Abstract
Lymphoma is a prevalent malignant tumor within the hematological system, posing significant challenges to clinical practice due to its diverse subtypes, intricate radiological and metabolic manifestations. Lymphoma segmentation studies based on positron emission tomography/computed tomography (PET/CT), CT, and magnetic resonance imaging represent key strategies for addressing these challenges. This article reviews the advancements in lymphoma segmentation research utilizing deep learning methods, offering a comparative analysis with traditional approaches, and conducting an in-depth examination and summary of aspects such as dataset characteristics, backbone networks of models, adjustments to network structures based on research objectives, and model performance. The article also explores the potential and challenges of translating deep learning-based lymphoma segmentation research into clinical scenarios, with a focus on practical clinical applications. The future research priorities in lymphoma segmentation are identified as enhancing the models’ clinical generalizability, integrating into clinical workflows, reducing computational demands, and expanding high-quality datasets. These efforts aim to facilitate the broad application of deep learning in the diagnosis and treatment monitoring of lymphoma.
Introduction
Lymphoma is a malignant tumor that occurs in the lymphatic system and is one of the most common malignancies of the hematological system globally. 1 Lymphoma exhibits complex classification (Figure 1), with each subtype demonstrating distinct biological behaviors and clinical manifestations. Due to the widespread distribution of the lymphatic system throughout the body, lymphomas can occur in various locations, including lymph nodes, spleen, liver, bone marrow, and other organs. 2 The heterogeneity of their radiological presentations further complicates matters; even within the same lymphoma type, differences in imaging characteristics such as size, shape, density, or signal intensity may exist. Imaging examinations serve as the primary noninvasive method for lymphoma assessment. However, the variability in radiological features poses significant challenges to the manual evaluation and interpretation of lymphoma images. 3 Furthermore, the lack of objective evaluation criteria results in assessment outcomes being influenced by subjective factors of the evaluating physicians. Therefore, research into computer-assisted lymphoma segmentation methods has emerged to address these issues.

Lymphoma typing and subtype schematic diagram.
Medical image segmentation is a fundamental technique in image analysis that aims to delineate specific anatomical or pathological structures from the background in medical images. 4 In the context of lymphoma, segmentation plays a pivotal role not just by outlining lesions or tumors, but more importantly by enabling robust and reproducible extraction of clinically meaningful quantitative features. Among these, total metabolic tumor volume (TMTV) has emerged as a key biomarker—crucial for staging, risk stratification, treatment planning, and response monitoring. The ability to automatically and consistently quantify such indicators underscores segmentation's value as a foundational tool for downstream clinical decision making, rather than an end goal itself. Recent studies, such as Albano et al., 5 have further emphasized the clinical relevance of TMTV and the necessity of accurate segmentation in achieving it. Additionally, segmentation supports treatment planning (e.g. defining radiation fields or surgical margins), tracking disease progression, assessing therapeutic outcomes, and informing prognosis. 6 Earlier approaches to lymphoma segmentation primarily relied on traditional machine learning methods, including thresholding based on pixel intensity, edge detection to identify boundary changes, 7 region-growing algorithms that expand from seed points based on predefined similarity criteria, 8 and clustering techniques that group pixels by feature similarity. 9 These methods offer interpretable models with relatively modest data and computational demands, and clear decision logic that aids understanding. 10 However, they depend heavily on handcrafted features and expert intervention, introducing subjectivity and potential variability across institutions and clinician expertise levels, which may lead to inconsistent segmentation results and limit reproducibility in clinical settings. In contrast, deep learning has increasingly become the dominant paradigm in lymphoma segmentation due to several key advantages beyond those already recognized—such as higher efficiency and end-to-end automation. Deep learning models are trained on large-scale, high-quality annotated datasets, enabling them to learn complex visual patterns and clinical decision strategies from experienced clinicians. This allows for encapsulation of expert-level knowledge and promotes more consistent, reproducible outcomes in practice.7,11 Moreover, unlike traditional approaches that rely on manual feature engineering and require substantial human involvement in multiple stages, deep learning enables direct learning from raw imaging data, supporting fully automated pipelines and achieving superior performance under standardized, operator-independent evaluation settings, as evidenced by results from international benchmarks such as BraTS, 12 HECKTOR, 13 and AutoPET. 14 Deep learning also facilitates automatic, hierarchical, and task-specific feature learning—a process known as automatic feature learning—which allows models to optimize feature representations in a data-driven manner. Empirical evidence confirms that such learned features are often more informative and discriminative than handcrafted alternatives, especially in complex medical imaging tasks. For instance, Shen et al. 15 reviewed deep learning applications in medical image analysis and highlighted the consistent superiority of Convolutional Neural Network (CNN)-derived features over manually designed ones across classification, detection, and segmentation. Similarly, Litjens et al. 16 concluded that deep features extracted by convolutional networks generally outperform traditional features across diverse imaging modalities and pathological conditions. Notably, Isensee et al. 17 demonstrated that the nnU-Net framework achieved state-of-the-art results in 23 different biomedical segmentation challenges without requiring any manual feature engineering. These findings collectively demonstrate that deep learning–derived features are not only more efficient to obtain—eliminating the need for time-consuming feature design—but also more effective in capturing high-level semantic information essential for accurate medical image interpretation.
The purpose of lymphoma segmentation research based on medical images is to serve clinical applications. Application-oriented deep learning methods in lymphoma segmentation research aim to integrate deep learning models into clinical workflows and application scenarios. This approach also addresses issues of model interpretability and generalization in clinical settings, thereby compensating for the shortcomings of deep learning methods in this regard. 18 To summarize various studies comprehensively, we conducted searches on Google Scholar and Web of Science using keywords “lymphoma segmentation” and “deep learning” to retrieve recent literature. We screened 45 studies, excluding non-English articles, studies outside the scope of deep learning-based segmentation, studies not related to lymphoma segmentation, and duplicate articles. We included studies published from 2018 to 2024, focusing on methods tested and validated, written in English, specifically addressing lymphoma segmentation using deep learning methods, ensuring all results were verified. Different from existing reviews, this review not only reviews the latest advances, strengths, and weaknesses of lymphoma segmentation from a deep learning perspective but also compares and summarizes relevant methods. Furthermore, it explores this issue from a clinical application perspective. The structure of this article includes an introduction section, a comprehensive overview of deep learning technology in the second section, a review of recent advances in deep learning applied to lymphoma segmentation in the third section, a discussion and outlook on clinical applicability and technical limitations in the fourth section, and finally, a summary of the entire article in the fifth section.
Fundamentals of deep learning
Introduction to deep learning
Deep learning methods are a class of algorithms inspired by the structure of the human brain, which construct multilayer neural networks to learn complex representations of data. 19 These methods have achieved significant successes in various fields such as image processing20,21 natural language processing, 22 and speech recognition. 23 In the field of medical image segmentation, deep learning methods particularly demonstrate their powerful capabilities. Deep learning models learn high-level features of data through their multilayered structures. Lower layers may learn basic features like edges, while higher layers may capture shapes, textures, and even semantic information. Neurons in each layer perform nonlinear transformations on input data, enabling the network to capture complex patterns. 15 Compared to traditional machine learning methods, deep learning models automate feature extraction, reducing the need for manual feature engineering. These models use backpropagation algorithms to compute gradients of the loss function with respect to network parameters, and optimization algorithms like gradient descent to update network weights. Deep learning models typically require large amounts of data for training to learn generalized feature representations. The application of deep learning methods in medical image segmentation continues to expand, enhancing the automation level and improving the accuracy and reliability of segmentation results.
Fundamentals of Convolutional Neural Networks
The CNN is a landmark model in deep learning that combines deep learning and image processing techniques, achieving significant advancements in image analysis. 24 Similarly, in medical image segmentation, CNN serves as the backbone network for the majority of segmentation models. CNN reduces the number of parameters and enhances training efficiency through weight sharing and leveraging spatial relationships, operating as a supervised learning model.24,25 Since the concept of visual receptive fields was introduced by Hubel et al. 26 in 1962, followed by Fukushima's proposal of the Neocognitron based on receptive fields in 1980, 27 and LeCun et al.’s 28 development of LeNet5 using the backpropagation algorithm in 1998, CNN has undergone a progression from theoretical exploration to experimental development. The introduction of AlexNet 29 marked CNN's breakthrough in the ImageNet competition in 2012, solidifying its core position in computer vision research and continuously driving further advancements in the field.
Advancements of deep learning in lymphoma segmentation research
In the field of medical image segmentation for lymphoma, specific tasks rely predominantly on deep learning models.
Lymphoma imaging encompasses diverse modalities that differ significantly from natural images, based on various physical properties and sources of energy. The resulting images span multiple scales. Therefore, training a unified multiscale base model from these multimodal images is currently impractical. All studies included in this article focus on specialized segmentation models for lymphoma. Next, we will elaborate on dataset characteristics, network structures, supervision methods, and model performance. The relevant content has also been summarized in Tables 1 and 2.
Basic information of lymphoma segmentation research based on deep learning.

2D: two-dimensional; 3D: three-dimensional; CT: computed tomography; PET: positron emission tomography.
Performance for lymphoma segmentation research based on deep learning.

AUC: area under the curve; DSC: dice similarity coefficient; HD: Hausdorff distance; MAD: mean absolute deviation; MAE: mean absolute error; MSE: mean squared error.
