Abstract
Objectives
To map global research on AI-driven histopathological image interpretation in oncology (2000–2024).
Methodology
Scopus-based bibliometrics using Boolean queries; data exported, cleaned, and analyzed in VOSviewer.
Results
The dataset included 1874 publications, surging after 2015. China (n = 561) and the United States (n = 467) led output; leading institutions included Harvard Medical School and the Chinese Academy of Sciences. Top journals were Scientific Reports, Cancers, and IEEE Access; the corpus H-index was 112. Collaboration networks intensified, especially U.S.–Asia. Keyword mapping showed four clusters: (1) deep learning for breast cancer/CNN diagnostics; (2) transfer learning/feature extraction; (3) prognostic modeling/tumor microenvironment; and (4) digital infrastructure/explainable AI. Overlay analyses traced a shift from classical machine learning to transformers and multimodal models integrating molecular and clinical data; emerging themes include semantic segmentation, self-supervised learning, and therapy-response prediction. Applications spanned breast, prostate, colorectal, head–neck, gynecologic, and gastrointestinal/liver cancers. Models primarily used whole-slide images (e.g., TCGA) and multi-omics; algorithms included CNNs, deep learning, classical machine learning, and weakly supervised approaches. Evidence ranged from proof-of-concept to multicenter validation and workflow integration; adoption remains constrained by data standardization, interpretability, and regulation. Clinically, AI improved diagnostic accuracy/efficiency and supported personalization via multi-omics. Bibliographic coupling revealed three clusters: clinical/translational journals; engineering/computational outlets; and interdisciplinary venues linking algorithmic innovation with digital pathology.
Conclusions
AI histopathology is advancing toward clinical-grade deployment, propelled by collaboration and methodological innovation, yet limited by data standards, explainability, and regulatory requirements.
Keywords
Introduction
The landscape of modern oncology is undergoing a profound transformation, driven by the convergence of artificial intelligence (AI), computational imaging, and digitized diagnostics.1,2 At the forefront of this transformation is histopathology, the microscopic examination of stained tissue sections, which remains the gold standard for cancer diagnosis, grading, and therapeutic guidance. While historically reliant on manual interpretation by expert pathologists, histopathology is increasingly being reimagined through the lens of digital and algorithmic technologies.3–5 The synthesis of AI with histopathological slide interpretation is not merely a technological augmentation; it represents a paradigm shift with the potential to enhance diagnostic accuracy, reduce interobserver variability, and introduce unprecedented levels of efficiency into clinical workflows.6–8
The digitization of pathology, commonly termed
Recent studies have documented a rapid and sustained increase in the number of publications focusing on AI applications in histopathology, reflecting the fields’ swift transition from foundational image processing work to high-impact, clinically oriented research and practice.13–16 However, despite the escalating volume of literature on AI in pathology, the field remains fragmented in its documentation and evaluation. The present study seeks to address this limitation through a focused and comprehensive knowledge mapping and bibliometric analysis that address AI-powered histopathological interpretation in cancer as a distinct research domain. Knowledge mapping and bibliometric analysis are two complementary methodologies used to systematically evaluate the landscape of scientific research on a specific topic. Knowledge mapping refers to the process of visualizing structure, development, key thematic areas within a body of literature, helping researchers identify how knowledge is generated, interconnected, and evolving over time. Meanwhile, bibliometric analysis quantitatively assesses publications using metrics such as average annual growth rate (AAGR), citation counts, and key contributors. 17 Unlike traditional narrative reviews, which are often subjective and selective, or systematic reviews, which focus narrowly on clinical outcomes or interventions using strict inclusion criteria, bibliometric analysis provides a broad, objective, and data-driven overview of the structure and dynamics of scientific research.18,19
Research gap
A focused literature scoping shows that while bibliometric work on AI in medicine is abundant, histopathology-specific research mapping and trend analysis studies remain scarce and fragmented. Six main publications centrally addressed digital pathology/histopathology: two disease-specific mappings (e.g., urological tumor histopathology, lung cancer pathology; 2024–2025),13,16 and four earlier,20–23 broad “digital pathology” overviews (2021–2022) that neither (i) provide an oncology-wide synthesis dedicated to whole-slide image (WSI) interpretation, nor (ii) capture the post-2022 transition toward transformers, self-supervised pretraining, multimodal fusion, and translation-oriented validation. Moreover, none stratifies findings by cancer site/subtype, WSI task family (classification, segmentation, prognostication), and model class, while giving transparent VOSviewer parameterization for full reproducibility. Our study fills this gap by delivering a comprehensive, oncology-wide bibliometric and knowledge-mapping analysis of AI-powered histopathology slide interpretation (2000–2024) with clinically meaningful stratification, explicit visualization parameters, and a translation-readiness perspective that connects algorithmic advances to validation pathways and adoption barriers. In short, whereas prior works either provide a broad navigation aid or an early-phase global snapshot, our contribution delivers a current, oncology-specific, decision-ready knowledge map that connects pixels to predictions and research activity to translation readiness. To achieve the objectives of the current study, the following bibliometric indicators and knowledge maps were sought and presented: (1) Volume of research output and AAGR; (2) The most prolific journals, countries, authors, institutions, and most cited articles; (3) Authorship, citations, and collaboration metrics; (4) Keyword co-occurrences and temporal map; (5) Journal coupling to map the scientific evolution and development of the field; and (6) Clinical insights in the field (cancer sub-types, proof-of-concept, clinical validation, and real-world deployment).
Methodology
Adherence to bibliometric standards
This current study was conducted in accordance with established protocols outlined in the BIBLIO methodology framework, which provides rigorous standards for designing, executing, and reporting bibliometric studies. 13 The 20-item checklist is shown in the attached checklist (Supplement 1). The BIBLIO approach emphasizes methodological transparency, replicability, and analytical depth, and has become a widely recognized set of guidelines for scholars conducting bibliometric inquiries across disciplines. The present study aligns with BIBLIO principles by clearly defining the search strategy, data sources, eligibility criteria, and analytical techniques, thereby ensuring the validity and reproducibility of the results. Key bibliometric indicators, such as citation frequency. Authorship productivity, institutional output, and keyword co-occurrence, were systematically analyzed and interpreted in line with these recommendations.
Data source and database selection
The primary corpus of literature for this study was extracted from Scopus, a multidisciplinary bibliographic database developed by Elsevier. Scopus was selected as the exclusive data source due to its expansive coverage of scientific journals, high-quality metadata, and robust citation tracking capabilities. 24 With over 25,000 peer-reviewed journals indexed across the life sciences, physical sciences, health sciences, and social sciences, Scopus provides an unparalleled breadth suitable for a bibliometric study of this scope. In contrast to other commonly used databases such as Web of Science and PubMed, Scopus offers several distinct advantages.25,26 Web of Science, while rich in citation data, maintains a narrower journal selection, and its search interface presents limitations in Boolean query flexibility. PubMed, although authoritative in biomedical sciences, is limited primarily to abstracts and lacks comprehensive citation analytic. Scopus, by comparison, allows full Boolean control, supports detailed field searches, and integrates citation counts, author affiliations, and funding data, essential elements for bibliometric mapping and network analyses. 24
In the current study, a single database was used to retrieve relevant literature. Bibliometric analyses are quite different from systematic or scoping reviews. Systematic reviews attempt to answer a quite specifically defined research question and therefore need to employ more than one database offering complete coverage but without any biasedness. 19 In contrast, bibliometric studies care about mapping and quantifying the general structure, dynamics, and evolution of scientific production, where the point is not to download all articles but to capture a significant enough and representative enough corpus. It is commonly known that published bibliometric analyses rely on a single massive multidisciplinary database, particularly when dealing with topics having large numbers of publications.27–30
Search strategy and query formulation
The search strategy was designed to capture literature at the nexus of AI, histopathological image interpretation, and oncology. Using advanced Boolean operators, truncation symbols, and field-specific syntax, the query was constructed to maximize both sensitivity (comprehensiveness) and specificity (relevance). The search query was carried out using “TITLE-ABS” function and the results of all queries were combined with the function “AND”. To increase relevance, provisional keywords such as (pathology* or histopathology* or cytopathology* or tissue biopsy) were used in TITLE-ABS as a must for all retrieved articles. Furthermore, the Boolean “AND NOT” was used as an exclusion for documents having any of the following terms: (“CT image” OR “NMR image” OR ultrasound OR radiolog*). The keywords used in the search query were based on terms found in previously published studies on related topics with slight modification and adaptation to the current study.13,15,16
The final search query was executed using the following keywords in the Scopus advanced search interface: (Supplement 2)
AI: “machine learning” OR “deep learning” OR “artificial intelligence” OR “neural network*” OR “artificial intelligence” OR “machine intelligence” OR “AI technology” OR “AI applications” OR “intelligent systems” OR “smart algorithms” OR “deep learning” OR “Convolutional neural network*” OR “deep network” OR “Bayes network” OR “deep learning” OR “neural network” OR “neural learning” OR “neural nets model” OR “artificial neural network” OR “fully supervised learning”. Histopathology slide image: “histopathology image” OR “digital pathology image” OR “whole slide image*” OR “tissue section image*” OR “biopsy image*” OR “microscopic image*” OR “cytopathology image*” OR “WSI” OR “patholog* imag*” OR “slide image” OR “digital patholog*” or “digital image”. Interpretation: “cancer”/ interpret* OR detect* OR grading* OR classif* OR diagnos* OR feature* OR identif* OR segment OR predict OR prognos*. Oncology: cancer OR oncology OR carcinoma OR malign* OR leukemi* OR lymphoma OR melanoma OR “multiple myeloma” OR sarcoma OR *blastoma OR neoplasm OR tumor OR tumour OR metast*.
