Abstract
Background
Neonatal early-onset sepsis (EOS), occurring within 72 h of birth, is a major cause of neonatal morbidity and mortality worldwide. The urgent need for rapid, accurate diagnosis is underscored by the condition's severity. Current diagnostic methods are hampered by non-specific clinical signs, leading to underdiagnosis or overtreatment. This highlights a crucial gap in neonatal care.
Methods
This retrospective study analyzed data from 1613 full-term pregnant women at a single center in Shenzhen, China (2022), including 69 EOS cases. Ten machine learning algorithms (e.g. Logistic Regression, Random Forest, XGBoost) were developed using maternal prenatal predictors. Data preprocessing involved imputation, standardization, and Lasso feature selection. Models were evaluated using 5-fold cross-validation, and the SMOTE technique was applied to address class imbalance.
Results
Among ten machine learning models, XGBoost and Random Forest demonstrated the highest discriminative ability (AUC=0.87). While a default threshold yielded low sensitivity, a threshold optimized for a clinical screening objective (0.04) achieved a sensitivity of 92.8% and a specificity of 73.1%. Key predictors identified included maternal temperature and inflammatory markers.
Conclusion
This study demonstrates that machine learning models based on maternal factors have the potential to serve as high-sensitivity screening tools for EOS. Tuning the decision threshold is a critical step to maximize clinical utility, which involves a necessary trade-off between sensitivity and specificity. This approach provides a framework for developing and evaluating clinically-oriented prediction models to improve neonatal care.
Introduction
Neonatal sepsis is a leading cause of neonatal morbidity and mortality, ranking third among the global causes of neonatal deaths, following prematurity and intrapartum-related complications. 1 Defined as a systemic infection occurring in infants less than 28 days old, neonatal sepsis manifests with symptoms such as respiratory distress, temperature instability, and lethargy. 2 It is categorized into early-onset sepsis (EOS) and late-onset sepsis based on the time of onset. EOS, which occurs within the first 72 h of life, is predominantly caused by pathogens acquired from the mother during delivery. 3 According to the 1990–2013 Global Burden of Disease Study, approximately 2.6 million neonates die annually, with three-quarters of these deaths occurring in the first week of life, predominantly in low- and middle-income countries. 4 In 2018, neonatal sepsis accounted for 15% of the 2.5 million neonatal deaths worldwide, with 42% of these deaths occurring within the first week. 5 For EOS, early diagnosis is crucial as outcomes are heavily dependent on the promptness of antibiotic treatment. 6 Current guidelines emphasize the importance of early identification, appropriate treatment, and minimizing unnecessary antibiotic use. 7 Despite advancements in neonatal care, EOS remains a significant challenge due to its rapid progression and high fatality rates. The importance of early detection and treatment cannot be overstated, as delayed diagnosis often results in poorer outcomes for affected neonates. 8
Diagnosing and predicting EOS are particularly challenging due to the non-specific clinical signs and symptoms, such as fever, lethargy, and respiratory distress, which overlap with other neonatal conditions. Current diagnostic methods, including blood cultures, are considered the gold standard but have notable limitations, such as low sensitivity, delayed results, and an invasive nature. Furthermore, the absence of standardized diagnostic criteria complicates timely and accurate diagnosis. These challenges underscore the urgent need for reliable, non-invasive, and rapid risk prediction models to assist clinicians in the early identification and intervention of EOS.
The emergence of artificial intelligence and machine learning (ML) offers promising avenues for enhancing the prediction and diagnosis of EOS. Various studies have demonstrated the potential of ML algorithms to improve diagnostic accuracy and efficiency. For example, Martin utilized random forest algorithms to develop a predictive model for EOS, revealing that biomarkers like C-reactive protein and white blood cell count significantly enhance prediction accuracy. 9 Samuel developed a clinical prediction model using multivariable logistic regression tailored for resource-limited settings, which included predictors such as maternal fever, foul-smelling amniotic fluid, prolonged rupture of membranes, neonatal temperature, respiratory rate, activity, chest retractions, and grunting. This model achieved an ROC of 0.74 but had high sensitivity and low specificity, limiting its utility in early diagnosis. 10 Fernando constructed a neural network model that identified key maternal and neonatal predictors for EOS, achieving an area under the curve (AUC) of 92.5%. However, existing models often lack external validation, are single-centered, and have limited generalizability. 11
Current research on EOS suggests that certain factors during maternal pregnancy and delivery may serve as high-risk factors and predictive indicators for neonatal EOS. Most existing studies focus on the postnatal period, emphasizing the progression and prognosis of neonatal sepsis, while fewer studies address the rapid early diagnosis of EOS, particularly those suitable for prenatal application.
In this study, we propose a novel ML-based approach to predict EOS risk using a comprehensive dataset from prenatal examinations. Our methodology involves rigorous data preprocessing, feature selection using Least Absolute Shrinkage and Selection Operator (Lasso) regression, and the application of ten different ML algorithms, including eXtreme Gradient Boosting (XGBoost), Support Vector Machine (SVM), and Random Forest (RF). To address the common issue of class imbalance in medical datasets, we employed the Synthetic Minority Over-sampling Technique (SMOTE), a resampling strategy that generates synthetic samples for the minority class. Additionally, SHapley Additive exPlanations (SHAP) analysis was integrated to interpret model decisions and identify key predictive features, enhancing the clinical interpretability of our findings.
The innovation of our approach lies in addressing class imbalance with SMOTE, ensuring model robustness through extensive cross-validation, and providing actionable insights into feature importance with SHAP analysis. This study aims to establish a reliable and accurate risk prediction model for EOS, thereby improving early diagnosis and clinical outcomes for neonates. The workflow of this study is illustrated in Figure 1.
