Abstract
Background
This study evaluated the performance of five Large Language Models based on actual cases to provide guidance for selecting appropriate models for clinical decision-making.
Objective
This study aimed to assess the performance of large language models (LLMs) in clinical decision-making for internal medicine and to provide evidence-based guidance for model selection in clinical practice.
Methods
We conducted a retrospective cross-sectional study with 405 cases across nine subspecialties: cardiovascular, respiratory, gastroenterology, nephrology, rheumatology, endocrinology, neurology, hematology, and infectious diseases. Two senior clinicians evaluated outputs on five dimensions: diagnosis, diagnostic criteria, differential diagnosis, examinations, and treatment. Statistical analyses were performed via the Kruskal‒Wallis tests and Pairwise comparisons were performed by Dunn's test with p-value adjusted by BH procedure.
Results
Overall, significant performance differences were observed among models (
Conclusion
GPT, O1, and Gemini demonstrated superior performance in clinical decision-making for internal medicine among all LLMs, whereas Claude showed the poorest performance. All LLMs demonstrated deficiencies in differential diagnosis and poor management for respiratory diseases. The complexity of subspecialty might be a performance differentiator for LLMs and O1 might have potential suitability for complex subspecialties like cardiology.
Keywords
Introduction
In recent years, the shortage of medical resources has become increasingly severe, driven by population aging and the increasing incidence of chronic diseases. Efficient use of clinical data to strengthen decision-making, improve diagnostic accuracy, guide medication choices, and ease resource constraints has become a pressing challenge. 1 With rapid progress, artificial intelligence (AI) has been widely applied in the medical field, especially for clinical decision-making in internal medicine, which has recently become a research focus. 2 Among the latest AI advances, generative large language models (LLMs) can quickly deliver diagnostic and therapeutic recommendations by processing basic information, symptoms, physical examination findings, and other clinical data. These capabilities are enabled by deep learning and natural language processing applied to extensive clinical datasets and expert-driven training. LLMs show great promise in improving diagnostic efficiency and optimizing clinical decisions.3,4 The application of AI not only relieves physicians from heavy workloads but also supports decision-making through precise data analysis. For example, in oncology, AI can provide individualized anticancer treatment plans by analyzing large-scale genomic and imaging data, improving patient survival rates and quality of life.5,6 Similarly, LLMs significantly improve the accuracy of lesion detection in gastroenterology. Research indicates that AI-assisted systems achieve excellent performance in endoscopic examinations, with a false-negative rate as low as 6%. 7 Moreover, clinical records in internal medicine often contain large amounts of complex, unstructured information that traditional analytical methods cannot fully interpret. LLMs can automatically extract features and recognize patterns from these datasets, revealing correlations and risk factors for disease and supporting the development of personalized medicine.8,9
Current LLMs widely used in the medical field include GPT-4.1 and O1, both developed by OpenAI; Gemini 2.5 Pro Think, developed by Google; Claude 3.7 from Anthropic; and DeepSeek, produced by the Chinese company DeepSeek. These models have become integral to various medical applications. For example, Cristina Madaudo et al. 10 reported that ChatGPT can assist physicians in documenting clinical cases, optimizing treatment plans, and providing timely clinical decision support. Angelica Cersosimo et al. 11 reported that integrating AI into cardiovascular nursing care can increase diagnostic accuracy and enable prompt adjustments to treatment plans. Additionally, research has proposed 12 that GPT-4 may be integrated with clinical data to create an AI-assisted, individualized disease assessment system, enabling treatment plans tailored to patient lifestyles and preferences. Chen J et al. 13 noted that DeepSeek has been implemented in nearly 90 tertiary hospitals across China, where it is extensively used to support clinical decision-making. A study by Lu W 5 indicated that AI systems can provide personalized treatment recommendations by analyzing genomic and imaging data, improving both survival rates and quality of life for patients. Furthermore, Kun W et al. 14 reported that the O1 model produces comprehensive and detailed responses to questions about programmed cell death in myeloma, achieving the highest Cohen’s kappa score and demonstrating strong concordance with assessments made by hematology-oncology experts.
However, the performance of LLMs in practical applications varies due to differences in their underlying algorithms. As a result, each LLM offers distinct advantages and limitations in clinical decision-making, which ultimately influences its clinical value.15,16 A bibliometric analyses revealed that novel cognitive computing algorithms is crucial for the precision of models in predicting cancer. 17 Most existing studies on LLMs in clinical practice focus on single diseases or specific tasks and lack comprehensive evaluations of LLM performance in complex, multidimensional clinical scenarios and horizontal comparisons between different models.18–20 Additionally, many current studies are limited by small sample sizes or simplistic evaluation metrics, which makes accurate assessment of the true capabilities and performance of these LLMs challenging. 21 Furthermore, a recent bibliometric analysis of machine learning in healthcare emphasizes that integrating machine learning applications into existing healthcare systems still remains a challenge. 22 This study evaluated the performance of five LLMs via large-sample clinical data drawn from multiple internal medicine subspecialties and applied comprehensive evaluation metrics. It followed the functional evaluation framework advocated in innovation capability research, 23 which emphasizes ease of interpretation and systematic assessment of distinct functional domains.
This study contributes to the literature by providing a large-sample, cross-subspecialty, multidimensional comparative evaluation of five leading LLMs using 405 real-world internal medicine cases and scoring performance across five core parts of clinical management (PCMs). It identified high-quality models suitable for clinical application and to clarify the strengths and limitations of each LLM across various aspects of clinical management, including diagnosis, differential diagnosis, examination, and treatment planning. These findings can inform future model optimization and provide valuable guidance for medical professionals selecting appropriate LLMs for clinical decision-making, thereby establishing a necessary pre-implementation foundation for integration into healthcare information systems and clinical workflows.
