal and fetal complications, including intrauterine growth restriction, preterm birth, and elevated maternal mortality risk.8,9 Therefore, early identification and precise diagnosis of thalassemia are essential for developing individualized treatment strategies that improve maternal and fetal health outcomes and reduce adverse events. However, conventional diagnostic methods for thalassemia primarily rely on genetic testing and sequencing technologies, which, although highly accurate, face limitations in widespread application within primary healthcare settings and low-prevalence regions due to high costs, the need for specialized personnel, and prolonged turnaround times. High-performance liquid chromatography (HPLC) provides an alternative method that is both efficient and cost-effective for screening. 10 However, the overlap between certain hemoglobins and HbA2, instrument variability, and post-translational modifications can affect the quantification and require expert interpretation.11,12
In the field of medical laboratory diagnostics, the application of machine learning (ML) techniques has been continuously expanding and deepening. With their robust capabilities in data processing and pattern recognition, these technologies offer novel approaches for achieving more precise disease diagnosis. 13 Recently, numerous studies have explored the use of red blood cell parameters from routine blood tests to differentiate thalassemia from other types of anemia, yielding promising results. 14 These studies have developed corresponding diagnostic models by analyzing variations in red blood cell morphology, count, and hemoglobin content, thereby providing valuable references for the classification and diagnosis of anemia. Nevertheless, most existing research focuses on the general population, with limited development of specific diagnostic models tailored for pregnant women. The physiological and hematological characteristics of pregnant women differ significantly from those in the non-pregnant state, potentially affecting the accuracy of anemia diagnosis. Hence, establishing a dedicated diagnostic model for thalassemia in pregnant women holds significant clinical relevance. This study aims to develop a thalassemia diagnostic model specifically for pregnant women using red blood cell parameters obtained from routine blood tests.
Methods
Study population
This study constitutes a retrospective investigation conducted in the Wenzhou region of Zhejiang Province, aimed at developing and validating a predictive model for thalassemia in pregnant women. A total of 523 pregnant patients who underwent initial evaluation at the Department of Laboratory Medicine, Wenzhou People's Hospital (Wenzhou Women's and Children's Hospital), between June 2016 and May 2025, were enrolled. The study population was strictly selected based on complete demographic and clinical information, including age, gestational status, complete blood count (CBC) parameters, and results from genetic testing for thalassemia. Exclusion criteria were systematically applied to eliminate male patients, non-pregnant females, underage females, cases with incomplete data, individuals with concurrent hematological disorders or acute/chronic conditions affecting hematopoietic function, as well as amniotic fluid, cord blood, chorionic villus samples, or duplicate test results from the same case. In summary, this research utilizes comprehensive hematological parameters and thalassemia screening results exclusively from adult female patients during pregnancy, ensuring data integrity and clinical relevance.
The entire dataset was randomly divided into a training dataset and a testing dataset in a 3:1 ratio using stratified sampling based on thalassemia status. The training dataset (n = 393; 180 positives, 213 negatives) was used to construct and optimize the predictive model, while the testing dataset (n = 130; 58 positives, 72 negatives) served to independently evaluate model performance.
Laboratory indicator measurements
Peripheral blood samples were collected from pregnant patients in the morning while fasting, using EDTA anticoagulant tubes. CBC tests were conducted within 2 h at room temperature using the Beckman Coulter Unicel DxH800 counter (Beckman Coulter, Inc., Brea, CA, USA). Thalassemia gene testing samples were stored at 4 °C and tested within one week. The thalassemia genotyping test was performed using the α- and β-thalassemia gene detection kit from Guangzhou Kapu Medical Technology Co., Ltd. This kit, employing polymerase chain reaction and reverse dot blot hybridization technology, can detect three common deletion types and three common mutation types of α-thalassemia, as well as 19 common mutation types of β-thalassemia prevalent in the Chinese population. All laboratory tests were conducted by trained technicians strictly in accordance with the operation manual to ensure accuracy and reliability.
