Abstract
Significant advances in the treatment of cancer have been achieved as reflected by the ever-expanding space of cancer therapeutics being available to cancer patients. Often, however, it is not clear which patient would respond to which drug and what combination of drugs will improve patient outcomes. Furthermore, while many of these drugs are initially effective, therapeutic resistance is often inevitable due to the evolving nature of cancer. Generative artificial intelligence (GenAI) powered by the increasingly large amount of accumulating clinical, molecular, and radiomics data about cancer patients and their treatments may serve as the kernel of rapid learning decision-support systems that could enable personalized cancer treatments to counter therapeutic resistance and overcome the shortcomings of the current standard of care. This perspective is explored in the context of current advances of AI applications in oncology and the potential of GenAI learning and inferencing capabilities to support patient-tailored dynamic cancer treatments. A discussion of this vision is elaborated with respect to issues pertinent to GenAI use in real-world clinical settings, including clinical validation, data curation, and sharing, large language model hallucinations as well as ethical concerns and considerations such as privacy, bias, transparency, and accountability.
Keywords
Introduction
Cancer is the leading cause of premature death in North America, Europe, and China, and ranks along with cardiovascular diseases (CVD) among the two or three top leading causes of premature death for the overwhelming rest of the world. 1 Although the universe of therapeutic options is continuously expanding for cancer patients, therapeutic resistance continues to be the most critical challenge in the fight against cancer. This challenge reflects the nature of cancer as a complex adaptive system2,3 continuously evolving under the thrust of its genetic, 4 immunological, 5 and eco-evolutionary 6 determinants. On the other hand, the increasingly large amount of accumulating clinical, molecular, and radiomics data about patients and their treatments hold the potential for data trained artificial intelligence learning systems that could recast these cancer multimodal data and the rapidly expanding therapeutic space into integrated dynamic knowledge base and inferencing systems to support more effective treatment decision-making. Indeed, applications of generative artificial intelligence (GenAI) and their underlying large language models (LLMs) in medicine are experiencing increasing research interests,7,8 owing in part to their demonstrated learning and inferencing capabilities in a vast array of cognitive tasks including natural language processing, language translation, text generation, speech and image recognition, and coding.9,10 Indeed, there is a noticeable growth in explorations of GenAI applications for healthcare such as medical diagnosis, clinical decision-support, drug discovery and development, patient education and self-management, clinical documentation, medical education, and healthcare administration.11,12 However, GenAI introduces new dimensions to health care delivery and related issues that need special attention, including LLMs’ hallucinations, 13 ethical considerations, and the need for large quality training datasets.
The challenges of treating cancer as a complex adaptive system are well appreciated, and so is the potential of exploiting the knowledge base embedded in the increasingly large datasets about cancer patients and their treatments to drive GenAI's learning and predictive capabilities in assisting clinical decision-making. The critical question is: how can GenAI's potential be exploited to assist in the development of treatments with long-lasting effectiveness? The proposed perspective elaborates on this question starting with an overview of combination and adaptive therapies as the two leading therapeutic strategies used to mitigate therapeutic resistance. This is followed by an overview of research and translation efforts devoted to AI applications in oncology. Through a critical reflection on cancer's treatment challenges and GenAI’s potential and vulnerabilities, a perspective is offered on reimagining cancer treatment through an integrated consideration of genetic, immunological, and eco-evolutionary dimensions of cancer whose dynamics can be reconstructed, from observations, into GenAI models that could assist adaptive combination therapy. The perspective considers cancer's unique challenges and GenAI vulnerabilities as the background for a discussion on pertinent issues regarding deployment in real-world clinical settings, including clinical validation, data collection and curation, privacy, bias, accountability, clinical environment, and clinical ethics.