Dataset
In lymphoma segmentation research based on deep learning methods, the critical initial step that determines the direction and content of the study is the construction of the dataset. A high-quality dataset forms the foundation for training effective models. 75 Firstly, dataset diversity and balance are crucial. The dataset needs to include images of different types and stages of lymphoma to ensure the model generalizes well across various scenarios. Some types of lymphoma may be more common than others, necessitating balance in the dataset. Secondly, annotation poses a unique challenge. Unlike natural image segmentation datasets, lymphoma segmentation annotations typically involve pixel or voxel-level annotations rather than scribble annotations or region of interest annotations. This is due to the high heterogeneity inherent in lymphoma imaging, which also exists within subtypes of the same lymphoma type. Accurate annotation serves as crucial supervision signal for training deep learning models. Errors or ambiguous annotations can lead to incorrect feature learning, affecting segmentation accuracy. 76 Thirdly, dataset scale is advantageous compared to traditional machine learning methods. Deep learning methods excel with large-scale datasets. Sufficient data volume helps models learn features better, enhancing their generalization capability. Insufficient data may lead to overfitting, where the model performs well on training data but poorly on new, unseen data. 77 Finally, constructing multimodal image datasets and multitask datasets including different information achieves the goals of segmentation models, such as lymphoma identification, prognosis prediction, and treatment monitoring. These datasets also contribute significantly to interpretable deep learning models.7,11
Among the 45 studies included in this article, the data modalities encompass positron emission tomography/computed tomography (PET/CT) with multimodal fusion and multitemporal data, different sequences of magnetic resonance imaging (MRI) data, and contrast-enhanced CT data. The proportions of the various modes are shown in Figure 2. The selection of different data modalities depends on specific medical issues in lymphoma segmentation research. PET/CT combines functional and structural imaging, providing information on lesion metabolism and anatomical location, playing a crucial role in lymphoma diagnosis, staging, treatment evaluation, and prognosis assessment. 78 Consequently, it is the primary modality type in this type of research dataset, comprising 38 studies. The majority of lymphoma types included are non-Hodgkin lymphomas, with a smaller portion being Hodgkin lymphomas. The lymphoma subtypes primarily studied include diffuse large B-cell lymphoma (DLBCL), primary mediastinal large B-cell lymphoma (PMBCL), mantle cell lymphoma, and follicular lymphoma, all clinically prevalent. The anatomical coverage in these PET/CT datasets varies; most cover the entire body excluding the head and neck. Two studies focus on the chest and mediastinum, one on nasopharynx, and one on abdominal organs. Another data modality is different sequences of MRI data. Unlike PET/CT, MRI offers higher soft tissue contrast and multiplanar imaging capabilities, addressing PET/CT's limitations in evaluating central nervous system lymphomas. 79 Six studies utilize this data modality, focusing mainly on PMBCL and primary central nervous system lymphoma (PCNSL). The third data modality is contrast-enhanced CT, which, due to its limited scanning range, primarily provides anatomical information and has limitations for lymphomas that can manifest throughout the body. It is an unconventional clinical assessment method for lymphoma but is chosen for its low radiation and rapid imaging characteristics, making it suitable for assessing lymphomas in children. 80 Two studies adopt this data modality, both sourced from pediatric lymphoma patients. Due to the ethical and privacy considerations of medical data, most high-quality datasets used are private. The openly accessible lymphoma datasets include the HECKTOR challenge dataset, 13 the BRATS dataset, 81 and AutoPET dataset. 14 The lack of high-quality annotated datasets to some extent constrains the development of lymphoma segmentation research.

The proportion of different modal data in lymphoma segmentation studies.
HECKTOR challenge dataset
The HECKTOR (Head and Neck Tumor Segmentation) dataset provides annotated PET/CT images for the task of segmenting primary tumors and lymph node metastases in patients with head and neck squamous cell carcinoma (HNSCC). 13 It includes multimodal imaging data from multiple institutions, reflecting real-world heterogeneity in acquisition protocols and patient anatomy. While the dataset mainly targets HNSCC cases, it also includes a limited number of patients with head and neck lymphoma, expanding its relevance to lymphoproliferative diseases in this anatomical region. This diversity enables research not only on carcinoma segmentation but also on modeling lesion appearance in lymphoma, where nodal involvement may differ in morphology and uptake. The challenge promotes the development of robust, generalizable models capable of handling the complex anatomy and variable lesion characteristics typical of head and neck cancers. Annotations are provided by expert radiologists and nuclear medicine physicians, enabling reliable benchmarking for automated segmentation methods.
BraTS dataset
The Brain Tumor Segmentation (BraTS) dataset is a widely used benchmark for evaluating the segmentation of gliomas from multimodal MRI scans, including T1, T1c, T2, and fluid attenuated inversion recovery sequences. 12 It provides manual segmentations of enhancing tumor, peritumoral edema, and necrotic core, supporting detailed subregion-level analysis. In addition to gliomas, the dataset has also been used in studies involving PCNSL cases, which may exhibit imaging patterns overlapping with high-grade gliomas but differ in biological behavior and treatment response. The inclusion of such cases broadens the dataset's applicability to lymphoid malignancies in the brain and enables comparative evaluation of segmentation methods across tumor types. The dataset spans several editions and includes both pre-operative and postoperative scans, with expert annotations curated for consistency. BraTS has driven innovation in brain tumor segmentation by encouraging methodological comparisons under standardized settings.
AutoPET dataset
The AutoPET dataset consists of whole-body flurodeoxyglucos (FDG)-PET/CT images with expert-annotated lesion segmentations from a wide range of oncological cases. 14 It is specifically designed to foster research on fully automated, end-to-end segmentation of metabolically active tumors. The dataset includes diverse malignancies, with multiple cases of lymphoma, making it especially relevant for developing models aimed at quantifying tumor burden in systemic cancers. AutoPET presents significant variability in lesion location, shape, and uptake patterns, posing challenges that mimic clinical reality. It supports the development of models aimed at calculating key quantitative biomarkers, such as TMTV, with potential applications in disease monitoring, treatment planning, and prognostic evaluation across cancer types.
Lymphoma segmentation network architecture
The network architecture is the core component of deep learning models, and its importance in lymphoma segmentation research is reflected in its impact on feature extraction, spatial resolution preservation, handling of multiscale and multimodal data, computational efficiency, generalization, and robustness. 19 Proper selection and design of network architecture can significantly enhance segmentation performance and the feasibility of clinical applications. This article specifically discusses the composition of network architecture in terms of backbone network selection and strategies for network structure adjustment.
Classification of backbones
In the network structure of deep learning models, “backbone” refers to the core structure that constitutes the main framework of the network. It is responsible for extracting features from input data, which serve as the basis for subsequent tasks. The backbone also plays a crucial role in achieving computational efficiency, automation, and scalability of the model. 82 In lymphoma segmentation research, all utilized backbones are based on CNN architectures and their variants. In this section, we will summarize and classify the backbones used in the 45 studies included in our analysis, providing specific explanations for each (Figure 3).

The main backbone species in lymphoma segmentation studies.
ResNet
ResNet, short for Residual Network, was proposed by He et al. 83 from Microsoft Research in 2015. It achieved first place in both classification and object detection tasks in the ImageNet competition of that year. The development of ResNet primarily aimed to address the issues of vanishing and exploding gradients that arise with increasing network depth. At the core of ResNet is the residual block, each of which contains two paths: a normal path of convolution layers and an identity shortcut connection that adds the input directly to the output. This design allows the network to learn identity mappings, where the input is propagated directly to the output without undergoing complex transformations. ResNet's network structure typically consists of stacking multiple residual blocks, enabling the construction of networks of varying depths such as ResNet-50, 84 ResNet-101, 85 etc. By using residual connections, gradients can propagate more effectively through the network, thereby mitigating the vanishing gradient problem during training of deep networks. This capability allows for the construction of deeper network architectures, enhancing the model's learning capacity. The design of residual blocks enables the network to learn more complex feature representations. Due to these characteristics, ResNet has been widely adopted as a backbone network in various applications, including lymphoma segmentation research. At present, ResNet continues to demonstrate strong performance and stability in various tasks due to its modular design, ease of optimization, and compatibility with other architectural innovations such as attention mechanisms and multiscale processing. However, its limitations include relatively high computational cost compared to lightweight alternatives, lack of adaptability to non-Euclidean data or dynamic input structures, and less efficiency in capturing global context compared to newer transformer-based models. Despite these drawbacks, ResNet remains a foundational and versatile architecture, particularly valuable when combined with other modern techniques in medical image analysis.
