Abstract
Objective
Accurate pathology reports are crucial for the diagnosis and treatment planning of cancer patients. However, these reports are prone to errors due to time pressures, subjective interpretation, and inconsistencies among professionals. Addressing these errors is vital for improving oncology care outcomes. Artificial intelligence (AI) systems, such as GPT-4, offer the potential to enhance diagnostic accuracy and efficiency.
Methods
A total of 700 malignant tumor pathology reports were collected from four hospitals. Of these, 350 reports had deliberate errors introduced by a senior pathologist, mimicking real-world reporting challenges. Error detection performance was evaluated by comparing GPT-4 to six human pathologists (two seniors, two attending pathologists, and two residents). Key metrics included error detection rates with Wilson confidence intervals and processing time per report.
Results
GPT-4 detected 88% of errors (350/400; 95% CI: [84, 91]), compared to a 95% detection rate by the top senior pathologist (382/400; 95% CI: [93, 97]). GPT-4 significantly reduced the average processing time to 4.03 seconds per report, compared to 65.64 seconds for the fastest human pathologist. However, GPT-4 exhibited a higher rate of false positives (2.3%; 95% CI: [1.52, 3.01]) compared to the best-performing senior pathologist (0.3%; 95% CI: [0.01, 0.91]).
Conclusions
GPT-4 demonstrates substantial potential in improving the efficiency and accuracy of pathology error detection, which could accelerate clinical workflows and enhance cancer diagnostics. However, its higher false-positive rate emphasizes the need for human oversight to ensure safe implementation in clinical practice.
Keywords
Introduction
In modern oncology, pathology reports play a pivotal role in the accurate diagnosis, staging, and treatment planning for cancer patients. However, errors in these reports can lead to misdiagnoses, inappropriate treatment strategies, and potentially harmful patient outcomes, highlighting the critical need for precise and reliable reporting. 1 Common challenges include clerical errors, data misinterpretation, and inconsistencies resulting from subjective judgment and workload pressures.2–4 As the volume of pathology reports continues to rise, particularly in oncology where timely and accurate diagnoses are crucial, the risk of human error escalates, especially under time constraints.2–5
Given these challenges, efficient methods for error detection in pathology reports are urgently needed. Traditional error detection approaches, though thorough, are labor-intensive, time-consuming, and prone to variability due to their reliance on manual review by pathologists.6–8 Recent advancements in artificial intelligence (AI), particularly the development of large language models (LLMs) like GPT-4, offer promising solutions to these challenges by automating error detection and improving the accuracy of medical documentation.9–11
GPT-4, with its advanced natural language understanding and generation capabilities, is capable of processing complex medical texts and identifying discrepancies within pathology reports. Its ability to recognize and correct errors, such as clerical mistakes and incorrect terminology, has significant potential to enhance diagnostic accuracy in cancer care.12,13 However, the effectiveness of GPT-4 can vary depending on the type of prompts used. While this study primarily employs zero-shot prompting, exploring alternative strategies such as one-shot or chain-of-thought prompting could further optimize its performance and reduce the incidence of false positives.12–14
The adaptability of GPT-4 across diverse medical contexts makes it a versatile tool for detecting errors in oncology-related pathology reports. Nevertheless, it is essential to benchmark its performance against traditional error detection methods to fully assess its capabilities and limitations. By integrating AI technologies with domain-specific medical expertise, this research aims to develop an efficient, accurate, and scalable approach to automating error detection, ultimately enhancing the reliability of cancer diagnostics.15,16
Furthermore, the use of GPT-4 for error detection could alleviate the workload of oncologists and pathologists, offering significant time savings while also serving as an educational tool for medical trainees. By identifying and correcting errors in real-time, AI technologies like GPT-4 could support faster and more accurate cancer diagnoses, contributing to improved patient outcomes.17–19
The primary objective of this study is to evaluate the efficacy of GPT-4 in detecting errors within oncology pathology reports and to explore its potential for integration into clinical practice. Through this work, we seek to contribute to the advancement of AI-driven solutions in cancer diagnostics, aiming to improve both the precision and efficiency of medical reporting, thereby positively impacting patient care outcomes.
Materials and methods
Prompt engineering
The study primarily utilized zero-shot prompts, where GPT-4 received instructions without prior examples to guide its error detection process. This approach was selected because it reflects a realistic clinical deployment scenario, especially where domain-specific annotated data are unavailable. Testing GPT-4 in its unmodified form allows for an unbiased evaluation of its baseline performance.
We recognize that this is one of several possible prompting strategies. Future studies should explore and compare other methods—such as few-shot prompting, chain-of-thought reasoning, or lightweight fine-tuning—to further enhance performance.
GPT-4 model configuration and training data
GPT-4, developed by OpenAI, is a large-scale language model used in this study. The specific version accessed via the OpenAI API was the default model available during the research period.
Model Configuration:
Architecture: GPT-4 is based on the Transformer architecture, consisting of multiple layers of self-attention and feed-forward networks. While OpenAI has not disclosed the exact number of parameters, GPT-4 is among the largest models available, with billions of parameters contributing to its robust language understanding capabilities. Input Configuration: The input consisted of pathology reports divided into sections such as “Macroscopic findings” and “Pathological diagnosis,” formatted for consistency. A zero-shot prompting approach was employed, with explicit instructions tailored for error detection.
Training Data:
Pretraining: GPT-4 was pretrained on a diverse text corpus from the internet, covering various domains, languages, and styles, enabling it to develop a broad understanding of language and contextual relationships. Domain Adaptation: Although no specific fine-tuning for pathology was conducted, GPT-4's general knowledge base includes medical literature, which aids in recognizing and understanding medical terminology and concepts relevant to this study.