Targeted and combination therapies
Targeted therapies are conceived to either suppress oncogenic pathways such as RAS/RTK (receptor tyrosine kinase) by specifically targeting one or more of their altered components using kinase inhibitors (KIs), and/or activate the killing functions of immune cells using monoclonal antibodies (mAbs) to target aberrant proteins. 14 One notable example of targeted therapy is osimertinib, a third generation EGFR (epidermal growth factor receptor) tyrosine kinase inhibitor (TKI), used to target altered HER2 and suppresses the oncogenic activity of downstream pathways (e.g., RAS/RTK, AKT/PI3K, and MAPK) in non-small cell lung carcinoma (NSCLC).15,16 HER2 is also targeted using the monoclonal antibody trastuzumab as a standard adjuvant therapy to lower recurrence risk for HER2+ breast cancer. 17 Despite their clinical success, targeted therapies are overwhelmed by tumor heterogeneity, the evolving genetic diversity of cancer cells and their phenotypic plasticity. For example, the treatment of NSCLC patients with EGFR TKIs lead to acquired resistance implicating MET amplification in about 20% of cases. 18 MET and ALK TKIs are also vulnerable to acquired resistance in lung cancer19–21 and so is BRAF and MEK inhibitors. 22 In an effort to thwart therapeutic resistance and improve patient outcome, combination therapies, which involve combining multiple drugs either sequentially or concomitantly, have become a mainstay of cancer treatments. 23 The combination of EGFR and MET TKIs is one such example currently in a phase II clinical trial (NCT03778229) involving the use of osimertinib plus savolitinib for advanced or metastatic NSCLC patients with EGFRm+ and MET+. 24 Also noteworthy is the recent FDA (U.S. Food and Drug Administration) approval of ipilimumab and nivolumab for advanced colorectal cancer. 25 These represent few among a very large number of possible drug combinations, thousands of which are undergoing clinical trials, underscoring the promising potential of combination therapy in hedging against therapeutic resistance.23,26 However, given the limited pool of patients that could be enrolled in clinical trials, the search for therapies that are efficacious under toxicity constraints will require rationalizing drug combinations. 27 Furthermore, optimizing doses of individual drugs in combination therapies and their administration schedules (i.e., order, duration, and time interval separating doses) is also needed to achieve efficacy while avoiding enhanced toxicity, 28 providing a rationale for adaptive dosage involving dose modulations and dynamic scheduling based on continuous treatment response monitoring.
Adaptive combination therapy
The eco-evolutionary dynamics of cancer is characterized by an evolving genetic and epigenetic diversity, phenotypic plasticity, and metabolic flexibility, which feed a near-bondless capacity for adaptation under targeted and systemic therapies, ultimately leading to the onset of therapeutic resistance.2,3,29,30 Adaptive therapy (i.e., modulation of doses and administration schedules) based on repeated monitoring of treatment response is increasingly viewed as an intrinsic necessity and a sound strategy in countering cancer evolutionary dynamics to delay or prevent the onset of therapeutic resistance.31–37 The integration of cancer evolutionary dynamics in the design of adaptive therapies has been explored through the use of mathematical models of clonal competition to inform the design of adaptation strategies.38–40 One such example is the clinical trial design of adaptive therapy (AT) for metastatic castration resistant prostate cancer (mCRPC) aided with mathematical models of evolutionary competition.34,41 The therapeutic strategy is to maintain a sufficient proportion of therapy sensitive cells to prevent the competitive release of resistant ones by intermittently stopping therapy when the PSA (prostate specific antigen) falls below 50% of its pre-treatment level and resuming it when the PSA reaches or surpasses the 50% of pre-treatment level.34,41 The corresponding mCRPC clinical trial (NCT02415621) of this evolutionary-informed adaptive therapy approach yielded 58.5 and 33.5 months as the median values for the overall survival (OS) and time to progression (TTP), respectively, asserting a significant improvement when compared to the standard of care (SOC) cohort for which the medians for OS and TTP were 31.3 and 14.3 months, respectively. 34 Other ongoing clinical trials of adaptive therapy include the study of adaptive BRAF and MEK inhibitor therapy guided by ctDNA (circulating tumor DNA) for late stage cutaneous melanoma, 42 and the study of adaptive chemotherapy (carboplatin) dosing based on changes in CA125 for ovarian cancer. 43 Adaptive combination therapy has also been explored using mathematical models of evolutionary competition 33 and cell adaptive fitness.36,44 Prospective studies of multi-drug adaptive therapy includes the clinical trial (NCT03511196) involving the combination of luteinizing hormone releasing hormone (LHRH) analog and new hormonal agent (NHA) such as abiraterone for metastatic castration sensitive prostate cancer (mCSPC), where the therapy adaptation is guided by PSA and testosterone levels as well as imaging. 35
The nascent clinical success of adaptive therapy and the strong rationale for its appropriateness in thwarting therapeutic resistance, support its consideration as a necessary component of effective cancer treatments. However, this will require key advances, such as the development of accurate and reliable treatment response biomarkers that can reveal the dynamics of tumor growth dynamics, and the development of algorithms that could reconstruct models of such dynamics to support the conception of adaptation schemes that are effective in thwarting therapeutic resistance.