U-Net
U-Net is a CNN architecture specifically designed for medical image segmentation tasks in deep learning. Originally proposed by Ronneberger et al. in 2015, 86 U-Net was developed to address the challenges of biomedical image segmentation where precise localization of anatomical or pathological regions is critical. Its architecture features a characteristic “U” shape composed of a contracting (encoder) path and an expansive (decoder) path. The encoder path captures context by progressively reducing spatial resolution and increasing feature channels, while the decoder path restores resolution through upsampling, enabling accurate delineation of segmentation boundaries. A key innovation in U-Net is the use of skip connections that link corresponding layers in the encoder and decoder, allowing fine-grained spatial information lost during downsampling to be preserved and reintegrated during reconstruction. This design enhances the model's ability to handle noisy inputs and maintain fine structural details—an essential capability in medical imaging tasks. Initially implemented as a two-dimensional (2D) architecture, U-Net processed slice-by-slice inputs and was widely applied to planar imaging modalities. However, this approach had limitations in capturing volumetric contextual information inherent in three-dimensional (3D) medical imaging data such as PET/CT scans used in lymphoma studies. To address this, researchers extended U-Net to 3D by replacing 2D convolutions and operations with their 3D counterparts, enabling the model to learn from volumetric input data directly. This transition from 2D U-Net to 3D U-Net 87 significantly improved performance in volumetric segmentation tasks, particularly for diseases like lymphoma where lesions exhibit irregular shapes, heterogeneous uptake, and complex spatial distributions in PET/CT volumes. By leveraging the full contextual information across slices, 3D U-Net architectures are better suited to accurately identify metabolically active regions, leading to more consistent and clinically relevant segmentation results. Furthermore, the open and modular design of U-Net makes it highly adaptable—researchers can modify the architecture by incorporating attention mechanisms, residual connections, or multiscale feature fusion modules to enhance task-specific performance. In lymphoma segmentation, such adaptations have been instrumental in improving delineation accuracy, especially in challenging cases involving small or low-contrast lesions. At present, U-Net retains several significant advantages, including architectural simplicity, flexibility for customization, and strong baseline performance across various segmentation tasks. Its extensive use and open-source implementations have fostered reproducibility and accelerated development in the medical imaging community. However, limitations remain: standard U-Net variants may struggle with long-range contextual dependencies due to their inherently local receptive fields, and the 3D version, while more powerful, imposes substantial memory and computational demands, particularly with high-resolution volumetric data. Additionally, U-Net may require further architectural tuning to address issues such as class imbalance and uncertainty estimation in clinical settings. Despite these challenges, U-Net and its evolving variants continue to form the backbone of state-of-the-art segmentation models, offering a robust and extensible foundation for deep learning applications in lymphoma imaging and beyond.
DenseNet
DenseNet, short for Dense Convolutional Network, was introduced by Huang et al. 88 from the University of Groningen in 2016. The core idea of DenseNet is feature reuse, achieved by connecting each layer to every other layer in a feed-forward fashion, enhancing the information flow and parameter efficiency of the network. In DenseNet, each layer receives the outputs of all preceding layers as inputs and passes its own output to all subsequent layers. This dense connectivity pattern facilitates more effective information flow within the network. Due to the dense connections, gradients can propagate directly from deeper layers to shallower ones, which helps alleviate the vanishing gradient problem and allows the network to be deeper. By reusing features through dense connections, DenseNet reduces the number of parameters compared to other deep networks, making it more parameter-efficient. After each dense block, DenseNet employs transition layers to reduce the size of feature maps and decrease the number of channels, facilitating the subsequent processing of the network. DenseNet typically replaces traditional fully connected layers with global average pooling layers, which further reduces the number of parameters and enhances the network's robustness to variations in input sizes. At present, DenseNet remains advantageous for its efficient use of parameters, improved gradient flow, and ability to promote feature reuse, making it suitable for tasks where model compactness and deep supervision are essential. It has shown effectiveness in medical image analysis, including segmentation and classification, particularly when computational resources are limited. However, its limitations include increased memory consumption due to the concatenation of feature maps from all preceding layers, which can lead to high graphics processing unit (GPU) usage during training, especially in deeper variants. Additionally, as the network grows, the feature concatenation can become computationally intensive and may introduce redundancy in the learned representations. Despite these challenges, DenseNet continues to serve as a strong backbone in various deep learning applications, offering a good balance between performance and efficiency, especially in scenarios where deeper models are required without significantly increasing the number of parameters.
V-Net
V-Net is another variant of CNN architecture, first proposed by Milletari et al. 89 in 2016. Similar to U-Net, V-Net's structure consists of an encoder path and a symmetric decoder path. However, V-Net is designed for 3D images, eliminating the need for slicing the input dataset. It introduces residual structures to accelerate convergence, replaces pooling layers with convolutional layers, and proposes a new objective function based on maximizing the Dice similarity coefficient (DSC). The DSC is a statistical metric used to quantify the spatial overlap between two binary segmentation results, typically the predicted and ground truth regions in medical image analysis, with values ranging from 0 (no overlap) to 1 (perfect agreement). In the encoder path, each stage consists of one to three convolutional layers to extract image features. Residual learning is applied at each stage, where the input feature map undergoes several convolutions and nonlinear activations, and the resulting feature map is added pointwise to the input feature map. After each stage, downsampling is performed using an appropriate stride to reduce the image resolution, halving the size of the feature map and doubling the number of feature channels. Similarly, the decoder path is symmetric to the encoder path, with each stage consisting of one to three convolutional layers. Parameter learning is applied in each stage of the decoding process, followed by transpose convolutions to double the size of the feature map. Finally, 1×1×1 convolutions produce two feature map outputs of the same size as the input volume, followed by softmax to output probabilities for each voxel belonging to foreground and background. V-Net utilizes skip-connection structures similar to U-Net to gather fine-grained details that may be lost in the compression path, thereby improving segmentation accuracy. In lymphoma segmentation research, V-Net's residual learning enables the model to learn more complex structures in tumor regions. Its capability with 3D convolutions is well-suited for clinical needs in lymphoma segmentation from multiple angles. Moreover, like U-Net, V-Net is an open backbone. Its modular design makes it easy to understand and modify, allowing researchers to adjust the network structure based on specific application scenarios to achieve better performance and meet diverse requirements. Currently, V-Net holds several advantages: its native 3D design allows it to fully utilize volumetric data such as PET/CT, enhancing spatial coherence in segmentation; the residual learning mechanism improves training efficiency and performance in complex lesion structures; and its Dice-based loss is especially suitable for medical segmentation tasks with class imbalance. However, its limitations include high computational and memory demands due to 3D convolutions and deeper architecture, making it challenging to deploy on resource-constrained hardware. Additionally, although residual connections help alleviate vanishing gradients, they may lead to feature redundancy when the number of layers becomes very large. Despite these constraints, V-Net remains a powerful and widely adopted architecture in 3D medical image segmentation, particularly when precise volumetric delineation is critical.
DeepMedic
DeepMedic is an open-source backbone designed specifically for medical image analysis, particularly excelling in neuroimaging-related tasks. Originally proposed by Kamnitsas et al., 90 DeepMedic utilizes 3D CNNs to perform segmentation and analysis of volumetric medical images, aiming to enhance diagnostic precision and inform treatment planning. One of its key contributions lies in its early and effective adoption of 3D CNNs over conventional 2D architectures, enabling more accurate modeling of spatial context and structural continuity in medical data, such as in brain tumor segmentation. Like most modern 3D deep learning segmentation frameworks, DeepMedic employs a patch-based training strategy, wherein large medical volumes are decomposed into smaller subvolumes or patches for model training. This approach addresses the substantial computational and memory demands inherent to full-volume 3D image processing, which typically exceed the capacity of contemporary GPUs. Thus, patch-based training has become a widely adopted and essential technique across 3D segmentation models, not limited to DeepMedic alone. The DeepMedic framework encompasses a modular workflow—including preprocessing, training, validation, and testing—that can be flexibly adapted to diverse imaging modalities and segmentation tasks. Network configurations, optimization strategies, and training parameters can be customized according to specific research or clinical requirements. Owing to its strengths in neuroanatomical analysis, DeepMedic has also been extended to central nervous system lymphoma segmentation tasks, with architectural adjustments made to accommodate the particular challenges of each dataset and clinical scenario. At present, DeepMedic offers several advantages: its deep multiscale architecture is adept at capturing both fine and coarse features through parallel pathways operating at different resolutions, enhancing segmentation accuracy, especially in small or irregularly shaped lesions; its patch-based strategy allows training on large 3D volumes even with limited GPU memory; and its modular, open-source design promotes adaptability and reproducibility in clinical and research settings. However, DeepMedic also has limitations. The patch-based approach, while memory-efficient, can lead to discontinuities at patch boundaries and requires careful tuning of patch size and sampling strategies. Additionally, the architecture is relatively rigid compared to newer transformer-based or attention-enhanced models, which offer more flexible and globally aware representations. Its reliance on hand-crafted preprocessing pipelines can further reduce generalizability across datasets with differing imaging characteristics. Despite these challenges, DeepMedic remains a robust and well-validated choice for 3D medical image segmentation, particularly in neuroimaging and lymphoma-related applications where volumetric context and precision are critical.