Eligibility criteria
A rigorous set of inclusion and exclusion criteria was applied to ensure that only high-quality, relevant publications were analyzed:
Only peer-reviewed original full research articles were retained. Reviews, editorials, commentaries, letters to the editor, preprints, conference proceedings, books, and book chapters were all excluded. We intentionally excluded conference proceedings and limited our corpus to peer-reviewed original research articles to maximize methodological comparability and clinical relevance. In oncology/histopathology, the primary vehicle for mature, fully documented evidence is the full-length journal article, whereas proceedings often contain preliminary findings with abbreviated methods, heterogeneous or lighter peer review, and inconsistent indexing/metadata. Including proceedings would (i) introduce versioning and double-counting when conference papers later appear as expanded journal articles, and (ii) impede reliable extraction of endpoints, validation schemes, datasets, and code/data availability, variables central to our analysis of rigor and translation readiness. Restricting to original research articles (and excluding reviews, editorials, and letters) also reduces citation inflation from secondary literature and focuses the bibliometric map on primary, reproducible contributions. This design choice improves internal validity of comparisons across institutions, countries, and model families, and aligns the dataset with our objective: to characterize the mature, clinically meaningful evidence base for AI-driven histopathology. Only publications that are finally published, but not those in press, were included to ensure metadata stability and repeatability. Language restriction was used and limited to English. Only articles published between 2000 and 2024 were included. Articles that addressed pathology image analysis outside of cancer contexts were not included.
Validation of the search strategy
To ensure the validity, precision, and reproducibility of the search strategy, a two-tiered validation process was employed. (1) Benchmark Article Recall Test: a curated list of seminal and highly cited articles (n = 30) known to the domain experts was used as a reference to test the sensitivity of the query. The selection of these seminal articles was based on the number of citations. Adjustments to the query syntax were made until the final version captured 100% of these benchmark articles, ensuring high recall. (2) False Positive Scan: a manual inspection of the 100 most recent articles in 2024 was conducted to assess relevance. Only a negligible proportion of false positives were found.
Data extraction and quantitative analysis
The finalized query was executed on June 19, 2025, and bibliographic data were exported from Scopus in CSV format. The metadata fields extracted included: article title, author names and affiliations, journal name, abstract and author keywords, citation count. Data were analyzed to generate the following indicators: annual scientific production and growth rate, authorship analysis, citation metrics, country-wise publication trends and collaboration metrics, keyword frequency and thematic evolution.
Visualization and knowledge mapping with VOSviewer
To uncover and visualize structural relationships within the bibliometric corpus, VOSviewer (version 1.6.20) was employed. 31 The software facilitated the generation of multiple bibliometric maps. Co-authorship networks map identifies relationships among authors or countries based on co-publication frequency. Node size represented publication volume, while edge thickness denoted collaboration strength. Keyword co-occurrence map group connected terms into major themes. The map enabled identification of intellectual structures and emerging domains. Overlay Visualization was conducted to illustrate the temporal evolution of keyword relevance. Earlier topics are contrasted with recent innovations using color gradient capturing the shifting technological frontier. Bibliographic coupling map identifies the relationship between two journals based on the number of references they share, i.e., the number of common sources they cite.
Visualization parameters
Bibliometric maps were generated in VOSviewer v1.6.20 using the following pre-specified settings. For keyword co-occurrence, we analyzed author keywords with a minimum occurrence threshold of 10, fractional counting, and association-strength normalization. The overlay visualization encoded the average publication year using VOSviewer's default layout. For co-authorship networks, a minimum of 5 documents per author (fractional counting; association strength) was required. The country collaboration map used a minimum of 10 documents per country. For journal bibliographic coupling, we set ≥20 documents per journal (fractional counting; association strength). Thresholds were chosen to balance noise reduction with structural fidelity on a corpus of 1874 items; fractional counting mitigates consortia-size bias and association-strength normalization stabilizes similarity estimation under uneven frequency distributions.
Ethical considerations
This study involves the analysis of publicly available scientific literature and does not include
human participants, clinical data or patient records. As such, it is exempt from institutional ethical review, in accordance with international standards for bibliometric research. All efforts were made to ensure accurate citation, data integrity, and adherence to academic publishing norms.
Results
Flow chart
Based on the search methodology, 1874 scientific documents were retrieved. Supplement 3 shows the steps implemented in the search strategy with the corresponding number of articles retrieved in each step.
Growth of publications
A total of 1874 original research articles were retrieved, revealing a substantial increase in research activity over the study period. The number of annual publications grew from less than 5 articles in 2014 to 479 in 2024. The growth of publications witnessed a slow emergence phase from 2015–2018, followed by a rapid growth phase from 2019 to 2024 (Table 1). The upward trend reflects a striking AAGR of 62.7% during the last decade.
Number of research articles published in the field over the study period (2000–2024).
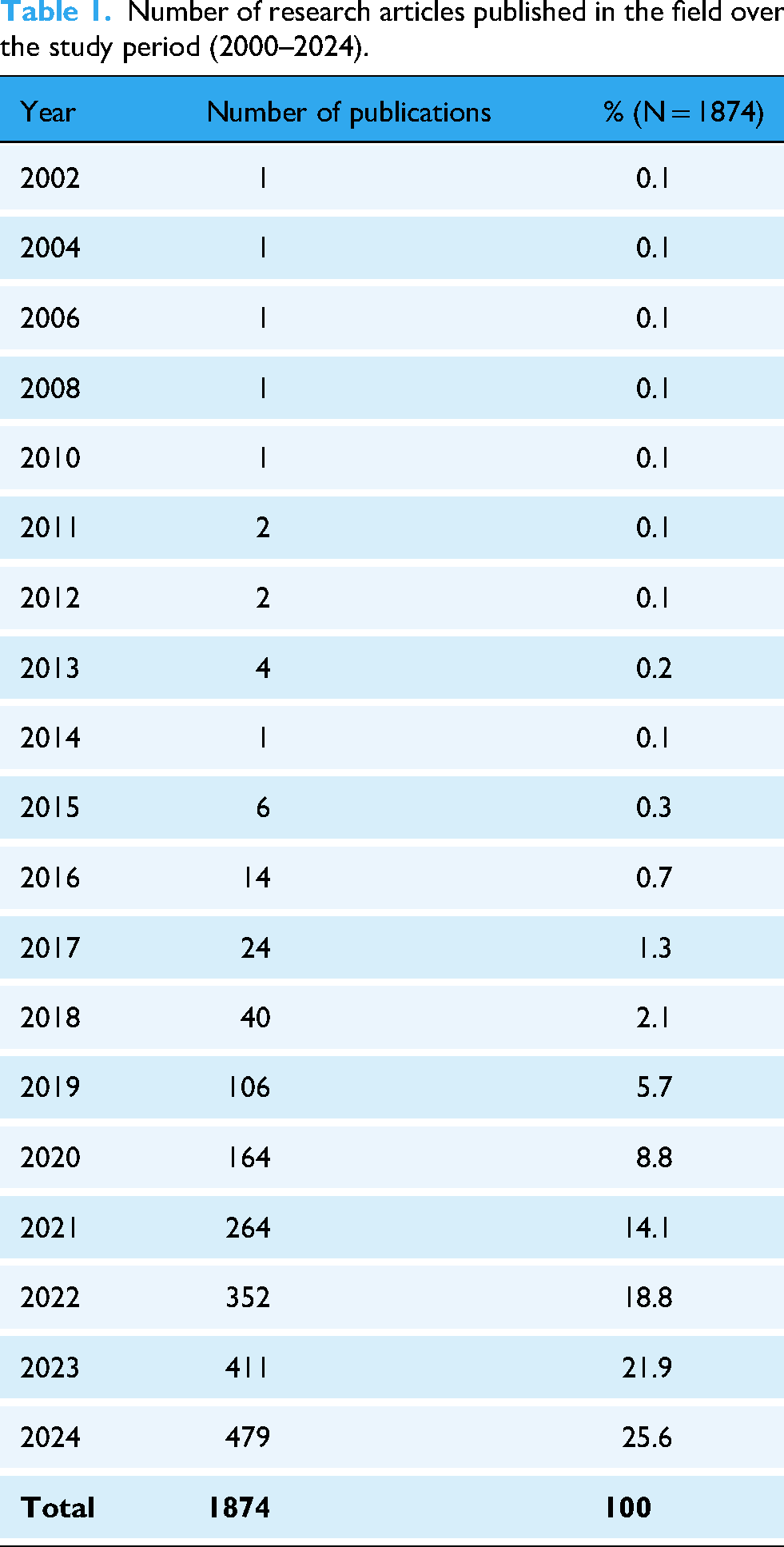
Key contributors
From a geographical perspective, China emerged as the most prolific contributor with 561 publications (29.9%), followed closely by the United States with 467 (24.9%), and India with 250 (13.3%) (Table 2). Institutionally, the
Top 10 prolific countries in publishing research articles in the field (2000–2024).