Artificial intelligence in oncology
Diagnosis, prognosis, and risk stratification applications of AI in oncology are steadily growing in number.7,8,45–52 So far, however, most studies of AI oncology applications are retrospective, with few prospective studies as rare exceptions.45,48,53 The lack of prospective studies and randomized clinical trials (RCTs) is a critical barrier to clinical translation of AI-assisted health care. 54 Nevertheless, the progress towards the adoption and integration of AI in real-world clinical settings is proceeding with difference cadences across various oncology applications. 55 In particular, AI-assisted cancer diagnosis is moving closer towards maturity and wider clinical adoption as demonstrated by an ever expanding list of AI/ML (artificial intelligence/machine learning) SaMDs (software as medical devices) approved by the FDA, 56 many of which are for oncology. AI-assisted prognostication, risk stratification, as well as outcome and survival predictions are also going through a wave of accelerated research interests.45,46,57,58 The same applies to treatment decision-making and treatment optimization using mathematical, AI and GenAI models.36,38,48,50,59,60 Despite the common objective of AI applications to improve cancer treatments, there is a distinction between traditional AI/ML algorithms which rest on task-specific models trained with structured and tabular data compared to GenAI models which are trainable on multimodal data for a broad spectrum of tasks. Given these and other differences regarding knowledge representation, interpretability, and adaptability, research focus on LLM applications in oncology and medicine in general is driven by additional emphasis on concerns such as transparency, ethics, and regulations.61–63
The complex and multifactorial clinical decision-making process,64–66 the evolving dynamics of cancer, and the “black box” nature of AI systems are among the factors conspiring to dampen the pace of convergence towards clinically feasible and effective AI-assisted treatment strategies. Despite these challenges and the lack of a decisive scientific understanding of why GenAI works so well,67,68 there is a flurry of studies exploring the utility of GenAI to support the tasks of clinical oncology, including treatment decision-making.7,69 However, the evaluation of general purpose LLMs applied to oncology have yielded mixed results, where some models have shown poor accuracy and lack of robustness, 70 challenging their acceptability for real-world clinical setting. 71 On the other hand, other GenAI models were found to be acceptable by oncologists, albeit lacking the specification of treatment order and exhibiting limitations with respect to intricate cases. 72 Performance limitations and uncertainty of general-purpose foundation LLMs and the lack of benchmarks for their clinical validation49,70–73 points to the need for more research, including the exploration of oncology-specific LLMs trained on quality cancer datasets to confine their learning, reasoning, and inferencing to the oncology knowledge space as a catalyst for improving performance and amenability to clinical validation.
Can generative AI transform cancer treatments?
Dynamic scheduling of treatments involving adaptation that accounts for cancer evolutionary dynamics is increasingly regarded as an appropriate strategy to thwart therapeutic resistance.31,32,34–36,41,74,75 However, treatments are unlikely to be effective in yielding a cure or long-term disease management without an integrated consideration of the genetic, immunological and eco-evolutionary dimensions of cancer complexity (see Figure 1). The expanding big data becoming available about cancer treatments open a plausible avenue to recast cancer treatment as a problem of controlling tumors as time-varying nonlinear dynamical systems, that implicitly account for cancer's biological dimensions, and whose nonlinear dynamics can be identified from observations of treatment response. 76

The genetic, immunological, and eco-evolutionary dimensions of cancer complexity and their targeting by various therapeutic modalities of the current standard of care.