Method
Baseline data
This cross-sectional study retrospectively included 405 clinically representative cases across 9 internal medicine subspecialties: cardiology (Card), respiratory (Resp), gastroenterology (GI), nephrology (Neph), rheumatology (Rheum), endocrinology (Endo), neurology (Neuro), hematology (Heme), and infectious disease (ID) (45 cases per specialty). The case inclusion criteria were as follows: (a) all cases were selected from the 10th edition of the Internal Medicine textbook edited by Professor Shu Zhang (People's Medical Publishing House, 2016) 24 ; (b) all cases were clinically representative; (c) case information was complete, including chief complaint, present illness history, past medical history, personal history, physical examination, and preliminary laboratory or imaging results; and (d) each case in the textbook contained the following components: diagnosis, supporting diagnostic evidence, differential diagnoses, required additional examinations, and treatment plans. The exclusion criteria 25 were as follows: (a) incomplete case information; (b) noninternal medicine cases or cases not from the nine internal medicine subspecialties; (c) cases lacking clinical representativeness; (d) cases providing only imaging pictures without textual reports; and (e) complex cases requiring multiple follow-up questions to complete evaluation.
This study employed an equal-allocation stratified sampling approach, extracting an equal number of typical cases from each subspecialty stratum to ensure balanced representation across all subspecialties. Potential cases meeting inclusion criteria were identified from the textbook. All cases were reviewed and structurally reconstructed by clinical specialists with at least attending physician qualifications from tertiary hospitals. A standardized deidentification process was applied to fully anonymize patient identity and treatment timestamps. Forty-five clinically representative cases were selected from the eligible case pool of each subspecialty, yielding a total sample size of 405 cases. This study was approved by the Institutional Review Board of The Affiliated Guangdong Second Provincial General Hospital of Jinan University (Approval Number: 2025-KY-KZ-216-01, Date: 2025-6-4).
The study employed a parallel testing design by inputting standardized case data into five generative artificial intelligence systems: GPT-4.1, DeepSeek-R1, Claude 3.7 Sonnet-Think, Gemini 2.5 Pro Think, and OpenAI O1. Each case included the chief complaint, present illness history, past medical history, personal history, physical examination, and preliminary laboratory or imaging results. Among the 405 cases, 257 contained imaging studies; however, only textual reports, not the images themselves, were provided to the models. All the case inputs were presented in Chinese, and each model's output was also required to be in Chinese. The cases were described via professional medical terminology. Each case allowed a single query, submitted by one researcher, with no follow-up questions or supplementary information permitted.
Scoring methods and thematic coding framework
The answers provided by LLMs were independently evaluated by two physicians with over five years of clinical experience and senior professional titles. A single-blind design was employed, so the physicians were unaware of which LLM the response was generated by. The evaluation covered five aspects: diagnosis (Dx), diagnostic criteria (DC), differential diagnosis (DDs), examination (Ex), and treatment (Tx). Each category was scored out of 10 points, for a total score of 50 points. The final score for each patient was the average of the scores given by both physicians.
To ensure systematic and reproducible scoring, a structured thematic coding framework was established through three steps: (a) Initial framework establishment: Key diagnostic elements and error types were identified based on clinical practice guidelines and standard answers from the textbook; (b) Pilot coding: Two senior clinicians independently coded 20 pilot cases (4.9% of the total sample) to refine coding categories and assess inter-rater reliability; and (c) Consensus building: Discrepancies identified during the pilot phase were resolved through discussion. The detailed scoring criteria and thematic coding approach were as follows:
(1) The diagnosis section employed a six-level thematic coding system based on accuracy: Complete accuracy (10 points): Diagnosis matches the standard answer exactly, including disease type and severity grading; (b) Substantial accuracy (eight points): Core diagnosis is correct with minor terminology variations; (c) Minor deficiencies (6 points): Primary diagnosis is correct but lacks specific elements (e.g., severity grading); (d) Moderate deficiencies (4 points): Diagnosis is partially correct (approximately 50% accurate); (e) Major deficiencies (2 points): Diagnosis contains significant errors with minimal correct elements; and (f) Complete inaccuracy (0 points): Diagnosis is entirely incorrect and unrelated to the actual condition. For example, in a patient diagnosed with “Graves’ disease with hyperthyreosis crisis,” the LLM response had to match the standard answer exactly to receive the full 10 points. Responses such as “Graves’ disease with thyrotoxic crisis” received 8 points, “Graves’ disease with hyperthyroidism” received 6 points, “Graves’ disease” received 4 points, “Hyperthyroidism” received 2 points, and answers not involving hyperthyroidism or thyroid disorders received 0 points.
(2) Coding for other PCMs: “DC,” “DDs,” “Ex,” and “Tx” sections were coded using a point-based system, calculated as follows: score = 10 points/number of standard answer points × number of correct points. When the number of points provided by the LLM differed from the standard answer, correct points were included accordingly, while additional points were scored on the basis of their clinical rationality. The case and its scoring results are illustrated in Figure 1.

Example of one case and its scoring results.
Two independent raters respectively scored all LLM responses. Both raters were blinded to model identity during scoring. Each rater independently scored all 2025 responses (405 cases × 5 models) across the five evaluation parts. We assessed inter-rater reliability using both Cohen's kappa for categorical agreement and intraclass correlation coefficients (ICC) for continuous scores. For the primary diagnosis accuracy (correct/incorrect), Cohen's kappa was 0.87 (95% CI: 0.84–0.90), indicating strong agreement.
Evaluation methods
After all the responses were scored, the researchers assessed the performance and clinical utility of each LLM in medicine across multiple dimensions on the basis of their respective scores. First, by comparing overall scores, the study identified which LLM performed best in internal medicine, providing evidence for model selection. Second, performance was compared across different subspecialties to evaluate each LLM's applicability and to identify the most suitable model for each subspecialty. Third, the study analyzed each model's scores in the five key parts (Dx, DC, DDs, Ex, Tx) to determine their relative strengths in different aspects of clinical management. Finally, subgroup analyses were conducted by grouping cases according to the 9 subspecialties and comparing the five LLM scores across the five PCMs enabling a comprehensive evaluation of each LLM in internal medicine.
Model setting
In this study, we utilized five currently popular generative large language models: GPT-4.1, DeepSeek-R1, Claude 3.7 Sonnet-Think, Gemini 2.5 Pro Think, and OpenAI O1. To ensure their ability for medical consultations, each model was pretrained on specialized medical datasets to strengthen their clinical reasoning and communication abilities before testing.