Statistical analysis
All statistical analyses were conducted using SPSS version 25.0. Continuous variables were presented as median (interquartile range). The Shapiro-Wilk test was applied to assess the normality of continuous variables. Normally distributed variables are compared using independent sample t-tests. Non-normally distributed data are analyzed using the Mann–Whitney U test. Categorical variables are expressed as frequencies (percentage), with comparisons made using the Chi-square test or Fisher's exact test where appropriate. A two-sided P-value of less than 0.05 was considered statistically significant.
Feature extraction and selection
Clinical and hematological parameters extracted from the eligible patients included age, red blood cell count (RBC), hemoglobin assay (HGB), hematocrit (HCT), mean corpuscular volume (MCV), mean corpuscular hemoglobin (MCH), mean corpuscular hemoglobin concentration (MCHC), red cells distribution width-standard deviation (RDW-SD), and red cells distribution width-coefficient of variation (RDW-CV). To identify the most informative features for predicting thalassemia, the Boruta algorithm was employed. As a wrapper based on Random Forest, Boruta 15 iteratively eliminates irrelevant features by comparing their importance against randomized shadow features. Selected features were then visualized using t-distributed stochastic neighbor embedding (t-SNE), 16 a nonlinear dimensionality reduction technique that maps high-dimensional data into a two-dimensional space. This visualization was performed using the R package Rtsne with parameters set to dimensions = 2 and perplexity = 100. Unsupervised hierarchical clustering based on the selected features was conducted using the R package pheatmap to assess the natural grouping of cases. Chi-square analysis was applied to examine the significance of differences in thalassemia status between the clustered groups.
Model construction
To construct predictive models, the selected features were input into eight widely used ML algorithms: CatBoost, LightGBM, Random Forest, K-Nearest Neighbors, Extreme Gradient Boosting, Extra Trees, and two neural network frameworks implemented using FastAI and PyTorch, respectively. Each algorithm was trained using 10-fold cross-validation, producing ten base classifiers per model. These were aggregated using bootstrap aggregation (bagging) to form a composite prediction. A greedy weighted ensemble algorithm 17 was subsequently applied to integrate the top-performing bagged models, optimizing predictive accuracy through weighted averaging. To prevent data leakage, all model evaluations were based on out-of-fold predictions. Classifier performance metrics were derived from these predictions, and the best-performing model was selected accordingly. All modeling processes were implemented using the AutoGluon framework. 18
Performance evaluation and benchmarking
To benchmark model performance, the proposed classifier was compared with several established hematological indices for thalassemia screening, including formulas developed by England and Fraser (E&F), 19 Mentzer, 20 Srivastava, 21 Shine and Lal (S&L), 22 Bessman, 23 Green and King (G&K), 24 Jayabose, 25 Sirdah, 26 and Ehsani. 27 Additionally, a recently published CatBoost model by Lai et al. 28 was also included in the performance comparison. Detailed definitions of all indices and formulas are provided in Table S1.
Model performance was assessed on the test dataset using a range of metrics, including accuracy, F1-score, Matthews correlation coefficient (MCC), precision, recall, area under the receiver operating characteristic curve (AUROC), and area under the precision–recall curve (AUPRC). Cut-off values were determined using the Youden index derived from the ROC analysis of the training dataset and then applied consistently during classification. Confusion matrices were generated to evaluate classification performance, including positive predictive value and negative predictive value. Calibration curves assessed agreement between predicted and observed probabilities, while decision curve analysis quantified clinical net benefit across a range of threshold probabilities.
Model interpretation
To improve clinical interpretability, two complementary model explanation methods were employed. First, permutation importance analysis was used to quantify each feature's impact on model performance by measuring the decrease in accuracy following random permutation. Features with the highest importance scores were identified as key contributors to predictive accuracy. Second, SHapley Additive exPlanations (SHAP) analysis 29 was conducted to estimate both the magnitude and directionality of feature contributions at the individual prediction level. The consistency between SHAP and permutation analysis provided robust interpretive insights into the model's decision-making process.