Fine-Tuning Procedures:
Zero-Shot Approach: GPT-4 was used in a zero-shot manner, with responses generated based on its pre-existing knowledge without further adaptation or training on the dataset used in this study.
Limitations and Implications for Reproducibility:
Model Reproducibility: While GPT-4's performance offers valuable insights, exact replication may vary due to updates to the GPT-4 model and API configurations. Researchers aiming to replicate this study should use the same version of GPT-4 and meticulously document model configurations and prompts.
Data collection and preprocessing
This was a retrospective, multi-center validation study conducted across four hospitals in China: Guizhou Provincial People's Hospital, the Third Affiliated Hospital of Sun Yat-sen University, the Third Xiangya Hospital of Central South University, and Jiangxi Cancer Hospital. The data collection took place between October and December 2023. All pathology reports were originally written in Chinese. A total of 700 original pathology reports on malignant tumors from 14 different organs were collected from these hospitals. These reports were randomized using a freeware research data randomization tool and divided into two equal groups: 350 correct reports and 350 manipulated reports.
In the manipulated group, a total of 400 errors were deliberately introduced by a senior pathology expert, with up to three errors per report (Figure 1). These errors were designed to reflect common real-world pathology reporting issues, with each of the five error categories—clerical errors, improper use of terminology, missing information, interpretation or diagnostic errors, and data inconsistency—represented by 80 instances, ensuring a balanced assessment across different error types (Figure 1). All reports were anonymized prior to analysis, and written informed consent was obtained from all patients or their legal guardians.
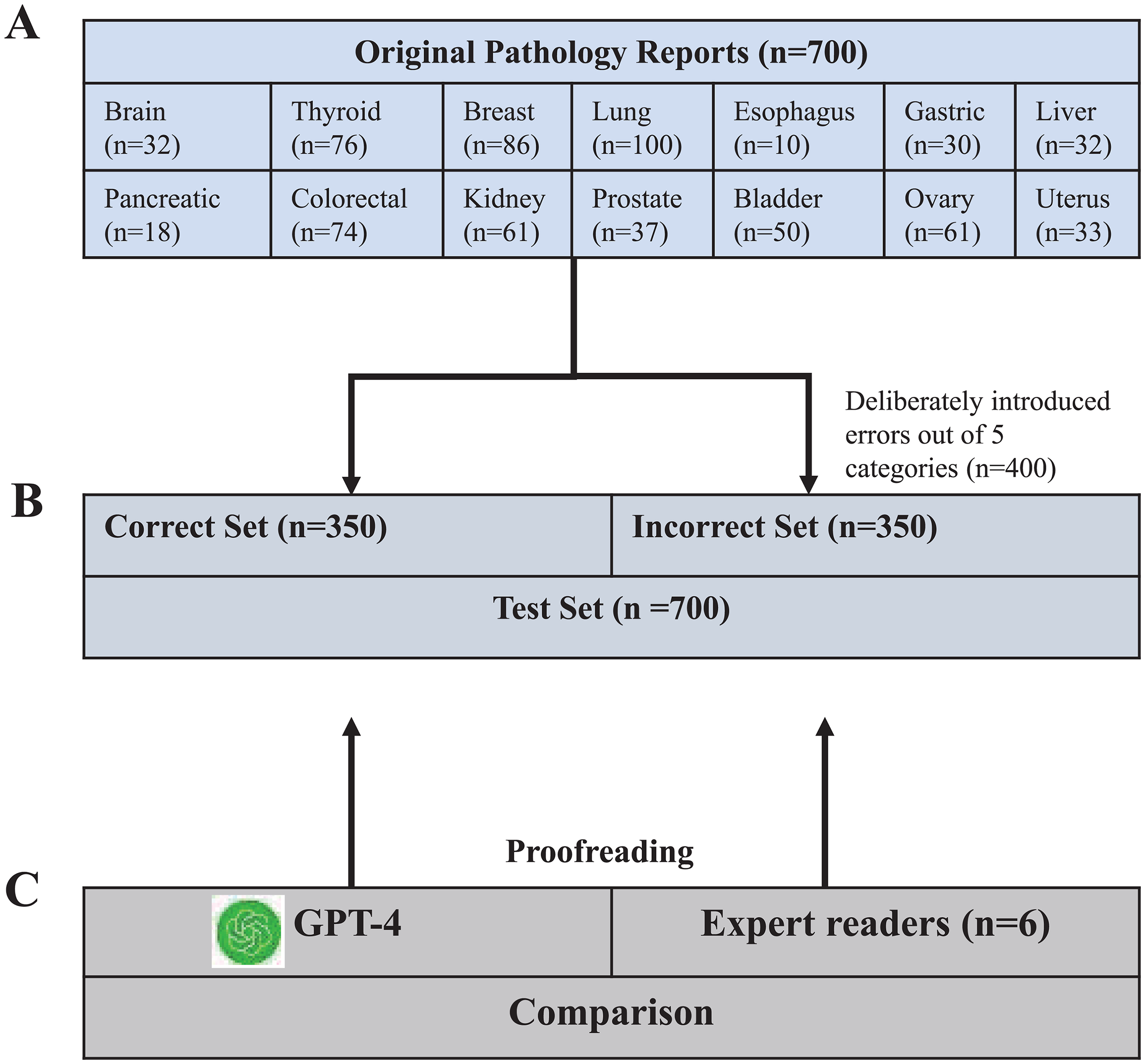
Study flowchart. (A) Initially, 700 original pathology reports from 14 organs were selected. (B) These were then randomized into two sets: a correct set and an incorrect set, each containing 350 pathology reports. Within the incorrect set, 400 errors across five categories (Clerical errors, Improper use of terminology, Missing information, explain or diagnose errors, Data inconsistency) were deliberately introduced, with a maximum of three errors per case. (C) GPT-4 and six doctors were tasked with evaluating each pathology report to identify potential errors, allowing for a comparative analysis of their performance.
