Abstract
Background
Diagnostic performance of generative artificial intelligences (AIs) using large language models (LLMs) across comprehensive medical specialties is still unknown.
Objective
We aimed to evaluate the diagnostic performance of generative AIs using LLMs in complex case series across comprehensive medical fields.
Methods
We analyzed published case reports from the
Results
Out of 557 consecutive case reports, 392 were included. The inclusion rates of the final diagnosis within top 10 DDx lists were 86.7% (340/392) for ChatGPT-4, 68.6% (269/392) for Google Gemini, and 54.6% (214/392) for LLaMA2 chatbot. The top diagnoses matched the final diagnoses in 54.6% (214/392) for ChatGPT-4, 31.4% (123/392) for Google Gemini, and 23.0% (90/392) for LLaMA2 chatbot. ChatGPT-4 showed higher diagnostic accuracy than Google Gemini (
Conclusions
This study demonstrated the diagnostic performance of generative AIs including ChatGPT-4, Google Gemini, and LLaMA2 chatbot. ChatGPT-4 exhibited higher diagnostic accuracy than the other platforms. These findings suggest the importance of understanding the differences in diagnostic performance among generative AIs, especially in complex case series across comprehensive medical fields, like general medicine.
Keywords
Introduction
Diagnosis is fundamental to the practice of medicine. Pursuing diagnostic excellence is crucial for enhancing the quality and outcomes of medical care. Diagnostic excellence encompasses a holistic approach that ensures diagnoses are not only accurate and precise, but are also delivered in a manner that is safe, effective, patient-centered, timely, efficient, and equitable. 1 This concept underscores the imperative of minimizing errors, 2 a critical step in optimizing the use of resources. Furthermore, it emphasizes the necessity to respect patients’ unique needs and experiences throughout the diagnostic journey. Achieving diagnostic excellence necessitates the implementation of various strategies. Promoting a team-based diagnostic process emerges as particularly critical among these strategies. This strategy stands out due to its collaborative nature, thereby improving diagnostic accuracy and comprehensiveness through collective reasoning. 3 This collaborative approach involves not only healthcare providers, patients, and their families, but also integrates an essential tool: Clinical Decision Support Systems (CDSSs). 4
The primary design goal of CDSSs is to enhance healthcare quality by refining the medical decision-making process. They play an impressive role in enhancing overall patient care. 5 CDSSs offer a multitude of capabilities, including diagnostic support. Representative examples of CDSSs underscore the diverse range of applications available. Examples of these tools include symptom checkers 6 and differential-diagnosis (DDx) generators, 7 such as DXplain. 8 Symptom checkers are online tools that yield a list of possible conditions from reported symptoms. DDx generators provide physicians with potential diagnoses based on collected medical information. DDx in the context of clinical practice denotes the comprehensive list of potential diagnoses compiled to account for a patient's symptoms, 9 a process streamlined by advanced DDx generators. DXplain, one of the DDx generators, employs artificial intelligence (AI) to analyze patient data and provide possible diagnoses based on clinical features. Despite their potential, several factors have limited the impact of CDSSs on clinical diagnostics. These factors include persistent negative perceptions among physicians and issues with accuracy. Another hurdle is the lack of seamless system integration, 5 which is essential for the effortless incorporation of CDSSs into existing healthcare workflows and electronic health record systems.