*The total number and % cannot be calculated due to overlap in publications between countries.
Top 10 prolific journals in publishing research articles in the field (2000–2024).

Citation and authorship metrics
A total of 14238 individual author mentions were recorded across the 1874 articles, yielding an average of approximately 7.6 authors per article. This relatively high average reflects the interdisciplinary nature of the field, which often involves joint efforts among computer scientists, biomedical engineers, pathologists, and clinical researchers. A total of 1293 articles (69.0%) featured five or more authors per article, underscoring the collaborative demands of developing, validating, and deploying AI systems in clinical pathology. These findings suggest that advancing research in the field not only requires technical innovations but also cooperation across diverse disciplines to ensure methodological rigor, clinical relevance, and translational potential.
The citation analysis of the 1874 retrieved articles indicates the significant academic and clinical impact of the field. Together, the retrieved articles have received a total of 72021 citations, an average of 38 citations per article. The calculated H-index of the retrieved articles at the time of analysis was 112, meaning that at least 112 articles have each been cited 112 times or more.
Top five most cited articles
A review of the top five most cited articles reveals foundational and transformative contributions that have shaped the trajectory of the field. The most cited work, published by Bejnordi et al. (2017) in
When citation impact is normalized by publication year, the most recently published studies exhibit a higher relative influence than their raw citation counts suggest. However, because the field is relatively young and most high-impact publications have appeared within a narrow time window, the year-normalized metric showed smaller citation differences when normalized by year of publication.
Author collaboration map
The author collaboration network map (Figure 1) comprising 129 researchers with a minimum of five publications and existed in a research cluster revealed a significantly fragmented yet evolving research landscape in the field. The map highlights both the key contributors in the field and the extent of intra- and inter-group collaboration, reflecting the field's increasing complexity, interdisciplinarity, and internationalization. Analysis of the map reveals the following:
At the center of the map is a prominent red cluster led by Nasir Rajpoot, a highly influential figure whose team is deeply engaged in computational pathology and weakly supervised learning approaches. This cluster includes collaborators such as Chen Hao, Jahanifar Mostafa, and Shaban Muhammad, showing strong internal collaboration and a wide-reaching influence that connects to multiple other clusters. Notably, Rajpoot group acts as a bridge between Western and Asian researchers, reflecting its pivotal role in promoting global research integration. In the bottom-left quadrant, Anant Madabhushi leads another major hub (green and blue clusters), anchoring a large and technically sophisticated cluster that includes Lu Cheng, Liu Zaiyi, Zhenhui Li, and others. This group is characterized by strong interdisciplinary work at the interface of artificial intelligence, radiology, and pathology. Their collaboration network is notably dense, reflecting a high degree of productivity and a leadership role in the development of clinical grade AI tools for histopathological analysis. The orange cluster in the top left is led by Yang Lin, focusing on breast cancer and convolutional neural network applications. This cluster's relatively contained structure suggests a strong institutional collaboration network, with key links to contributors such as Rong Ruichen and Amgad Mohamed. Adjacent to this is a green cluster featuring Wang Xiang, Zhang Ning, and other Chinese researchers in Chinese institutions. On the right side of the map, the yellow cluster highlights Jakob Nikolas Kather, whose work is influential in spatial biology, multimodal integration, and explainable AI. His collaborations with Lu Ming Y. and Brobeli Alexander signal a focus on precision oncology and the integration of AI with molecular pathology, establishing a strong European research presence. The cyan cluster contains researchers like Shao Wei and Fan Richard E, forming smaller but focused groups engaged in method development, model explainability, and digital infrastructure. Their separation from the central clusters suggests niche research themes, possibly with less global collaboration but significant technical depth. Reaching influence that connects to multiple other clusters. Notably, Rajpoot group acts as a bridge between Western and Asian researchers, reflecting its pivotal role in promoting global research integration. The presence of Pantanowitz Liron in the pink cluster and Kurc Tahsin in the purple cluster further expands the map's disciplinary reach, representing researchers involved in translational digital pathology, clinical deployment, and education-focused AI systems. These peripheral clusters maintain limited but strategic connections to central hubs, signaling their relevance in shaping practical applications and standards in AI assisted pathology.

Network visualization map of author collaboration created by VOSviewer program. The map included authors (n = 129) with a minimum of five publications and existed in a research cluster. Center (red): Nasir Rajpoot's hub bridging Western–Asian networks. Bottom-left (green/blue): Anant Madabhushi's dense, interdisciplinary cluster driving clinical-grade tools. Top-left (orange ± adjacent green): Yang Lin's breast/CNN focus; nearby Chinese institutional cluster (e.g., Wang, Zhang). Right (yellow) & periphery (cyan/pink/purple): Jakob N. Kather's spatial biology/multimodal/XAI; smaller method/infra & translational/education nodes (e.g., Shao, Fan, Pantanowitz, Kurc) linking selectively to core hubs.
Overall, the map reveals a moderately centralized but growingly interconnected research ecosystem, with a few dominant research hubs facilitating collaboration across regions, institutions, and disciplines. While dense internal connections within each cluster suggest strong intra-team productivity, the emergence of cross-cluster links points to a maturation of the field moving from siloed technical work toward a globally integrated, translational science. Importantly, the map reflects the field's interdisciplinary nature, involving computer scientists, clinicians, biomedical engineers, and pathologists all working collaboratively to transform histopathology with artificial intelligence.
Country collaboration map
The country level collaboration map depicts a globally distributed, yet hierarchically structured network, with a few dominant countries acting as international hubs of collaboration in the field (Figure 2). The US stands as the central global connector, exhibiting strong co-authorship ties with nearly all major contributors including China, India, Germany, United Kingdom, Canada, and Australia. This confirms the US research leadership and high collaborative ties with a large number of countries. In contrast, China has a large research output but has lesser collaborative ties with other countries. China has collaborative ties with US, European countries, and certain Asian countries such as Japan, Singapore, and South Korea. Both India and Germany function as secondary hubs; Germany is well connected across Europe, while India shows bilateral connections with Middle Eastern and Anglophone countries. South–South collaboration exists but remains sparse compared to North–South ties, highlighting opportunities to broaden global equity and leadership.

Network visualization map of country collaboration. The map included 39 countries, each with a minimum contribution of 10 publications. US is the dominant hub; China has high output but fewer international links (mainly US/Europe/East Asia). India and Germany are secondary hubs; South–South ties remain limited versus North-South.
Keyword co-occurrence analysis
The author keyword co-occurrence map visualizes the intellectual landscape of the field (Figure 3). At the center of the map lies the keyword “deep learning”, the most prominent and widely connected node, symbolizing its foundational role in the computational analysis of histopathological slides. Deep learning is directly linked to broader concepts like “artificial intelligence”, “whole slide images”, and “digital pathology”, revealing its integration across data formats, systems, and analytical pipelines. Surrounding this core are distinct clusters, each reflecting a specific subdomain.