The use of GenAI in cancer treatments exploits the property of deep neural networks as universal approximators 77 to yield models of cancer dynamics that are learned from big data about cancer in order to provide patient-personalized predictions based on repeated monitoring of treatment response. 8 The flexibility of GenAI models to integrate and learn from new data and their capacity to yield patient-adapted predictions offer an unprecedented potential to support adaptive treatments based on an integrated attention to known levers of therapeutic actions (see Figure 2).
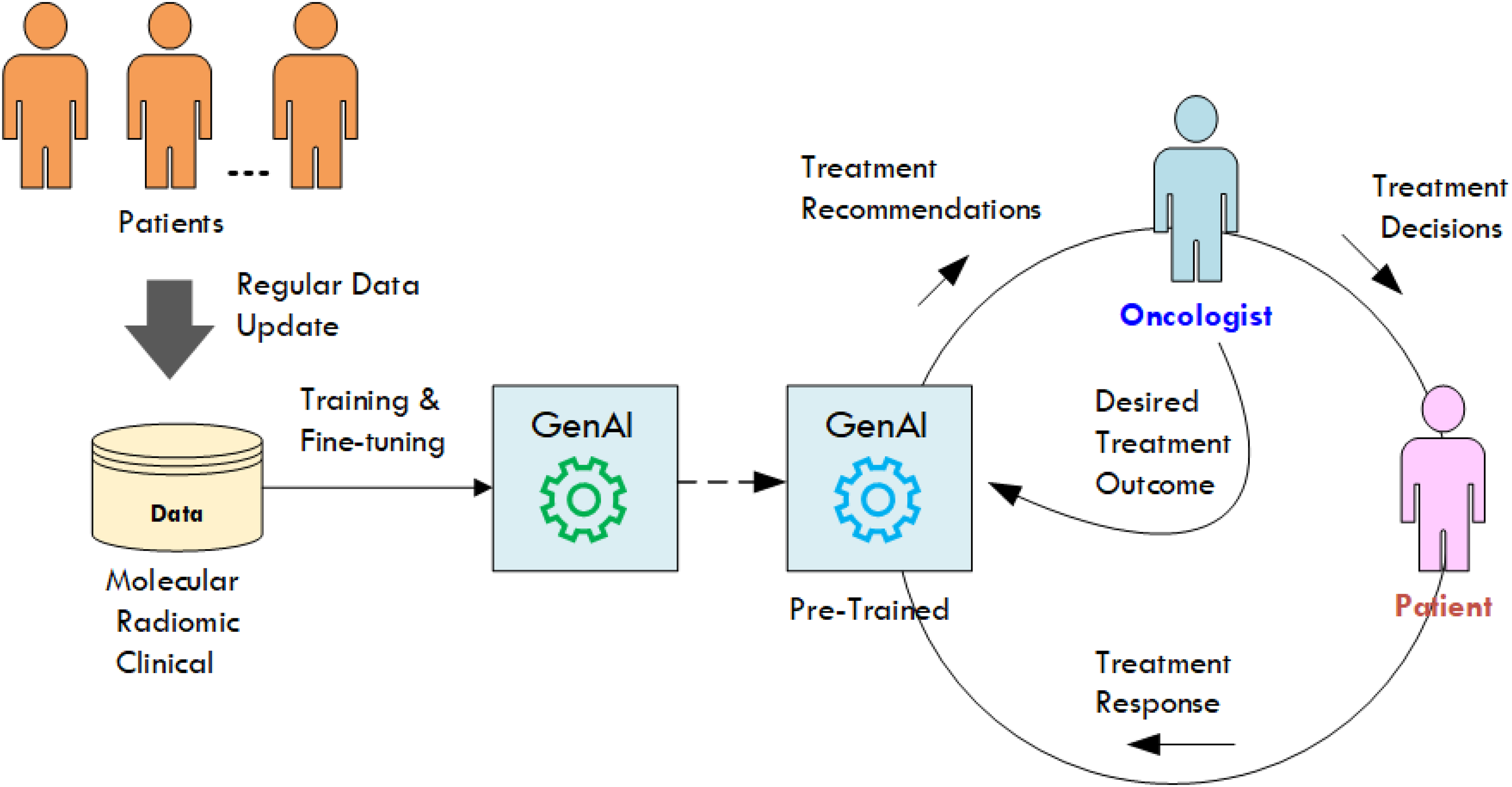
GenAI as the engine of rapid-learning systems for precision oncology. GenAI: generative artificial intelligence.