Error categorization and definition
The introduced errors were categorized based on existing literature and real-world pathology practices into five main categories2–4:
Clerical errors: This category comprises the most common errors, such as typos, punctuation errors, misspellings of words, etc. For instance, the term “carcinoma” may be incorrectly transcribed as “cardinoma.” Additionally, inaccuracies may arise from improper copying and pasting of previous reports, leading to the inclusion of irrelevant or incorrect information in the report. Improper use of terminology: Pathologists may inaccurately employ technical terms or use inappropriate language when describing histological features. An example of this would be incorrectly labeling a “benign” lesion as “malignant,” which could lead to incorrect treatment decisions. Missing information: Pathology reports may omit crucial diagnostic details, such as tumor size, grade, stage, etc., which are pivotal for subsequent treatment planning. For instance, failing to mention the margin status of a resected tumor could result in inadequate patient management. Interpretation or diagnostic errors: This type of error may stem from misinterpretation of tissue samples or inadequate understanding of pertinent clinical information. An example would be incorrectly diagnosing a benign lesion as malignant, or vice versa, based on histopathological findings. Data inconsistency: The data within the report may not align with information present in laboratory records or other medical documentation. For example, a pathology report might list the wrong patient demographic information, or there could be discrepancies between the reported findings and the documented clinical history.
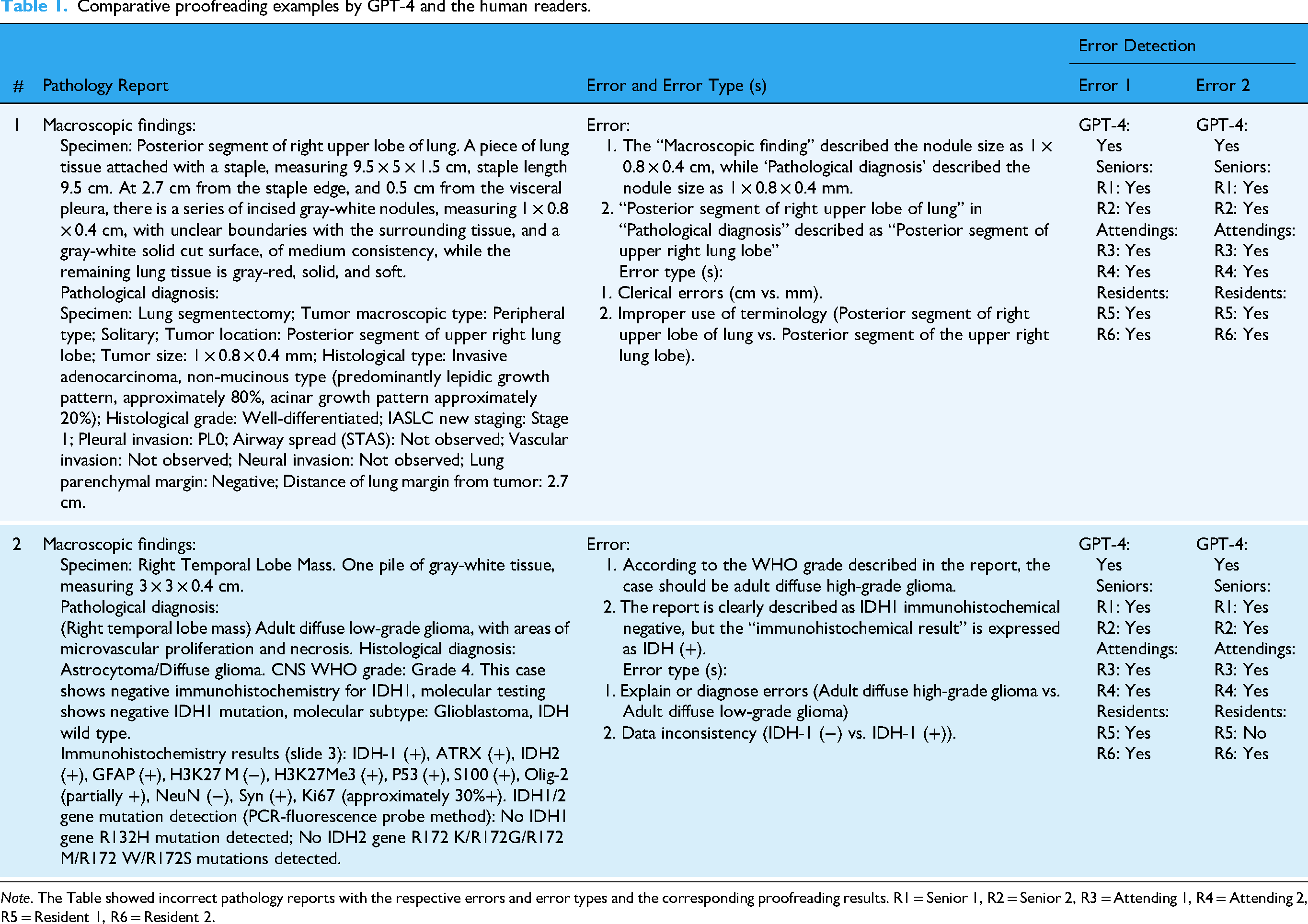
These categories were defined to capture the most common and clinically significant types of errors that occur in pathology reports. To ensure clarity and consistency, these error categories were consistently referenced throughout the study, with specific examples provided to enhance understanding. The role of the senior pathology expert was crucial in both the introduction and categorization of these errors to ensure they accurately represented realistic challenges faced in pathology reporting (Table 1).