Blue Cluster (Disease-focused applications): This cluster is organized around keywords such as: “breast cancer”, “classification”, “histopathology images”, “convolutional neural networks” (CNNs). It reflects a mature and well–studied application area where AI is used to automate tumor grading, subtype classification, and biomarker prediction in breast cancer diagnostics. The appearance of “invasive ductal carcinoma” highlights the focus on real-world pathological subtypes. Red Cluster (Methodological innovation): Key terms include “transfer learning”, “data augmentation”, “feature fusion”, “image processing”. This cluster points to the technical backbone of AI modeling in histopathology. It reflects efforts to optimize model performance, reduce reliance on large labeled datasets, and improve generalizability across tissue types and image modalities. These techniques are critical for domains like colon cancer and oral cancer, which appear in close proximity. Green Cluster (Cross-cancer and Prognostic modeling): Notable terms include “colorectal cancer”, “gastric cancer”, “prognosis”, “tumor microenvironment”, “pathomics”. This cluster demonstrates the expanding application of AI from diagnosis to prognosis and treatment planning, using deep learning to extract high-dimensional features (e.g., pathomics) that can predict outcomes, immune infiltration, and therapy response. Purple Cluster (Digital infrastructure and Model explainability) includes “digital pathology”, “image analysis”, “computational pathology”, “interpretability”, “semantic segmentation”. This group reflects the convergence of software tools, data platforms, and explainable AI (XAI) techniques essential for clinical deployment. Terms like “U-Net”, “nuclei segmentation”, and “self-supervised learning” point toward cutting-edge methods used to enhance accuracy and reduce annotation burden in large-scale WSI analysis.
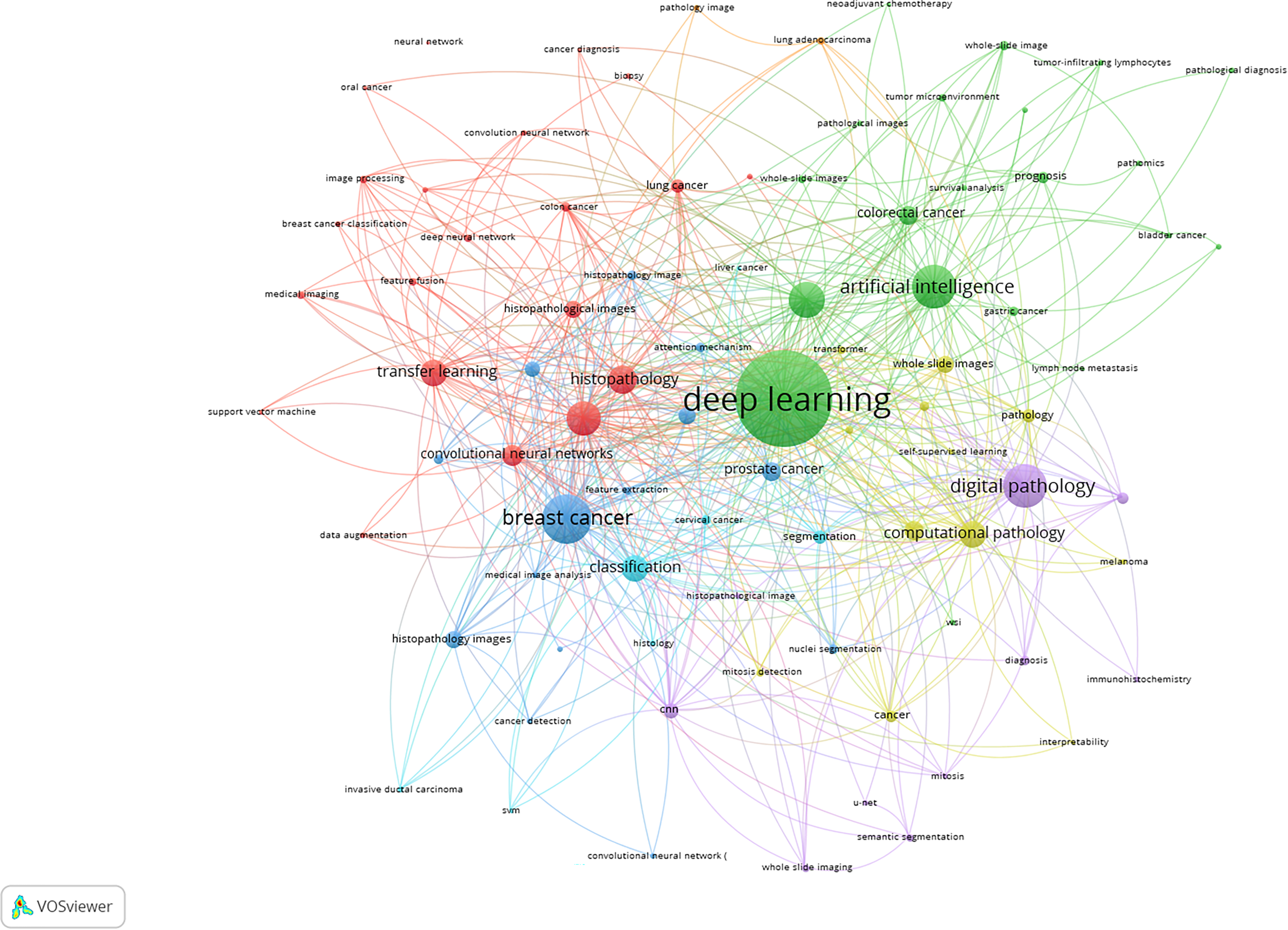
Network visualization map of author keywords co-occurrence map. The map included 86 author keywords, each with a minimum occurrence of 10 times. Four subdomains emerge: (Blue) disease-focused applications (breast cancer, CNNs, classification); (Red) methodological innovation (transfer learning, augmentation, feature fusion); (Green) cross-cancer/prognostic modeling (colorectal/gastric cancer, prognosis, tumor microenvironment, pathomics); (Purple) digital infrastructure & explainability (computational pathology, segmentation/U-Net, self-supervised learning).
Overlay visualization map
Figure 4 is an overlay visualization map generated by VOSviewer based on author keywords. This type of map is not just a co-occurrence network, it is temporally coded using a color gradient to show the average publication year of keywords. The color bar at the bottom right indicates the time line: Purple to blue nodes represent older terms (before 2021), Green to yellow nodes represent terms that are more recent/emerging terms (after 2022).
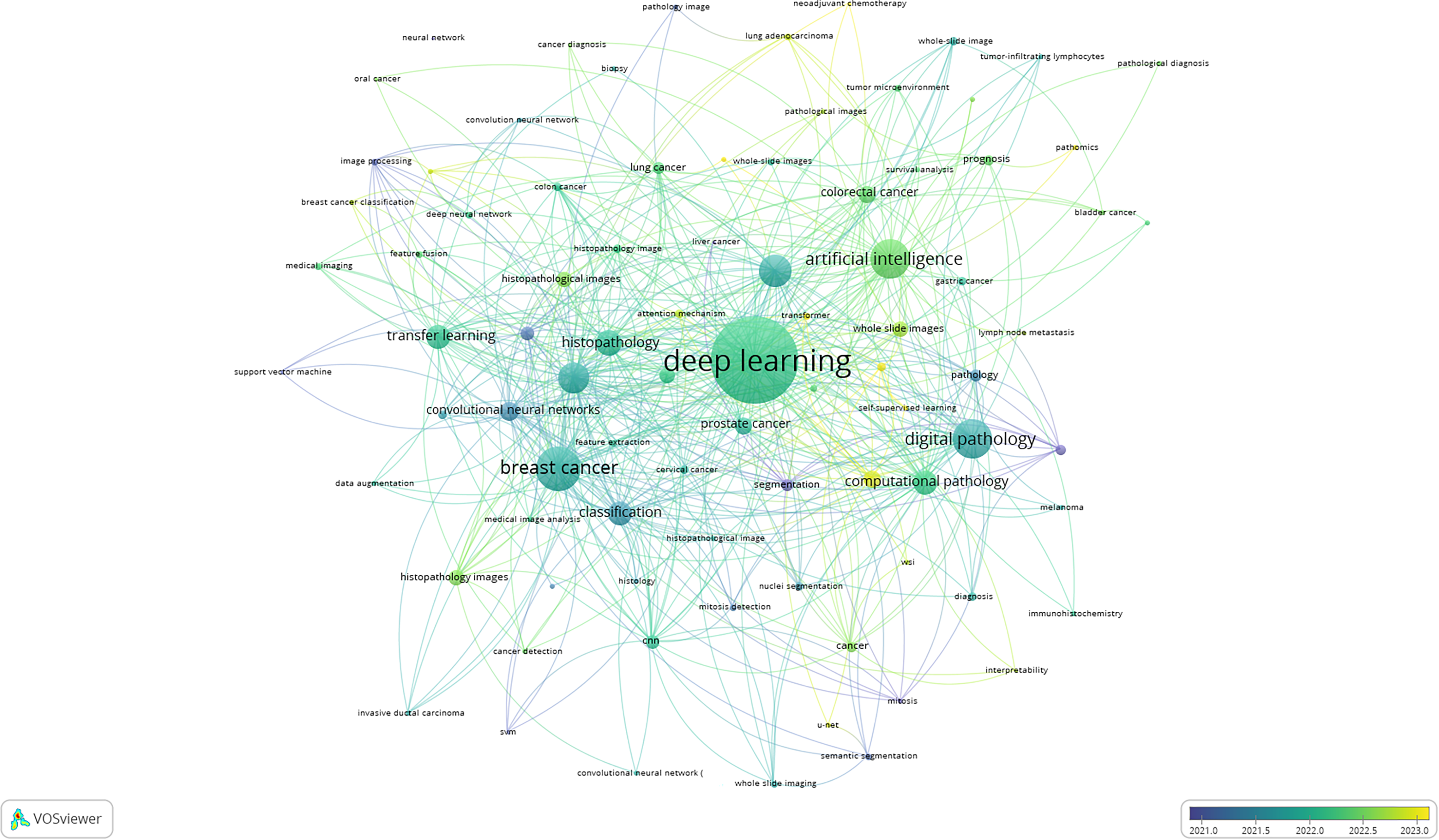
Network overlay visualization map showing the temporal dynamics of author keywords. Dark colored nodes appeared in literature before 2021 while light colored nodes appeared after 2022. Purple/blue = earlier (≤2021); green/yellow = recent (≥2022). Central, well-connected foundations: “deep learning,” “AI,” “whole-slide images,” “digital pathology,” “CNNs are blue-green; older methods (SVM, image processing, feature fusion, breast-cancer classification) are purple/blue. Newer trends in green–yellow highlight “self-supervised learning,” “transformer,” “U-Net/semantic segmentation” and expansion beyond breast cancer to colorectal/gastric/HCC/lung adenocarcinoma. Clinical/translation terms (“prognosis,” “TILs,” “pathomics,” “interpretability,” “immunohistochemistry,” “multiple-instance learning,” “neoadjuvant chemotherapy”) signal a shift from classification toward prognostic, integrative applications.