Owing to its high performance, the GPT-4.1 enables healthcare professionals to quickly summarize patient information, analyze clinical data, and generate treatment or medication recommendations that can improve the curative effect. Claude 3.7 Sonnet-Think, developed by Anthropic, is an advanced conversational AI system that helps physicians identify correlations in complex medical histories and improves diagnostic accuracy. Its strong semantic analysis and reasoning skills enable it to generate clear, structured intervention plans, which enhance clinical communication and support effective decision-making. Gemini 2.5 Pro Think, created by Google DeepMind, is a multimodal AI platform that integrates medical records, imaging, and biochemical data through cross-modal analysis. It supports personalized health assessments and precise diagnoses, making it well suited for disease evaluation. OpenAI O1, developed by OpenAI, is an advanced reasoning model that synthesizes medical information and applies logical analysis to evaluate health status. This study provides trustworthy treatment recommendations and contributes to more precise clinical decision-making. DeepSeek-R1, an independently developed LLM from China, uses advanced deep learning techniques to extract and integrate critical information from complex medical data. It is particularly effective in disease prediction, diagnosis, and analysis while delivering rational treatment suggestions. DeepSeek-R1 has become a prominent focus in current medical AI research.
LLM Implementation and Technical Parameters
All LLM evaluations were conducted between 2025-4-30 and 2025-9-30. The following model versions and parameters were used: (1) GPT-4.1: Version released on 2025-4-15, accessed via OpenAI API (model identifier: gpt-4.1-turbo). Temperature = 0.3, top_p = 0.9, max_tokens = 2048. (2) DeepSeek-R1: Version released on 2025-1-20, accessed via DeepSeek API Temperature = 0.3, top_p = 0.9, max_tokens = 2048. (3) Claude 3.7 Sonnet-Think: Version released on 2025-2-15, accessed via Anthropic API (model identifier: claude-3.7-sonnet). Temperature = 0.3, max_tokens = 2048. (4) Gemini 2.5 Pro Think: Version released on 2025-3-25, accessed via Google AI Studio API. Temperature = 0.3, top_k = 40, max_tokens = 2048. (5) O1:Version released on 2025-4-15, accessed via OpenAI API . Temperature = 0.3, top_p = 0.9, max_tokens = 2048.