The learning capability of GenAI models can be exploited to assist in virtually all aspects of cancer treatments, including: (a) treatment response predictions, (b) treatment planning, (c) treatment adaptation, and (d) toxicity management. Ultimately, the improvement of cancer treatments will depend on the extent to which GenAI can facilitate the use of the full spectrum of available therapeutic agents to decisively thwart therapeutic resistance without causing unbearable levels of toxicity that compromise patient's long-term quality of life (QoL).
Treatment response predictions
Cancer treatment can be framed as a problem of controlling a nonlinear dynamical system36,78 where the timing and doses of administrated drugs are adapted based on real-time treatment response monitoring with the objective of steering the disease to a desirable end-state. Effective treatments of cancer require disease state feedback to track the evolving dynamics of cancer progression under treatment. In this respect, imaging 79 combined with liquid biopsy (LB) 80 and next generation sequencing (NGS) can be used for repeated monitoring of treatment response to estimate the evolving genotypic and phenotypic characteristics of the tumor based on radiomic data and the information carried by ctDNA, circulating tumor cells (CTCs), circulating tumor RNA (ctRNA), extracellular vesicles (EVs), cancer-associated stromal cells, cytokines, proteases, and tumor-educated platelets. 81 For example, ctDNA LB can be used for the detection of microsatellite instability (MSI) and the tracking of the evolution of tumor mutational burden (TMB) during immunotherapy. 82 The longitudinal monitoring of cell-free DNA (cfDNA) using LB was also used to track sub-clonal evolution based on low pass whole genome sequencing of somatic copy number alterations (SCNA). 83 These approaches, while ingenious in their design, they nevertheless rely on the explicit consideration of a limited number of treatment response variables, leaving out a massive wealth of untapped information in the sampled radiomic and molecular data about the evolving dynamics of the tumor and its microenvironment. Herein lies the potential of exploiting the learning capability of GenAI to identify the dynamic nonlinear relationship between a near-infinite number of treatment response variables and disease state variables. 84 The clinical feasibility of this GenAI approach hinges on the availability of training data from a significantly large cohort of patients. This is a universal challenge facing the exploitation of GenAI in oncology and healthcare in general as will be discussed in later sections. Notwithstanding their vulnerability to hallucinations, 85 which may be mitigated using retrieval-augmented generation (RAG), 86 GenAI is endowed with the advantage of rapid-learning from the expanding space of cancer treatment data, making it uniquely positioned to assist in reimagining cancer treatments by providing the all critical prediction models of cancer evolutionary dynamics necessary for an effective adaptive cancer therapy.
Treatment planning and adaptation
GenAI-supported dynamic treatment may be conceived along two possible modalities. 84 First, GenAI models of disease state and treatment response dynamics can be integrated into dynamic decision-support systems that enable oncologists to undertake offline treatment planning and modifications by exploring different “what if” treatment scenarios and regularly adjusting treatment schedules based on feedback provided by GenAI predictions of disease progression. This application falls in the category of GenAI-enhanced traditional clinical decision-support systems used to inform rather than directly drive the treatment decision-making process. In the second scenario, GenAI models could enable three possible support configurations to adaptive therapy, corresponding to the three essential components of adaptive control, 78 namely: (a) disease state trajectory planning, (b) disease state estimation, and (c) treatment decision synthesis. 84 For the first configuration, the desired disease state trajectory is planned by the oncologist. The treatment adaptation could, on the other hand, be synthesized by one of the many possible adaptive control algorithms 36 driven by the differential between the oncologist planned disease state trajectory and GenAI estimated disease state. The second configuration is similar to the first one except that oncologists only specify the desired disease end-state and delegate the planning of the desired disease state trajectory to a trained GenAI model. The third configuration goes further by using a GenAI treatment recommender model to synthesize treatment adaptations, leading to a fully GenAI-driven adaptive treatment system, where the input of the oncologist is limited to the desired disease end-state. These three possible approaches to GenAI-assisted treatment decision-making vary in their degree of automation and reliance on the learning and predictive capabilities of LLMs. Hence, for these to maintain the desired performance over time, they would have to be regularly fine-tuned using newly acquired data about patient treatments, cancer knowledge and new approved cancer drugs.