DeepLabv3
DeepLabv3, short for “”DeepLab: Deep Labelling for Semantic Image Segmentation,” is a deep learning model designed for semantic image segmentation. 91 The DeepLab series models hold a significant position in the field of image segmentation by integrating deep convolutional neural networks and conditional random fields (CRFs) to enhance segmentation accuracy. Key features of DeepLabv3 include: multiscale feature fusion, which leverages deep convolutional networks (such as ResNet) to extract and integrate features across various levels, enabling the model to recognize both global structures and local details; Atrous (dilated) convolution, which expands the receptive field without increasing the number of parameters or reducing resolution, allowing broader contextual understanding while maintaining spatial precision; an Encoder-Decoder structure that facilitates efficient feature extraction and reconstruction for fine-grained segmentation; a Pyramid Pooling Module that captures contextual information at multiple spatial scales to improve segmentation performance in scenes with complex structures; and CRF-based refinement in the final stage to enhance the spatial coherence of the segmentation outputs. Currently, DeepLabv3 exhibits several advantages: it achieves state-of-the-art accuracy on many benchmark datasets due to its powerful multiscale context aggregation and spatial precision mechanisms; its modular design allows integration with various backbone networks, enhancing adaptability to specific tasks such as lymphoma segmentation in medical imaging. Moreover, the use of atrous convolutions and pyramid pooling equips the model to handle objects of varying sizes and shapes—a critical advantage in medical contexts where lesion morphology is diverse. However, DeepLabv3 also has limitations. Its architecture is computationally intensive, especially when using high-resolution inputs and deep backbones, which may hinder deployment in real-time or resource-constrained environments. Additionally, its performance is heavily reliant on large annotated datasets, limiting its effectiveness in medical scenarios where expert-annotated data are scarce. The reliance on CRFs, while beneficial for refinement, introduces additional computational overhead and complexity in postprocessing. Furthermore, DeepLabv3 was originally developed for natural images, and direct application to 3D medical data requires significant architectural adaptations. Despite these challenges, DeepLabv3 remains a powerful and widely used model in semantic segmentation, and its core components—especially atrous convolutions and multiscale context fusion—continue to influence the development of advanced segmentation networks for both natural and medical image domains.
nnU-Net
nnU-Net is an adaptive deep learning segmentation framework proposed by Isensee et al. 92 in 2018. It is based on the original 2D and 3D U-Net architectures and specifically designed for medical image segmentation tasks. The design philosophy of nnU-Net is to automate the entire segmentation process, including preprocessing, network structure selection, hyperparameter tuning, model training, and postprocessing, without the need for manual intervention. Essentially, it is a collection of U-Net models. Each U-Net structure remains largely unchanged, inheriting the robustness of the U-Net network. Depending on different data modalities, nnU-Net adapts different processing strategies; the 3D U-Net handles 3D data, while the structure of multiple U-Net cascades enables the model to handle multimodal and multiviewpoint data. This feature aligns well with the requirements of lymphoma data, making nnU-Net a crucial backbone for lymphoma segmentation research. Additionally, nnU-Net is a modular open-source framework. At present, nnU-Net exhibits several notable advantages: it eliminates the need for manual architecture and hyperparameter engineering, enabling researchers to focus on dataset preparation and model evaluation; it provides strong baseline performance across diverse datasets and tasks, making it a reliable first choice for segmentation problems; and it emphasizes reproducibility and fairness in model evaluation, promoting standardized benchmarking in the medical image analysis community. However, despite these strengths, nnU-Net also has limitations. Its performance is highly dependent on the quality and quantity of annotated training data, which remains a bottleneck in many clinical applications such as lymphoma segmentation. Moreover, although it automates much of the pipeline, it is computationally intensive, requiring substantial time and GPU resources for training, especially on large 3D volumes. Additionally, its automated design may not always be optimal for very specialized tasks, where domain-specific architectural innovations or training strategies might yield better results. Overall, nnU-Net represents a significant advance in medical image segmentation by streamlining model development and establishing a high-performance, generalizable baseline, but it still faces challenges in efficiency, customization, and adaptation to data-scarce or highly specialized clinical scenarios.
Adjustments of network architecture based on research
Objectives
Adjusting and modifying the network architecture of deep learning models based on research objectives refers to making targeted modifications to the model's network structure according to specific research or application needs, aiming to achieve better performance, higher efficiency, or improved adaptability. These adjustments and changes are made with the goal of effectively addressing specific problems or optimizing particular tasks, rather than altering the network structure arbitrarily or without purpose.
For lymphoma segmentation studies, these adjustments to network structure can be categorized into four main types based on research objectives (Figure 4).
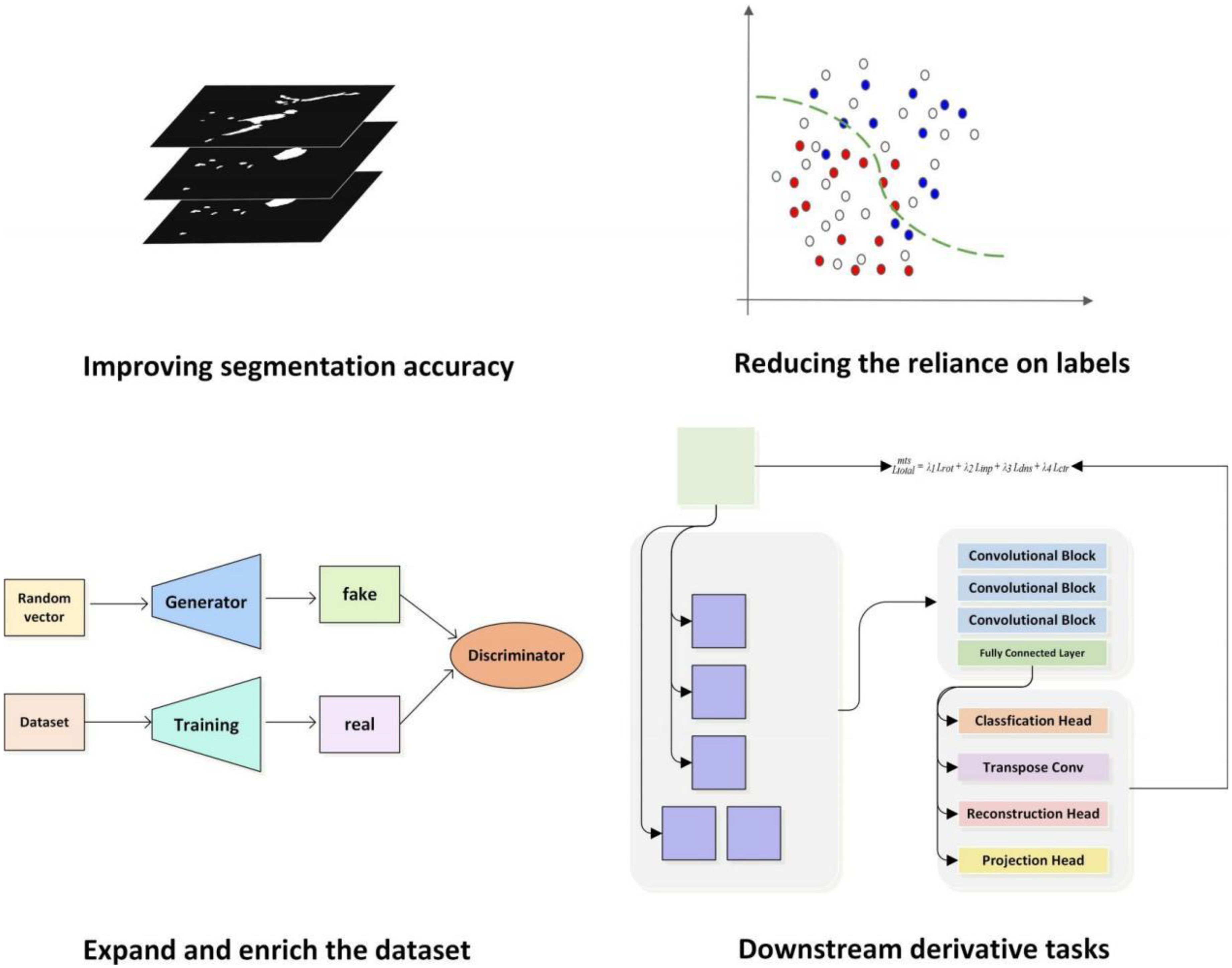
Adjustments of network architecture based on research objectives.
Improving segmentation accuracy
In lymphoma segmentation research, enhancing segmentation accuracy remains a central objective, as it underpins image-based diagnostic assessment, treatment planning, and response monitoring—thereby helping to reduce clinical errors and improve patient outcomes. While histopathological biopsy remains the definitive standard for lymphoma diagnosis, the term “diagnosis” in this study is used in a broader clinical context, encompassing radiological evaluation based on imaging modalities such as PET/CT. In this setting, segmentation plays a pivotal role in visualizing and quantifying tumor burden, thus contributing meaningfully to clinical staging and therapeutic decision making. To support this goal, numerous studies have proposed adjustments to network architectures, which can be categorized into five primary types (Figure 5).