The limitations inherent in current CDSSs have necessitated the exploration of innovative solutions. Emerging as a promising response to these challenges is generative AIs, especially large language models (LLMs). 10 Generative AIs’ ability to process extensive amounts of data with human feedback, understand nuanced human language, and generate precise, contextually relevant responses overcome some of the critical drawbacks of earlier systems. These AIs hold the potential to enhance diagnostic accuracy, personalized treatment recommendations, and improve patient engagement, addressing the gaps in existing CDSSs’ accuracy and adaptability. In this innovative domain, several LLM-based chatbots have gained prominence as notable generative AI tools, 11 including the chat generative pre-trained transformer (ChatGPT) developed by OpenAI, 12 Google Gemini,13,14 and LLM meta AI 2 (LLaMA2), a product of meta AI. 15
These new generative AI technologies are poised to address pivotal questions in DDx. 16 Several studies have extensively researched the clinical potential of generative AI tools like the fourth-generation ChatGPT (ChatGPT-4).17–19 Our investigation, among others, has shown that ChatGPT-4 exhibits more impressive diagnostic accuracy than its predecessor, the third-generation ChatGPT, ChatGPT-3.20,21 Other studies indicated ChatGPT-4's superiority over other generative AIs in specialized fields, such as neurosurgery question banks, 20 respiratory medicine questions, 22 and general internal medicine (GIM) case series. 23
Despite the promise shown by ChatGPT-4 and other generative AIs, few studies have comparatively analyzed their diagnostic performance. This gap exists because of certain tools, including Google Gemini and LLaMA2 chatbot utilizing the LLaMA2 model, remain under-evaluated. Additionally, their diagnostic performance across comprehensive medical specialties is still unknown. Although some studies have demonstrated their efficacy in comprehensive medical exams,19,24 evaluations based on case reports, which are closer to clinical applications, have yet to be fully understood. This can be attributed to the intrinsic complexity and specialized nature of different medical fields, 25 which pose challenges for conducting broad-scale evaluations.
Therefore, this study aims to fill this gap in the current literature by comparing the diagnostic performance of ChatGPT-4, Google Gemini, and LLaMA2 chatbot for case series across a comprehensive range of medical specialties. By offering a comprehensive assessment across these diverse areas of medicine, this study not only seeks to fill a current gap in the literature but also to guide future development and applications of generative AIs toward enhancing diagnostic excellence.
Materials and methods
We conducted an experimental study to assess diagnostic accuracy of generative AI systems using data from published case reports. This study was conducted by the Department of GIM (Diagnostic and Generalist Medicine) at Dokkyo Medical University in Tochigi, Japan. As the research utilized published case reports, ethics committee approval was not applicable. The study involved preparing case materials, generating DDx lists utilizing AIs, and evaluating these lists. Figure 1 illustrates the study flow.

Flowchart in the study.
Preparing case materials
To provide a detailed understanding of the selection criteria for case reports, we included specific inclusion and exclusion criteria to ensure the generalizability of our findings. The case reports were sourced from the
We edited these reports to highlight the case descriptions and final diagnoses. Typically, the case descriptions were extracted from the case report section of each article, which often detailed patient history, physical examinations, investigation results, and management prior to the final diagnosis. Other sections like titles, introductions, discussions, conclusions, and assessments present in the case report section, and any accompanying tables or figures were omitted. Any disagreements between TH and YH were discussed until a resolution was reached.
As an example, consider the case reports titled “A 75-Year-Old Woman with a 5-Year History of Controlled Type 2 Diabetes Mellitus Presenting with Polydipsia and Polyuria and a Diagnosis of Central Diabetes Insipidus,” 27 which was referenced as case number 1 in Supplementary Tables 1–3. From this article, we extracted case descriptions: from “A 75-year-old Japanese woman visited a primary care doctor due to a 2-month history of thirst, polydipsia…” to “The plasma AVP level was relatively low (0.6 pg/mL).” The final diagnosis for this case was central diabetes insipidus.
Generating differential-diagnosis lists utilizing artificial intelligences
In this study, we employed three different generative AIs: ChatGPT-4, Google Gemini, and LLaMA2 chatbot. None of the generative AIs, including ChatGPT-4, Google Gemini, and LLaMA2 chatbot, received additional training or reinforcement for medical diagnoses. Table 1 provides the details of the generative AIs employed in this study.
Details of generative artificial intelligences employed in this study.

ChatGPT-4: fourth-generation chat generative pre-trained transformer; LLaMA2: large language models by Meta AI 2.
