Abstract
In recent times, appropriate diagnosis of brain tumour is a crucial task in medical system. Therefore, identification of a potential brain tumour is challenging owing to the complex behaviour and structure of the human brain. To address this issue, a deep learning-driven framework consisting of four pre-trained models viz DenseNet169, VGG-19, Xception, and EfficientNetV2B2 is developed to classify potential brain tumours from medical resonance images. At first, the deep learning models are trained and fine-tuned on the training dataset, obtained validation scores of trained models are considered as model-wise weights. Then, trained models are subsequently evaluated on the test dataset to generate model-specific predictions. In the weight-aware decision module, the class-bucket of a probable output class is updated with the weights of deep models when their predictions match the class. Finally, the bucket with the highest aggregated value is selected as the final output class for the input image. A novel weight-aware decision mechanism is a key feature of this framework, which effectively deals tie situations in multi-class classification compared to conventional majority-based techniques. The developed framework has obtained promising results of 98.7%, 97.52%, and 94.94% accuracy on three different datasets. The entire framework is seamlessly integrated into an end-to-end web-application for user convenience. The source code, dataset and other particulars are publicly released at https://github.com/SaiSanthosh1508/Brain-Tumour-Image-classification-app [Rishik Sai Santhosh, “Brain Tumour Image Classification Application,” https://github.com/SaiSanthosh1508/Brain-Tumour-Image-classification-app] for academic, research and other non-commercial usage.
Introduction
Brain Tumour is the abnormal growth of cells in the brain. The lesions are identified through advanced medical equipment such as magnetic resonance imaging (MRI) and computed tomography (CT). MRI uses magnets and radio waves to produce the images on computer, 1 whereas CT scan uses X-rays to create cross-sectional images of the body. 2 These lesions are benign in most cases and can be treated with medications. Glioma and meningioma account for 95% of all brain tumours. Glioblastoma, a subtype of glioma, accounts for 45.2% of all malignant tumours. Glioblastomas are malignant brain tumours classified as a grade IV tumour by World Health Organisation (WHO). 3 The 5-year survival rate of glioblastoma is less than 5%, indicating the significant challenges encountered in the diagnosis of this aggressive cancer. Meningiomas are the second most common brain tumour in cancer diagnosis systems. It accounts for approximately 53.8% of non-malignant tumours, 4 Pituitary tumours account for 10%‒17% of primary tumours. They have a very high five-year survival rate, indicating that pituitary tumours are easily diagnosed.
Several researchers have been working on solutions for classifying brain tumour's with higher precision. Many researchers have proposed various solutions for image classification. Traditional machine learning techniques, such as support vector machine (SVM), decision tree, and Multi-Layer Perceptron, have been utilized for classification.
However, traditional approaches often fail to produce accurate results in medical diagnoses, which can lead to fatal outcomes. Therefore, accurate identification of tumor cell from medical images indeed crucial and non-trivial task. Hence, after the advent of deep learning models, performance of state-of-the-art (SOTA) methods improved remarkably. Convolutional Neural Networks (CNNs) is one of the extensively applied deep model in medical image segmentation and classification due to their automatic and intrinsic feature extraction ability from complex image patterns. The limitations of traditional models, such as SVM, compared to CNNs in image classification tasks are clearly outlined in 5 .The paper compares brain tumor MRI classification using linear SVM, polynomial SVM, and CNN, concluding that CNNs are significantly more effective for image classification Over the past few decades, numerous state-of-the-art (SOTA) models, such as AlexNet, 6 GoogleNet, 7 and NASNet, 8 have been developed, significantly advancing image classification tasks. The concepts of transfer learning and fine-tuning have enabled researchers to leverage these SOTA models, adapting them to specific requirements and improving their performance. Notably, several of these models have demonstrated remarkable results on benchmark datasets, including ImageNet and CIFAR-100, which are widely utilized to evaluate image classification methodologies. More recently, Vision Transformers (ViTs) have emerged as a cutting-edge architecture, employing the attention mechanism originally introduced for Natural Language Processing (NLP) tasks. ViTs decompose images into sequences of patches, treating them as input tokens for the transformer architecture. While these advancements have propelled image classification techniques, applying them to medical imaging poses unique challenges. Medical imaging data, such as grayscale MRI scans, differ substantially from the high-resolution RGB datasets for which these models were initially designed. Consequently, their performance in medical applications is often suboptimal without domain-specific adaptations. Additionally, medical datasets are characterized by limited sample sizes, higher complexity, and domain-specific nuances, further complicating model development and generalization.