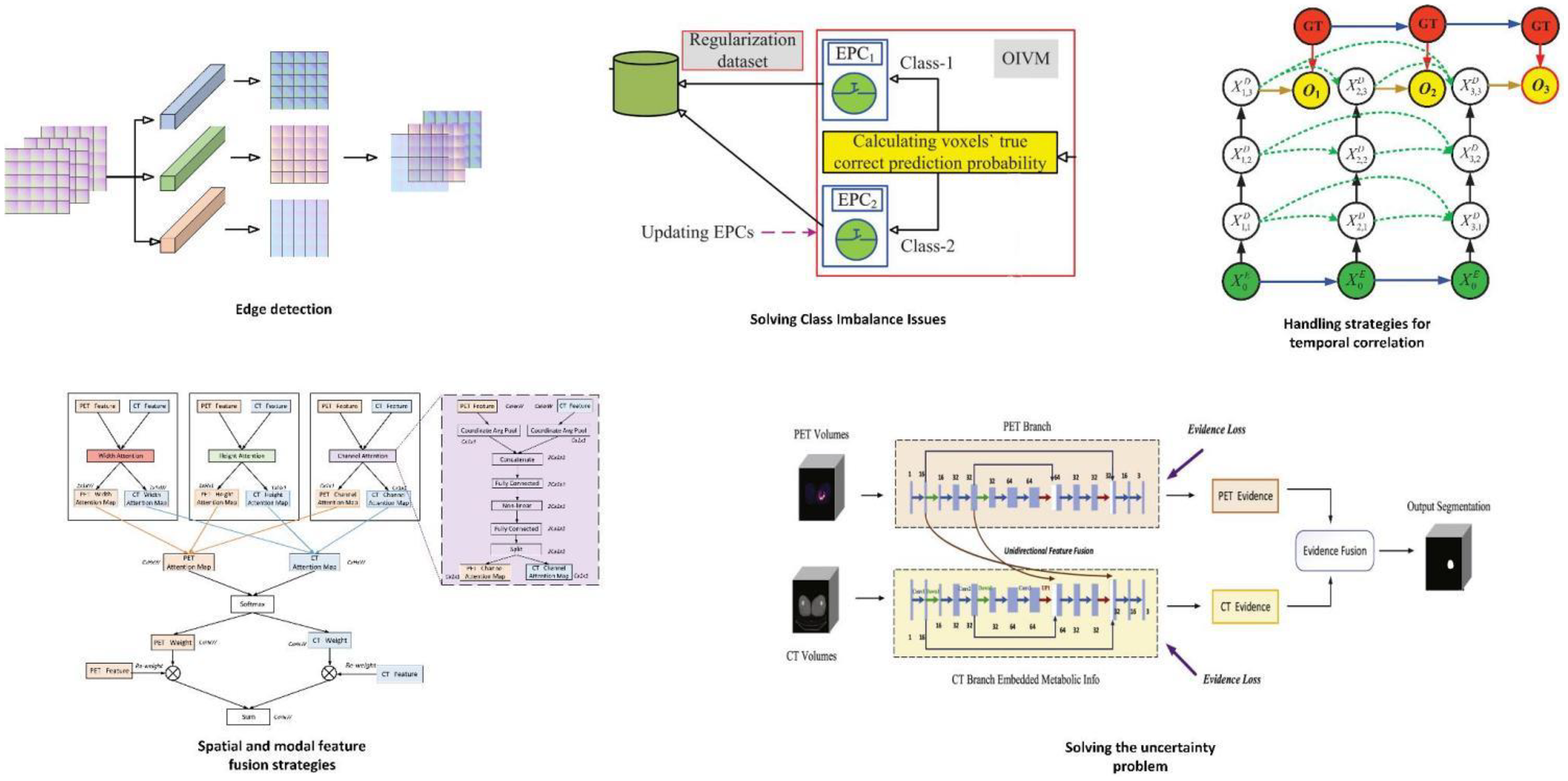
The key components of methods aimed at enhancing segmentation accuracy. In edge detection, Luo et al. 40 have integrated a Multi-Atlas Boundary Awareness module into the backbone network. This module operates on gradient maps, uncertainty maps, and level set maps. To address class imbalance issues, Wang et al. 64 introduced the Prior-Shift Regularization module to enhance the model's sensitivity toward minority classes. In the context of time-series problems, Wang et al. 57 incorporated a Recursive Dense Decoder (RDS-Decoder), which emulates the behavior of a Recurrent Neural Network (RNN) within the decoder. This facilitates the dense reuse of feature information between decoder feature maps at the same scale, capturing temporal dependencies between feature maps. In the process of leveraging spatial and multimodal feature fusion, Yuan et al. 45 designed dual encoder branches, each processing a different modality of images (positron emission tomography (PET) or computed tomography (CT)). Each encoder branch consists of multiple convolutional layers to extract features at various scales for each modality. A fusion learning component generates spatial fusion maps from the features obtained by the two encoder branches, quantifying the contribution of information from each modality. In tackling issues of uncertainty, the methods of Huang et al., 51 Diao et al., 52 and Huang et al. 53 showcase the integration strategy and functionality of the Dempster–Shafer (DS) theory within the model framework.
Edge detection
Improving tumor boundary information is an effective approach to enhancing lymphoma segmentation accuracy. Lymphoma boundaries are inherently uncertain, making it difficult to discern changes in various information classes in medical imaging by naked eye alone. Zhu et al. 43 proposed the use of a cross-shaped structure guidance and boundary optimization strategy to enhance boundary segmentation accuracy. The cross-shaped structure extraction (CCE) method is based on axial context, extracting the cross-shaped structure from annotations of the largest tumor slice, which serves as additional input to assist in segmenting other tumor slices. The researchers also defined a new boundary gradient change-based loss function. This loss function measures the gradient change between predicted boundary pixels and true boundary pixels, extracting boundary gradients using the Sobel operator and using the mean pixel-wise gradient difference as the loss function value. The Boundary Gradient Change Loss Function collaborates with the true tumor boundaries in the decoder to form a Boundary Optimization Module, supervising boundary segmentation. Luo et al. 40 integrated the multi-Atlas boundary awareness (MABA) module into the backbone. This module is based on gradient maps, uncertainty maps, and level set maps. The gradient map utilizes gradient information of pixel intensity changes in the image to detect tumor boundaries. In medical images, the junction between tumors and surrounding normal tissue often exhibits significant intensity changes, which can be captured by computing the image gradient. The uncertainty map focuses on uncertain regions between the tumor and surrounding tissues, which may be challenging to classify during segmentation. By analyzing the predicted probability maps of the network output, the uncertainty map identifies pixels with predicted probabilities close to the classification threshold. The level set map employs a method based on level set evolution to capture the geometric contour and distance information of the tumor. In the level set method, the tumor boundary is viewed as a dynamically evolving curve that gradually approaches the real tumor edge over time. This module focuses on uncertain regions between the tumor and adjacent tissues to obtain potential tumor boundaries. Jurdi et al. 39 further optimized their study based on a boundary loss function, the Boundary Irregularity Index (BI). They defined a Dice coefficient-based BI Index to measure dissimilarity between smoothed segmentation maps and true segmentation maps, directly implementing a differentiable and smooth gradient loss function. During training, minimizing the difference between the BI Index and the true BI Index optimizes the network parameters. Thus, the network tends to generate smoother and more regular boundaries during segmentation, thereby improving segmentation accuracy.
Solving class imbalance issues
In lymphoma research, the pixels occupied by tumors are a minority. This imbalance in foreground-background pixel ratio may bias model predictions toward the majority class (background), leading to insufficient prediction of tumors and thereby reducing segmentation accuracy. Addressing class imbalance is thus another strategy to improve segmentation accuracy. Liu et al. 41 proposed a novel loss function called Class Balanced Dice Loss (CBDL), which tackles class imbalance by considering the effective number of samples. This method protects loss and gradient computations from being dominated by simple majority negative samples. During training, CBDL is integrated into the backbone as the loss function, calculating the CBDL between network outputs and ground truth labels to measure segmentation accuracy. Additionally, they employed a Class Balanced Coefficient Module (CBCM) to compute balance coefficients for each class based on the effective number of samples. Wang et al. 64 introduced the Prior-Shift Regularization (PSR) module to address this issue. The PSR module comprises three main parts: firstly, the Prior-Shift Layer (PSL) performs Online Informative Voxel Mining (OIVM) based on Expected Prediction Confidence (EPC) for each class, extracting informative voxels from the original training dataset for regularization. Secondly, the Regularizer Generation Layer (RGL) generates a Generalized Informative-Voxel-Aware (GIVA) regularizer based on EPC, computed on the regularization dataset created by PSL for model fine-tuning. Lastly, the Expected Prediction Confidence Updating Layer (EPCUL) utilizes L2 loss to update EPC, ensuring adaptive adjustment during training. These steps enable the model to continually adjust attention to informative voxels, thereby enhancing the model's ability to recognize minority classes and addressing class imbalance issues.