At the center of the map, there are keywords such as “deep learning”, “artificial intelligence”, “whole slide images”, “digital pathology”, “histopathology”, and “convolutional neural networks”, which are predominantly coded in blue-green hues. These represent established concepts and technologies that have formed the backbone of this research field before 2021. Their central placement and dense interconnectivity indicate their foundational role in shaping both the theoretical and applied landscape of AI in computational pathology. Several classical terms such as “support vector machine (SVM)”, “image processing”, “feature fusion”, and “breast cancer classification” appear in blue or purple suggesting that they were used more frequently in earlier studies (pre-2021). These methods, although pioneering, are gradually being supplanted by more sophisticated and flexible approaches such as deep learning and transformer architectures. Conversely, recent methodological advances like “self-supervised learning”, “transformer”, U-Net”, and “semantic segmentation” are shaded in green to yellow, indicating they have emerged in publications from 2022 onward. These reflect a shift toward data-efficient and interpretable models, especially relevant in histopathology where annotated datasets are limited.
Disease-specific terms like “breast cancer”, “lung cancer”, “prostate cancer”, and “colorectal cancer” remain heavily interlinked with technical terms but vary in temporal coding: “Breast cancer” appears in blue, confirming it as a long-established testbed for AI applications. “Colorectal cancer”, “gastric cancer”, “hepatocellular carcinoma”, and “lung adenocarcinoma” appear in green-yellow, suggesting emerging attention in more recent years. This trend reflects the broadening scope of histopathology-AI research toward less-explored malignancies. Clinical outcome-focused keywords like “prognosis”, “tumor-infiltrating lymphocytes”, and “pathomics” are also recent additions (green), suggesting a transition from simple classification tasks to predictive and prognostic modeling in oncology.
Keywords such as “interpretability”, “immunohistochemistry”, “Multiple instance learning”, and “Neoadjuvant chemotherapy” highlight recent expansion into explainable AI, molecular-pathological integration, and treatment response prediction. Their recent emergence (green/yellow) signifies the field's evolution from algorithmic novelty to clinical applicability, targeting translational utility in personalized medicine and integrative diagnostics.
Cancer sub-types
Keyword co-occurrence and overlay analysis of the retrieved documents showed that AI-powered histopathology has been applied to various cancer subtypes, including: Breast cancer,37–40 prostate cancer,41,42 colorectal cancer,43,44 head and neck cancer,45–47 gynecological cancers, 48 and gastrointestinal/liver cancer.48–50
Data sources and algorithms for AI models
Keyword mapping indicated that AI models utilize various data sources, including WSIs (Digitized slides from multiple medical centers and large-scale databases like The Cancer Genome Atlas (TCGA),51–53 Another important source is multi-omics that include integration of transcriptomics, proteomics, and other molecular data to enhance diagnostic accuracy and treatment personalization.48,54 AI algorithms being employed in histopathology include (1) CNN, a widely used for image classification, tumor identification, and segmentation43,45,55; (2) deep learning models, applied for complex tasks like tumor grading, molecular profiling, and prognostic predictions50,56; (3) machine learning models, utilized for tissue classification, mutation detection, and predictive modeling41,45; and finally (4) weakly supervised learning, a techniques like EWSLF address issues of high-resolution images and lack of manual annotations. 57
Clinical validation
The retrieved literature indicated that AI applications in histopathology have undergone several stages of clinical validation. As a Proof-of-Concept, initial studies demonstrated AI's potential in enhancing diagnostic accuracy and efficiency.38,43 AI models were validated on multi-center datasets and integrated into clinical workflows.42,51,52 Challenges and future directions discussed in literature pertaining to implementation of AI in histopathology included issues like data standardization, model interpretability, and regulatory considerations need to be addressed for broader clinical adoption. 50
Clinical insights and practice
Highly cited articled and keyword co-occurrence analysis showed that AI models (1) achieve high accuracy in tissue classification and cancer diagnosis, often surpassing traditional methods38,43,51; (2) enhances diagnostic efficiency, reducing pathologists’ workload and improving patient outcomes41,58; and (3) allow for personalized treatment strategies through integration of multi-omics data. 54 Clinically, data standardization, model interpretability, and regulatory hurdles remain significant barriers. 59
Bibliographic coupling map
Figure 5 is a bibliographic coupling map generated using VOSviewer. The bibliographic coupling map offers an insightful visualization of the intellectual structure of journals publishing literature on AI powered histopathology slide image interpretation in oncology. In this network, each node represents a journal, with the size reflecting the volume of relevant publications and the proximity and linkage indicating shared citation pattern, i.e., how often the same references are cited across journals. Three dominant clusters emerge, each capturing a distinct thematic domain within the field.
The green cluster represents a clinically focused and translational research domain, anchored by journals such as Scientific Reports, Cancers, Modern Pathology, American Journal of Pathology, and Nature Communications. These journals bridge computational research with clinical application and often publish studies involving AI assisted cancer diagnosis, prognostic modeling, and digital pathology workflows. Their dense interconnectivity and central positioning underscore their importance in disseminating interdisciplinary, clinically grounded AI research. The red cluster is dominated by engineering and computational science journals such as IEEE Access, Biomedical Signal Processing and Control, Computers in Biology and Medicine, and Multimedia Tools and Applications. This group primarily supports methodological innovations, including the development and benchmarking of deep learning algorithms, signal processing techniques, and image analysis frameworks. The strong bibliographic coupling within this cluster indicates a shared foundation in computational methodologies tailored to biomedical imaging.