Results
Patient characteristics
A total of 1501 anemic patients who underwent thalassemia genetic testing between June 2016 and May 2025 were initially screened. To ensure cohort homogeneity, only pregnant women were included in the final analysis. Exclusion criteria led to the removal of 812 male patients, underage females, and adult non-pregnant females. An additional 166 cases were excluded due to incomplete clinical information or a history of blood transfusion prior to sample collection. The final dataset comprised 523 pregnant women, including 238 patients diagnosed with thalassemia based on genetic testing and 285 non-thalassemia patients. The dataset was randomly divided into a training set and a test set in a 3:1 ratio. The overall study workflow is illustrated in Figure 1. Table 1 summarizes the distribution of hematological parameters and thalassemia types across both datasets. Statistical comparisons indicated no significant differences between the training and test sets (all P-values > 0.05), confirming that the stratified sampling preserved data balance and representativeness.
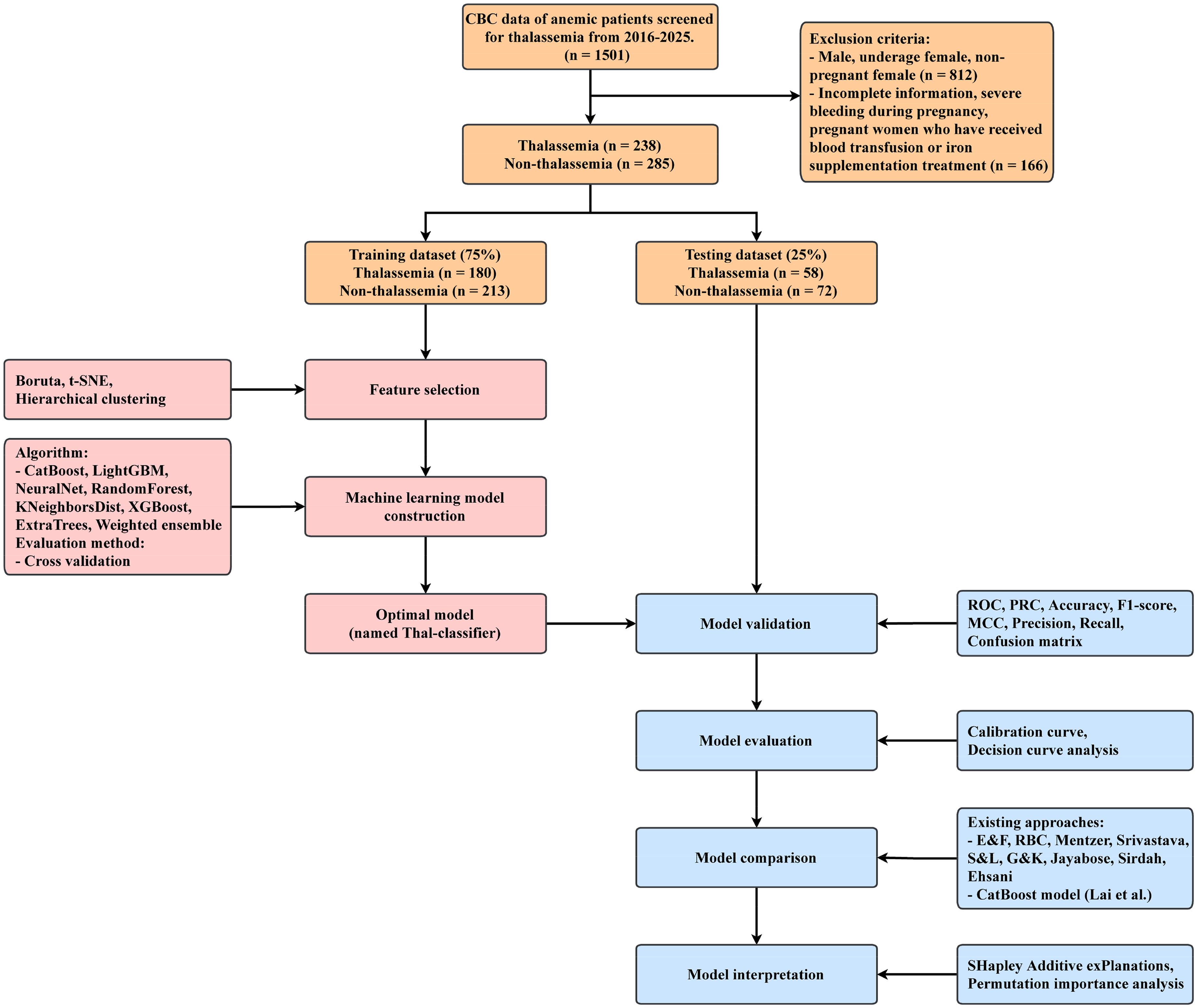
Schematic overview of the study workflow.