Handling strategies for temporal correlation
In lymphoma segmentation research, the datasets predominantly used are PET/CT scans. PET/CT imaging mode does not provide rapid imaging but records metabolic informations of lesions at different time points, fused with plain CT data. Metabolic informations of lesions can vary over time. Normal and diseased tissues may exhibit similar metabolic activities in PET images at a single time point. By comparing images from different time points, physiological high metabolism areas (e.g., brain and bladder) can be distinguished from pathological ones. Additionally, biological characteristics of lesions like metabolic rate and heterogeneity may manifest different patterns over time. Incorporating temporal correlations from image sequences better reflects the morphological nature of lymphoma. Wang et al. 57 introduced the Recurrent Dense Siamese Decoder (RDS-Decoder), simulating recurrent neural network (RNN) behavior within the decoder. This allows dense reuse of feature information among decoder feature maps of the same scale, capturing temporal dependencies between feature maps. Wang modified the backbone network to a multibranch GCU-HGCU encoder based on Unet. GCU, a modified Conv-LSTM unit with ReLU activation replacing Tanh, generates high-resolution cell memory maps accumulating uncompressed information from feedforward GCUs, preserving more image details and contextual information. HGCU combines GCU with hyper-dense connections (HDC), selectively aggregating multimodal information and enhancing the model's capability to handle diverse data modalities. Each branch processes specific modal inputs, with cell memory from encoding branches utilized not only within but across all branches, achieving efficient multimodal information fusion. The decoder is a Recurrent Dense Siamese Decoder (RDS-Decoder), making the model sensitive to sequential information and utilizing potential time correlations between data. Pang et al. 72 introduced concepts of metabolic variance fea-tures (MVF) and metabolic heterogeneity features (MHF). MVF quantifies metabolic differences between tissues and organs at different time points using PET images, aiding in evaluating metabolic differences between lymphoma and other tissues. MHF describes metabolic distributions in high metabolism areas, subdividing these areas at 80 seconds and calculating MVF across different time points to assess metabolic distribution heterogeneity. Integrating MVF and MHF with traditional lymphoma features forms a feature statistical analysis module, guiding lymphoma segmentation and eliminating false positives and negatives. In clinical practice, PET and CT images are generally acquired simultaneously during a single imaging session, with CT primarily used for attenuation correction and inherently aligned with PET data. While acquiring multiple PET scans in rapid succession immediately after radiotracer injection is uncommon due to clinical and logistical challenges. Temporal correlations in lymphoma segmentation research most often pertain to images obtained at distinct clinical time points, such as baseline and interim PET/CT scans during the course of treatment. For instance, in the study by Tie et al., 58 the dual-branch network processes PET/CT images acquired at baseline (PET1) and at an intermediate treatment stage (PET2), leveraging information from the earlier scan to inform segmentation of the later scan. Their model incorporates a longitudinal cross-attention mechanism, where the longitudinal attention window attention (LAWA) module in the encoder applies windowed multihead cross-attention to dynamically focus on correlated regions between PET1 and PET2, while the longitudinal attention gate attention (LAAG) module in the decoder integrates attention coefficients from both time points to enhance feature fusion across longitudinal data. This approach respects clinical imaging protocols and effectively utilizes temporal information to improve lymphoma segmentation.
Spatial and modal feature fusion strategies
PET/CT itself, aside from its temporal features, represents a spatio and multimodal dataset combining metabolic and density informations, and it features multiple perspectives such as coronal, sagittal, and axial views. Given these rich characteristics, strategies for feature fusion have been developed. By employing multiple classifier heads, including auxiliary and target heads, synergistic operations capture and integrate information across different scales and modalities within the fused images. This diversity contributes to a more comprehensive understanding of the data, thereby enhancing segmentation accuracy. Hu et al. 46 proposed a strategy based on multiview and 3D fusion. Initially, three 2D networks based on 2D ResU-Net were trained separately to capture lymphoma image information from different directions. Subsequently, a 3D network based on 3D ResU-Net processed volumetric data. The preliminary results from these approaches were fused using a Conv3D fusion strategy, which not only preserved initial segmentation results from each model but also compensated for spatial information loss in single 2D methods by integrating information from different views. Yuan et al. 45 designed two encoder branches, each handling a modality image (PET or CT). Each encoder branch consisted of multiple convolutional layers to extract features at different scales for each modality. A hybrid learning component generated spatial fusion maps from features obtained from both encoder branches, quantifying the contribution of each modality's information. Through 3D convolution operations, the hybrid learning unit integrated features from different modalities, producing new fused feature maps. These fused maps were subsequently concatenated with PET- and CT-specific feature maps to obtain final fused feature representations at different scales for lymphoma prediction. Diao et al. 47 proposed a method based on spatial compression and multimodal feature fusion attention network. Initially, 3D PET/CT volume data underwent coronal plane compression, generating m 2D images. Using these 2D images, a 3D coronal spatial compression attention volume was obtained, leveraging whole-body anatomical information to generate spatial attention maps guiding precise segmentation. In PFAS-Net, input comprised 2D axial PET/CT slices, guided by coronal spatial compression attention maps obtained during the CSAE-Net phase for segmentation. Additionally, a Multi-Modal Fusion Attention (MFA) module fused PET metabolic information and CT anatomical information across three-dimensions in the network layer. Through attention mechanisms, learned fusion weights weighted the feature maps of different modalities and positions according to learned weights. In the decoding stage's skip connection, the MFA module fused PET features extracted during the encoder phase with CT image features while using spatial compression attention maps obtained during the CSAE-Net phase to guide accurate segmentation.
Wang et al. 66 introduced a Structured Collaborative Learning (SCL) approach to enhance segmentation accuracy. It employed two main components: the Context-Aware Structured Classifier Population Generation (CA-SCPG) module and the Knowledge-Aware Structured Classifier Population Supervision (KA-SCPS) module. CA-SCPG used a lightweight High-Level Context-Aware Dense Connection (HLCA-DC) mechanism to directly enhance feature propagation in target classifier paths. HLCA-DC strengthened feature propagation by densely reusing previous classifier outputs’ prediction maps as inputs for subsequent classifier paths. The KA-SCPS module introduced a novel Knowledge-Aware Dice Loss (KA-DL) to supervise auxiliary classifiers. KA-DL broadcasted knowledge learned by the target classifier to other classifier heads, coordinating the learning process of the classifier group. KA-DL defined positive samples as voxels predicted positively by both the target classifier head and the current classifier head, ensuring optimization of auxiliary classifier heads could explicitly lead to improvements in the target classifier's segmentation performance.
Solving the uncertainty problem
The uncertainty problem in image segmentation encompasses various factors that can affect the accuracy and reliability of segmentation results. In lymphoma segmentation studies, these factors primarily include variability in imaging quality, blurred boundaries between lymphoma and normal tissues, challenges in multimodal data fusion, subjective nature of annotations, and uncertainties in model predictions. To address these challenges, researchers have introduced the Dempster–Shafer (DS) theory, also known as evidence theory or belief function theory, which is a mathematical framework for handling uncertain information. Proposed by Arthur P. Dempster and Glenn Shafer in the 1970s, DS theory aims to provide a more flexible way than traditional probability theory to represent and combine evidence from different information sources. 93 It does not require the additivity condition of probabilities and allows assigning belief to single elements or subsets of the hypothesis space, resembling the evidence gathering process at various levels of abstraction. Moreover, DS theory allows uncertainty to be represented in mass functions and preserves this information during evidence synthesis. The enhancement of segmentation accuracy through DS theory stems from several aspects: it provides a finer representation of uncertainty, including data uncertainty, model uncertainty, and distribution uncertainty. 94 This is crucial in medical image segmentation where data often exhibit noise, blur, and incompleteness. 95 Using Dempster's combination rule, evidence from different modalities (such as PET and CT images) can be effectively fused, addressing information deficiencies or ambiguities that may exist in single-modal data. When conflicting segmentation results arise from different modalities, DS theory offers a mechanism to quantify and resolve these conflicts, rather than relying on simple voting or linear combinations. DS theory also offers a way to quantify the model's confidence in its predictions, thereby improving interpretability and enabling medical professionals and researchers to better understand and trust the model's outputs. Huang et al., 51 Diao et al., 52 and Huang et al. 53 have all utilized DS theory to construct evidence fusion layers, using prototypes in feature space to compute belief functions for each voxel, quantifying the uncertainty of whether lymphoma is present or absent at that location. By leveraging the advantages of PET and CT images, these methods are capable of maintaining high resolution while improving recognition and segmentation accuracy of tumor regions.
Reducing the reliance on labels
In lymphoma segmentation, labels refer to pixel-level annotations of lymphoma regions provided by expert clinicians based on imaging data. These annotations are essential for guiding and optimizing image segmentation algorithms to accurately identify and delineate lymphoma tissues. In supervised learning, such labels serve as ground truth, enabling the model to learn distinctions between lymphoma and surrounding tissues. However, due to the clinical complexity of lymphoma diagnosis—which involves over 100 distinct pathological subtypes with varying radiological and metabolic presentations—obtaining accurate and consistent annotations is particularly challenging. High-quality segmentation requires not only careful interpretation of imaging features but also consideration of underlying pathological results, as different lymphoma subtypes may present differently on PET/CT. This heterogeneity underscores the importance of precise lesion segmentation in supporting the diagnostic and monitoring processes, which are inherently complex. In practice, the scarcity of accurately annotated medical imaging data limits the scalability of supervised learning approaches in medical image analysis, especially in lymphoma-related tasks. To address this issue, researchers have increasingly turned to weakly supervised and semisupervised learning methods as promising alternatives that reduce dependence on extensive manual annotations while preserving or even enhancing model performance. Weakly supervised learning refers to scenarios where the supervision signals are incomplete, inexact, or noisy. In the context of lymphoma segmentation, this may involve using partially labeled images, coarse annotations (e.g. bounding boxes or partial masks), or image-level tags rather than pixel-wise labels. For example, Huang et al. 38 proposed a weakly supervised approach where only a portion of the lymphoma volume in each patient was manually annotated, while the rest of the tumor region remained unlabeled. To compensate for the lack of dense annotation, they introduced multiscale feature consistency constraints, enforcing the network to learn consistent representations across predicted tumor regions and the sparse labeled regions. The model also employed cosine similarity to compute feature-level distances between tumor and normal tissue, improving discriminative learning in the absence of dense labels. Their training framework incorporated a combination of supervised loss, deep supervision loss, and a regularization loss—where the latter enforced spatial and semantic consistency between weakly labeled and predicted regions, facilitating robust training from partially annotated data. On the other hand, semisupervised learning leverages both labeled and unlabeled data during training. It assumes that even in the absence of manual labels, meaningful patterns can still be extracted from the data distribution. Yousefirizi et al. 37 developed a novel semisupervised segmentation framework tailored for medical imaging by incorporating a loss function derived from the Fuzzy C-Means (FCM) clustering objective. Unlike traditional Dice or cross-entropy loss, which rely on crisp class assignments, FCM captures the inherent fuzziness and uncertainty in medical image classification, making it more suitable for modeling soft boundaries in tumors. Their method enabled the model to use soft cluster memberships of unlabeled voxels in conjunction with labeled samples to better learn decision boundaries, leading to improved generalization. By fusing supervised and unsupervised loss components, the model was able to maintain high segmentation accuracy even when trained on a limited number of annotated cases. In summary, both weakly and semisupervised learning strategies offer valuable solutions to the data annotation bottleneck in lymphoma segmentation. Weak supervision allows models to benefit from coarse or partial annotations by introducing auxiliary constraints and consistency mechanisms. Semisupervised learning, meanwhile, utilizes unlabeled data to refine model representations and decision boundaries through hybrid loss functions and regularization. These approaches reduce reliance on exhaustive manual annotation while enabling the development of accurate, scalable, and clinically meaningful lymphoma segmentation systems.