We used relatively low temperature settings (0.3) to minimize response variability and increase reproducibility. Each case was evaluated in an isolated session with no conversation history to prevent cross-contamination between cases. No system-level prompts were used beyond the standard default configurations for each model. For models with chain-of-thought capabilities (DeepSeek-R1), we did not explicitly request reasoning traces; responses represent the models’ standard output format.
Before each model input, the following prompt was included in the case text: "The patient's medical history and physical examination will be provided below. As a clinician of (specific subspecialties), please evaluate a patient on the basis of the provided clinical information and provide the following: (a) the primary diagnosis; (b) the diagnostic criteria for the primary diagnosis; (c) differential diagnoses; (4) further auxiliary examinations are needed for the patient; and (5) the treatment plan and medications for this patient, ensuring adherence to clinical guidelines and retaining professionalism. Please ensure your answers are professional, based on clinical guidelines, and as detailed as possible.”
Data analysis
All the data were recorded via Excel and imported into IBM SPSS Statistics 27.0 for statistical analysis. The Shapiro‒Wilk test was used to assess the normality of the data distribution. For normally distributed data, one-way analysis of variance (ANOVA) was used to compare scores among groups. For non-normal distribution data, the Kruskal-Wallis H test was used to compare scores among groups. For significant results, post-hoc pairwise comparisons were performed using Dunn's test. To account for multiple comparisons and control the false discovery rate (FDR),
Results
Overview
The scores of the five LLMs in the analysis of internal medicine cases are presented in Table 1 and Figure 2. Since the data did not follow a normal distribution, the Kruskal‒Wallis test was employed, revealing significant differences among the five LLMs (

Total scores of the five LLMs.
Total scores of the five LLMs.

Compared with Gemini,
Performance of five LLMs across nine internal subspecialties
The performance of the five LLMs across nine internal subspecialties is shown in Table 2, Table 3 and Figure 3, Figure 4. When grouped by model, significant differences were observed among nine subspecialties (all
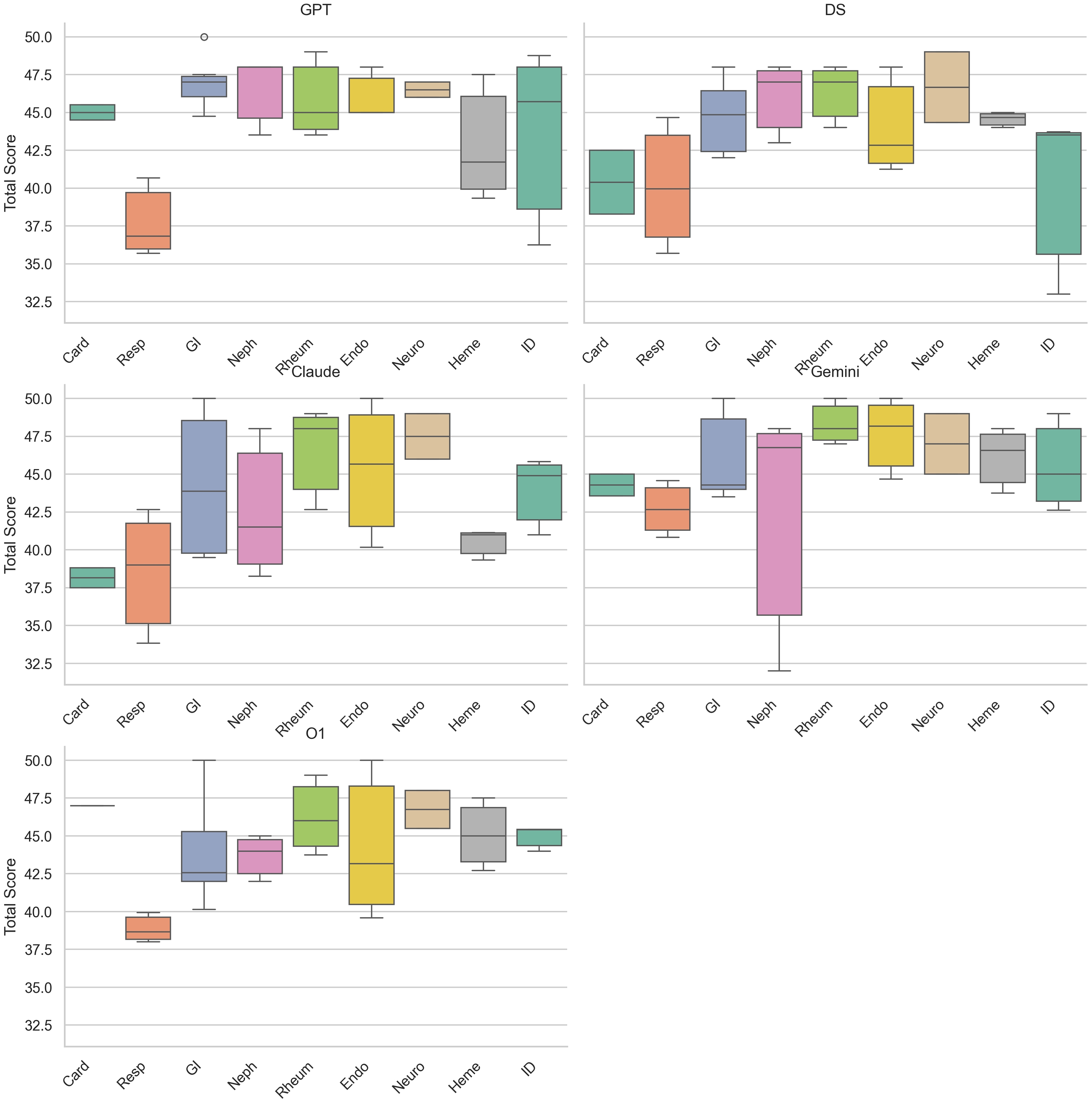
Scores of nine internal subspecialties across five LLMs.
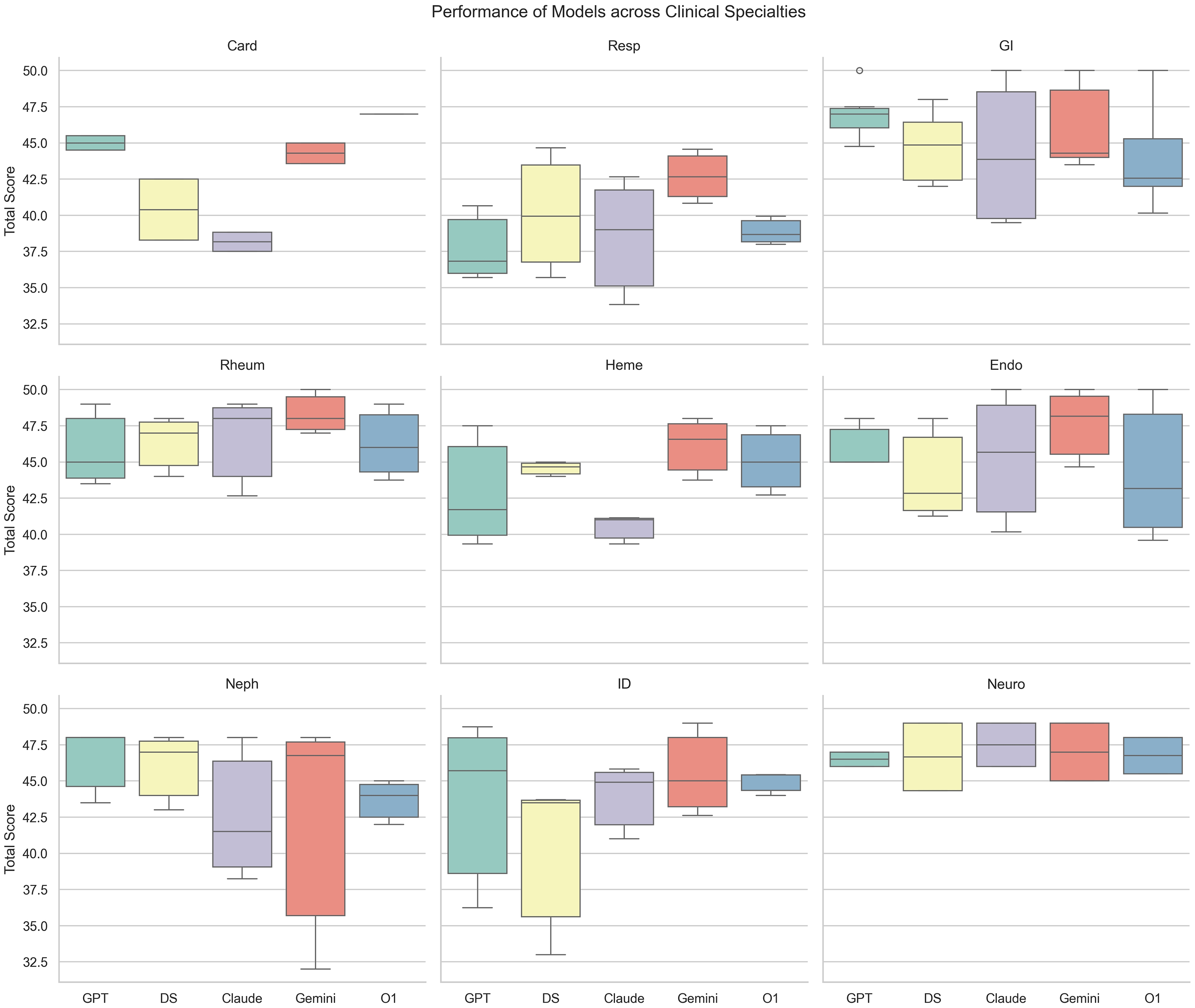
Scores of the five LLMs across nine internal subspecialties.

Scores of the five LLMs across the five PCMs.
Scores of nine internal subspecialties across five LLMs.