Baseline demographic characteristics of the training and testing datasets.

Identification of predictive features
To optimize model performance, feature selection was conducted using the Boruta algorithm. Nine candidate variables were initially considered, including patient age and eight red blood cell parameters. Boruta identified eight informative features, ranked by importance as follows: RDW-SD, RBC, MCV, MCHC, MCH, RDW-CV, HGB, and HCT (Figure 2(A)). Age was excluded due to its negligible contribution to classification accuracy.

Construction of ML models for thalassemia prediction using the training dataset. (A) Ridge plot illustrating feature importance scores generated by the Boruta algorithm. (B) Two-dimensional t-SNE visualization of sample distribution based on the eight selected features, with red and blue dots representing thalassemia and non-thalassemia cases, respectively. (C) Unsupervised hierarchical clustering heatmap based on the same feature set, revealing diagnostic groupings within the training dataset. (D) ROC curves and (E) PR curves for nine ML algorithms evaluated via 10-fold CV. (F) Comparison of classification metrics across all models. A star indicates the algorithm with the best performance.
The predictive value of these features was further evaluated using t-SNE. Dimensionality reduction yielded a clear separation between thalassemia and non-thalassemia cases in two-dimensional space, suggesting strong discriminative capacity (Figure 2(B)). In parallel, unsupervised hierarchical clustering revealed that patients with the same diagnosis were predominantly grouped together, a pattern supported by Chi-square testing (P < 0.001; Figure 2(C)). These findings demonstrate that the selected features can serve as a robust signature for thalassemia prediction.
Construction of the thalassemia classifier
Eight ML algorithms were implemented to construct predictive models, including KNN, RF, ET, XGB, LGBM, CB, NN-Torch, and NN-FastAI. Each algorithm was trained using 10-fold cross-validation to generate ten base classifiers, which were subsequently aggregated via bagging. A weighted ensemble model was then developed by integrating the bagged classifiers from LGBM, CB, and both neural networks. Cross-validation results showed that all models achieved strong classification performance, with AUROC values exceeding 0.88 (Figure 2(D)) and AUPRC values above 0.86 (Figure 2(E)). The ensemble model exhibited the highest overall performance, with an AUROC of 0.945, AUPRC of 0.940, accuracy of 0.875, F1-score of 0.866, and MCC of 0.750 (Figure 2(F)). These results indicated the ensemble model was the most suitable and was thus designated as the optimal classifier.
Validation and performance evaluation
The ensemble model was evaluated on the entire training dataset and the held-out test dataset. In the training set, the model achieved an AUROC of 0.949, AUPRC of 0.945, and classification accuracy of 0.875 (Figure 3(A) and (C)). When applied to the independent test set, the model maintained robust diagnostic performance with an AUROC of 0.899 and an AUPRC of 0.909 (Figure 3(B)). Using the optimal threshold value of 0.439, determined by the Youden index from the training set, the model achieved an accuracy of 0.869, F1-score of 0.850, MCC of 0.735, precision of 0.873, and recall of 0.828 on the test set (Figure 3(B) and (C)). Derived from the confusion matrix of the 130-patient test set, the model classified 55 subjects as positive and 75 as negative, identifying 48 true positives and 65 true negatives with 10 false negatives and 7 false positives, corresponding to a positive predictive value of 87.3% and a negative predictive value of 86.7%. Calibration analysis showed excellent concordance between predicted probabilities and observed outcomes (Figure 3(D)). Decision curve analysis demonstrated clear net clinical benefit across a wide range of threshold probabilities (45–95%) compared to the “treat none” strategy (Figure 3(E)). Based on these results, the ensemble model was formally named the Thal-classifier for subsequent analysis.