We utilized ChatGPT-4, a generative AI developed by OpenAI. This model, known for its advanced generation and comprehensive capabilities, is the GPT-4 series. We employed the 24th May version of ChatGPT-4 to generate DDx lists. In the version, there was no Internet connection option. ChatGPT-3.5 was excluded from this study due to its comparatively lower medical performance as observed in prior research.28,29 Specifically, ChatGPT-4 demonstrated a significantly higher accuracy, surpassing ChatGPT-3.5. Given our aim to evaluate the most advanced generative AI systems, we selected ChatGPT-4. Figure 2A presents an example output of DDx list generated by ChatGPT-4 for a sample case description. Figure 2B explains Figure 2A. Supplementary Table 1 lists the DDx generated by ChatGPT-4 alongside the final diagnosis.

Example output from the fourth-generation chat generative pre-trained transformer (ChatGPT-4) for a sample case description (A) and the explanation (B).
We also employed Google Gemini (previously Bard), an experimental AI application from Google, known for being updated daily. While it does not possess a specific version, Google Gemini is recognized for its robust language processing and generation skills. Figure 3A presents an example output of DDx list generated by Google Gemini for a sample case description. Figure 3B explains Figure 3A. Supplementary Table 2 lists the DDx generated by Google Gemini and the final diagnosis.
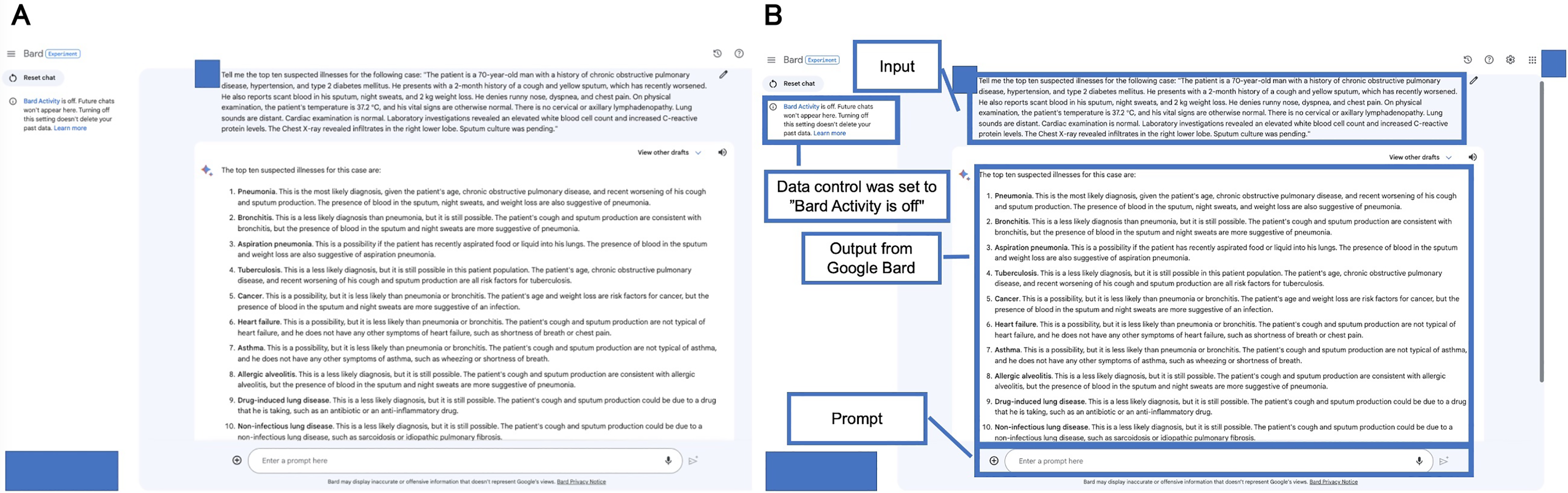
Example output from Google Gemini (previously Bard) for a sample case description (A) and the explanation (B). The date was accessed before the system transitioned to Google Gemini.