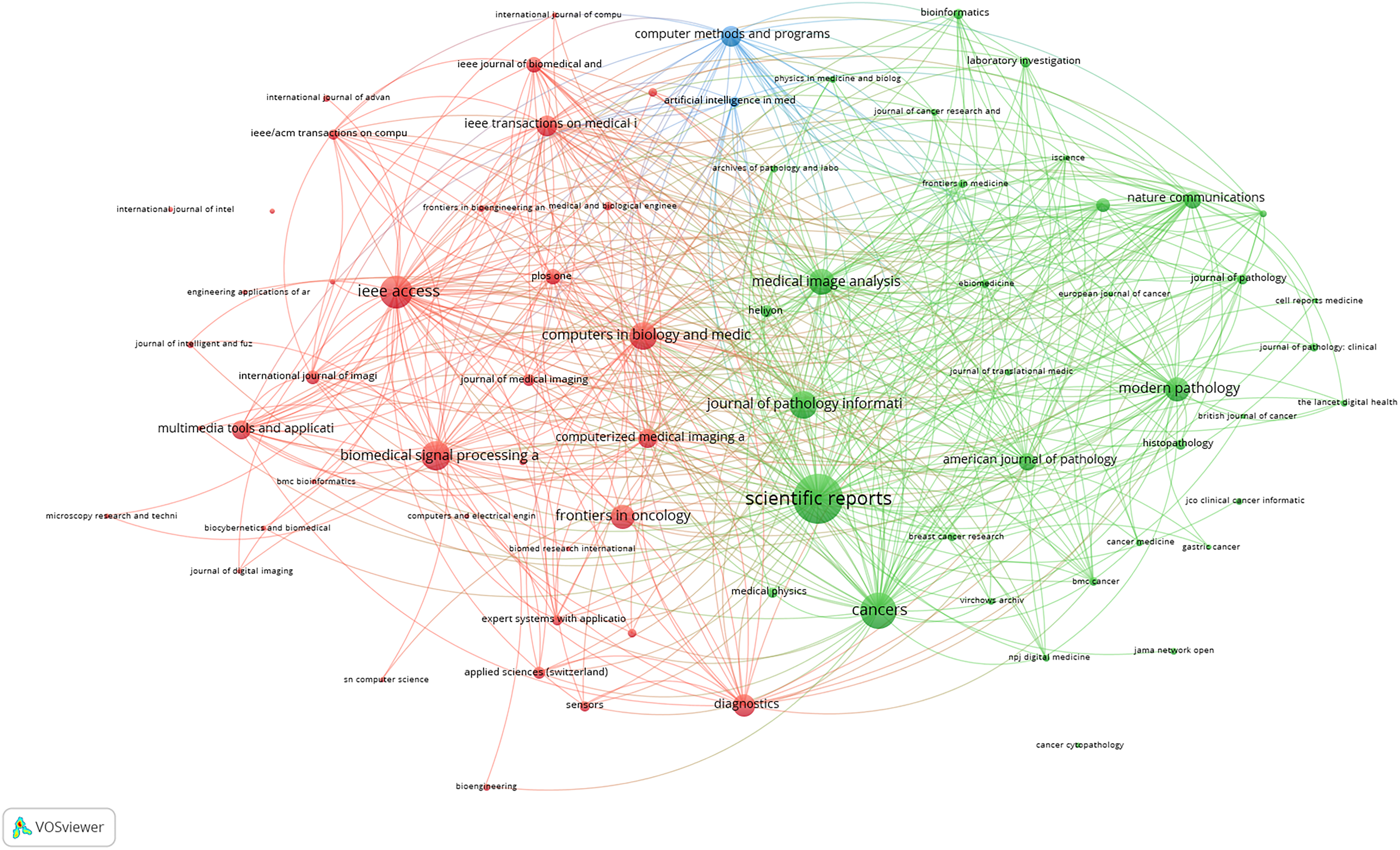
Bibliographic coupling map of journals with a minimum contribution of five publications. Three clusters emerge: Green (clinical/translational)
Discussion
Growth and key contributors
Between 2000 and 2024, global research on AI-powered histopathological image interpretation expanded from only a handful of papers per year to 479 papers in 2024, yielding an average annual growth rate of 62.7% in the last decade and a clear inflection after 2015. This surge coincided with two enabling advances: (1) breakthroughs in deep learning architectures such as U-Net,60,61 Res-Net,62,63 and transformer model, 64 alongside the proliferation of open-access whole slide imaging datasets. 65 The surge in the number of publications mirrors (1) the global initiatives to automate diagnostic processes, improve efficacy, and handle increasing pathology workloads 34 ; (2) community datasets/challenges (e.g., CAMELYON16 for metastasis detection) that standardized benchmarking; and (iii) rapid digitization of pathology and maturing regulatory/validation frameworks for clinical WSI use.66,67 Together these factors compressed the bench-to-bedside cycle and explain the step-change in publication volume after 2015.
Research impact
The h-index of 112 indeed reflects a substantial body of highly cited work in the field of AI-powered histopathology. To contextualize this finding, it is important to note that bibliometric studies in other domains of medical AI often report considerably lower h-indices. For example, bibliometric analyses of AI in radiology and medical imaging typically report h-indices in the range of 70–90, while fields such as AI in digital pathology and computational oncology are usually lower, ranging from 40–80 depending on the scope and time frame.13,21,2368–71 The comparatively high h-index in our study underscores both the maturity and the rapidly growing impact of AI in histopathology relative to other medical AI specialties. This indicates that the field is not only producing a large volume of publications but also generating sustained scholarly influence through highly cited contributions. By situating our result against benchmarks from parallel AI-driven medical disciplines, we highlight that the h-index of 112 is a strong indicator of the field's centrality, visibility, and influence in shaping the broader landscape of medical AI research.
Top-cited articles
The five top-cited papers in our corpus collectively moved the field from proof-of-concept toward regulated clinical deployment and guideline-supported practice. Several of these highly cited studies have contributed to shaping clinical practice guidelines, particularly by providing evidence that underpins the integration of artificial intelligence tools into histopathological workflows in oncology. 12 For instance, landmark studies reporting the diagnostic accuracy and reproducibility of AI-based histopathology models have been referenced in consensus statements and professional guidelines, highlighting their role in augmenting pathologist decision-making. 72 Other influential works have provided proof-of-concept evidence that facilitated subsequent regulatory approvals of AI-enabled diagnostic systems,32,73,74 reflecting their transition from research to real-world clinical deployment. Moreover, some of the top-cited contributions demonstrated prognostic utility and biomarker discovery, which have informed ongoing clinical trials and translational studies, thereby bridging the gap between computational innovation and therapeutic decision-making.75–78 The landmark studies provided the empirical foundation that helped catalyze key regulatory milestones and professional guidance for digital pathology and AI: the first FDA authorization for a WSI platform, which enabled primary diagnosis on digitized slides and opened a regulatory pathway for downstream AI tools; the first FDA De Novo authorization of an AI system in pathology, explicitly cleared to flag cancer-suspect foci on prostate biopsies in support of pathologist review; and subsequent clearances expanding AI safety-net use cases. 79
Geographic distribution
The geographic distribution in our analysis, China (29.9%), the United States (24.9%), and India (13.3%) leading output, mirrors macro-level differences in national investments, infrastructure, and AI policy for oncology diagnostics. The United States remains at the forefront due to its longstanding leadership in both biomedical research and artificial intelligence driven by institutions like Harvard, Stanford, and the National Institute of Health (NIH), which provide substantial funding and foster interdisciplinary research collaboration between different scientific disciplines.16,80 Additionally, US-based companies have significantly advanced digital pathology tools, accelerating publication output in this domain. 8 China's emergence is powered by major governmental initiatives such as Healthy China 2030 Plan, and the AI Development Plan, which promote AI integration into healthcare and fund large scale precision medicine efforts. 81 India, on the other hand, has seen rapid growth in AI-related publications due to surge in government-backed AI for healthcare programs as well as the strong collaborations between academic institutions and health-tech start-ups targeting cancer diagnostics, reflecting both a rising disease burden and the potential for scalable AI solutions in low-resource environments.82,83 While geographical and institutional analysis highlights the dominance of China, the United States, and India, it is important to acknowledge potential biases inherent in the database used. Scopus, despite being one of the largest and most comprehensive multidisciplinary databases, has comparatively broader coverage of English-language journals and those published by major international publishers. Consequently, research outputs disseminated in non-English or regionally indexed journals may be underrepresented, which could partly explain the relative visibility of countries with strong traditions of publishing in English. However, this limitation is intrinsic to bibliometric studies based on any single database and has been recognized in prior literature.84–86 We have therefore interpreted the geographical and institutional findings with this caveat in mind, and we emphasize that the dominance observed for China, the United States, and India reflects both their actual strong research capacity and, to some extent, database-related visibility patterns. Acknowledging this nuance enhances the robustness and transparency of our analysis.