The data challenge
Although synthetic data generated from clinically calibrated models of cancer may be used in explorative projects of GenAI applications to cancer treatment optimization, the development of clinical grade GenAI-supported treatments will require systematic collaborative efforts across diverse clinical and research institutions to collect and curate the necessary training data. National and international organizations and initiatives such as NCI (National Cancer Institute) cancer research data common (CRDC), NCI-PDXNET (Patient Derived Xenograft Network), the Cancer Moonshot Biobank, the Prostate Cancer Clinical Trials Consortium (PCCTC), and the European Organization for Research and Treatment of Cancer (EORTC) are all potential avenues for international collaborations to collect and share clinical data. However, since the sole purpose of such data curation is to support the training of GenAI models, a dedicated cancer data consortium similar to TCGA (The Cancer Genome Atlas) may be needed in order to address the unique challenges associated with the training of GenAI models for oncology. Among these are privacy concerns and regulations which hamper the sharing and integration of patient data in sufficient quantity to facilitate a robust training of LLMs for oncology applications. The quality of clinical data in terms of completeness and representativeness of medical and demographic diversity of the patient population is yet another challenge to the training of GenAI models for clinical oncology. In this respect, future efforts towards consortium-led collection and curation of oncology data would have to include the development of governance frameworks for big multimodal data ownership, protocols for data use and sharing, ethical considerations around the perpetual embedding of knowledge from collected data in trained GenAI models, and institutional rights of access to the parameters of GenAI models trained using contributed data. The reliance on frameworks of LLM federated learning across multi-sites, each keeping their patient data private, could also be an effective approach to mitigate the challenges of LLM training using big and diverse quality datasets while safeguarding patient privacy. 87
Transparency and the black box problem
The use of LLMs in cancer treatment decision-making entails a tacit recognition of the associated black box problem whereby LLMs’ outputs (e.g., predictions and recommendations) are difficult to interpret and to explain by clinicians. Such opacity creates significant hurdles for regulatory oversight, constraining efforts to assert reliability, and ultimately undermine clinicians trust in GenAI-driven treatment decision-support systems. Such trust is critical to GenAI acceptance and adoption in real-world clinical setting. Opacity or lack of transparency of GenAI systems is rooted in their underlying DL models and is manifested along multiple dimensions. First, despite the empirical success of GenAI in a dizzying number of application domains, a definitive scientific understanding of the underlying reasons for their impressive generative performance is yet to be developed.67,68 Second, the internal complexity of LLMs and the expansive scope and size of their training data make it challenging to interpret and explain their predictions and recommendations for any application context. 88 Third, also owing to the complexity of LLMs, it is not clear how to effectively peer in such black boxes to fully tease out the reasoning process by which specific predictions or recommendations are generated. Although chain-of-thought explanations provide a multi-step breakdown of an output, this does not expose the actual process by which the final output is generated, only creating an impression of transparency. 89 On the other hand post-hoc explanations, such as highlighting input features with the highest contribution to the generated output, constitute a promising strategy to improve transparency.90,91 However, it has been argued that explanation of GenAI outputs alone is not sufficient to engender trust and that efforts to improve transparency should be focused on the socio-technical system (e.g., treatment delivery system) that embed the AI models. 92 In this respect, applying explainable AI (XAI) approaches under the guidance of a bioethical framework for the responsible use of GenAI 93 may help mitigate the inevitable challenges of LLM opacity. Ultimately, the sum effect of these challenges on GenAI-assisted provision of safe and effective treatments must be assessed through clinical evaluations.