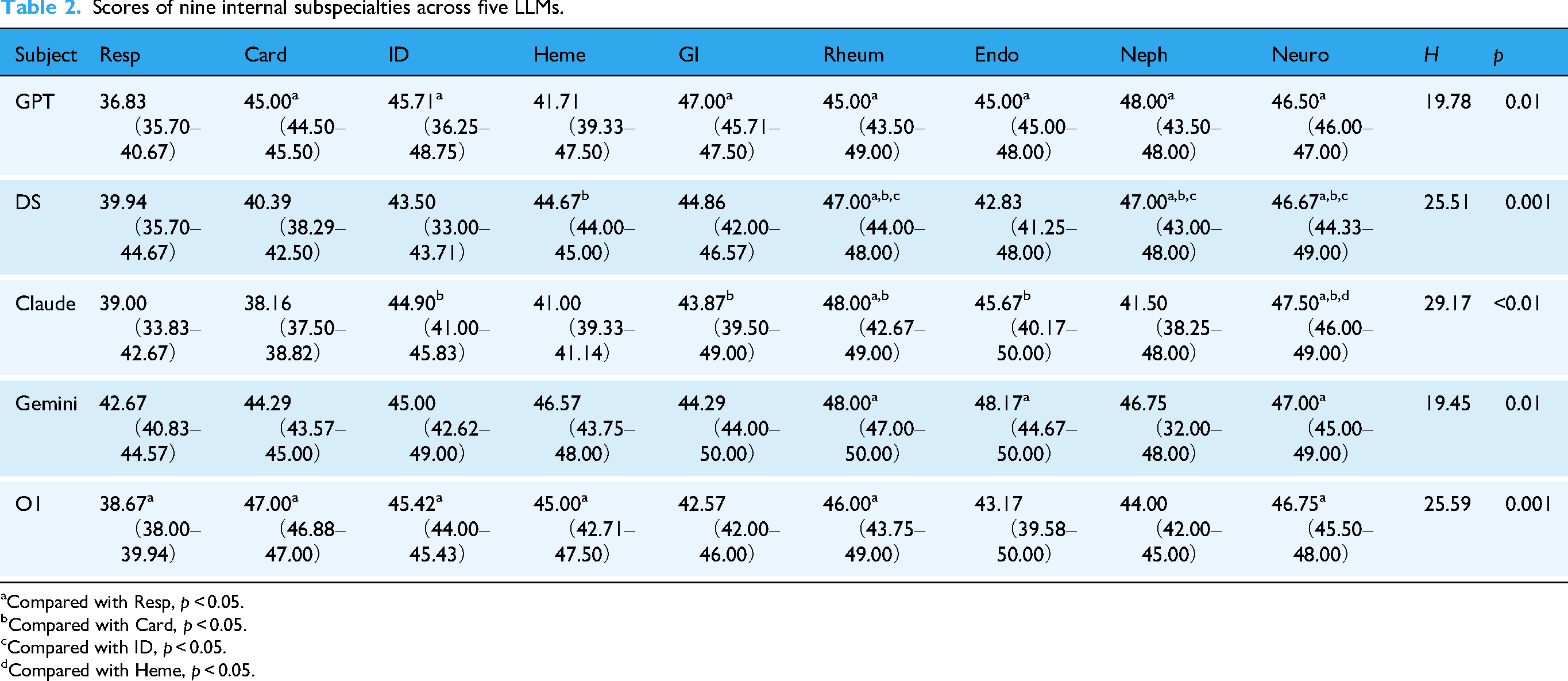
Compared with Resp,
Compared with Card,
Compared with ID,
Compared with Heme,
Scores of five LLMs across nine internal subspecialties.

Compared with Claude,
Compared with Gemini,
Compared with DS,
Scores of the five LLMs across the five PCMs.

Compared with Gemini,
When grouped by subspecialties, comparisons revealed statistically significant differences among the five models in Card and Heme (card:
Performance of the five LLMs across the five PCMs
The performance of the five LLMs across the five PCMs is shown in Table 4 and Figure 5. In the “DC,” “DX,” “Ex,” and “Tx” parts, no statistically significant differences were observed among the five models (
Results of the subgroup analysis
We further conducted a subgroup analysis of the performance of the five LLMs across the five PCMs by grouping with nine subspecialties. Kruskal–Wallis tests indicated significant differences among models in several subspecialties. Post-hoc Dunn's tests with Benjamini–Hochberg FDR correction retained the key significant contrasts while filtering out marginal findings. The results are illustrated in Figure 6.

Performance of the five LLMs across the five PCMs in subgroup analysis.
The most pronounced performance disparities were observed in the Card subgroup. Significant differences among LLMs were detected across all five PCMs: Dx (H = 14.61, adjusted
Discussion
Research on AI applications in clinical decision-making within internal medicine is a key direction for the advancement of intelligent medicine. It offers effective solutions to current medical challenges and can improve the quality of healthcare. The evaluation of LLM performance not only provides clinical evidence for improving AI but also promotes its development in the medical field. Therefore, conducting such research has significant academic value and clinical importance.26,27 In this study, we assessed the diagnostic and management capabilities of five LLMs—GPT-4.1, DeepSeek-R1, Claude 3.7 Sonnet-Think, Gemini 2.5 Pro Think, and O1—using clinical cases from nine internal medicine subspecialties. The evaluation covered five aspects of clinical management: Dx, DC, DDs, Ex, and Tx.
Principal findings and overall performance distribution
According to the results, GPT achieved the highest median score, closely followed by Gemini and O1, highlighting their superior overall performance in internal medicine cases. In contrast, Claude recorded the lowest median scores, as well as the widest interquartile range, indicating lower performance and stability. With respect to the score distribution, the maximum scores of all the models approach full points, but considerable variation exists in the minimum scores. This may indicate that performance differences still emerge when dealing with certain subspecialty cases. Therefore, we conducted analysis of the five LLMs performance across the nine subspecialties.