Research collaboration
Global collaboration networks, especially United States–Asia ties, are well positioned to move the field from proof-of-concept to validated, regulatory-cleared tools by enabling larger, multi-ethnic training cohorts, external validation across scanners and workflows, and pragmatic clinical studies. The co-authorship/country analyses and abstract-level synthesis emphasize this maturation and the role of cross-regional data and expertise exchange; pairing such networks with transparent lifecycle practices has already produced regulatory milestones in digital pathology AI (e.g., De Novo for Paige Prostate in 2021 and recent 510(k) clearance for Ibex Prostate Detect), signaling a viable translation pathway for future models. 87
Keyword clusters
The four dominant keyword clusters we identified—breast-cancer diagnostics; methodological innovations; prognostic modeling/tumor microenvironment; and digital infrastructure/explainable AI, capture the field's breadth and its deeply interdisciplinary nature. The clusters couple disease-focused applications (e.g., breast cancer pipelines) with advances in representation learning, weak supervision, and outcome modeling, while a parallel stream tackles data standard, WSI platforms, and interpretability tooling needed for clinical deployment. Overlay analysis shows these strands increasingly intersect through multimodal learning and domain-specific explainability, underscoring how engineering progress is now tightly braided with clinical questions. In the current study, keywords mapping showed that breast cancer dominated the AI-histopathology literature. The most plausible reasons for this dominance are (i) disease burden and screening-driven slide volume, (ii) early availability of large, open benchmarks, and (iii) task fit with methods that matured first in this field. Epidemiologically, breast cancer remains among the highest-incidence cancers worldwide, ensuring abundant biopsy material and routine high-volume workflows (e.g., screening and sentinel lymph-node evaluation) that naturally produce digitized slides for training and validation. 88
Research themes and temporal development
The introduction of digital WSI in the late 2000 marked a turning point, allowing entire glass slide to be scanned at high resolution and analyzed digitally, thus catalyzing large scale computational histopathology. 89 By the mid-2010, the field witnessed a surge with the application of deep learning, particularly CNN. Ronneberger et al.'s development of the U-Net architecture in 2015 revolutionized biomedical image segmentation by enabling pixel-level annotation of complex tissue structures. 90 Meanwhile, Coudray et al. (2018) demonstrated CNNs; diagnostic power, achieving pathologist-level precision in identifying tissue types and mutations from WSIs. 65 Subsequently, the widespread adoption of transfer learning allowed pre-trained models to be fine-tuned on smaller pathology datasets, significantly lowering the barrier to entry and enhancing performance across diverse tissue types. 91 Large scale initiatives, such as CAMELYON16, provided standardized datasets and benchmarks, accelerating uptake in tasks like lymph node metastasis detection.65,92 More recently, the introduction of transformer-based architectures in 2021 further sharpened capabilities in capturing global tissue context, enabling better modeling of spatial dependencies across the whole slide.93,94 Integrative multi-modal pipelines now combine histological images with genomics and proteomic data, ushering in an era of pathomics and AI-informed precision oncology. 95
The shift from classical machine-learning pipelines to transformer-based and multimodal models has three practical implications. 96 First, accuracy: transformers expand the receptive field and capture long-range histomorphologic dependencies across gigapixel WSIs, improving slide-level classification and prognostic tasks compared with patch-local models; multimodal models integrate histology with clinical/molecular data to boost predictive power. 97 Second, interpretability: attention maps and modern MIL frameworks (e.g., CLAM) provide human-auditable evidence regions, though transformer attention remains a research-grade explanation rather than a validated clinical rationale—reinforcing the need for transparent reporting. Third, translation: agencies now expect Good ML Practice, transparency, and lifecycle change-management (e.g., predetermined change control plans) for AI devices, which favors architectures with stable behavior and auditable update.36,64,98
Trends toward semantic segmentation, self-supervised learning (SSL), and therapy-response prediction create both challenges and opportunities. Segmentation (revitalized by U-Net) improves pixel-level understanding and supports explainability but is annotation-intensive; SSL mitigates annotation burdens by pretraining on vast unlabeled WSIs and has recently produced foundation-scale pathology models, which can then be fine-tuned for prognosis or response prediction. The opportunity is clear—larger, more diverse pretraining corpora and multi-task heads can lift performance while improving robustness, yet translation depends on high-quality curation (harmonized staining/scanners), public benchmarks (e.g., PANDA), and regulatory pathways that accommodate model updates under PCCPs.
Bibliographic coupling
Journal bibliographic-coupling map stratifies the literature into (i) technical development, (ii) clinical application, and (iii) translational informatics venues, with increasing cross-linking among them, evidence that engineering advances are being pulled toward clinical workflows. This structure argues for stronger, earlier integration between algorithm development and practice environments: co-design with pathologists, multi-site external validation, and deployment studies embedded within LIS/WSI systems. Such coupling is precisely what regulators and guideline bodies emphasize when evaluating clinical utility and safety of AI tools.
Finding implications
The findings of the current study carry far-reaching implications across clinical practice, healthcare policy, and academic research. From a practical standpoint, the study underscores the transformative potential of AI in histopathological workflows and enhancing diagnostic precision, particularly in oncology, where timely and accurate tissue classification is critical for prognosis and treatment planning. 56 The rising publication trend reflects increased clinical confidence in AI's ability to process WSIs with high sensitivity and specificity, enabling pathologists to focus on complex or ambiguous cases while delegating routine pattern recognition tasks to algorithms. 54 Moreover, this bibliometric mapping identifies leading institutions and countries that can cat act as models for healthcare systems aiming to integrate digital pathology, thereby facilitating international benchmarking and policy formulation around digital infrastructure and AI governance. 99 These findings should inform hospital administrators and regulatory authorities about prioritizing AI-readiness, investing in computational resources, and training medical professionals in AI- assisted diagnostics.
On a social level, the study highlights the democratizing potential of AI in histopathology by facilitating equitable access to diagnostic expertise in underserved or rural areas that lack specialized pathologists.1,56 If guided properly, AI can bridge disparities in cancer diagnostics across regions, particularly in low-resource settings where histopathology turnaround times are prohibitively long. 100 Furthermore, the prominent inclusion of keywords such as “explainable AI” and “ethics” in recent literature clusters implies increasing attention to public concerns about algorithmic bias, transparency, and the accountability of AI in medical decisions. 101 Thus, researchers, clinicians, and developers must work collaboratively to ensure that AI systems are not only efficient but also trustworthy and ethically deployable.
In terms of research implications, the current study provides a roadmap for scholars seeking to contribute to this rapidly evolving field. By identifying underexplored intersections, such as immune-oncology integration, spatial transcriptomics, and multi-omics fusion, the study directs attention to promising yet understudied domains ripe for scientific innovation. 102 The co-occurrence and bibliographic coupling maps guide early-career researchers in identifying core journals, influential contributors, and emerging research fronts, thereby aiding strategic publication and collaboration planning. Moreover, by highlighting the dominance of certain institutions and the skewed citation patterns, this study also raises questions about the need for inclusive funding policies that support diverse geographic and institutional contributions to AI pathology scholarship.
Clinical performance and cost considerations
While bibliometrics cannot directly estimate diagnostic accuracy or cost, several highly cited translational studies in our corpus collectively indicate that AI-assisted computational pathology achieves clinical-grade performance on key WSI tasks (e.g., metastasis detection, slide-level cancer detection) and can operate at health-system scale under weak supervision.33,34,103 Such results support workflow triage in which algorithms pre-screen high-volume tasks and flag suspicious foci for expedited review, a pattern that is associated with reduced turnaround times, greater inter-observer consistency, and reallocation of expert time to complex cases.32,104 In practice, these effects underpin cost-effectiveness mechanisms (labor efficiency, avoided repeat work, throughput gains) that warrant prospective, multi-site economic evaluations.105–107 Our field-level maps, in particular the surge in segmentation, self-supervised pretraining, and multimodal integration, point to continued improvements in performance, robustness, and explainability, all prerequisites for sustainable clinical adoption and regulatory alignment.
Challenges and future research
Despite the rapid advancement of AI technologies in digital pathology, several challenges continue to hinder their full-scale integration into clinical practice. One of the foremost issues is the lack of standardization in slide preparation, staining protocols, and image acquisition across institutions, which affects the generalizability and robustness of AI models. Moreover, many models are trained on single-center or small-scale datasets, limiting their external validity when applied to diverse populations. Another major concern is the black-box nature of deep learning algorithms; the absence of explainability undermines clinician trust and complicates regulatory approval. Additionally, AI-enabled histopathology also raises salient ethical and legal barriers that must be addressed for responsible translation. The secondary use of WSIs for model development requires a clear legal basis and, where applicable, patient authorization; even when direct identifiers are removed, WSIs can retain protected health information (e.g., on slide labels) and may carry re-identification risks, necessitating robust de-identification, label cropping, and privacy-preserving data governance.108,109 Accountability for AI-assisted diagnostic errors is an evolving policy arena: in the European Union, clinical AI used for diagnosis falls under the AI Act as a high-risk system, which assigns concrete obligations to providers and deployers (risk management, high-quality data, human oversight, logging, post-market monitoring) that operationalize responsibility and support auditability in the event of misdiagnosis.110,111 The scarcity of prospective, multi-center clinical trials evaluating AI performance against or alongside pathologists also limits the evidence base necessary for institutional and regulatory acceptance. Lastly, computational infrastructure requirements and lack of integration into laboratory information systems present practical hurdles, especially in resource-limited settings
Future research in AI-driven histopathology should prioritize the development of explainable AI (XAI) frameworks to foster clinical trust and support regulatory compliance. There is also a pressing need for federated learning approaches that allow training across decentralized, privacy-protected datasets, thereby improving model generalizability while safeguarding patient confidentiality. Researchers should focus on multimodal integration, combining histopathological features with genomic, transcriptomic, and radiological data to enhance diagnostic and prognostic accuracy. In parallel, efforts must be directed toward benchmarking and validation, with large-scale, prospective, and multi-institutional studies to assess real–world effectiveness and cost-efficiency. Furthermore, creating open-access annotated datasets and AI development toolkits can democratize research and foster reproducibility, Finally, future investigations should focus on learning models capable of evolving with new data and clinical feedback, enabling continuous improvement of diagnostic performance over time.