Expand and enrich the dataset
Characteristics of lymphoma image data and the requirements for training deep learning models exhibit conflicting situations: firstly, medical images face ethical and acquisition challenges, limiting the scale of constructed datasets. Moreover, due to the difficulty of annotation and the diverse types of lymphoma, datasets often do not cover all types comprehensively, leading to limitations. On the other hand, deep learning models demand large-scale datasets with diverse content to achieve better clinical generalization and robustness. Therefore, expanding and enriching datasets without additional case collection is a research direction. One approach to address this issue is integrating generative adversarial networks (GANs) into the backbone network of models. GANs, introduced by Goodfellow et al. 96 in 2014, consist of two key components: the generator and the discriminator. These components competitively train during the process, forming an adversarial training mechanism to enhance model performance. The generator aims to produce realistic data (e.g., images, videos, or audio) from random noise input via neural networks. Meanwhile, the discriminator's task is to distinguish between fake data generated by the generator and real data from the dataset, maximizing its ability to correctly classify them. Throughout training, the generator and discriminator compete against each other: the generator attempts to deceive the discriminator, while the discriminator strives not to be deceived. This adversarial training drives the generator to produce higher-quality data. Conte et al. 54 and Shi et al. 69 have both employed this approach to enrich limited lymphoma data content and verified the feasibility of generated images.
Downstream DerivativeTasks
In lymphoma segmentation research, analyzing segmentation results for downstream tasks is also a research direction closely related to clinical applications, often involving disease diagnosis, treatment, and prognosis. The metrics formed from such studies are not exclusive to deep learning models; these tasks can also be accomplished using traditional machine learning or statistical methods, providing supervised signals for model training. This training strategy based on clinical practical issues enhances deep learning models’ clinical generalization ability. Downstream derivative tasks in these studies include identifying and detecting small lymphoma lesions, calculating TMTV and MTV, 97 and linking them to disease prognosis. From the perspective of network structure adjustment, there are two main approaches to achieve these goals: multitask learning and cascade learning. Both are common techniques in the fields of deep learning and machine learning, where certain parts of the model can be reused to solve multiple problems or tasks across different stages. Both methods leverage the concept of feature sharing, where features learned for one task can provide useful information for others. By sharing model components or features, both approaches can improve learning efficiency and reduce computational resources. They can be implemented within the framework of deep learning, using neural networks to tackle complex tasks. However, they differ fundamentally, which will be elaborated based on specific research in the following sections.
Multitask learning
Multitask learning (MTL) typically refers to the simultaneous learning of multiple related or unrelated tasks, where these tasks may share underlying feature representations but have separate objective functions optimized jointly to improve overall performance across tasks. In medical image analysis, particularly in lymphoma segmentation, MTL has emerged as a powerful paradigm, allowing a single network to handle tasks such as lesion segmentation and prognosis prediction in a unified framework. A common approach involves using a shared encoder for joint feature extraction, followed by task-specific decoder branches, and training with a composite loss function that balances multiple task-specific objectives. For example, Liu et al. 35 proposed a U-Net-based MTL framework where shared features supported both lesion segmentation and the prediction of 2-year event-free survival (2y-EFS), with a weighted sum of segmentation and classification losses facilitating joint optimization. Despite its potential, MTL faces several challenges, including task conflicts and negative transfer due to divergent learning objectives, imbalanced task difficulty and label availability (pixel-level segmentation vs patient-level survival data), and limited interpretability, which can hinder clinical trust. Solutions such as dynamic loss weighting, curriculum learning, task-specific modules, and integration of explainable AI (artificial intelligence) techniques like attention visualization are being explored to mitigate these issues. Nevertheless, the benefits of MTL are significant: it enhances feature representation through shared learning, improves data efficiency when annotated data is limited, and aligns with real-world clinical workflows by enabling end-to-end models capable of both anatomical analysis and prognostic insight. Future directions include incorporating uncertainty-aware loss functions, integrating multimodal data such as clinical or genomic information, and developing continual learning mechanisms to adapt to evolving clinical knowledge. Overall, MTL offers a promising strategy for advancing lymphoma segmentation and related tasks, though it requires careful design and validation to ensure robustness, fairness, and clinical usability.
Cascade learning
Cascade learning focuses on sequentially addressing a series of tasks where the output of each task serves as the input for the next task, forming a processing pipeline. 98 Networks in cascade learning consist of multiple independent models, each dedicated to a specific task, connected sequentially. Each model may be trained independently, and the training process can be conducted in stages where the output of each stage is used as the input for the next stage. This approach is also a primary strategy for achieving downstream derivative tasks in lymphoma segmentation research, including tasks such as TMTV and MTV computation, where the segmentation results of the previous part are fed into the next part of the network. Jemaa et al. 30 cascaded Unet and Vnet to sequentially achieve whole-body-region-organ-lesion segmentation and TMTV computation goals. Yousefirizi et al. 34 in their study cascaded two 3D U-Nets using a soft voting module, achieving the above tasks under a semisupervised setting. In summary, multitask learning emphasizes solving multiple tasks simultaneously within a single model, whereas cascade learning emphasizes sequentially solving a series of tasks, where each task may be handled by different models or model components. Both approaches are effective methods for enhancing learning efficiency and performance but are applicable in different scenarios and for different requirements.
Model performance and included metrics
In the task of lymphoma segmentation based on deep learning, evaluation metrics are crucial for quantifying segmentation performance. They reflect the algorithm's efficacy and the accuracy of segmentation results. The following section will summarize the evaluation metrics applied in current research and compile them, along with experimental results, in Table 2.
Dice similarity coefficient
DSC is a statistical tool used to evaluate the similarity or overlap between two samples, particularly in medical image analysis for assessing the accuracy of image segmentation. The DSC value ranges between 0 and 1, where a value closer to 1 indicates a higher overlap between the two samples and thus a more accurate segmentation result.
Here, A represents the ground truth, B the algorithm's predicted segmentation region, |A ∩ B| denotes the intersection of A and B, which is the size of the overlapping area between the two regions. |A| and |B| represent the total area or volume of A and B, respectively.
Jaccard Index
The Jaccard Index is a measure of similarity between two sets. In fields such as medical image segmentation, the Jaccard coefficient is commonly used as an evaluation metric to quantify the overlap between segmentation results and ground truth annotations. The expression is given by: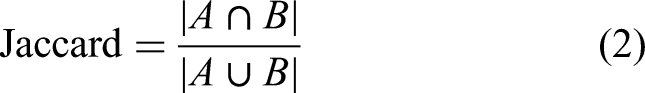
Here, A represents the set of ground truth annotations, and B the set of predicted segmentation results by the algorithm. |A ∩ B| denotes the size of the intersection of sets A and B, which is the number of elements common to both sets. |A U B| denotes the size of the union of sets A and B, which is the total number of elements in both sets combined. The Jaccard coefficient ranges between 0 and 1, where a value closer to 1 indicates a higher degree of overlap between the sets, reflecting better consistency between the segmentation result and the ground truth annotation. The Jaccard coefficient provides a simple and intuitive method to assess the performance of segmentation algorithms, particularly suitable for binary image segmentation tasks where sets can be simplified to sets of pixels.
Accuracy
ACC is a commonly used evaluation metric in classification tasks, used to measure the model's correctness in classifying all samples. It is defined as the proportion of correctly classified samples to the total number of samples. The formula for accuracy is given by: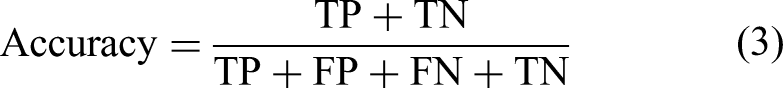
Here, TP (True Positives) represents the number of samples that are correctly identified as positive samples. TN (True Negatives) denotes the number of samples that are correctly identified as negative samples. FP (False Positives) indicates the number of negative samples incorrectly classified as positive, and FN (False Negatives) indicates the number of positive samples incorrectly classified as negative. Accuracy values range from 0 to 1, with higher values typically indicating better classification performance. However, in the context of medical image segmentation—particularly in modalities such as PET, CT, and MRI where there is often a severe class imbalance between background and lesion voxels—accuracy may not serve as a reliable performance metric. Even models with suboptimal segmentation performance can yield deceptively high accuracy scores due to the predominance of background voxels. While accuracy is sometimes reported, it provides limited insight in such scenarios. Therefore, overlap-based metrics like the Dice coefficient and Jaccard index, as well as distance-based metrics such as the Hausdorff distance, are more informative and widely accepted for evaluating segmentation quality, especially in the presence of class imbalance.