Comparative proofreading examples by GPT-4 and the human readers.
Role of pathologists in the study
Three experienced pathologists (Z.Y., J.Y.J., and S.R.N.B., with 20, 20, and 12 years of experience, respectively) were involved in the validation and classification of the errors. Initially, each pathologist independently verified the reports to ensure that their evaluations were unbiased and not influenced by the others’ judgments. After these independent assessments, the pathologists convened to discuss any discrepancies in their findings. Each pathology report's final error classification was determined through consensus among these three pathologists, and any disagreements during this process were resolved through thorough discussion until consensus was reached, ensuring consistency in error categorization.
Error detection process using GPT-4
The time taken by GPT-4 to correct each pathology report was measured from the moment the prompt was sent to when the final response was received. This duration was assessed for 70 randomly selected reports of varying lengths.
Statistical analysis
All analyses were conducted using software including R software (version 4.3.2) and Python (version 3.12). Outcome measures included the number of correctly detected errors, precision, recall, F1-score, Wilson confidence intervals, and time taken per report.
To compare the performance of GPT-4 with that of the pathologists, confidence intervals were the primary formal inferential tool used, supplemented by McNemar's test for paired comparisons. Wilson confidence intervals were calculated for all proportion estimates due to their better performance compared to Wald intervals, especially with smaller sample sizes. This approach ensures more accurate interval estimates for the error detection rates.
Precision, recall, and F1-score were calculated to provide a more comprehensive understanding of GPT-4's performance, each accompanied by their respective confidence intervals to assess the range of expected performance. Precision was calculated as the number of true positive errors detected by GPT-4 divided by the sum of true positives and false positives. Recall was calculated as the number of true positive errors detected by GPT-4 divided by the sum of true positives and false negatives. The F1-score, which balances precision and recall, was calculated as the harmonic mean of precision and recall.
The performance of GPT-4 and the pathologists was summarized using 95% Wilson confidence intervals (CIs) for the detection rates, precision, recall, and F1-score. Comparisons between GPT-4 and the pathologists were primarily descriptive, focusing on confidence intervals to provide insight into the reliability and precision of the results, rather than relying heavily on
The average time to process a GPT-4–corrected pathology report was compared with those of the pathologists using paired-sample
Results
Performance of GPT-4 and human pathologists in error detection
GPT-4 demonstrated strong performance in detecting errors within pathology reports, with a detection rate of 88% [350 of 400; 95% CI: 84, 91], a precision of 0.975, recall of 0.875, and F1-score of 0.922 (Tables 1 and 5). McNemar's test results were considered supplementary to the primary focus on confidence intervals, which provide a more comprehensive understanding of the variability in error detection rates between GPT-4 and each pathologist. In comparison, the best-performing senior pathologist achieved a higher detection rate of 95% [382 of 400; 95% CI: 93, 97], highlighting the expertise of experienced human pathologists (Table 1). The resident pathologists had the lowest detection rates, with the worst-performing resident achieving 81% [325 of 400; 95% CI: 77, 85]. Moreover, GPT-4 exhibited a higher rate of incorrectly flagged reports at 2.3% [9 of 400; 95% CI: 1.5, 3.0], compared to the best-performing senior pathologist's rate of 0.3% [1 of 400; 95% CI: 0.01, 0.9].
Comparison of detection rates and error metrics