Additionally, LLaMA2 chatbot, an open-source application based on the LLaMA2 model, was available in versions 7B, 13B, and 70B. We employed the 70B version, the largest and most complex model among the options. The application allowed adjustments in settings such as temperature, top P, max sequence length, and prompt setting. The temperature parameter, which influences the model's randomness and creativity, ranged from 0.01 to 5.00. The top P parameter option adjusts the randomness of the model's predictions, ranging from 0.01 to 1.00. The max sequence length parameter specifies the maximum number of word segments, tokens that can be generated by the model, ranging from 64 to 4096. The prompt setting was configured according to the role defined in the LLaMA2 model. Parameters set included a temperature of 2.49, a top

Example output from large language model Meta AI 2 (LLaMA2) chatbot for a sample case description (A) and the explanation (B).
For all generative AIs, TH utilized a consistent prompt for each case description: “Tell me the top 10 suspected illnesses for the following case: (copy and paste each case description).” This prompt aimed to encourage the generative AIs to output a DDx list. The selection of the prompt was based on preliminary testing. The initial lists generated by the AIs served as the DDx lists. We cleared previous chat histories before inputting new case descriptions. To preclude any interference from previous interactions, we ensured that ChatGPT-4 and Google Gemini did not retain chat histories. Similarly, the LLaMA2 chatbot was used without a data control setting, preventing carryover effects from prior sessions.
Evaluating the differential-diagnosis lists
Two GIM expert physicians (KM and TS; Tetsu Sakamoto), working independently, reviewed each final diagnosis and the DDx lists generated by AIs. The physicians assigned a binary code to each item. An item was coded “1” if it either accurately matched the final diagnosis with an acceptable specificity or was sufficiently close to the final diagnosis such that proper treatment would be initiated without compromising patient safety. A “0” was assigned if the items were those that were significantly different from the final diagnosis, such that treatment based on the responses would not have been effective. 30 Discrepancies between reviewers were resolved through consultation with an additional GIM expert physician (KT). All physicians in this evaluation process were blind to the specific generative AI that produced each list.
Measurement
Our primary outcome was the rate of the final diagnosis within the top 10 DDx lists generated by the AIs. The rate of the final diagnosis within the top 5 DDx lists and the rate of the final diagnosis as a top diagnosis were measured as secondary outcomes. Additionally, interrater reliability between the physicians’ evaluation for the DDx lists was calculated as Cohen's kappa coefficient.
Statistical analysis
We analyzed the data using R version 4.2.2 (The R Foundation for Statistical Computing, Vienna, Austria) with the stats library (version 4.2.2). Categorical or binary variables were presented as numbers (percentages), and comparisons were made using the chi-square test. Given the multiple comparisons performed in this study, we applied the Bonferroni correction to adjust the significance levels and control the familywise error rate,
31
It adjusts the alpha or significance level by dividing it by the number of comparisons (three generative AIs in this study: ChatGPT-4, Google Gemini, and LLaMA2 chatbot) being made thereby setting a more stringent threshold for individual tests to be considered significant. The Bonferroni-corrected significance level was defined as a
Results
Included case reports
A total of 392 case reports were included. Three generative AIs, including ChatGPT-4, Google Gemini, and LLaMA2 chatbot, answered all 392 case descriptions. Table 2 displays the representative DDx list by three generative AIs. The DDx lists for all case descriptions generated by ChatGPT-4, Google Gemini, and LLaMA2 chatbot are shown in Supplementary Tables 1–3, respectively.
Representative differential-diagnosis list by generative artificial intelligences and final diagnosis.

ChatGPT-4: fourth-generation chat generative pre-trained transformer; LLaMA2: large language models by Meta AI 2.
Diagnostic performance
Table 3 and Figure 5 present the diagnostic accuracy of three generative AIs. Regarding the rate of final diagnoses within the top 10 DDx lists, ChatGPT-4 demonstrated 86.7% (340/392), which was statically higher than those of Google Gemini (269/392, 68.6%,

Diagnostic performance of generative artificial intelligences.
Diagnostic accuracy of generative artificial intelligences.

ChatGPT-4: fourth-generation chat generative pre-trained transformer; DDX: differential-diagnosis; LLaMA2: large language models by Meta AI 2.