Sensitivity
Sensitivity, also known as True Positive Rate (TPR) or Recall, is a crucial metric for evaluating the performance of classification models, particularly in fields like medical image segmentation and disease diagnosis. It measures the model's ability to correctly identify positive samples (e.g. vascular regions). The formula for sensitivity is defined as:
Sensitivity values range between 0 and 1, where higher values indicate a stronger ability of the model to recognize positive samples. In medical imaging, higher sensitivity implies that the model can effectively detect relevant features or regions of interest, which is critical for accurate segmentation and disease diagnosis.
Specificity
Specificity, also known as True Negative Rate (TNR), is a critical metric for evaluating the performance of classification models, particularly in fields such as medical image segmentation and disease diagnosis. It measures the model's ability to correctly identify negative samples (such as nonlesion areas). The expression is given by:
Specificity values range from 0 to 1, where a higher value indicates stronger capability of the model to recognize negative samples. For instance, in medical image segmentation, a high specificity means that nearly all true nonlesion areas are correctly identified, thereby reducing the likelihood of misdiagnosis.
Precision
Precision is a crucial evaluation metric in classification tasks, used to measure the proportion of correctly predicted positive samples among all samples predicted as positive by the model. A high precision indicates that a higher proportion of positively predicted samples are indeed true positives, implying fewer false positives (FP). The formula for precision is given by:
Precision values range from 0 to 1, where a higher value signifies greater accuracy of positive predictions by the model. In practical applications like medical image segmentation, precision helps us understand the model's accuracy in predicting regions of interest, thereby evaluating its performance.
Mean squared error
Mean squared error (MSE), an index measuring the discrepancy between predicted values and actual values, represents the average of the squared prediction errors. The formula for calculating MSE is as follows:
Herein, yi denotes the true value,
Mean absolute deviation
Mean absolute deviation (MAD) is a statistical measure that quantifies the amount of variation or dispersion from the mean in a set of values. It represents the average distance of each data point from the mean. MAD is calculated by taking the absolute value of the difference between each data point and the mean, summing these absolute values, and then dividing by the number of data points. The formula for MAD is:
Hausdorff distance
Hausdorff distance (HD) is used to measure the maximum distance between two sets, and here it is employed to assess the maximum inconsistency between segmentation boundaries and true boundaries.
99
The formula for HD is:
Area under the curve
Area under the curve (AUC) is a widely used metric to evaluate the performance of binary classification models, particularly in medical image segmentation and diagnostic tasks. It represents the area under the receiver operating characteristic (ROC) curve, which plots the TPR(or sensitivity) against the FPR at various threshold settings. AUC quantifies the model's ability to distinguish between positive and negative classes, where a value of 1.0 indicates perfect discrimination and 0.5 corresponds to random guessing. The ROC curve is defined by the following:
The AUC is then computed as the integral of the ROC curve: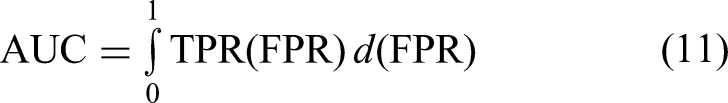
In practice, AUC is often approximated numerically using the trapezoidal rule based on discrete threshold points.
Discussion
Clinical prospects of deep learning strategies for lymphoma segmentation
Research in medical image processing derives its materials from clinical scenarios, driven by specific clinical problems. Throughout this article, most studies focus on evaluation metrics related to natural image segmentation, while some researchers translate clinical issues into technical problems for computer-based research (Figure 6). In this section, we will discuss these two aspects: the application of lymphoma segmentation research based on deep learning strategies in clinical settings, and its potential translational prospects.

Workflow of a clinically application-oriented integrated lymphoma segmentation method.
Diagnosis and staging
The preliminary evaluation of lymphoma through imaging, particularly PET/CT, plays a pivotal role not only in diagnosis but also in disease staging, which is central to treatment planning. Contemporary research on lymphoma segmentation has decomposed this task into multiple subproblems, such as enhancing segmentation accuracy and refining automated decision-support pipelines. As discussed in previous sections, numerous efforts have been made to improve network architectures to address these objectives. Recent studies have explored challenges including temporal and spatial correlations in imaging data, feature fusion strategies, class imbalance, and uncertainty estimation. While these theoretical advancements hold promise for improving lymphoma-related imaging analysis, large-scale clinical validation remains limited. This limitation is primarily due to the difficulty of acquiring comprehensive lymphoma datasets and the presence of over one hundred histological subtypes, 2 each exhibiting significant heterogeneity in radiological and metabolic features. Moreover, current public and private datasets fail to represent the full spectrum of lymphoma subtypes, thereby constraining the clinical translation of automated segmentation methods. Accurate imaging-based staging—particularly using PET/CT—is critical for guiding treatment strategies, such as determining therapy intensity or eligibility for clinical trials, highlighting the necessity for robust, generalizable segmentation models.
Disease monitoring and prediction
Recent research has increasingly integrated lymphoma segmentation into multitask and cascade learning frameworks to support downstream tasks such as disease monitoring and prognostication. In multi-task learning settings, segmentation is jointly optimized with related objectives—such as classification, out-come prediction, or survival estimation—allowing the model to benefit from shared representations and improve overall performance. Cascade learning, on the other hand, typically involves using the output of the segmentation network—such as lesion boundaries or volumetric measurements—as input to subsequent modules, including radiomic feature extraction or risk stratification algorithms. A key quantitative parameter in this context is TMTV, derived from PET/CT imaging. TMTV quantifies tumor metabolic activity based on FDG uptake, 97 and its accurate computation relies on precise segmentation of lymphoma lesions. 73 ROC curve analysis can be used to determine optimal TMTV thresholds for patient risk stratification. Baseline PET/CT scans acquired before therapy are used to assess TMTV and predict prognosis, while interim scans help monitor treatment response and guide therapeutic adjustments. TMTV also serves as an independent prognostic factor in multivariate Cox regression models. Additional imaging-derived indicators—such as tumor dissemination patterns, 100 metabolic heterogeneity, 101 and total lesion glycolysis 102 —further contribute to patient stratification. The development of segmentation-guided, multi-indicator, and multitask models presents a promising direction for advancing personalized lymphoma management.
Limitations of technology
Despite the promising translational potential of deep learning-based lymphoma segmentation, several technical challenges must be addressed to enable reliable and widespread clinical deployment. A primary concern is the limited generalizability of deep learning models across different clinical sites. Models trained on data from a single institution or scanner often underperform on external datasets due to domain shifts arising from variations in imaging protocols, scanner manufacturers, and reconstruction parameters. To overcome this, recent research has explored strategies such as domain adaptation, 103 multicenter collaborative training, and federated learning 104 to enhance cross-site robustness. Another significant challenge is the severe class imbalance inherent in lymphoma segmentation, where lesions often occupy only a small proportion of the total imaging volume. This imbalance can bias models toward the background class, reducing their sensitivity to lesion regions. To mitigate this, approaches such as specialized loss functions 105 and lesion-aware sampling strategies have been proposed and are actively being refined. Furthermore, the high computational demands of deep learning models—particularly for processing high-resolution 3D volumetric PET/CT data—limit their practical deployment in real-time and resource-constrained clinical environments. Although methods such as model compression, quantization, and network pruning offer potential solutions, achieving efficiency without compromising accuracy remains a key bottleneck. Interpretability also poses a major barrier to clinical adoption. Most deep learning models operate as black boxes, providing little insight into their decision-making processes. Since clinicians require transparent, explainable outputs to validate and trust automated suggestions, recent efforts have focused on integrating attention mechanisms, saliency maps, and feature attribution methods. However, the clinical relevance and robustness of these interpretability techniques remain under investigation. Finally, reproducibility is a longstanding issue in medical image analysis. Variations in preprocessing steps, model architectures, training settings, and evaluation metrics often make it difficult to reproduce and fairly compare results. The adoption of standardized frameworks—such as nnU-Net—and public bench-marks and challenges are crucial for advancing reproducible and comparable research in this field. Together, these limitations highlight the need for continued technical innovation and standardization to enhance the robustness, fairness, and clinical utility of deep learning approaches for lymphoma segmentation.
Conclusion
In summary, deep learning strategies are advancing the progress of lymphoma segmentation research. The clinical motivation underlying this research provides new directions and perspectives for lymphoma segmentation studies. Future research in lymphoma segmentation will increasingly align with medical realities, aiming for the transformation of technology into practical applications.
Footnotes
Contributiorship
WL was involved in conceptualization, methodology, investigation, and writing—original draft; FY in writing—review and editing and formal analysis; PT in writing—review and editing, methodology, and investigation; TZ in writing—review and editing and conceptualization; andWS in writing—review and editing, supervision, and project administration.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Jilin Province Health Talent Special Fund (grant no. JLSWSRCZX2023-14) and the Natural Science Foundation of Jilin Province (grant no. YDZJ202301ZYTS065).
Guarantor
Prof. Weizhang Shen takes responsibility for the integrity of the work as a whole, from inception to published article.
Informed consent
Not applicable.
Peer review
This manuscript was externally peer reviewed. All peer review was conducted independently and anonymously.