In the overall analysis, while GPT-4's detection rate of 88% was lower than that of the best-performing senior pathologist (95%), its performance was still notable and comparable to that of attending pathologists, with no significant difference observed (
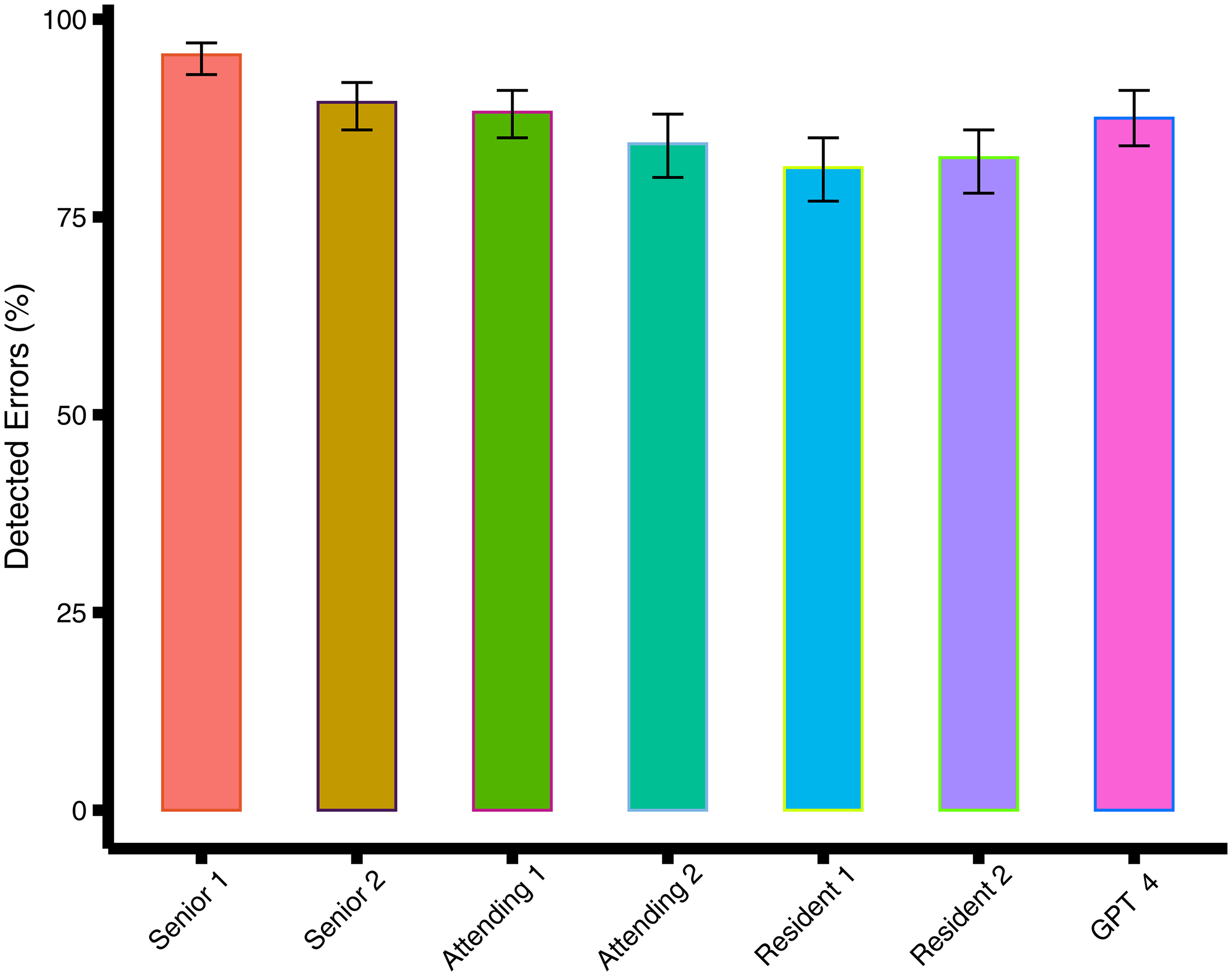
Bar graph shows the percentage of detected errors for GPT-4 and the doctors. The error bars are 95% CIs.
Subgroup analysis by anatomic site
In the subgroup analysis of different anatomic sites, GPT-4 detected fewer errors compared to the best-performing senior pathologist in pathology reports of the abdomen (detection rate, 88% [207 of 236; 95% CI: 85, 97] vs. 95% [224 of 236; 95% CI: 85, 97]; χ²=6.8, df = 1,
Comparison of error detection rates between GPT-4 and doctors using McNemar's test.

Bonferroni correction was used to correct
Validation of specific error types
We further analyzed the performance of GPT-4 and pathologists across different types of introduced errors, including clerical errors, missing information, improper use of terminology, interpretation or diagnostic errors, and data inconsistency. The performance of GPT-4 in detecting ‘improper use of terminology’ was relatively weaker compared to other error categories, with a detection rate of 79% [63 of 80; 95% CI: 68, 87]. This suggests that while GPT-4 is generally effective in error detection, the identification of improper use of terminology remains a challenging area for the model, requiring further refinement. In contrast, the best-performing pathologist detected these errors with a higher rate of 91% [73 of 80; 95% CI: 83, 96], indicating a potential area where human expertise still outperforms the AI model. Regarding the other error categories, there was no significant evidence of GPT-4 detecting fewer errors compared to the pathologists (
Comparison of detection rates for different error types in pathology reports using McNemar's test.

GPT-4's false positives
Among the 700 reports examined, GPT-4 erroneously identified nine reports as containing errors when, in fact, they did not contain any errors. In comparison to the most proficient senior pathologist, GPT-4 displayed a higher rate of incorrectly flagged pathology reports (2.3% [9 of 400; 95% CI: 1.5, 3.0] vs. 0.3% [1 of 400; 95% CI: 0.01, 0.9]; χ² = 5.0, df = 1,
Comparison of incorrectly flagged pathology reports between GPT-4 and the doctors.

The number of correctly detected errors by GPT-4 was compared with the doctors by using χ2 test.
Performance metrics of error detection in pathology reports.

The statistical analysis supports the conclusion that GPT-4 is less effective in identifying complex errors compared to experienced human pathologists. Examples of false positives included cases where GPT-4 flagged minor inconsistencies that did not constitute actual errors, such as slight formatting deviations between sections or the use of uncommon but clinically acceptable terminology. In one instance, GPT-4 incorrectly identified the phrase “no tumor cells seen” as inconsistent due to its variation from the more common phrasing “tumor-free margins,” despite both conveying the same clinical meaning. Another example involved GPT-4 mistakenly flagging missing tumor grading when the case involved a benign lesion that did not require grading.
Interrater agreement
The interrater agreement between GPT-4 and the pathologists, as well as among the pathologists themselves, was observed to be only slight to fair, with Cohen's κ ranging from 0.01 to 0.39 (Table 6). While perfect concordance may not be ideal in this context, the variability in agreement highlights the complementary nature of GPT-4 in the error detection process. Low concordance suggests that GPT-4 could serve as a valuable supplementary tool, potentially identifying different errors that human readers might miss. This emphasizes GPT-4's role in enhancing sensitivity in error detection, making it a useful adjunct to human review rather than a replacement.
Interrater agreement between GPT-4 and the doctors.