When grouped by model, significant differences in scores of nine subspecialties were identified across the five LLMs (all
We further compared the scores of five LLMs across five PCMs. All the models obtained high scores in four parts(Dx, DC, Ex, and Tx), and there were no significant differences between the groups. These findings suggest that LLMs may have potential advantages in quickly recognizing diseases, providing immediate diagnoses, guiding clinicians in further examinations, and recommending appropriate treatment plans. Previous research
28
has shown that LLMs possess a high level of core medical knowledge and can substantially assist in clinical diagnosis. The five LLMs only showed differences in DDs. Gemini achieved the highest median score of 9.17 (8.00–10.00), with significant differences compared with other models(all adjusted
To further explore the performance of the five models across the five PCMs, a subgroup analysis was conducted by nine subspecialties. Within the Card subgroup, statistically significant differences were detected among the five LLMs in all five PCMs (all
The universal “respiratory valley”: A multi-dimensional challenge
Perhaps the most striking and consistent finding across all five LLMs is their underperformance in Resp. For GPT, Resp scored significantly lower than seven subspecialties (Card, ID, Rheum, Endo, Neph, Neuro, GI; all adjusted
Collectively, these factors underlie the underperformance observed in LLMs within the Resp. To overcome this fundamental bottleneck, future development of LLMs for respiratory clinical decision-making should place greater emphasis on advancing multimodal integration capabilities.
Differential diagnosis: Gemini's unique advantage in diagnostic breadth
An interesting phenomenon emerged in the analysis of clinical management. While no significant differences of LLMs were observed in DC, Ds, Ex, Tx, Gemini significantly outperformed all other models in DDs (H = 29.52, adjusted
This divergence likely reflects differences in model training philosophy and safety calibration. Gemini appears to prioritize recall over precision, favoring a “cast a wide net” strategy that maximizes coverage of plausible diagnoses, even at the risk of including less likely options. This design aligns with Google DeepMind's principles of medical AI safety. In contrast, models like GPT and DS may be optimized for rapid convergence to a single best answer, which improves efficiency but may sacrifice diagnostic breadth.
From a cognitive psychology perspective, this behavior exhibited by Gemini represents a debiasing mechanism, serving to mitigate the cognitive error of “premature diagnostic closure”—the tendency of clinicians to focus on a single diagnosis before adequately excluding other possibilities. Therefore, this capacity has considerable value in disrupting cognitive rigidity and supplementing the blind spots inherent in physicians’ assessments. This finding suggests that different LLMs may assume complementary and synergistic roles in clinical practice: models such as GPT can function as “precision diagnostic engines,” well-suited for rapid, high-confidence decision-making in time-sensitive settings such as the emergency department; whereas Gemini can serve as a “diagnostic safety net,” safeguarding diagnostic completeness in complex or diagnostically ambiguous cases. Healthcare institutions may benefit from deploying a multimodel framework that leverages the differentiated strengths of each model, thereby maximizing overall diagnostic efficacy.
Subgroup analysis: Complexity of subspecialty as a performance differentiator
In our comparison of the five LLMs across nine subspecialties, significant inter-model differences were identified exclusively in Card and Heme. Subgroup analysis further reinforced the distinctive status of Card, which emerged as the only subspecialty demonstrating significant intermodel differences across all five PCMs: Dx (H = 14.61, adjusted
The subgroup analysis indicates that the difficulty levels of subspecialties may act as a “magnifying glass” for model performance differences. In subspecialties with low-to-moderate complexity, diagnostic pathways are often standardized; consequently, all models achieved near-ceiling performance, masking subtle disparities in reasoning depth. This implies that for clinical management in less complex specialties, mainstream LLMs are largely interchangeable.
Conversely, Card demands the integration of ECG interpretation, hemodynamic calculations, multi-drug interactions, and comorbidity management, thereby imposing substantially greater demands on the models’ deep reasoning capabilities. This multidimensional, highly integrative reasoning requirement constitutes a genuine “stress test” of model capability, underscoring that in high-complexity subspecialties. In such subspecialties, it is more appropriate to select the best-performing model based on their performance in clinical management. Post-hoc pairwise comparisons within the Card subgroup indicated that O1 consistently achieved higher scores and significantly outperformed other models in DC and DDs (all adjusted
Therefore, future benchmarks for medical LLMs must intentionally incorporate “high-stress” subspecialties (e.g., cardiology, multisystem critical care) to effectively differentiate model capabilities; otherwise, evaluations risk lacking discriminative power.
Improvement and future development of LLMs
On the basis of the observed performance differences among LLMs in internal medicine clinical decision-making, we propose the following strategies for improvement and future development.
1. Model enhancement strategies: First, given the consistently poor performance of all LLMs in respiratory diseases, we recommend building a specialized pulmonology knowledge base. This database should comprehensively cover clinical manifestations, imaging features, and the differential diagnosis of complex respiratory conditions. Additionally, data collection efforts should be strengthened for rare respiratory diseases and atypical presentations by the acquisition of high-quality multicenter clinical datasets. Differential diagnosis remains a common limitation across all models. To address this, future research should prioritize strengthening disease association reasoning capabilities in LLMs. This could be achieved by enriching training datasets with expert-annotated differential diagnosis examples that explicitly integrate diagnostic and pathophysiological relationships among diseases, thereby constructing more robust frameworks for disease differentiation, improving both the systematic approach and accuracy of differential diagnosis performed by LLMs.
To optimize model performance, multimodal fusion strategies should be implemented to integrate clinical symptoms, laboratory results, imaging data, and patient information, resulting in more comprehensive clinical decision-making systems.
29
Hybrid model architectures based on ensemble learning should be developed; these can leverage the complementary strengths of different LLMs across subspecialties through dynamic weight allocation.
2. Evidence-based model selection: This study reveals that Gemini has unique advantage in diagnostic breadth. GPT and O1 offer significant potential for medical AI development. They can serve as foundational architecture for creating models tailored to various medical subspecialties. By making targeted adjustments according to the features of each subspecialty, an intelligent medical system covering general medicine can be established. Furthermore, applying supervised and reinforcement learning, especially under the guidance of internal medicine experts, can enhance the clinical management capabilities of LLMs. For models with relatively weak performance and poor stability, such as Claude, their use may be considered cautiously and limited to auxiliary, nondecisional support, pending further validation.