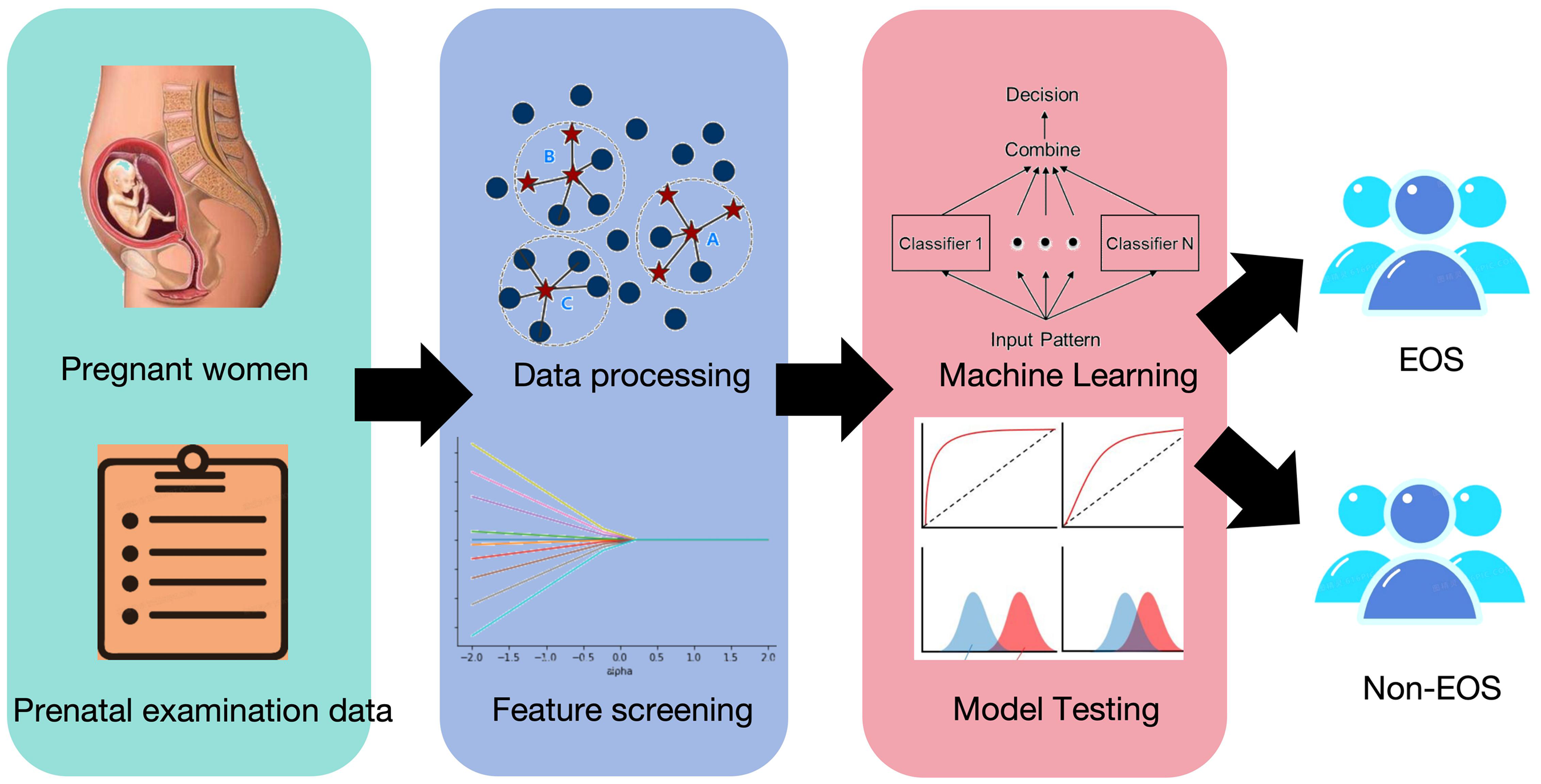
Workflow diagram of this study.
While existing studies have developed predictive models for neonatal EOS, they often focus on preterm neonates or rely on single predictive algorithms, which limits their generalizability and accuracy. Furthermore, few studies address the specific challenges associated with EOS prediction in term neonates, a population with unique risk factors and clinical presentations. Our study bridges this gap by:
Developing and comparing ten ML algorithms to identify the most robust and accurate model for EOS prediction. Utilizing the SMOTE technique to address class imbalance, a common limitation in sepsis datasets. Focusing exclusively on predicting EOS in newborns born to full-term mothers undergoing a trial of labor.
Methods
Patients
Study design and population
This retrospective study was conducted using prenatal examination data from 1613 full-term pregnant women who attempted trial labor, collected at the Baoan Women's and Children's Hospital in Shenzhen, China, from January 2022 to December 2022. Among these women, 69 neonates were diagnosed with EOS. The inclusion criteria included all full-term pregnant women who attempted trial labor at our hospital during the study period, with complete prenatal examination, intrapartum records, and delivery information. Exclusion criteria were as follows: (a) incomplete medical records, (b) fetal malformations, chromosomal abnormalities, (c) pregnancy complications such as chronic infectious diseases, heart disease, and immune system diseases. This study was approved by the Ethics Committee of Baoan Women's and Children's Hospital (ethics number: LLSC-2024-01-04-33-KS). Informed consent was waived for this retrospective analysis, as all data were anonymized and the study posed no risk to the participants. The study was conducted in accordance with the principles outlined in the Declaration of Helsinki.
Data collection
All data used in this study were extracted retrospectively from the hospital information system (HIS) at Baoan Women's and Children's Hospital, Shenzhen, China. The HIS database integrates electronic medical records, laboratory test results, and clinical observations. Data were systematically extracted using predefined inclusion and exclusion criteria, focusing on maternal prenatal, perinatal, and neonatal clinical records. To ensure data accuracy and consistency, clinical variables such as maternal CRP and PCT levels, neonatal clinical manifestations (e.g. respiratory distress, lethargy), and laboratory findings were cross-verified by two independent clinicians. Key variables, including diagnosis and treatment timelines, were manually reviewed to confirm their alignment with the established criteria for EOS.