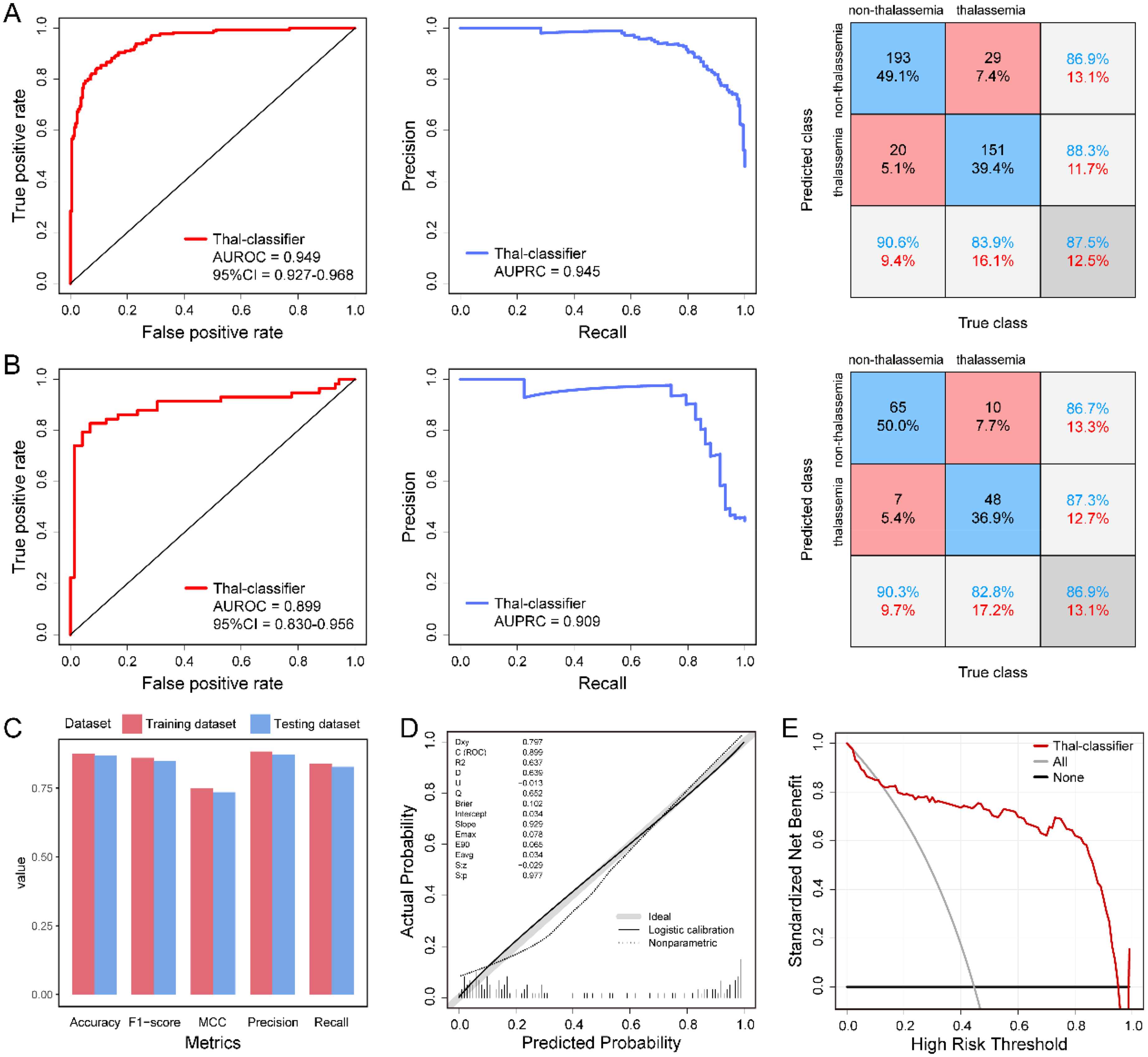
Validation and performance evaluation of the Thal-classifier. (A) ROC curves (left), PR curves (middle), and confusion matrices (right) for the Thal-classifier on the training dataset. (B) Corresponding ROC curves, PR curves, and confusion matrices on the independent testing dataset. (C) Summary of classification metrics for both training and testing sets. (D) Calibration curve of the Thal-classifier on the testing dataset. (E) Decision curve analysis showing the net clinical benefit of the Thal-classifier across a range of threshold probabilities.