We also propose developing an evidence-based LLM decision-making system to provide model recommendations tailored to different subspecialties and clinical scenarios. O1 may be prioritized in cardiology, particularly for diagnosis and differential diagnosis. For other subspecialties, the choice of LLM has limited impact on overall performance. However, in cardiology and hematology, the use of Claude might need to avoided. Additionally, for differential diagnosis tasks in internal medicine, Gemini can be recommended as the preferred model.
3. Policy and Regulation: The integration of LLMs into clinical decision-making require the establishment of comprehensive policy and regulatory mechanisms.
30
Policymakers should develop standardized evaluation protocols for AI-assisted diagnostic systems, ensuring that LLMs meet rigorous safety and efficacy standards before clinical deployment. Continuous performance monitoring, clinician feedback, and patient safety reporting systems must be incorporated.
31
Data governance policies are equally critical, requiring strict adherence to patient privacy protection regulations while facilitating secure data sharing for continuous model improvement. 4. Implications for Stakeholders: This study provides actionable insights for all stakeholders in the healthcare system. For clinicians, we recommend developing subspecialty-specific LLM selection guidelines. Training programs should be established to enhance clinician proficiency in interpreting LLM outputs and recognizing model limitations, particularly in differential diagnosis. For hospital administrators, they should focus on integrating high-performing models (such as GPT, Gemini) into electronic health record systems while implementing quality assurance mechanisms to monitor AI-assisted decision outcomes. For patients, transparent communication protocols must be developed to ensure informed consent regarding AI involvement in their care, clearly explaining both the benefits and limitations of LLM-assisted diagnosis. For AI developers, this study identifies critical improvement areas: enhancing differential diagnosis capabilities across all models, developing specialized respiratory disease modules, and improving performance consistency across diverse clinical scenarios. 5. Integration with Digital Health Technologies: The integration of LLMs with Internet of Things (IoT) devices and e-health platforms has the potential for enhancing quality of life (QoL), particularly for patients with chronic internal medicine conditions.
32
Patient monitoring through IoT-enabled wearable devices (such as continuous glucose monitors, cardiac monitors, and pulse oximeters) can generate continuous physiological data that enable proactive clinical decision-making after processing by LLMs. For instance, combining cardiovascular IoT devices with O1 models could enable real-time arrhythmia detection and automated alert systems for high-risk patients. E-health platforms can leverage LLMs to provide 24/7 virtual consultation services, particularly valuable in resource-limited settings or for patients with mobility constraints. Telemedicine systems integrated with subspecialty-optimized LLMs can deliver preliminary diagnostic assessments, triage patients efficiently, and recommend appropriate specialist referrals. To realize this vision, technical challenges must be addressed, including ensuring interoperability between IoT devices and LLM platforms, developing robust data security protocols for real-time health data transmission, and creating user-friendly interfaces that accommodate diverse digital literacy levels among patients and healthcare providers. 6. Ethical Considerations: We should examine the privacy implications of integrating LLMs with clinical data systems, electronic health records, and IoT-enabled medical devices. Informed consent and patient autonomy must be preserved in AI-mediated healthcare.
33
Patients have the right to know when AI systems are involved in their care, understand the capabilities and limitations of these systems (such as this study's finding that all LLMs perform poorly in respiratory disease management), and choose whether to accept AI-assisted recommendations. Healthcare institutions must develop clear informed consent processes that explain in accessible language how LLMs analyze patient data, what role they play in clinical decision-making, and what human oversight mechanisms are in place.
With the development of AI technology, LLMs will continuously learn and improve from clinical practice while preserving patient privacy. AI technological development will make reasoning processes more transparent and precise, thereby enhancing clinician trust. 34 Research in the future should focus on prospective validation of LLMs in real-world clinical environments, establishing standardized evaluation systems and quality-control mechanisms to ensure the safety and efficacy of AI-assisted diagnosis and treatment, laying a solid foundation for widespread intelligent healthcare applications.
Limitations
This study has several limitations. First, all the evaluations and model interactions were conducted in Chinese, which may introduce language-related bias. Most LLM training data are primarily based on English medical literature and clinical materials, and their performance in Chinese medical contexts may differ from that in English environments. The expression patterns of Chinese medical terminology, linguistic habits in disease descriptions, and cultural differences in clinical thinking may affect the understanding and reasoning capabilities of LLMs. Additionally, the capabilities of Chinese language processing differ among LLMs, which may lead to underestimation or overestimation of the performance of certain models, affecting the fairness of intermodel comparisons. A US study 25 revealed that the GPT-4 may produce biased outputs when processing clinical problems because of factors such as language, ethnicity, and gender. Future research should consider conducting multilingual comparative studies to evaluate the specific impact of language factors on LLM medical performance.
Second, this study focused mainly on internal medicine and did not involve other specialties, such as surgery, pediatrics, or obstetrics and gynecology. This will have a certain impact on the universality of the results and their clinical application. Owing to significant differences in disease spectra, diagnostic thinking, and decision-making patterns across medical specialties, the results from internal medicine may not be directly adopted in other specialties. For example, surgery focuses more on surgical indication assessment and perioperative management; pediatrics require the consideration of age-related physiological characteristics and medication dosage adjustments; and obstetrics and gynecology involve special physiological states and hormonal influences.35,36 Therefore, the clinical application of the results from this study is limited primarily to internal medicine, and to construct a comprehensive medical AI evaluation system, similar systematic research needs to be conducted in other specialties.
Third, the rapid development and updates of LLMs affected the timeliness of this study. During the implementation of this study, some AI companies released model updates that may have included performance improvements, knowledge base expansion, and algorithm optimization. This somewhat limits the practical value of the research results.
Fourth, an important limitation of this study is the use of textbook-based reconstructed cases rather than actual cases from real patients. While this approach ensures standardization and reproducibility, textbook vignettes typically present information in a more structured format with less clinical ambiguity, incomplete data, and atypical presentations than encountered in actual clinical practice. This may inflate LLM performance and reduce generalizability to real-world clinical decision-making.