The study model included 36 features, encompassing maternal demographic characteristics, prenatal clinical indicators, and intrapartum factors. Key variables and their definitions are as follows:
Vaginal examination number: Total number of vaginal examinations performed during labor, recorded as an integer value from clinical documentation, which was treated as a continuous variable to capture incremental risk. Maternal temperature: Maternal temperature was measured as part of routine clinical assessments at two time points: upon admission and at delivery. Temperature was recorded in degrees Celsius and treated as a continuous variable in the feature selection and modeling process. Elevated maternal temperature (≥38°C) was noted as a clinically significant marker of fever and used for additional clinical interpretation. These measurements provided insights into maternal inflammatory responses during labor. Induction method: Mode of labor induction, categorized as mechanical (e.g. Foley catheter) or pharmacological (e.g. oxytocin or prostaglandins). Labor duration: Labor duration in this study specifically defined as the time from cervical dilation >3 cm to full cervical dilation (10 cm), consistent with established Chinese Obstetric and Gynecology. For modeling purposes, labor duration was treated as a continuous variable. Pregnancy BMI: Maternal body mass index (BMI) calculated as weight (kg) divided by height squared (m2), measured at the end of pregnancy. Pre-pregnancy BMI: Pre-pregnancy BMI was calculated as weight (kg) divided by height squared (m2) and recorded during the first prenatal visit. Maternal BMI at delivery was calculated using weight measured immediately prior to delivery. For feature selection and modeling, BMI was treated as a continuous variable. For clinical interpretation, thresholds of ≥30 kg/m2 (indicating obesity) were applied based on WHO criteria. Painless to fever time: Time interval (in minutes) between the initiation of epidural analgesia and the onset of maternal fever. HBV status: Binary variable indicating maternal hepatitis B virus infection (positive or negative), based on routine prenatal screening results. Gravidity and parity: Gravidity refers to the total number of pregnancies a woman has had, while parity indicates the number of pregnancies carried to viable gestational age, recorded as integer values. GBS status: Binary variable indicating the presence or absence of Group B Streptococcus colonization, determined through third-trimester vaginal and rectal swabs. PROM (Premature Rupture of Membranes): A binary variable indicating whether premature rupture of membranes occurred (Yes/No), defined as the rupture of the amniotic sac before the onset of labor. PROM is associated with an increased risk of neonatal EOS. CRP and PCT levels: Serum levels of C-reactive protein (CRP, mg/L) and procalcitonin (PCT, ng/mL) measured at two time points: maternal admission and in labor, which were used to indicating the presence of systemic inflammation or bacterial infection. The thresholds for CRP and PCT are set at ≥10 mg/L and ≥0.5 ng/mL, respectively.
Other variables included maternal age, hypertension status, diabetes status, anemia, thyroid dysfunction, and so on. These features were chosen based on their clinical relevance and availability in the dataset.
Definition
The diagnosis of neonatal EOS was based on the expert consensus for the diagnosis and treatment of neonatal sepsis. The diagnosis of neonatal EOS required the presence of clinical manifestations along with a positive blood culture or positive culture from cerebrospinal fluid (CSF) or other sterile cavity fluids. Blood culture was used as the gold standard for confirming EOS, as it remains the definitive method for identifying causative pathogens. However, blood culture negativity does not exclude the diagnosis of EOS. Therefore, clinical diagnosis of EOS in this study also required clinical abnormal manifestations combined with meeting any of the following criteria: (a) Two or more positive results in non-specific blood tests (e.g. elevated C-reactive protein, procalcitonin, or white blood cell abnormalities). (b) CSF examination showing changes consistent with purulent meningitis. (c) Detection of pathogenic bacterial DNA in blood or CSF samples. Based on these diagnostic criteria for EOS, neonates were divided into the EOS and non-sepsis group. The diagnosis of neonatal EOS was based on a combination of clinical manifestations, laboratory findings, and microbiological results, as recorded in the HIS database. EOS cases included both culture-positive and culture-negative (biomarker-positive) sepsis to capture the broader spectrum of clinical presentations. Among the 69 EOS cases, 75% were culture-positive (51/69).
Data labeling was performed retrospectively by experienced clinicians who reviewed the electronic records to identify cases meeting the following criteria:
Clinical manifestations: Clinical signs prompting sepsis screening included respiratory distress (e.g. tachypnea, nasal flaring), lethargy, fever (≥38°C), hypotension, and feeding intolerance. These signs were recorded by attending neonatologists during routine clinical assessments. Laboratory findings: Data on maternal and neonatal CRP, PCT, and other inflammatory markers were extracted from HIS laboratory modules. Thresholds for CRP (≥10 mg/L) and PCT (≥0.5 ng/mL) were applied based on neonatal diagnostic guidelines. Microbiological results: Blood and CSF culture results recorded in the HIS were used to confirm EOS cases.
Data preprocessing
During the model construction process, data cleaning and noise reduction were first performed. The proportion of missing values was calculated for each feature, and features with more than 30% missing data were excluded. For features with less than 30% missing data, mean imputation was used. Next, data standardization was conducted using the StandardScaler method, which calculated the mean and standard deviation of the training data and applied these parameters to both the training and testing datasets to ensure consistent feature scaling. To address class imbalance, the SMOTE method was applied to the training data, generating new samples for the minority class to improve the model's ability to recognize EOS. Finally, the target variable was converted into a one-dimensional array to ensure compatibility with the model training functions.
Feature selection
Feature selection was conducted using Lasso regression (Least Absolute Shrinkage and Selection Operator) in combination with cross-validation (LassoCV). The Lasso algorithm employs an L1 regularization term, which effectively mitigates overfitting by prioritizing the most relevant features while shrinking less important ones toward zero. LassoCV automatically adjusts the regularization parameter (\lambda) and uses cross-validation to assess model performance across different parameter settings, ensuring the model's generalizability and the stability of the selected features. To guarantee that the algorithm had ample opportunity to converge on the optimal solution, we set the maximum number of iterations to 10,000.
All ten ML models evaluated in this study—such as XGBoost (eXtreme Gradient Boosting), SVM, and Random Forest—were trained using the feature set selected by Lasso regression. This approach ensured that only the most informative predictors were included in the models, reducing dimensionality and enhancing performance while maintaining interpretability. By limiting the feature set, we minimized the risk of overfitting and improved computational efficiency during model training and evaluation.