Benchmarking against existing diagnostic indices
To evaluate the Thal-classifier's practical utility, we conducted a comprehensive comparison against ten existing thalassemia discrimination formulas, including the E&F index, RBC, Mentzer index, Srivastava index, S&L index, Bessman index, G&K index, Jayabose index, Sirdah index, and Ehsani index, as well as a CatBoost-based model proposed by Lai et al. 28 As shown in Figure 4, the Thal-classifier outperformed all other methods in terms of AUROC (Figure 4(A)), AUPRC (Figure 4(B)), accuracy, F1-score, and MCC (Figure 4(C)). The Green and King index demonstrated the second-best performance, with an AUROC of 0.879, AUPRC of 0.890, accuracy of 0.846, F1-score of 0.808, MCC of 0.695, precision of 0.913, and recall of 0.724. While most formulas achieved acceptable discrimination (AUROC ≥ 0.80), the Jayabose and Bessman indices exhibited suboptimal performance. These findings confirm the Thal-classifier's superiority in identifying thalassemia among pregnant women.

Comparative performance of the Thal-classifier and existing diagnostic approaches for thalassemia using the testing dataset. (A) ROC curves and (B) PR curves for the Thal-classifier and 11 established diagnostic formulas or models. (C) Summary of classification metrics for all 12 methods. The star denotes the method with the best performance.
Model interpretation of the Thal-classifier
To ensure clinical transparency, two complementary interpretability analyses were performed. Permutation importance analysis revealed that RDW-SD contributed most significantly to the model's predictive performance, followed by MCHC, MCH, MCV, and RBC (all P < 0.01, Figure 5(A)). While HCT, HGB, and RDW-CV showed relatively lower scores, all selected features exhibited positive importance values, indicating their benefits to the model. SHAP analysis provided a more granular understanding of feature contributions at both the global and individual levels (Figure 5(B)). Ranking by mean absolute SHAP value identified RDW-SD as the most influential predictor, corroborating the permutation results. The SHAP beeswarm plot further demonstrated that elevated RDW-SD, MCV, and MCH were associated with a lower probability of thalassemia, whereas decreased MCHC and RBC values contributed to a lower predicted risk. The agreement between these two interpretability methods supports the model's clinical transparency and utility.

Global explanation of the Thal-classifier on the testing dataset. (A) Feature importance scores derived from permutation analysis, indicating the overall contribution of each feature to the model's predictions. (B) SHAP summary plot showing the impact and relative importance of each feature. The left panel displays a dot plot illustrating the effect of individual feature values on model output, while the right panel presents a bar plot ranking features by mean absolute SHAP value.
Discussion
During pregnancy, due to physiological blood dilution, relatively insufficient red blood cell production, and the increased demand for hematopoietic raw materials by the fetus, anemia becomes a high-incidence period. 30 Routine prenatal examinations often attribute anemia to nutritional factors, such as iron deficiency anemia. 31 However, in high-incidence areas, such as southern China, thalassemia, as a hereditary hemoglobin disorder, is an important cause that is easily overlooked during pregnancy. 32 Thalassemia carriers may experience exacerbation or manifestation of their inherent mild anemia under the load of pregnancy, and the symptoms are similar to those of nutritional anemia, which can lead to misdiagnosis.3,33 Prenatal thalassemia screening is essential for identifying high-risk couples likely to have children with life-threatening severe thalassemia. 34 Beyond routine anemia management, gene screening accurately diagnoses anemia causes, guides treatment, and, through genetic counseling and prenatal diagnosis, prevents severe thalassemia births, reducing birth defects and improving outcomes. 35
In this study, we developed and validated the Thal-classifier, an interpretable ensemble learning model tailored for thalassemia detection in pregnant women. Based solely on routine hematological parameters, the model demonstrated excellent classification performance, achieving high accuracy, discrimination, and calibration in both internal cross-validation and independent testing. The clinical utility of the model was further supported by decision curve analysis, which demonstrated a clear net benefit across a wide range of decision thresholds. These results indicate that the Thal-classifier has strong potential as a practical, low-cost decision support tool in prenatal screening.