Additionally, we evaluated each case using single responses without assessing output variability. Given the probabilistic nature of LLMs, responses can vary across multiple runs, and without repeated sampling, we cannot determine consistency or reliability—metrics essential for clinical decision support.
To address these limitations, we are currently conducting a follow-up study using real clinical cases from actual patients. This subsequent research will directly compare LLM performance between textbook-based and real-world clinical cases, allowing us to quantify potential performance differences and identify specific areas where real-world complexity poses greater challenges to LLMs. Importantly, to assess response consistency and reliability, we have implemented a repeated-sampling protocol in which each case is evaluated with at least three independent LLM responses, enabling analysis of output variability and identification of cases where models produce inconsistent recommendations. This comparative approach will provide more robust evidence regarding the clinical applicability of LLMs and help reduce bias in assessing their true capabilities for clinical decision support.
These limitations suggest that future research should adopt larger-scale multicenter collaborative research designs, expand them to multiple medical specialties, and establish dynamic evaluation mechanisms to adapt to the rapid development of AI technology. Moreover, we recommend establishing standardized AI medical application evaluation frameworks to ensure the comparability and timeliness of research results, providing more reliable evidence-based support for the safe and effective application of AI in clinical medicine.
Conclusion
This study systematically evaluated the performance of five LLMs in internal medicine clinical decision-making, revealing the potential of applying AI in the medical field. The results demonstrated that GPT exhibited optimal performance in terms of comprehensive capabilities, whereas the Claude model showed relatively poor performance and stability. All five LLMs demonstrated high performance in four PCMs, indicating their significant value in assisting in clinical decision-making for physicians. However, all the models showed deficiencies in differential diagnosis, particularly poor performance in managing respiratory diseases, reflecting the limitations of current LLMs in understanding complex diseases. The complexity of subspecialty might be a performance differentiator for LLMs and model such as O1 could perform better in complex subspecialties such as cardiology.
This study provides initial, textbook-based evidence for LLM applications in the clinical practice of internal medicine subspecialties, offering reference on how to select appropriate models on the basis of different subspecialties. The findings suggest that we should focus on optimizing the weak areas of models, improving algorithms for differential diagnosis, and establishing standardized evaluation systems for the medical application of AI to promote the safe and effective application of AI technology in clinical practice.
The integration of LLMs into clinical practice requires co-ordinated efforts from multiple stakeholders to ensure safe and effective implementation. Policymakers should establish comprehensive evaluation frameworks for medical AI, mandating rigorous safety and efficacy standards before clinical deployment, along with continuous performance monitoring and patient safety reporting systems. Healthcare administrators should develop evidence-based implementation strategies, prioritizing selecting most appropriate model for subspecialties while implementing quality assurance mechanisms and integrating high-performing models into electronic health record systems. Physicians must maintain primary responsibility for clinical decisions, using LLMs as decision support tools. Training programs should enhance clinician proficiency in interpreting AI outputs and recognizing model limitations. Patients should be informed about AI involvement through transparent communication protocols that explain both benefits and limitations, preserving their autonomy to accept or decline AI-assisted recommendations. AI developers must prioritize addressing identified weaknesses by enhancing differential diagnosis capabilities, developing specialized respiratory disease modules, and improving cross-subspecialty performance consistency through collaboration with clinical experts. Future development should include establishing national LLM evaluation standards, creating evidence-based guidelines for LLM use in each subspecialty, creating AI training programs for physicians, establishing patient safety reporting systems and integrating LLMs with IoT devices and e-health platforms.
Supplemental Material
sj-doc-1-dhj-10.1177_20552076261433073 - Supplemental material for Performances of five large language models in clinical decision-making for internal medicine: A comparative study
Supplemental material, sj-doc-1-dhj-10.1177_20552076261433073 for Performances of five large language models in clinical decision-making for internal medicine: A comparative study by Dan Wu, Jingjing Lu and Danghan Xu in DIGITAL HEALTH
Supplemental Material
sj-docx-2-dhj-10.1177_20552076261433073 - Supplemental material for Performances of five large language models in clinical decision-making for internal medicine: A comparative study
Supplemental material, sj-docx-2-dhj-10.1177_20552076261433073 for Performances of five large language models in clinical decision-making for internal medicine: A comparative study by Dan Wu, Jingjing Lu and Danghan Xu in DIGITAL HEALTH
Footnotes
Abbreviations
Acknowledgments
Not applicable.
Ethics approval and consent to participate
This study was approved by the Institutional Review Board of The Affiliated Guangdong Second Provincial General Hospital of Jinan University (Approval Number: 2025-KY-KZ-216-01, Date: 2025-6-4). The IRB classified this research as minimal risk because it involved evaluation of publicly available educational materials rather than patient data. No patient consent was required as no human subjects were involved.
Consent for publication
Not applicable.
Authors’ contributions
DW conducted the investigation, collected the data, and drafted the original manuscript. JJL designed the methodology, coordinated resources, and carried out the data analysis. DHX conceived the study and provided overall supervision.
All authors contributed to interpreting the findings, revised the manuscript, and approved the final version.
Competing interests
Author Danghan Xu is affiliated with Guangzhou Yuansheng Ruihang Medical Information Technology Co., Ltd, a medical technology company. However, this affiliation is unrelated to the current study. The company had no role in study design, data collection, analysis, interpretation of results, manuscript preparation, or the decision to submit for publication. The company does not develop or market any of the large language models evaluated in this study, and the author received no financial support from the company for this research. All other authors declare no conflicts of interest. None of the authors have financial relationships with OpenAI (GPT-4.1, O1), DeepSeek (DeepSeek-R1), Anthropic (Claude 3.7 Sonnet-Think), Google (Gemini 2.5 Pro Think), or any other entities involved in the development of the evaluated models.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The original clinical cases were derived from the 10th edition of Internal Medicine textbook (People's Medical Publishing House, 2016), which we have obtained permission to use the data. However, all cases were substantially restructured for this study to create clinically realistic presentations suitable for LLM evaluation. The complete dataset, including both the original textbook cases and our restructured versions used for LLM evaluation, is provided in Supplemental Appendices 1 and 2. These materials allow full reproduction of our study.
Supplemental material
Supplemental material for this article is available online.