Our study included 1613 participants with 69 EOS events. The final predictive model was developed using a set of 36 predictors identified through LASSO regression. We acknowledge that the resulting events per variable (EPV) ratio is approximately 1.9, which is below the traditional guideline of 10 often recommended for conventional logistic regression models to avoid overfitting. However, it is recognized that this guideline is less critical for modern ML methods, particularly those that employ regularization techniques like LASSO. Regularization inherently penalizes model complexity and reduces the risk of overfitting, even with lower EPV ratios. The robustness of our findings is further supported by our rigorous internal validation strategy, which includes both 5-fold cross-validation and a 200-iteration bootstrap optimism-adjustment, confirming that the model's performance is reliable and not substantially over-estimated.
Machine learning model construction
In recent years, the application of ML in predicting neonatal sepsis has made significant progress. Masino et al. developed models using random forests and gradient boosting machines based on electronic health records from NICUs. Although these models demonstrated high predictive accuracy, they faced challenges such as potential overfitting and a lack of external validation. 12 Li et al. utilized LASSO and SVM-RFE to identify biomarkers from gene expression data, constructing robust predictive models. However, these methods typically require large sample sizes and complex preprocessing steps. 13 Mani et al. compared various ML methods, such as SVM, Naive Bayes, and decision trees. While these techniques have advantages in early diagnosis, they pose challenges in terms of model interpretability and computational resource demands. 14 Barghi assessed six ML algorithms, among which the Boosted Tree algorithm excelled in handling imbalanced data, but it required optimization to avoid overfitting and long training times. 15
In this study, we employed 10 different ML algorithms, including AdaBoost, Logistic Regression, KNN, SVM, Random Forest, Decision Tree, Gradient Boosting, LDA, Naive Bayes, and XGBoost, to compare and select the best predictive model. These algorithms were chosen based on their performance on various datasets and their adaptability to different problems. For instance, XGBoost stands out for its ability to handle high-dimensional data and prevent overfitting, while Random Forest and Gradient Boosting are known for their robustness and good generalization ability. AdaBoost and Logistic Regression were included for their simplicity, ease of use, and low computational cost. While advanced models like LSTM and ElasticNet have shown promise in neonatal sepsis studies, they were not included in this study for several reasons. LSTM, a deep learning model, requires large datasets and intensive computational resources, which may not align with our dataset's size and retrospective design. ElasticNet, although useful for feature selection, often necessitates extensive preprocessing and is less robust in imbalanced datasets compared to ensemble methods like XGBoost. Future studies incorporating larger datasets may explore these models to evaluate their comparative performance. This approach ensures a balance between predictive performance and clinical applicability, enabling the identification of the most suitable model for EOS prediction in term neonates.
Regarding model parameter settings, key parameters for the XGBoost model included a learning rate of 0.1, a maximum tree depth of 6, a minimum child weight of 1, and 100 boosting rounds. The Random Forest model was configured with 100 trees and a maximum depth of 60 to ensure sufficient data fitting. For the SVM model, we used an RBF kernel and employed cross-validation to determine the optimal regularization parameter C and kernel parameter γ. The other models were also fine-tuned according to the characteristics of the data to improve prediction accuracy and robustness. Additionally, we incorporated SHAP analysis to interpret the model's decision-making process, helping to identify the clinical features that play key roles in EOS risk prediction, thereby enhancing the model's transparency and interpretability.
Model evaluation
The performance of the predictive models was comprehensively evaluated using four key metrics: accuracy (Acc), the area under the receiver operating characteristic curve (AUC), calibration curves, and decision curve analysis (DCA). AUC assessed each model's ability to distinguish between EOS and non-EOS cases, with higher values indicating better discrimination. Calibration curves evaluated the accuracy of predicted probabilities against actual outcomes, with a well-calibrated model closely following the diagonal line. DCA measured the clinical utility of the models by calculating the net benefit across various probability thresholds, helping to determine the practical value of each model in a clinical setting. These metrics were applied to all models to identify the best-performing algorithm for predicting neonatal EOS. To facilitate clinical applicability, a risk cut-off threshold of 0.5 was applied for the algorithm's predicted probabilities. Predictions with a probability of ≥0.5 were classified as high risk, while probabilities <0.5 were considered low risk. This threshold was chosen based on standard ML practice and the need for binary classification in clinical decision-making. Given the severe class imbalance in our dataset and the critical clinical need to minimize false negatives for EOS, the default classification threshold of 0.5 is suboptimal for screening purposes. Therefore, we performed a post-hoc threshold-moving analysis on the cross-validated probabilities generated by the model. We evaluated a range of thresholds from 0.01 to 0.99 to assess the trade-off between sensitivity and specificity. A new threshold was selected to achieve a clinically acceptable sensitivity of approximately 90%, balancing the need for high detection rates against the increase in false positives. All reported performance metrics, including the confusion matrix, were re-calculated based on this optimized threshold. To further assess model generalizability and correct for potential performance overestimation (“optimism”), we conducted a bootstrap internal validation procedure with 200 iterations to calculate an optimism-corrected estimate of the AUC.
Experimental setup
The experiments utilized a five-fold cross-validation approach to evaluate model performance, where the dataset was divided into five parts, each serving as a test set once, with the remaining parts used for training. This method ensured robust and generalizable results, with performance metrics averaged across the folds. Model parameters were optimized using grid search and random search methods to prevent overfitting. The computational environment included an NVIDIA GeForce RTX 3080 GPU and an Intel Core i9-11900K CPU, running on Ubuntu 20.04 LTS. Models were implemented using Python 3.8 with libraries such as scikit-learn 0.24.2 and XGBoost 1.4.2, providing the necessary computational power for high-dimensional data processing and intensive model training.
Results
Maternal and neonatal characteristics in the EOS group
A total of 69 cases of EOS were analyzed. The maternal mean age was 31.04 ± 3.03 years, with a mean pre-pregnancy BMI of 21.87 ± 3.11 kg/m2, which increased to 27.78 ± 3.35 kg/m2 during pregnancy. Among the deliveries, 44.93% were vaginal, 39.19% were cesarean sections, and 15.94% were instrumental deliveries. Induction of labor was performed in 49.28% of cases, predominantly using oxytocin (17.39%) and prostaglandins (8.70%). Pregnancy complications included gestational diabetes mellitus (27.54%), gestational hypertension (17.39%), and premature rupture of membranes (23.19%).