Advances in knowledge transfer techniques such as transfer learning and fine-tuning have facilitated the application of SOTA architectures, such as ResNet-50, Inception-v3, and EfficientNet, to complex medical imaging tasks. For brain tumor classification, these fine-tuned models have demonstrated significant improvements in accuracy and generalizability compared to traditional machine learning approaches. Ensemble learning strategies, which aggregate predictions from multiple models, have further enhanced the robustness and reliability of automated systems in clinical settings. Beyond CNN-based methods, hybrid architectures and advanced techniques, such as Generative Adversarial Networks (GANs), have been investigated to address the limitations of small medical datasets by generating synthetic data for augmentation. Although transformer-based models are gradually gaining traction in medical imaging tasks, their reliance on large datasets and substantial computational resources poses challenges for implementation in resource-constrained environments.
To address these challenges, this study proposes a weight-aware decision framework, which combines predictions from multiple models and assigns higher weights to those with superior performance. This approach aims to enhance classification accuracy and reliability in clinical applications. The key contributions of this study are summarized as follows:
Weight computation of employed deep models: The framework utilizes VGG19, DenseNet169, Xception, and EfficientNetV2B2 to obtain model-wise validation scores. These scores are normalized and considered as model-specific weights, which significantly enhance the final classification performance in multi-class problems. Weighted-aware decision Strategy: developed framework implements weight-aware decision module into the framework to generate more accurate and conclusive results as compared to traditional majority-based voting technique in multi-class problem. Stream lit-Based Web Application: The developed framework has been seamlessly incorporated into a user-friendly web application called ‘NeuroVision’ built using the Streamlit library. This application aids medical professionals to efficiently classify and analyse tumour cells in the cancer affected areas of input image. High Performance in Multi-Class Classification: Our framework achieves superior accuracy in predicting Grade 4 glioma tumours, which significantly outperforms the current state-of-the-art results. Utilization of Callbacks for Optimized Training: During the training process, the deep model utilizes LRScheduler and checkpoint methods for optimum hyperparameter tuning. The LRScheduler dynamically adjusts the learning rate to enhance model accuracy, whereas the checkpoint callback ensures that the best-performing weights are saved throughout the training.
The rest of the paper is organized as follows: Related work section provides a comprehensive review of the related works, highlighting the shortcomings of existing research. Methodology section discusses the overall framework of the proposed work, employed deep models architectures and the components of the framework. Experiment results and analysis section details the results obtained from the experiments along with insightful analysis. The section also analyses the performance of the proposed framework on external data, the ablation study details the impact of each model in the framework by examining the overall inference time and the accuracy.
Conclusion and future directions section presents the conclusive remarks and discusses the future scope of the research.
Related work
In recent years, significant advancements have been made in the field of brain Tumour classification. This section reviews and highlights the methodologies employed and the results by examining previous studies to establish a foundation for our proposed approach. Gajula et al. 9 proposed an approach Logistic regression with threshold segmentation that achieves superior performance in medical imaging classification. Sandhiya et al. 10 have utilized Inception V3 and DenseNet201 to extract the basic features and integrate them with (PSO-KELM) algorithm to produce a state-of-the accuracy. In the study by Dheepak et al., 11 the researchers proposed a SVM classifier ensemble that employs feature extraction techniques like grey level co-occurrence matrix and local binary patterns combined with four different SVM along with a different kernel function. Khan et al. 12 have developed a voting classifier ensemble by utilising DenseNet169 as a feature extractor, and the extracted features were fed into three machine learning classifiers (RandomForest, SVM, and XGBoost) to achieve high accuracy on par with state-of-the-art models. Sarada et al. 13 have proposed a modified version of ResNet50V2, 14 the model utilised ResNet50v2 as the backbone and incorporates BatchNormalization, Dropout and MaxPooling layers to reduce the number of parameters and over-fitting, achieving high accuracy with fewer parameters. Rasheed et al. 15 have designed a framework utilizing image enhancement techniques such as Gaussian-blur-based sharpening and Adaptive Histogram Equalization combined with the proposed architecture and achieved remarkable accuracy with just 1 million parameters – significantly fewer compared to the state-of-the-art models.