Chi-squared test.
Discussion
Principal results
Several key findings emerged from this study, providing insights into the diagnostic performance of different generative AIs. ChatGPT-4 demonstrated superior diagnostic accuracy compared to Google Gemini and LLaMA2 chatbot. The primary distinction lies in the LLMs powering these AIs, yet this superiority could be attributed to ChatGPT's earlier introduction to the public, allowing more time for reinforcement. The timing of release may have provided more time for iterative improvements based on user feedback and real-world performance data. 33 Additionally, ChatGPT-4's fee-based application model could contribute to its superior performance. The fee-based model could influence the frequency of model updates and the extent to which user feedback is incorporated. The superiority of ChatGPT-4 highlighted that generative AIs have the potential of rapid improvement in diagnostic performance, even without specific training within the diagnostic domain.
In contrast to ChatGPT-4's marked success, our analysis revealed a difference in the performance of the other AI platforms tested. Google Gemini performed better diagnostically than LLaMA2 chatbot. This result indicates that LLaMA2 chatbot requires more refinement in the diagnostic domain. While the fundamental difference was the underlying LLMs of these AIs, the variability in performance could be due to the adjustable parameters unique to the LLaMA2 chatbot in the current study, an aspect of flexibility absent in ChatGPT-4 and Google Gemini. For example, LLaMA2 chatbot allows adjustments to parameters including temperature, which influences randomness of the generated responses. Therefore, LLaMA2 chatbot might improve with the optimization of these parameters.
We found that these generative AIs exhibited impressive diagnostic accuracy when presented with complex case descriptions, even without specialized medical training and tuning. These results highlight the necessity of recognizing the differences in diagnostic performance among generative AIs, especially for complex case series across comprehensive medical fields like GIM. Generative AIs could serve as a form of collective intelligence, pooling knowledge and insights from various sources, proficient in verifying medical diagnoses made by healthcare professionals.
The differences in diagnostic performance among ChatGPT-4, Google Gemini, and LLaMA2 chatbot can be attributed to the underlying technologies, training data, and iterative improvements specific to each AI. ChatGPT-4's early introduction, extensive training data, and commercial model likely contribute to its superior performance. In contrast, Google Gemini benefits from Google's advanced AI infrastructure, while LLaMA2's performance indicates the need for further refinement and parameter optimization. Understanding these factors is essential for developing more effective generative AI systems in the medical diagnostic domain.
Limitations
A primary limitation of this study is its reliance on case materials from a single-case report journal. While this ensured consistency in the data, it introduced selection bias, limiting the generalizability to a broader range of medical scenarios. Different journals may emphasize distinct aspects in their case descriptions. This diversity could impact the diagnostic performance of generative AIs. Therefore, our results may not fully represent the diversity of real-world clinical situations. It is imperative for healthcare professionals to evaluate the efficacy and safety of generative AI systems in real-world cases across various settings to ascertain their capabilities comprehensively. 34 Furthermore, we excluded pediatric cases and those primarily focused on management from our analysis. This exclusion may have influenced the results and introduced additional selection bias.
Additionally, we did not compare the diagnostic performance of physicians directly with generative AIs. This decision stemmed from the view that generative AIs are intended to supplement, not replace, physicians. 35 A comparative study could provide deeper insights into how generative AIs perform relative to medical professionals and highlight areas where AI can most effectively augment clinical decision making.
Moreover, our study focused solely on textual data, excluding tables and figures, which could be crucial for assessing the diagnostic performance of generative AIs. While our previous study revealed that visual information did not improve the diagnostic accuracy of ChatGPT-4 for case report series, 36 visual information often plays a significant role in medical diagnostics, and its exclusion may have limited the scope of our findings.
As for the limitations of generative AIs, it is crucial to note that these AI tools lack formal medical approval. Additionally, optimal prompt settings, such as temperature, Top P, max sequence length, the setting of prompt, and prompt text, remain to be established. Moreover, our study did not encompass all available generative AI platforms. 10 Future research should comprehensively explore these generative AIs.