Neonatal data revealed a mean birth weight of 3377.1 ± 434.33 g, with 56.52% of neonates being male. Most neonates were born between 39 and 40 weeks of gestation (31.88% and 49.28%, respectively). Low Apgar scores were observed in 5.80% of neonates (≤7 at 1 or 5 min) and in 1.45% of neonates (≤3 at 1 min or ≤5 at 5 min). Common clinical symptoms upon NICU admission included:
Fever: Observed in 40 cases. Respiratory distress: Reported in 34 cases, characterized by tachypnea and retractions. Lethargy: Documented in 18 cases, indicative of systemic infection. Hypotension: Diagnosed in 17 cases, suggesting compromised hemodynamic stability. Feeding intolerance: Noted in 45 cases, highlighting potential systemic or gastrointestinal complications. Laboratory findings: CRP levels were present in 89.86% of cases, and PCT levels were elevated in 85.51%.
The baseline maternal and neonatal characteristics for both the EOS and non-EOS groups are detailed in Table 1. Statistically significant differences between the two groups were observed for several key characteristics. Notably, mothers in the EOS group had significantly higher parity, weight at delivery, and pre-pregnancy/pregnancy BMI. Furthermore, the EOS group had significantly higher rates of complications, including hypertension, antiphospholipid syndrome, and anemia. For neonates, there were significant differences in the distributions of mode of delivery, gestational age at birth, and APGAR scores (all p < 0.01).
Maternal and neonatal characteristics.

Table 1 provides detailed maternal and neonatal characteristics.
Feature selection results
During the feature selection stage, we identified key predictors based on the coefficients derived from training the Lasso model. Features with non-zero coefficients were retained, and by ranking these coefficients by their absolute values, we pinpointed the 36 most influential features that significantly impacted the model's performance (Figure 2 shows the visualization of the feature screening process and the arrangement of important features). These features encompassed a range of prenatal and intrapartum clinical indicators, including peak temperature, number of vaginal examinations, parity, total labor duration, admission and recheck CRP levels at delivery, induction method, pregnancy BMI, recheck CRP and PCT levels at delivery, admission PCT, admission Lymphocytes, admission neutrophils, recheck neutrophils at delivery, recheck lymphocytes at delivery, recheck neutrophils lymphocytes at delivery, PROM (premature rupture of membranes), thyroid dysfunction, diabetes, painless to fever time, fever to delivery time, cervical dilation fever, fever duration, HBV, Gravidity, GBS, ICP, Prepregnancy BMI, painless delivery, hypertension, weight, admission temperature, amniotic fluid, pregnancy methods, age, antibiotics, uterine scar, anemia, and membrane-to-delivery time. A larger Lasso coefficient indicated a greater contribution of the feature to the model.

Visualization of LASSO training iterations (A and B) and the distribution of selected feature importance rankings (C).
Among these, peak temperatures emerged as one of the strongest predictors of EOS risk, highlighting the critical role of febrile responses in the early identification of sepsis. The occurrence of PROM was also a significant predictor, as it increases the risk of infection and, consequently, the likelihood of EOS. CRP levels at delivery were vital indicators, elevated CRP levels (mean: 17.02 mg/L, median: 15.2 mg/L) with a threshold of ≥10 mg/L suggested an active inflammatory response, which is crucial in detecting EOS. Similarly, elevated PCT levels (mean: 0.74 ng/mL, median: 0.05 ng/mL) with a threshold of ≥0.05 ng/mL were highly relevant for predicting EOS, as they serve as reliable markers of bacterial infection. These findings underscore the importance of CRP and PCT as sensitive indicators for EOS risk. Lastly, pregnancy BMI was another key feature, with higher BMI values correlating with an increased risk of EOS, possibly due to complications related to maternal health.
Model evaluation results
Based on the ROC curve results shown in Figure 3(A) and the accuracy results presented in Table 2, the XGBoost and Random Forest models stand out as the best-performing models with an AUC of 0.87 and 0.86, respectively. Both models demonstrate strong discriminative power, indicating their effectiveness in identifying EOS cases. Gradient Boosting also performed well with an AUC of 0.85, highlighting its robustness. Furthermore, a bootstrap validation was performed to provide a more conservative estimate of performance. The apparent AUC on the full dataset was 1.000. After 200 bootstrap iterations, the average optimism was calculated to be 0.022. This resulted in an optimism-adjusted AUC of 0.978. This result suggests that the model's performance is robust and the estimate from 5-fold cross-validation is reliable and not substantially inflated by optimism.

ROC curves and corresponding AUC values of machine learning prediction models (A), calibration curves (B), and DCA curves (C) of the best performing models (XGBoost).
Comparison of verification accuracy results of various machine learning models.

However, the AUC only measures discriminative ability across all thresholds. For clinical application, a specific decision threshold must be chosen. As reported in our initial analysis and consistent with the challenges of imbalanced data, applying a default 0.5 classification threshold to the model's output resulted in a clinically insufficient sensitivity of 5.9%. This highlighted the need for a more clinically-oriented evaluation approach to develop a useful screening tool.
To address this, we performed the threshold-moving analysis detailed in the methods section. By selecting an optimized threshold of 0.04 to meet a clinical screening target of high sensitivity, we achieved a powerful set of performance metrics. At this new cutoff, the model obtained a sensitivity of 92.8%, a specificity of 73.1%, a positive predictive value (PPV) of 13.4%, and an extremely high negative predictive value (NPV) of 99.6%. These results demonstrate that the model can be tuned to effectively identify the vast majority of EOS cases, making it a viable candidate for a screening tool. The updated confusion matrix reflecting this performance is presented in Supplemental Figure 5.