Comparison of Reading time
The aggregate reading time for all 700 reports by GPT-4 amounted to 0.78 h. In contrast, the swiftest pathologist completed the review of 700 reports in 12.76 h, while the slowest pathologist took 21.35 h (Figure 3 and Table 7). The mean reading time per pathology report for GPT-4 was significantly shorter compared to the fastest pathologist (mean reading time, 4.03 seconds ± 0.8 [SD] versus 65.64 seconds ± 18.63;
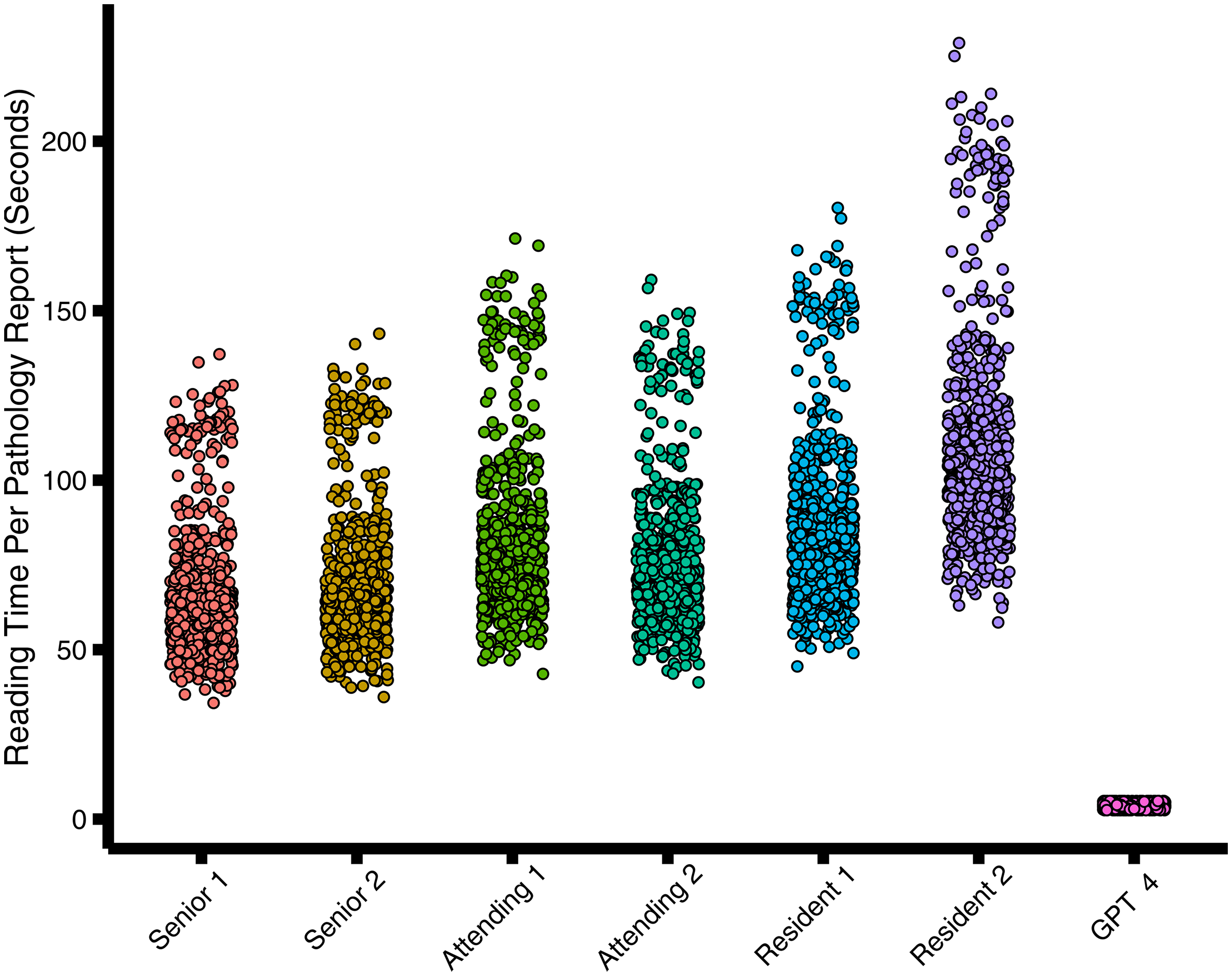
Scatter diagram shows reading time per radiology report in seconds.
Mean reading time per pathology report.

Discussion
This study aimed to evaluate the performance of GPT-4 in detecting errors in oncology-related pathology reports, focusing on key metrics such as precision, recall, and F1-score. By using confidence intervals instead of solely relying on p-values, we were able to offer a more robust understanding of GPT-4's reliability and variability in clinical applications. Although GPT-4 was employed in a zero-shot manner without specific fine-tuning for pathology, our findings suggest that fine-tuning on pathology-specific datasets could enhance its ability to manage complex medical terminology and further reduce error rates. Future improvements in precision and recall through model fine-tuning would make GPT-4 a more reliable tool for clinical settings, particularly in oncology, where diagnostic accuracy is paramount.
Our study demonstrated that GPT-4 performs comparably to attending pathologists in error detection but was outperformed by the best senior pathologists. However, GPT-4 surpassed resident pathologists, especially in detecting clerical errors. Despite these promising results, GPT-4's higher rate of false positives raises concerns about its practical application in clinical oncology settings. False positives can lead to unnecessary diagnostic tests or treatments, adversely affecting patient care and resource management. Therefore, further refinement of GPT-4's algorithms is critical to reducing these false positives and improving its reliability in cancer diagnostics. The clinical implications of false positives—whether introduced by GPT-4 or human reviewers—warrant careful consideration. In a high-stakes environment such as oncology, unnecessary alerts or misinterpretations could lead to additional diagnostic testing, prolonged review time, or patient anxiety. While none of the false positives in this study resulted in simulated management errors, the potential for downstream impact highlights the importance of incorporating AI as a decision-support tool rather than a standalone diagnostic authority. Human oversight remains essential to contextualize AI-generated flags and ensure safe, patient-centered care.