Another interesting aspect that was not fully explored in the current study is the comparison of different AI systems’ performance with regard to their training data. Different AI systems are trained on varied datasets, which can significantly influence their diagnostic capabilities and performance. Future research should delve into how the nature and diversity of training data affect the diagnostic accuracy and generalizability of these systems.
Lastly, the rapid evolution of AI technology means that our findings may quickly become outdated. The update from ChatGPT-4 to ChatGPT-4o, from Google Bard to Google Gemini, 13 and from LLaMA2 to LLaMA3 exemplifies the fast-paced advancements in this field. Ongoing research is necessary to keep up with these developments and to continually reassess the performance and safety of new generative AI iterations.
Comparison with prior work
Diagnostic performance of the DDx lists for ChatGPT-4 and Google Gemini is summarized in Table 4. Relative to our previous study
21
on the GIM case series, the differences in diagnostic accuracy by ChatGPT-4 were no more than 6% within the top 10 DDx list (340/392, 86.7% vs 43/52, 83%, respectively), within the top 5 DDx list (307/392, 78.3% vs 42/52, 81%, respectively), and as top diagnosis (214/392, 54.6% vs 31/52, 60%, respectively). In contrast, compared to another study
17
of clinicopathologic conference from
Diagnostic performance of differential-diagnosis lists for ChatGPT-4 and Google Gemini.

ChatGPT-4: fourth-generation chat generative pre-trained transformer; DDX: differential-diagnosis; GIM: general internal medicine.
JMIR Med Inform. 2023 Oct 9:11:e48808.
JAMA. 2023;330(1):78–80.
Regarding Google Gemini, 23 the present finding revealed higher diagnostic accuracy within the top 10 DDx list (269/392, 68.6% vs 22/52, 42.3%, respectively), within the top 5 DDx list (235/392, 59.9% vs 21/52, 40.4%, respectively), and as top diagnosis (123/392, 31.4% vs 15/52, 28.8%, respectively). This enhancement can be partly attributed to the continuous updates to the model. The timeframe for accessing the AI in the current study spanned from July to August, while the previous study was drawn in June.
In the context of radiological differential diagnoses, a study 37 found acceptance rates of 85% and 69% for the top five DDx generated by ChatGPT-3.5 and Google Bard (currently Gemini), respectively. Our findings for Google Gemini (60%) show a similar level. Additionally, our finding for the next-generation model, ChatGPT-4 (78%), indicates that while improvements are not significant, variations in case types and evaluation methods likely influence these results.
Compared to the symptom checkers, 6 ChatGPT-4 showed higher rates of final diagnoses within the top 10 DDx lists (340/392, 86.7% vs 60.9–76.9%, respectively), while Google Gemini performed at a similar level (269/392, 68.6% vs 60.9–76.9%, respectively) and LLaMA2 chatbot lower (214/392, 54.6% vs 60.9–76.9%, respectively). Regarding the DDx generators, 38 ChatGPT-4 demonstrated higher rates (340/392, 86.7% vs 63–77%, respectively), while Google Gemini performed at the same level (269/392, 68.6% vs 63–77%, respectively) and LLaMA2 chatbot lower (214/392, 54.6% vs 63–77%, respectively). These findings underscore ChatGPT-4's potential as a reliable adjunct in medical diagnostics, while Google Gemini maintains comparable performance, and LLaMA2 chatbot lags behind.
Future directions
Based on this study, future research should explore real-world cases in various settings—such as different medical specializations, varying levels of healthcare, or diverse geographical locations—after securing the necessary regulatory approval for medical use. This approval, which includes clearance or authorization from health authorities, ensures that the generative AI systems adhere to patient safety and efficacy standards.
Additionally, research should investigate the fine tuning of AI models for medical applications and explore ensemble methods that integrate multiple generative AIs. Specifically, future studies should focus on establishing optimal prompt settings for these AIs, including parameters such as temperature, Top P, max sequence length, and prompt text. Understanding the impact of these settings on diagnostic accuracy is crucial for optimizing AI performance in clinical practice.