Visualization and clinical interpretability analysis
The SHAP analysis offered valuable insights into the clinical significance of various features influencing the XGBoost model's predictions for EOS. As highlighted in the SHAP summary plot in Figure 4, temperature-related factors, such as peak and admission temperatures, showed a strong positive impact on EOS risk. This finding underscores the critical role of elevated temperatures as predictors, highlighting the need for vigilant temperature monitoring in newborns. Inflammatory markers like CRP and PCT levels at delivery also emerged as key contributors, indicating the body's inflammatory response, which is essential for diagnosing EOS. Furthermore, labor and delivery factors, including PROM, total labor duration, and the number of vaginal examinations, were found to significantly influence EOS risk, emphasizing the importance of careful management during these processes to mitigate risks. The SHAP analysis not only reinforces the robustness of the XGBoost model in predicting EOS but also provides actionable insights for clinicians, aiding clinicians in understanding the key maternal drivers of EOS risk. This interpretability, combined with the high sensitivity achieved at an optimized threshold, forms a strong foundation for developing a trustworthy clinical screening tool, pending future external validation.

SHAP analysis visualization of XGBoost.
Comparison of SHAP and LASSO-selected features
Feature selection using Lasso regression identified 36 key predictors of neonatal EOS, including maternal CRP, peak temperature, PROM, total labor duration, and amniotic fluid characteristics. These features were selected based on their Lasso coefficients, which reflect their linear contribution to EOS risk.
To interpret the XGBoost model, SHAP analysis was employed to assess feature importance within a non-linear framework. Among the top 10 features identified by SHAP, 5 overlapped with the features selected by Lasso. These included peak temperature, PROM, total labor duration, vaginal examination number, and pregnancy BMI, all of which were strongly associated with EOS risk. However, SHAP analysis also highlighted recheck CRP and PCT at delivery, which was not selected by Lasso. This suggests that SHAP is capable of capturing non-linear interactions and feature dependencies that may not be evident in Lasso's linear constraints.
The overlap between SHAP and Lasso demonstrates the robustness of certain key predictors, while the additional features identified by SHAP highlight the nuanced contributions of less obvious predictors. These results underscore the complementary nature of the two methods: Lasso ensures dimensionality reduction and model stability, whereas SHAP provides a deeper understanding of feature importance within complex, non-linear relationships.
Discussion
The global burden and clinical challenges of EOS
The global incidence of neonatal EOS varies significantly between regions, reflecting disparities in healthcare resources and monitoring practices. In developed countries, the incidence of EOS is relatively low due to advanced medical systems and effective preventive measures. For example, in the United States, the incidence of EOS among neonates with a birth weight greater than 2500 g is approximately 0.57 per 1000 live births, rising to 1.38 per 1000 live births in neonates weighing between 1500 and 2500 g. 16 Conversely, in developing countries and resource-limited settings, the incidence of EOS is markedly higher. Recent global burden data indicate an estimated 6.31 million cases of neonatal sepsis and related infections in 2019, representing a 12.79% increase since 1990. 17 These regions also experience significantly higher mortality rates, with EOS accounting for a substantial proportion of neonatal deaths. This stark contrast highlights the urgent need for targeted interventions, including improving access to prenatal care, enhancing infection prevention strategies, and ensuring timely treatment in resource-constrained areas to reduce the burden of EOS globally.
EOS in term neonates, while relatively rare (approximately 0.6 per 1000 live births in high-resource settings), poses significant diagnostic and management challenges due to its non-specific clinical signs and overlap with other neonatal conditions. These challenges often result in overtesting and unnecessary empirical antibiotic use, which can lead to increased healthcare costs and antimicrobial resistance. By focusing specifically on term neonates undergoing vaginal trial labor, our study addresses this clinically significant subgroup, where risk factors such as maternal infections and perinatal complications play a predominant role. This tailored approach provides a more precise framework for risk stratification, reducing unnecessary interventions and improving neonatal outcomes.
Development and validation of the prediction model
In this study, we developed and evaluated 10 ML models for predicting EOS in neonates, utilizing comprehensive prenatal and perinatal data from a cohort of 1613 full-term pregnant women. Among the models tested, XGBoost demonstrated superior performance, achieving the highest AUC of 0.87, indicating strong discriminative ability. The SHAP analysis further highlighted the clinical relevance of key predictive features, such as temperature-related factors, inflammatory markers (CRP and PCT levels), and labor and delivery characteristics (e.g. PROM and total labor duration). These findings underscore the potential of ML models, particularly XGBoost, to enhance the early identification of EOS in full-term neonates. By focusing on a well-defined population and utilizing advanced modeling techniques, our study provides a robust framework for integrating ML into clinical practice, potentially improving early diagnosis and neonatal outcomes.
We made a deliberate choice to exclude preterm cases, a decision driven by the need to minimize variability within the dataset and to ensure that the resulting predictive model is both precise and highly applicable to full-term neonates. This exclusion is significant because the mechanisms and infection risks associated with EOS differ considerably between preterm and full-term infants. Preterm neonates, whose immune systems are still maturing, are more susceptible to a wider array of pathogens, including those acquired in utero or in a hospital setting, and are prone to complications such as respiratory distress syndrome, which can complicate the diagnosis of sepsis. 18 Conversely, EOS in full-term neonates typically results from pathogens transmitted from the mother during birth, with Group B Streptococcus and Escherichia coli being the most frequent culprits. 16 Additionally, the incidence of EOS in term neonates is relatively low, and its clinical manifestations are often non-specific, which poses significant challenges for timely and accurate diagnosis. These challenges frequently result in overtesting and the empirical overuse of antibiotics, both of which can contribute to increased healthcare costs and the risk of antibiotic resistance. Existing predictive models often target preterm neonates or mixed populations, which do not account for the unique of EOS in term neonates. By narrowing our focus to full-term mothers undergoing a trial of labor, our model aims to mitigate these issues by providing precise risk stratification that aligns with the specific clinical presentations and risk factors pertinent to this group, thereby enhancing predictive accuracy and supporting more targeted clinical decision-making.