Asiri et al. 16 have proposed a Generative Adversarial Networks(GAN) based architecture, this approach utilised a generator and discriminator, the discriminator was trained on the real-time MRI image dataset, and the generator takes in a random vector of fixed dimension and generates images similar to the images in the dataset. This helps the discriminator to classify images with better performance. Khan et al. 17 have proposed a multi-scale deep neural network for script identification. The research proposed a three-module framework – (a) multi-scale-based CNN prediction, (b) scale-wise weight computation and (c) weight-aware decision mechanism for final classification, the framework produced excellent results surpassing the existing state-of-the art with fewer trainable parameters. Arora et al. 18 have provided a comparative analysis of CNN based pre-trained deep learning models on image classification tasks, the study concluded that VGG16 performs way better than the other utilised models, especially when utilised with transfer learning. Shaik et al. 19 have developed multilevel attention network (MANet) incorporates spatial and cross-channel attention obtained from the Xception backbone. Sevli 20 utilised transfer learning approach on VGG-16,ResNet50, 21 Inception V3 22 and performed an extensive comparison of these pre-trained models on brain tumour image classification. Srinivas et al. 23 have presented in their study provided a comparative analysis of performance of pre-trained models on Brain Tumour classification of MRI images and concluded highlighting that VGG-16 was the most effective.
Paul 24 et al. have proposed an ensemble of SCNN and EfficientNetB1,the results demonstrated that the performance of the weighted ensemble was higher than the state-of-the-art models with significantly fewer parameters. Anantharajan et al. 25 have developed an Ensemble Deep Neural Support Vector Machine (EDN-SVM),the approach also utilises image processing techniques – Adaptive Contrast Enhancement Algorithm (ACEA) and median filter. The pre-processed images were segmented using the Fuzzy c-means based segmentation. Gray-level co-occurrence matrix (GLCM) was used to extract the features. The combined pre-processing, feature extraction techniques, and the proposed model achieved remarkable accuracy, outperforming certain state-of-the-art models.
Khan et al., 26 have performed a comprehensive study on various hand-crafted feature descriptors and deep feature descriptors by performing extensive experiments on the AUTNT dataset to understand the best performing method in image classification tasks. Ramtekkar et al. 27 have developed an architecture for accurately detecting brain tumours using optimized feature selection-based technique involving a four phases, the first phase involves pre-processing steps- grayscale conversion and filtering, the second phase involves segmentation of the tumour region with the threshold and histogram method, the third phase involving utilisation of Gray-level co-occurrence matrix (GLCM) feature extraction technique to extract feature from grayscales by analysing spatial relationship between neighbouring pixels. The fourth phase involves utilising an optimization algorithm, the study evaluates the various existing optimization algorithms for brain tumour detection tasks. Throughout the study, the various available optimization algorithms were experimented on brain tumour detection and concluded that utilization of threshold and histogram for segmentation, GLCM for feature extraction paired with the whale and grey wolf optimization algorithm achieved the state-of-the-art accuracy. Khan et al. 28 have exploited the text-stroke information for visual object detection from camera captured natural scene images. Ramakrishnan et al. 29 proposed a hybrid CNN architecture composed of InceptionV3, ResNet-50, VGG-16 and DenseNet, the work also optimized the architecture through the oneAPI to compare the model performance and achieve significant accuracy. Mugdha et al. 30 compared four state-of-the-art pre-trained deep learning models – VGG16, ResNet-50, AlexNet and Inception-V3 on the compilation of three distinct datasets, the proposed study utilised the transfer learning approach to train the deep models, after extensive data pre-processing and augmentation the models were trained on the augmented datasets and on evaluation it was reported that VGG-16 outperformed the other models and achieve the state-of-the-art accuracy. The short-comings of the existing state-of-the-art models are reported in Table 1.