A key innovation of this study lies in the development of a diagnostic model specifically designed for pregnant populations. Previous studies have proposed numerous indices20,24 and ML-based models28,36 for thalassemia screening in general populations. However, they are often not applicable when used for pregnant women due to physiological changes in red cell indices during gestation. By incorporating a pregnancy-specific dataset and performing targeted feature selection, the Thal-classifier effectively captures the hematological alterations unique to pregnancy. The model's superior performance relative to conventional indices and existing ML models underscores its clinical relevance and diagnostic robustness. Several factors contributed to the model's success. First, feature selection using the Boruta algorithm enabled identification of the most predictive and physiologically relevant hematological markers. Second, ensemble modeling allowed integration of diverse ML algorithms, enhancing classification robustness and generalization. Third, the model's interpretability, achieved through permutation importance and SHAP analysis, facilitates transparency in clinical application, allowing clinicians to understand and trust the model's decision-making process.
Despite these strengths, the study has limitations. The model was developed using only CBC parameters, intentionally excluding advanced diagnostic indicators such as hemoglobin electrophoresis or genetic markers. This design choice was made to ensure applicability in resource-limited settings. Nonetheless, future research could adopt a tiered diagnostic strategy, where the Thal-classifier serves as a first-line screening tool, followed by confirmatory testing in ambiguous cases. Additionally, the current analysis was based on a single-center dataset with a moderate sample size, which may limit generalizability. Thalassemia prevalence and genotype distribution vary across geographic and ethnic populations, potentially influencing model performance. To address this, future multicenter studies with larger, more diverse cohorts are planned to further validate and refine the model.
Conclusion
This study presents a clinically applicable and interpretable ML model for thalassemia detection in pregnant women. The Thal-classifier demonstrated superior performance over traditional indices and recently proposed models, offering a reliable, accessible, and efficient screening tool that can be integrated with routine CBC analyzers to assist pathologists. Its application in prenatal care may facilitate early identification of thalassemia, and improve maternal and fetal outcomes. However, it should be noted that this model is intended for screening purposes only, and positive results must be confirmed by HPLC and genetic testing.
Supplemental Material
sj-xlsx-1-dhj-10.1177_20552076251396982 - Supplemental material for Development of an interpretable ensemble learning model for thalassemia detection in pregnant women using routine hematological parameters
Supplemental material, sj-xlsx-1-dhj-10.1177_20552076251396982 for Development of an interpretable ensemble learning model for thalassemia detection in pregnant women using routine hematological parameters by Qian Wang, Xianning Dai, Kai Xu, Mengjie Xu and Jianchao Ying in DIGITAL HEALTH
Footnotes
Acknowledgements
We are grateful to the staff of the Department of Clinical Laboratory, Wenzhou People's Hospital, for their assistance in data collection.
Ethical approval
The protocol of this study was carried out according to the Declaration of Helsinki and was approved by the Ethics Review Committee of Wenzhou People's Hospital (Approval Number: Y20210099). As this was a retrospective study using anonymized clinical and laboratory data, the requirement for written informed consent was waived by the committee. An English version of the ethics approval is available upon request.
Author contributions
Q.W.: data curation, formal analysis, funding acquisition, and writing—original draft. X.D.: validation, and resources. K.X.: visualization, and supervision. M.X.: investigation. J.Y.: conceptualization, project administration, and writing—review and editing. All authors read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the Science and Technology Planning Project of Wenzhou (Y20210099).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
Guarantor
Qian Wang is the guarantor of this article and accepts full responsibility for the integrity of the work, including the data, analysis, and manuscript content.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