Among the models, XGBoost demonstrated the highest performance, achieving an accuracy of 92.57% and an AUC of 0.87. The SHAP analysis further highlighted the clinical relevance of key predictive features, such as temperature-related factors, inflammatory markers (CRP and PCT levels), and labor and delivery characteristics (e.g. PROM and total labor duration). To complement these findings, we combined LASSO regression and SHAP analysis to enhance feature selection and model interpretation. LASSO regression effectively identified 36 predictors by prioritizing linear relationships and ensuring dimensionality reduction. SHAP analysis provided a global interpretation of the XGBoost model, capturing both linear and non-linear relationships among predictors. The overlap between these two methods validated high-priority features like maternal CRP, peak temperature, and PROM, while SHAP uniquely identified additional contributors such as maternal WBC count, reflecting nuanced interactions often missed by LASSO. This dual-method approach emphasizes the importance of integrating interpretable ML tools to balance model robustness and clinical applicability, enabling actionable insights for EOS risk stratification.
Advancing beyond existing prediction tools
Our machine-learning-based study provides several advantages over existing methods, including traditional tools like the Kaiser Permanente EOS calculator and recent predictive studies. The Kaiser Permanente EOS calculator, while widely used, relies on outdated data (1993–2007) and subjective clinical assessments, which introduce variability and risk of underdiagnosis in certain populations. Specifically, while the calculator is an invaluable tool based on a robust logistic regression model, its framework is constrained to a pre-defined set of predictors. In contrast, our ML approach explores a much wider feature space of 36 maternal variables, allowing the model to uncover complex, non-linear relationships and interactions that simpler statistical models may miss. Our study therefore represents a complementary, data-driven methodology that seeks to identify novel risk patterns beyond the established predictors used in current calculators. Similarly, studies such as An and Neal rely on conventional statistical models, achieving moderate performance (e.g. AUC of 0.74 in Neal et al.) without the advanced flexibility and predictive accuracy provided by ML.10,19 But et al. demonstrated the potential of ML with neural networks but faced challenges in interpretability and scalability. Unlike the widely used Kaiser Permanente EOS calculator, which relies on a limited set of predictors within a traditional logistic regression framework, our approach employs ML to explore complex, non-linear relationships within a much larger feature space (36 maternal variables). While the EOS calculator provides a valuable baseline, our study demonstrates the potential of data-driven methods to uncover more nuanced risk patterns, offering a complementary and potentially more granular approach to risk stratification. In contrast, our study systematically evaluates 10 ML algorithms, with XGBoost achieving the highest AUC (0.87) and accuracy (92.57%), supported by SHAP analysis for interpretable outputs. In addition to leveraging ML, our study incorporates modern techniques such as SMOTE to address class imbalance, a common limitation in EOS datasets. 20 This innovation contrasts with Van Der Hoeven et al., which focused on retrospective analysis of CRP, blood culture, and clinical signs of infection to guide early antibiotic discontinuation. 21 While their approach reduced unnecessary antibiotic use, it lacked predictive adaptability and generalizability. Our model not only incorporates CRP, blood culture results, and clinical data but also uses advanced algorithms to enhance real-time prediction and minimize missed diagnoses. Our study further distinguishes itself by targeting a specific population—term neonates born following trial labor—where risk factors such as maternal fever, prolonged rupture of membranes, and labor duration are crucial. This contrasts with broader studies like Stoll et al. and Flannery et al., which primarily address preterm or mixed populations. Our tailored approach improves the precision and clinical relevance of EOS prediction for term neonates.16,18
By integrating contemporary data and advanced techniques, our model offers a robust, clinically actionable tool for EOS prediction. It surpasses conventional methods and existing tools by providing enhanced accuracy, reducing unnecessary interventions, and addressing the unique challenges associated with EOS in term neonates. Future work should focus on external validation and multi-center deployment to ensure broad applicability and further optimize neonatal outcomes.
Our models utilized a range of features, including maternal age, BMI, clinical indicators, and inflammatory markers. The SHAP analysis provided crucial insights into the key features contributing to EOS risk prediction. Among these, maternal peak temperature emerged as the most influential predictor. Elevated maternal temperature during labor is a well-recognized risk factor for neonatal EOS, often indicative of conditions like chorioamnionitis. By integrating SHAP analysis with LASSO regression, we leveraged the complementary strengths of these methods to refine feature selection and enhance interpretability. LASSO ensured model simplicity and stability by selecting predictors based on their linear contribution to EOS risk, while SHAP captured complex, non-linear interactions among features. For instance, maternal WBC count, identified by SHAP but not LASSO, underscores the importance of considering nuanced inflammatory responses during labor. This integrative approach provides clinicians with both robust predictive accuracy and an interpretable framework for understanding EOS risk factors, fostering trust in ML-driven decision-making.
A study conducted in Hangzhou, China, highlights that maternal fever (≥38.0°C) significantly predicts neonatal EOS, emphasizing the importance of monitoring maternal temperature to prevent sepsis. 18 CRP and WBC count are both critical inflammatory markers that play a significant role in the early detection of neonatal EOS. Studies have demonstrated that combining CRP levels with WBC counts can enhance the predictive accuracy for EOS, as both markers respond to different aspects of the inflammatory process. Elevated CRP levels, particularly when measured serially, provide insight into the progression of the inflammatory response, while an increased WBC count before delivery can serve as an early warning sign of potential neonatal infection.7,21 Our study focused on developing a predictive model for neonatal EOS using maternal prenatal and perinatal factors. This approach aims to enable early identification of high-risk neonates before the onset of clinical symptoms or postnatal laboratory results, facilitating timely interventions. By relying exclusively on maternal data, the model provides a unique perspective on risk stratification that complements traditional diagnostic methods. The exclusion of neonatal CRP, PCT, and temperature data reflects the study's design to assess maternal factors as primary predictors. While neonatal biomarkers can provide direct evidence of infection, their utility is often limited to postnatal evaluation, whereas maternal data allow for earlier risk prediction. Future studies could integrate neonatal data to enhance predictive accuracy and provide a more comprehensive assessment. Maternal BMI and maternal age have been linked to pregnancy complications, including gestational diabetes and hypertension, which can predispose neonates to infections like sepsis. A study in Ethiopia found that maternal obstetric factors, such as BMI and maternal age, are significant predictors of EOS, highlighting the need for comprehensive maternal health monitoring.7,22 This finding suggests a need for heightened vigilance and proactive management in older pregnant women. The model also identified total labor duration as an important factor, with prolonged labor, especially when accompanied by prolonged rupture of membranes, increases the risk of neonatal infection. One study highlighted that extended labor, especially when accompanied by a longer duration of membrane rupture, is strongly associated with higher rates of EOS in neonates. This finding underscores the necessity for careful management and timely intervention during labor to reduce the risk of infection transmission to the neonate. 23
Clinical utility and interpretability of the model
The primary clinical utility of our model, when tuned to an optimized threshold of 0.04, lies in its potential as a first-line screening tool. A high sensitivity of 92.8% ensures that the vast majority of at-risk neonates are identified for closer observation or further evaluation, directly addressing the critical need to avoid missing EOS cases. However, this improved sensitivity comes at the cost of a lower specificity of 73.1%, leading to a higher number of false positives. This trade-off is fundamental in clinical screening: the model is designed to cast a wide net to flag a larger cohort of neonates as “potentially at-risk,” necessitating a second stage of clinical assessment, rather than serving as a definitive diagnostic test.