A brief overview of state-of-the-art methods for brain tumour detection, along with their key features and limitations.

Based on the shortcomings reported in Table 1, several limitations in the existing research methodologies and frameworks have been identified. These shortcomings are summarized as follows:
Evaluation on small-sized datasets: It is observed that, some existing methods have been evaluated on datasets of few number of samples. Such evaluations often fail to reflect the model's generalizability for large and complex datasets that resemble real-world scenarios. Imbalanced datasets: Brain tumor datasets used in prior research are frequently imbalanced, with certain classes containing significantly more samples than others. This imbalance can bias the model, resulting in suboptimal performance for underrepresented classes. Limited evaluation of model performance: Most studies evaluate model performance solely on one dataset. This approach may lead to misleading results due to factors such as data leakage and insufficient robustness testing.
In our proposed framework, we address these shortcomings through the following approaches:
Data Augmentation: To increase the sample size and introduce complex features, basic data augmentation techniques such as image rotation and flipping were applied. Class Imbalance Mitigation: To counteract the effects of imbalanced datasets, class weights were assigned based on the proportion of samples in each class relative to the total dataset. This ensures that classes with fewer samples are assigned higher weights. The class weight calculation formula is provided below: Extensive Model Evaluation: To assess the real-world performance of the proposed framework, evaluations were conducted using three distinct datasets from multiple sources. This comprehensive evaluation approach ensures a robust analysis of the framework's effectiveness.
Methodology
The framework employs an open-source python framework to deploy the proposed model on the web, facilitating access to medical professionals. The web application enables users to upload the images. Upon successful image uploading, the application performs essential pre-processing steps to transform the image into a format compatible with the model. Once the image is pre-processed, it is provided as an input to each fine-tuned model. Each model generates predictions based on the input image. The inputs are subsequently channelled into the weighted prediction algorithm, in this algorithm each model's predictions are combined with its respective weights, effectively contributing to the final decision, returning the highest aggregated score as final output class. For example, if model 1 predicts the image as class 1, the weight associated with model 1 is added to the cumulative score of class 1. The weighted aggregation process is continued for all models, and the class with the label with the highest accumulated score is returned as the output. The complete illustration of workflow is provided in Figure 1.

The working pipeline of the NeuroVision framework. At first, images of training dataset pass through various deep learning models, which yield validation accuracies. The obtained accuracies are then input to a weight computation module to determine model-specific weights. For each model, if the predicted class matches one of the possible output classes, the weight of that model is iteratively aggregated into the corresponding class bucket. Finally, the class with the highest aggregated weight score is selected as the final output.
Data pre-processing and distribution
The models incorporated in the framework are trained on the data obtained from ‘Brain Tumour MRI Dataset’ available on Kaggle. 36 The dataset is a combination of Figshare, Br35H, and SARTAJ datasets. The dataset consisted of 7023 gray-scale images, which consisted of 5712 images across four classes: glioma (1321), meningioma (1339), no tumor (1595), and pituitary (1457). The testing dataset consisted of 1311 images equally distributed across the four classes.
The dataset splits are pre-processed into images of 128 × 128,converted to RGB colour format to make images compatible with the models utilized. Rescaling is used to normalize the pixel values for better computation. The steps followed in the image pre-processing are depicted in Figure 2. Data augmentation is employed to enhance the model performance and enabling the model to learn more features from the images. The dataset creation utilized the following basic data augmentation techniques: flipping, rotation, and a combination of both. This contributed to an increase in the dataset size, with a total image count of 22,848 images and the class distribution of each dataset in illustrated in Figure 3. The extended version of the experimental dataset after data augmentation is publicly released in our GitHub repository. 37

Image pre-processing workflow. This figure illustrates the image pre-processing techniques applied to enhance performance and computational efficiency. The images are resized to 128 × 128 pixels, converted to the RGB colour format, and normalized to scale the pixel values, facilitating faster processing during model training.

Class distribution over dataset splits. This figure provides a comprehensive view of the class distribution within the dataset splits, highlighting the significance of class balance in optimizing model performance and ensuring effective training.