To address these issues, future studies should conduct case-by-case analyses of false positives to identify underlying causes, whether related to medical terminology, context misinterpretation, or other factors. Understanding and mitigating these errors will be essential for GPT-4 to become a more accurate and reliable support tool for pathologists in oncology. Furthermore, the lack of consistency in error detection between GPT-4 and human pathologists, as well as variability among human pathologists themselves, warrants further investigation. These discrepancies could arise from differences in training, experience, and diagnostic interpretation. Addressing these factors is crucial to improving both AI and human performance in cancer-related pathology error detection.
The study also highlighted a noticeable lack of consistency between GPT-4 and the human pathologists in terms of error detection, as well as among the pathologists themselves. This variability warrants further attention to understand its underlying causes, which may include differences in the interpretation of pathology reports, the subjective nature of certain diagnostic decisions, and the distinct methodologies used by GPT-4 versus human readers. Addressing this variability is crucial for improving both the AI's reliability and human experts’ accuracy. Future research will focus on a more detailed analysis of these discrepancies, with the goal of identifying specific factors contributing to these differences. By exploring these causes, we aim to enhance the standardization of error detection processes across different settings, thereby reducing variability and ensuring more consistent results
Our research builds upon prior studies that have explored the potential of LLMs like GPT-4 in medical error detection and structuring of reports.14,22 While our study primarily used zero-shot prompting, future research should explore other prompting strategies, such as one-shot or chain-of-thought prompts, which may enhance GPT-4's ability to detect nuanced errors. By refining the prompt strategies, GPT-4's performance can be more closely aligned with the expectations and judgments of human pathologists in oncology settings.
Additionally, our findings are consistent with studies like Gertz et al., who demonstrated the utility of GPT-4 in detecting errors in radiology reports. 14 This reinforces the broad applicability of LLMs across different medical fields, including oncology. Moreover, prior research has shown that integrating AI with human expertise, as done in this study, can further enhance the performance of AI models in clinical practice. Moreover, Sun et al. (2023) have shown how GPT-4 can effectively generate impressions in radiology reports, indicating a broad applicability of LLMs across different medical fields. 23 Additionally, Zhu et al. (2024) emphasized leveraging professional radiologists’ expertise to enhance LLMs’ evaluation, which parallels our approach of combining human expertise with AI in pathology. 24 Our study advances this understanding by quantifying GPT-4's performance in pathology reports, a domain characterized by highly specialized language and complex diagnostic information, and identifying specific strengths and weaknesses in its application.
The dual benefit of GPT-4 in both error detection and data structuring for future machine learning applications holds promise for the oncology field. 22 By improving the accuracy of pathology reports and ensuring more reliable data for downstream AI applications, GPT-4 has the potential to streamline oncology workflows and improve diagnostic accuracy. However, its role should be seen as complementary to human pathologists, ensuring a balance between AI's efficiency and human expertise's clinical judgment.
This study contributes to the field by highlighting the practical implications of integrating GPT-4 into clinical workflows. While previous studies focused on the ability of LLMs to process and structure large amounts of data, our research emphasizes the importance of error detection in maintaining the integrity of clinical documentation. By ensuring that errors are promptly identified and corrected, GPT-4 can support pathologists in delivering more accurate diagnoses, which is crucial for effective patient care.
Notably, GPT-4 exhibited shorter processing times compared to human readers, further highlighting its efficiency in pathology report proofreading. The findings from our study highlight specific strengths and weaknesses of GPT-4 in error detection across different categories. Notably, identifying the improper use of terminology emerged as a relative weak point for GPT-4, with a detection rate of 79%, compared to higher performance in other categories. This indicates that while GPT-4 enhances overall report accuracy and efficiency, there are areas that require further refinement to match the proficiency of human pathologists. However, its role may be best utilized as a supplementary tool to human pathologists, maximizing detection sensitivity while requiring human sign-off to ensure specificity. These results align with previous studies indicating the potential time and cost savings associated with the implementation of LLMs in pathological workflows.22,25,26 Given that errors in pathology reports are prevalent across all experience levels, the proficiency of GPT-4 underscores its suitability for real-world clinical environments.2–4 By serving as a cost-effective proofreading tool, GPT-4 offers a viable solution for enhancing pathology report generation processes.
In addition to our primary analyses based on error types, we also conducted a subgroup analysis based on anatomic sites. However, we acknowledge that this analysis has limited clinical relevance due to the artificial nature of the error distribution. These errors were uniformly introduced across different organs and do not reflect natural patterns observed in real-world pathology reporting. Therefore, this analysis was included primarily for completeness and should be interpreted with caution. Our core focus remains on performance differences across error types, which offer more practical insights into GPT-4's capabilities in real-world pathology applications.