This approach aligns with clinical practice where initial screening tests prioritize sensitivity. While a PPV of 13.4% may seem low, it signifies that within the group flagged as high-risk by our model, the prevalence of EOS is substantially enriched compared to the baseline prevalence of 4.3% in the total population. This could help clinicians allocate monitoring resources more effectively. Conversely, the extremely high NPV of 99.6% provides strong reassurance, suggesting that neonates classified as low-risk are highly unlikely to have EOS, which could potentially help reduce unnecessary interventions in this group, pending prospective validation.
Study limitations and future perspectives
While our study presents promising results, there are several limitations that need to be addressed in future work. Firstly, our models focused solely on predicting the presence or absence of EOS. Future research could aim to develop models that also predict the severity of sepsis, enabling more nuanced clinical decision-making. The dataset included 1613 maternal-neonatal pairs, with only 69 EOS cases, resulting in a prevalence of 4.2%. This low prevalence reflects the rare nature of EOS but also poses challenges for ML model development, particularly in achieving robust generalizability. Although we employed the SMOTE to address class imbalance during model training, the inherent limitations of oversampling methods, such as the potential introduction of synthetic noise, cannot be overlooked. Additionally, the low NPV and the high FPR observed in our results are a direct consequence of this class imbalance. The low NPV indicates challenges in reliably identifying true negative cases, while the high FPR suggests a tendency to misclassify some true negatives as false positives. This highlights the need for careful interpretation of model predictions, especially for low-prevalence conditions such as EOS. Future studies should consider employing alternative strategies, such as cost-sensitive learning or threshold adjustments, to mitigate these limitations and improve overall performance. Furthermore, the DCA demonstrated the clinical utility of the XGBoost model, showing a higher net benefit compared to the “treat all” and “treat none” strategies across most threshold probabilities. However, it is important to note that the net benefit briefly dips below the “treat none” line in the threshold range of 0.6–0.8, suggesting reduced clinical utility in this specific range. This phenomenon may be attributed to the low prevalence of EOS in our dataset, which can impact the model's predictive reliability at certain thresholds. Despite this limitation, the overall DCA results support the model's potential for clinical application, particularly in thresholds outside this range. Future studies with larger, more balanced datasets and refined threshold calibration are warranted to address this issue and ensure more consistent performance across all probability thresholds. The small sample size of EOS cases may also limit the model's stability and increase the risk of overfitting, particularly in high-dimensional feature spaces. While ensemble algorithms such as XGBoost and Random Forest are well-suited to handle such challenges, other approaches tailored for small datasets, such as few-shot learning techniques, could be explored in future studies to enhance robustness and performance. Secondly, the dataset used in this study is derived from a single center, which may limit the generalizability of the findings to other populations and healthcare settings. The absence of external validation using multi-center datasets is a significant limitation, as it precludes the assessment of the model's robustness and performance in different clinical environments. Future studies should prioritize multi-center collaborations to validate the model and ensure its broad applicability. Additionally, while the model incorporates a range of maternal and neonatal factors, certain variables such as genetic or microbiome data, which may influence EOS risk, were not included. The integration of such data in future studies could improve the model's accuracy and provide deeper insights into EOS pathogenesis. Despite these limitations, our study provides a robust foundation for future research, offering a tailored and interpretable model that addresses a critical clinical need. The next steps should include prospective validation in multi-center settings, deployment of the model into clinical workflows, and ongoing updates as new data become available to enhance its clinical relevance and utility. While SMOTE was employed to address the inherent class imbalance in the dataset, we acknowledge that the introduction of synthetic samples may introduce biases that do not fully reflect the complexity of real-world EOS presentations. Synthetic cases generated from a limited number of positive samples (n=69) may fail to capture the true heterogeneity of the minority class. This could potentially lead to overfitting or reduced generalizability when applied to external populations. Future studies should consider the inclusion of larger, multi-center datasets and prospective data collection to validate model performance in more diverse and clinically representative scenarios.
Potential for clinical implementation
A potential implementation pathway for this model would involve its integration into the hospital's Electronic Health Record (EHR) system. Upon a full-term mother's admission for labor, the required maternal variables could be automatically extracted to calculate an EOS risk score in real-time. If the model's output probability exceeds the optimized threshold of 0.04, a non-intrusive flag or alert could be presented to the attending clinician within the EHR interface. This alert would not dictate a diagnosis but would serve as a clinical decision support tool, prompting heightened surveillance or consideration for early screening tests for the neonate. Such a workflow could potentially streamline the identification of high-risk infants without inducing significant alert fatigue.
Conclusion
In conclusion, this study demonstrates the potential of ML to identify significant maternal risk factors for EOS. Crucially, we demonstrated that by tuning the classification threshold, the model can be optimized for a high-sensitivity screening task, though this necessitates accepting a higher false-positive rate. Our findings represent a foundational step, highlighting both the potential of using AI in this domain and the challenges that must be overcome for clinical implementation. Future work must focus on external multi-center validation and prospective studies to determine the real-world clinical utility and optimal implementation of such a screening tool.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251376968 - Supplemental material
Supplemental material, sj-docx-1-dhj-10.1177_20552076251376968
Footnotes
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the 2024 Bao’an District Medical and Health Science Research Project (grant number 2024JD097).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