Employed deep learning models
The deep learning models used in this research are chosen due to their demonstrated effectiveness in handling complex image classification challenges, with each model presenting distinct architectural advantages suited for analysing medical images. These models underwent specific fine-tuning and optimization for classifying brain tumours, utilizing their ability to recognize complex patterns within MRI scans. By integrating a variety of architectures—from densely interconnected layers to streamlined convolutional blocks—this research seeks to assess the relative performance of these models when trained on the identical dataset, offering insights into their respective strengths and weaknesses. The following sections describe the architecture of each model, the rationale behind their design, and their relevance to the task, in addition to an evaluation of their performance within our framework.
DenseNet-169
The DenseNet inherits the baseline CNN architecture that connects each layer to every other layer in a feed-forward fashion. 38 The architecture provided in Figure 4 comprises of multiple combinations of dense block and transition layers where each network layer is connected to ensure maximum information flow between layers. This architecture has also significantly reduced the vanishing gradient issue, and the number of parameters compared to other state-of-the-art models.

Layer-wise architecture of denseNet-169. 38
VGG -19
VGG-19 is the state-of-the-art model achieving high accuracies with just 16–19 weighted layers. The model utilizes the convolutional layers with filters with a small receptive field: 3 × 3 (the smallest size to capture the notion of left/right, up/down, and centre). 39 The combination of lower depth of the model and small hyperparameters like kernel size, and padding allowed the model to identify more feature patterns with fewer parameters. Figure 5 illustrates a detailed overview of the architecture of VGG-19.

VGG-19 architecture. 39
EfficientNetV2
EfficientetV2 is a convolutional network architecture that has significantly reduced the number of parameters and improved training speed while achieving the same state-of-the-art model results. The architecture in Figure 6 uses a smaller kernel size and additional layers to compensate for the reduced receptive field. 40 This model incorporates MBConv 41 and FusedMb-Conv layers to ensure that accuracy is not compromised for computation.

Layer-wise architecture of EfficientNetV2. 40
Xception model
The Xception stands for Extreme Inception, introduced by François Chollet. The architecture consists of 36 convolutional layers structured into 14 modules, all of which have linear residual connections around them, except for the first and last modules. 42 The model uses a depth-wise-separable convolutional layer to reduce the number of parameters and computation without compromising efficiency and accuracy. Figure 7 provides a complete flow of the input image in the Xception architecture.

Xception Architecture. 42
Training phase of deep models
Our study employed fine-tuning techniques by unfreezing a specific number of layers in each model, determined by their ability to adapt to the dataset. Fine-tuning was chosen as part of our proposed framework due to the inefficiencies of directly applying transfer learning. While pre-trained models were initially trained on large datasets such as ImageNet and CIFAR-10, which consist of high-resolution RGB images, our MRI brain tumor dataset features noisy, low-resolution grayscale images. These substantial differences in data characteristics and feature complexities render the pre-trained weights less effective for this specific task.
For our experiments, we utilized Google Colab equipped with T4-GPU hardware to enable faster and more efficient model training. The frameworks adopted for the study include TensorFlow, Keras API, scikit-learn, and python. The augmented dataset was downloaded and loaded into data loaders to transform the data into a compatible format. Data loaders batched the dataset with a batch size of 32 and an image size of 128 × 128 pixels to optimize computation on the GPU. Subsequently, an extensive exploratory data analysis was conducted to understand the class distribution and image-specific characteristics. A class imbalance was identified in the dataset, and to counteract its effects, class weights were assigned based on the proportion of samples in each class relative to the total dataset. This approach ensures that classes with fewer samples are assigned higher weights. The class weights are calculated using the mathematical formula presented below in equation (1).
After data exploration, pre-trained models were loaded from the TensorFlow Applications module. To tailor each model for our use case, the following layers were added:
An input layer A pre-processing layer to transform the input to a compatible format for the specific pre-trained model A pooling layer A dropout layer with a dropout rate of 0.4 A fully connected layer with four neurons, each representing a class in the dataset.