Several limitations of our study should be acknowledged. First, GPT-4's higher rate of incorrectly flagged pathology reports highlights the need for improved specificity. These false positives, if unchecked, could lead to unnecessary interventions, impacting patient care. Maintaining human oversight is essential to mitigate these risks and ensure that AI systems enhance rather than undermine clinical judgment. Second, the variability in error detection between GPT-4 and human pathologists underscores the subjective nature of pathology interpretation. Understanding these differences is vital to improving both AI performance and human diagnostic accuracy in cancer pathology. Third, one important limitation of our study is the use of intentionally manipulated pathology reports, rather than naturally occurring reports with real-world errors. Although this design enabled us to control the type, frequency, and distribution of errors across a large sample, it may not fully capture the complexity and variability of actual reporting mistakes. In real-world settings, errors are more heterogeneous and context dependent. Therefore, future studies should incorporate prospective or retrospective analyses of naturally occurring errors to validate and extend our findings in clinical practice. It is also worth noting that the pathology reports were in Chinese, and GPT-4's performance may be affected by language-specific limitations, as the model is primarily trained on English data. Finally, while our study focused on a specific sample of 700 pathology reports, broader validation with diverse datasets is required to assess the generalizability of these findings to various clinical settings and cancer types.
Future research should prioritize the continued optimization of AI models such as GPT-4 to ensure their safety, reliability, and applicability in real-world clinical oncology settings. Collaborative efforts among AI developers, oncologists, and regulatory bodies will be essential for establishing standardized guidelines for the clinical deployment of AI technologies in cancer care. While this study focused on zero-shot prompting to assess GPT-4's baseline performance, future investigations should incorporate direct comparisons with alternative approaches, such as few-shot prompting or lightweight domain adaptation using small datasets. These strategies may enhance the model's ability to detect subtle or context-dependent errors and support more effective and reliable integration into clinical workflows.
Conclusion
GPT-4 shows considerable potential as a tool for improving the accuracy and efficiency of pathology report generation in oncology. However, the integration of AI models into clinical workflows requires careful oversight, refinement, and ongoing evaluation to ensure their reliability and safety. By combining AI's strengths in error detection with human expertise, GPT-4 can become a valuable asset in cancer diagnostics, ultimately enhancing patient care. Future research should focus on establishing robust frameworks for the continuous monitoring and improvement of AI systems like GPT-4 in clinical environments.
Declarations
Ethics approval and consent to participate
All procedures involving collection of tissue were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. This retrospective compliance study was approved by the Ethics Review Committee of Guizhou Provincial People's Hospital (Ethics Number: 2024004), the Third Affiliated Hospital of Sun Yat-sen University (Ethics Number: B2023074), the Third Xiangya Hospital, Central South University (Ethics Number: 2024011), and Jiangxi Cancer Hospital (Ethics Number: JC2024006). Written informed consent was obtained from all participants or their legal guardians prior to data collection. Ethical approval was obtained to ensure that the study adhered to the highest standards of patient confidentiality and data protection.
To maintain patient data privacy, several measures were implemented:
Data Anonymization: All patient identifiers were removed before the data was accessed by the research team. This included the anonymization of names, dates of birth, medical record numbers, and any other potentially identifiable information. The anonymized data was assigned unique study identifiers to ensure that patient identities could not be traced. Secure Data Storage: The anonymized data was securely stored on encrypted servers with restricted access. Only authorized personnel involved in the study had access to the data, and all data transfers were conducted using secure, encrypted channels to prevent unauthorized access. Compliance with Ethical Standards: The study complied with all relevant regulations and guidelines regarding the use of patient data in research. Informed consent was obtained from all patients whose reports were included in the study. This consent process was conducted in accordance with the guidelines established by the IRB, ensuring that patients were fully informed about the study's purpose, procedures, and their rights.
These measures were rigorously followed to ensure that patient data was handled with the utmost care, respecting their privacy and maintaining compliance with ethical standards throughout the research process.
Footnotes
Acknowledgments
During the preparation of this work the authors used GPT-4 (powered by OpenAI's language model; ![]() ) in order to generate parts of this article. After using this GPT-4, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication. Thanks to the colleagues in the department of pathology for their help in this paper, your excellent work has made our research more efficient.
) in order to generate parts of this article. After using this GPT-4, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication. Thanks to the colleagues in the department of pathology for their help in this paper, your excellent work has made our research more efficient.
Authors’ contributions
Xiongwen Yang contributed to conceptualization, methodology, software, validation, formal analysis, investigation, resources, data curation, and writing‒original draft. Yun Zhang contributed to validation, formal analysis, investigation, resources, data curation, and writing‒original draft. Jinyan Jiang contributed to validation, formal analysis, investigation, resources, data curation, and writing‒original draft. Zhijun Chen contributed to validation, formal analysis, investigation, resources, data curation, and writing ‒ original draft. Rinasu Bai contributed to validation, formal analysis, investigation, resources, data curation, and writing‒original draft. Zihao Yuan contributed to formal analysis, investigation, data curation, and writing‒original draft. Longyan Dong contributed to formal analysis, investigation, data curation, and writing‒original draft. Yi Xiao contributed to resources, data curation, and writing‒original draft. Di Liu contributed to validation, formal analysis, investigation, resources, data curation, and writing‒original draft. Huiyin Deng contributed to data curation and writing‒original draft. Jian Huang contributed to data curation and writing‒original draft. Huiyou Shi contributed to formal analysis, investigation, resources, data curation, and writing‒original draft. Dan Liu contributed to conceptualization and project administration. Maoli Liang contributed to conceptualization and project administration. WeiJuan Tang contributed to conceptualization, methodology, writing‒review and editing. Chuan Xu contributed to conceptualization, methodology, writing‒review and editing, visualization supervision, project administration, and funding acquisition.
Funding
This study was supported by the Talent Fund of Guizhou Provincial People's Hospital (grant number: [2014-16]), and additionally by the Basic Research Program of the Guizhou Provincial Department of Science and Technology (Qian Ke He Ji Chu) [Grant No.QKH-JC-MS (2025)492].
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