Clinical practice
To translate computational pathology from promising prototypes to routine care, we identify three actionable infrastructure gaps. (1) Public, well-curated WSI benchmarks need sustained funding and expansion beyond CAMELYON/PANDA/PAIP to include diverse scanners, staining protocols, and under-represented populations, with long-term stewardship rather than ad-hoc projects.32,104,112,113 (2) Federated/decentralized learning at scale requires funded networks of secure ‘edge’ training nodes (audited logging, privacy-preserving compute, shared SDKs) and subsidized access to cloud and orchestration services so multi-institution model training becomes practical outside single-center studies.114,115 (3) Standards-first tooling and compliance should be resourced to accelerate clinical adoption: continued uptake of DICOM-WSI and CAP-aligned validation workflows; maintenance of open-source stacks used in labs (OpenSlide, QuPath, Digital Slide Archive/Histomics UI, MONAI); and interoperable governance aligned with FAIR/GA4GH to ease regulatory readiness (e.g., EU AI Act). Finally, privacy deserves targeted investment, de-identification methods for gigapixel WSIs and secure, brokered viewing layers, to enable data sharing that is ‘as open as possible, as closed as necessary.116–119
The adoption of AI in histopathology requires coordinated efforts at the managerial and institutional levels. Healthcare administrators must invest in digital infrastructure, including high-resolution scanners, secure data storage, and integrated AI platforms compatible with laboratory information systems. Institutional leaders should establish interdisciplinary teams that bridge pathology, data science, and clinical informatics to facilitate the development and validation of AI tools. Policies must be created to support ethical AI deployment, including guidelines for data sharing, informed consent, and accountability. Training programs should be introduced to enhance the digital literacy of pathologists and laboratory technicians, ensuring smooth human-AI collaboration. Furthermore, academic and research institutions should prioritize collaborative grant funding and international partnerships to accelerate innovation and address global disparities in AI access. From a strategic standpoint, emphasizing value-based care models that integrate AI performance metrics into diagnostic quality assessments will further align institutional goals with technological advancement.
Limitations
A few certain structural features of this bibliometric design might constrain interpretation. Most fundamentally, we relied on a single data source (Scopus): although its breadth, metadata quality, and citation tracking motivated this choice, a Scopus-only corpus can omit items published in un-indexed databases or in emerging repositories and thus shift counts, networks, and topic boundaries. Second, excluding reviews, conference proceedings, preprints, and “in-press” items likely underrepresent very early innovations and the newest outputs; future updates could include a narrative sensitivity scan of major preprint servers and flagship conferences to confirm that cluster structure and leading themes remain stable when such sources are considered. Third, the search was executed in TITLE-ABS fields using a finite, hand-crafted keyword set with explicit radiology-related “NOT” filters; even with two-tier validation (curated benchmark re-call and a manual scan of the 100 most recent 2024 records), false negatives, topical drift, and synonym variability remain possible, limitations intrinsic to keyword-based methods. Fourth, our analyses draw solely on exported bibliographic metadata (titles, authors/affiliations, journals, keywords, citation counts); we did not analyze full texts, funding statements, or clinical outcomes, and any disambiguation errors in affiliations/author identifiers in the source can propagate into productivity and collaboration maps. Fifth, all maps (co-authorship, keyword co-occurrence/overlay, and bibliographic coupling) were generated in VOSviewer; such visualizations are sensitive to parameter choices (e.g., thresholds, counting methods) and should be read as indicative structures rather than definitive taxonomies. Sixth, citation-based indicators reported here (e.g., total citations and H-index) are susceptible to time-lag and field-size effects that privilege older, highly cited works and larger communities, potentially down-weighting cutting-edge contributions at the analysis time. Finally, as a bibliometric mapping that involves no patients, clinical datasets, or prospective evaluations, the study can describe who publishes, collaborates, and cites whom, but it cannot establish diagnostic accuracy, clinical effectiveness, or cost-effectiveness of any AI system in practice.
Conclusions
This bibliometric study provides a panoramic view of the evolving field of AI-powered histopathology slide image interpretation in oncology, highlighting its remarkable growth, shifting research themes, and key contributors. The exponential rise in publications since 2015 underscores the increasing clinical and scientific interest in leveraging AI, particularly deep learning and CNN, for automated, accurate, and scalable cancer diagnostics. This shift is catalyzed by advancements in digital pathology infrastructure, increasing computational capabilities, and the availability of large annotated datasets. The findings demonstrate how AI technologies are not only augmenting but, in some cases, rivaling traditional diagnostic workflows. Seminal works have shown that AI can predict genomic mutations, identify tumor subtypes, and classify cancer grades from histological slides with precision comparable to human experts. These achievements are paving the way for precision oncology, where AI-powered tools could support pathologists in rendering quicker and more consistent diagnoses, particularly in high-volume or resource-constrained settings. The bibliometric landscape is characterized by growing interdisciplinary collaboration-with engineering, computer science, pathology, and clinical oncology converging to create real-world applications. Countries like China and India are rapidly advancing due to strategic investments in AI infrastructure and public health digitization, while institutions in the United States and Europe continue to lead in translational and clinical-grade AI research. Thematic analysis confirms a temporal shift from early efforts in image classification to more sophisticated models addressing segmentation, prognosis, treatment response prediction, and explainability. The emergence of transformers, self-supervised learning, and multi-modal AI signifies a move towards flexible, generalizable, and clinically explainable models- a necessary evolution for regulatory approval and physician trust. From a policy and clinical perspective, this study emphasized the need for standardization, clinical validation, and ethical deployment. Hospitals, health systems, and governments must proactively invest in digital pathology infrastructure, train personnel, and enact regulatory frameworks that ensure safe and equitable Ai adoption.
In conclusion, this study serves not only as a knowledge map of current research but as a strategic guide for stakeholders seeking to advance AI in oncology diagnostics. The next frontier lies in fostering explainable, multimodal, and globally inclusive AI solutions that transcend technological novelty to deliver genuine improvements in patient care and health equity.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251393286 - Supplemental material for AI-Powered histopathology slide image interpretation in oncology: A comprehensive knowledge mapping and bibliometric analysis
Supplemental material, sj-docx-1-dhj-10.1177_20552076251393286 for AI-Powered histopathology slide image interpretation in oncology: A comprehensive knowledge mapping and bibliometric analysis by Moutaz W Sweileh in DIGITAL HEALTH
Supplemental Material
sj-docx-2-dhj-10.1177_20552076251393286 - Supplemental material for AI-Powered histopathology slide image interpretation in oncology: A comprehensive knowledge mapping and bibliometric analysis
Supplemental material, sj-docx-2-dhj-10.1177_20552076251393286 for AI-Powered histopathology slide image interpretation in oncology: A comprehensive knowledge mapping and bibliometric analysis by Moutaz W Sweileh in DIGITAL HEALTH
Supplemental Material
sj-pptx-3-dhj-10.1177_20552076251393286 - Supplemental material for AI-Powered histopathology slide image interpretation in oncology: A comprehensive knowledge mapping and bibliometric analysis
Supplemental material, sj-pptx-3-dhj-10.1177_20552076251393286 for AI-Powered histopathology slide image interpretation in oncology: A comprehensive knowledge mapping and bibliometric analysis by Moutaz W Sweileh in DIGITAL HEALTH
Footnotes
Acknowledgements
The author acknowledges the use of AI in language editing only. No results or data were obtained or generated by any type of AI.
Ethics approval
The current manuscript doesn’t involve humans or parts of humans or animals and no IRB approval was required
Funding
The author received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
All data are available through Scopus using the detailed search query
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