Clinical evaluation
GenAI utility towards a successful transformation of cancer treatments will depend on the extent to which it can help address the challenges of cancer care and facilitate the clinical realization of effective treatments. In particular, the clinical validation of GenAI's capacity to yield accurate and reliable treatment response predictions is a critical step towards building the clinical maturity of adaptive cancer therapy as a viable treatment paradigm. Reimagining cancer treatment must be accompanied with a systematic approach to clinical evaluation not only to generate the necessary evidence for clinical validation but also to shed light on the factors that would impact the capacity of hospitals and clinics to deploy GenAI-supported cancer therapy in real-world settings. These factors include the characteristics of clinical workflows and target patient populations, staff training, infrastructure cost, operational efficiency and the overall environment of clinical care delivery.94,95 More research focus on these implementation factors would be instrumental in providing much needed clarity about the impact of AI in healthcare delivery. 95 This would in turn help promote the acceptance and ultimate adoption of GenAI-supported dynamic treatment scheduling as a clinically feasible strategy that can address the shortcomings of current standard of cancer care.
Clinical workflow integration
Workflow integration of data-driven GenAI rapid-learning systems for precision oncology faces the traditional barrier of interoperability with existing information technology infrastructure which includes electronic health records (EHR) systems. System usability and clinicians training are other integration aspects that require adequate attention. Beyond these traditional clinical workflow integration issues, GenAI-driven systems present unique challenges that are germane to the dynamic nature of learning and inferencing of LLMs and the perpetually changing data that feed their capabilities. The “dataset shift” 96 is one particular challenge that epitomizes the inherent dynamism that characterizes the working of GenAI-assisted decision-support systems within the oncology workflow. Dataset shift is caused by changes spanning various aspects, including patient demographics, standard of care and treatments, and technology. 96 These lead to performance drift of LLMs and a deterioration of their clinical reliability, necessitating regular monitoring of their predictions/recommendations and corrective actions such as fine-tuning or retraining using up to date data that are more representative of the current patient population and care standards. This is particularly critical for the integration of GenAI-driven rapid-learning systems which are inherently bound to dynamically expanding spaces of data, therapeutics and standards of care in order to support the provision of the most effective, tailored cancer treatments for patients. Indeed, data-driven learning throughout the lifecycle of GenAI-driven therapeutic decision-support systems would necessarily entail dataset shifts that may lead to AI models’ instability (i.e., poor performance and harmful recommendations) if unmitigated. 97 In this respect, mitigating the risks of LLM instability and consequent lapses in treatment guidelines and protocol adherence, may require a governance committee 96 to oversee the continuous maintenance of the system's capabilities in supporting the provision of safe and effective patient treatments.
Ethical concerns
The deployment of LLMs in clinical settings raises numerous ethical concerns, including: transparency, interpretability, explainability, bias, privacy, accountability, and responsibility.93,98,99 The training of LLMs using multi-site clinical data through a federated learning framework 87 would be a significant step towards mitigating the risks of bias and compromised patient privacy. Indeed, the multiplicity of sites collaborating in federated learning would mean that patient diversity and variance in healthcare delivery environments will be better represented in the training data. This would improve the universal efficacy of treatment recommendations across patient populations. Furthermore, future data collections should be undertaken with explicit mandates to minimize bias related to demographics, care delivery environments, and patient medical conditions. In addition, since clinical ethics assert that the quality of decision-making is determined by the achieved degree of accuracy and transparency, 100 trained LLMs may need to be frequently re-trained or fine-tuned within the deployment sites using data from the target patient population in order to maintain model accuracy in the face of evolving care practices and an expanding space of therapeutics. In addition, despite the “black box” nature of LLMs, patients must be provided with explanations of treatment decisions that are made based on GenAI recommendations, in order to achieve the goal of transparency. 100 Advances in XAI 101 may provide additional tools to facilitate transparency through well informed doctor-patient communications about the use of LLMs. The inevitable LLM hallucinations 102 must also be mitigated to facilitate the adoption of GenAI-assisted clinical decision-support systems in real-world clinical setting. Although many techniques have been proposed to address this problem, 103 the use of RAG 86 to condition LLMs’ outputs on factual knowledge is most promising.