As the integration of technology and healthcare deepens, there is also a pressing need to evaluate the efficacy of collaborative efforts between healthcare professionals and generative AIs in clinical settings. Future research should investigate the potential benefits and challenges of human–AI collaboration in the diagnostic process, aiming to enhance the overall quality of care.
Moreover, fostering effective collaboration between generative AIs, healthcare professionals including GIM physicians, and patients warrants attention for a team-based approach going forward. 39 This includes exploring how these systems can be integrated into daily clinical practice, ensuring seamless interaction between AI tools and medical professionals. Additionally, regulatory considerations must be underlined to ensure that AI integration in clinical practice adheres to safety, ethical, and legal standards, thereby protecting patient welfare and maintaining trust in AI-assisted healthcare.
Conclusions
In summary, this study highlighted the diagnostic performance of generative AIs, including ChatGPT-4, Google Gemini, and LLaMA2 chatbot. ChatGPT-4 exhibited higher diagnostic accuracy than the other platforms across comprehensive medical fields, like GIM. These findings emphasize the importance of understanding the differences in diagnostic performance among generative AIs, especially in complex case series across comprehensive medical fields. Additionally, generative AIs, especially those exhibiting superior accuracy like ChatGPT-4, possess the potential to be pivotal diagnostic support tools in the near future. However, the variability in performance among the different AIs tested indicates the necessity for continued refinement and validation of these tools, especially in real-world clinical contexts.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076241265215 - Supplemental material for Diagnostic performance of generative artificial intelligences for a series of complex case reports
Supplemental material, sj-docx-1-dhj-10.1177_20552076241265215 for Diagnostic performance of generative artificial intelligences for a series of complex case reports by Takanobu Hirosawa, Yukinori Harada, Kazuya Mizuta, Tetsu Sakamoto, Kazuki Tokumasu and Taro Shimizu in DIGITAL HEALTH
Supplemental Material
sj-docx-2-dhj-10.1177_20552076241265215 - Supplemental material for Diagnostic performance of generative artificial intelligences for a series of complex case reports
Supplemental material, sj-docx-2-dhj-10.1177_20552076241265215 for Diagnostic performance of generative artificial intelligences for a series of complex case reports by Takanobu Hirosawa, Yukinori Harada, Kazuya Mizuta, Tetsu Sakamoto, Kazuki Tokumasu and Taro Shimizu in DIGITAL HEALTH
Supplemental Material
sj-docx-3-dhj-10.1177_20552076241265215 - Supplemental material for Diagnostic performance of generative artificial intelligences for a series of complex case reports
Supplemental material, sj-docx-3-dhj-10.1177_20552076241265215 for Diagnostic performance of generative artificial intelligences for a series of complex case reports by Takanobu Hirosawa, Yukinori Harada, Kazuya Mizuta, Tetsu Sakamoto, Kazuki Tokumasu and Taro Shimizu in DIGITAL HEALTH
Supplemental Material
sj-doc-4-dhj-10.1177_20552076241265215 - Supplemental material for Diagnostic performance of generative artificial intelligences for a series of complex case reports
Supplemental material, sj-doc-4-dhj-10.1177_20552076241265215 for Diagnostic performance of generative artificial intelligences for a series of complex case reports by Takanobu Hirosawa, Yukinori Harada, Kazuya Mizuta, Tetsu Sakamoto, Kazuki Tokumasu and Taro Shimizu in DIGITAL HEALTH
Footnotes
Acknowledgments
This study was made possible using the resources from the Department of Diagnostic and Generalist Medicine, Dokkyo Medical University.
Contributorship
TH and YH researched literature and conceived the study. TH was involved in protocol development, gaining ethical approval, participant recruitment, and data analysis. TH wrote the first draft of the manuscript. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
Not applicable.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by JSPS KAKENHI [Grant Number 22K10421].
Guarantor
TH
Informed consent
Not applicable for studies not involving humans.
Trial Registration
Not applicable.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