One of the thorniest concerns about the deployment of GenAI-supported treatment decision-making is accountability. For instance, it is argued that disclosure of AI use in clinical decision-making does not have a strong support in the legal tenets of patient informed consent except for some specific cases. 104 Although the use of LLMs to assist rather than replace clinicians in decision-making is viewed as unlikely to disrupt the physician–patient relationship 105 and hence would not affect traditional accountability, it is unclear how to draw the boundaries around legally acceptable LLM use in cancer treatment decision-making. 106 In all cases, bioethical frameworks for the responsible use of LLMs such as the one proposed by Ong et al. 93 are needed to guide future explorations of GenAI-driven innovations in cancer care.
The road ahead
GenAI-assisted adaptive cancer therapy approaches are in part conceptionally analogous to the real-world successful “Digital Twin” paradigm in engineering, whereby integrated multi-scale, multi-physics models of physical systems such as aerospace vehicles are simulated with continuous real-time sensor data to predict future operational states. 107 This approach has also been explored to frame biological complexity 108 and is experiencing a resurgence under the umbrella of digital twins for precision oncology,109,110 providing further support for GenAI-driven reimagining of cancer treatments.
Big cancer data curation and sharing, transparency, clinical validation, and cancer care delivery environments represent critical dimensions that require systematic attention in reimagining cancer treatments in the era of generative artificial intelligence. Beyond these dimensions, there are catalysts that will undoubtedly energize the vision underlying the proposed perspective as well as challenges that may dampen its progression course. First, therapeutic resistance and the underlying evolutionary dynamics of cancer represent a thorny and still open challenge to patient survival. Human cognitive limitations regarding multivariate decision-making 111 and the challenges of reasoning about the treatment of cancer as a time-varying dynamical nonlinear system while attempting to leverage an expanding big multimodal cancer data and an ever increasing number therapeutic agents becoming available, are undoubtedly among the factors frustrating progress towards more effective cancer treatments. These challenges are also embodiments of strong catalysts for enlisting the support of data-driven GenAI learning and inferencing capabilities in reimagining a more effective cancer treatment paradigm. Such paradigm would address the multivariate decision-making problem at hand and would be open and responsive, through rapid learning, to the expanding knowledge about cancer and to the evolving space of drugs and therapies becoming available. Rapid learning would be realized through regular retraining and fine-tuning of GenAI models using data curated by the institutions where the therapy systems are deployed in order to consider new drugs and treatment protocols, address changing care practices and needs of different patient populations, as well as to maintain the institutions’ standards of safety and quality assurance. This will require clinical institutions to build rapid learning infrastructures (e.g., data collection systems, computing resources and expertise, clinicians’ training in GenAI use, processes of GenAI integration and use, and relevant policies and guidelines), which would ultimately support the long-championed vision of rapid learning healthcare systems,112–114 whose realization will likely be driven by GenAI advances.
Conclusions
Despite the expanding space of therapeutic options being available to cancer patients, the current standard of care is often limited in delivering effective treatments. This constitutes a compelling rationale to rethink cancer treatments, given the impressive learning and inferencing capabilities of GenAI models and the expanding universe of collected clinical, molecular and radiomic data about cancer patients and their treatments. In this respect, GenAI-driven rapid learning has the potential to support effective adaptive treatments that improve patient outcomes in the face of cancer's evolving nature. However, the realization and deployment of such GenAI-assisted decision support systems in real-world clinical settings will require clinical validation and the establishment of rapid learning infrastructures in addition to the need for further advances in addressing the myriad of issues that are intrinsic to GenAI and deep learning, including data collection and sharing, hallucinations, transparency, explainability, privacy, bias, accountability, and other ethical concerns.
Highlights
The current standard of cancer care is often ineffective in dealing with therapeutic resistance as the most critical challenge in the treatment of cancer
Generative artificial intelligence enables rapid learning from continuously updated patient data to support adaptive treatments and counter the evolving nature of cancer.
The use of generative artificial intelligence in treatment decision-making will require addressing a myriad of issues that are intrinsic to the use artificial intelligence in medicine, including explainability, hallucinations, accountability, bias, data privacy, and ethical concerns.
Footnotes
Ethical approval
This article does not contain any studies with human or animal participants. There are no human participants in this study and informed consent was not required.
Funding
The author received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Declaration of generative AI and AI-assisted technologies in the writing process
No AI tools were used in the preparation of the manuscript.