The top layers of each model are progressively unfrozen until the model achieved the desired performance. The number of layers unfreezed vary from one model to another due to architectural differences and can only be determined by experimentation .The learning rate, a critical hyperparameter, was carefully optimized. To decide the initial learning rate of the model training, three initial learning rates (0.01, 0.001, 0.0001) has been tested on, utilising the article 43 as the reference. The value of 0.0001 was identified as the optimal initial learning rate, as it minimized fluctuations in the training curves, reducing overfitting risks. The impact different learning rates on the fine-tuning process are visually represented in Figure 8.

Effect of learning rate in fine-tuning. The figures provide insights into how the learning rate is affecting the loss curves of the model fine-tuning. For convenience the start of the learning rate at any point is specified with a uniquely identifiable markers.
To address the challenges posed by a static learning rate we have employed the ‘ReduceLROnPlateau’ callback from the Keras API, which adjusts the learning rate during training based on the provided parameters. The callback requires parameters such as an initial learning rate (0.0001), patience (3), a monitoring metric (validation loss), and a factor of 0.1. When the monitoring metric showed no improvement over the specified patience period, the callback adjusts learning rate using the mathematical notation mentioned in equation (2).
44

The hyperparameters and associated values of different deep models during training are reported in Table 2.
Initial hyper-parameters and associate values of the deep models during of training.
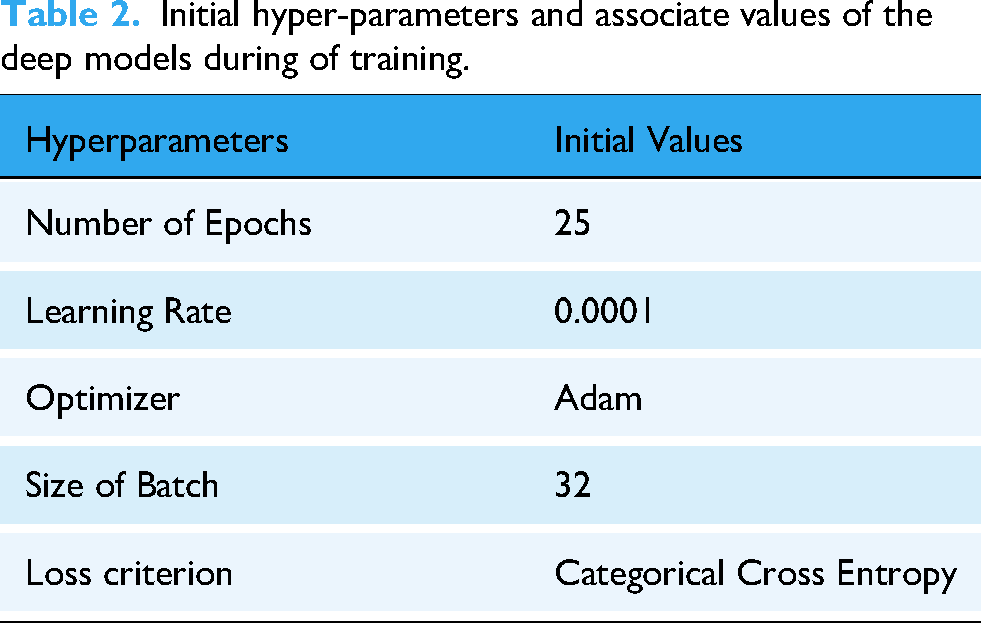
To ensure optimal performance, we utilized the ‘ModelCheckpoint’ callback, which monitors a specified metric (validation accuracy) and saves the model weights that achieve the highest validation accuracy during training. After fine-tuning, the models were incorporated into the framework. Each model's output predictions were processed to determine the predicted class. For weight-aware decision-making, each class was represented by a variable to store cumulative weights. When a model predicted a sample as belonging to a specific class, the model's weight was added to the respective class's variable. After processing predictions from all models, the class with the highest aggregated weight was selected as the final output. The weight computation mechanism and the weight-aware decision process are discussed in detail in Weight computation of employed model section. The next step involves developing a weighted-decision architecture by assigning calculated weights to each model. To enhance computational efficiency, an input layer is added to the ensemble, ensuring that the input image is simultaneously sent to all models for parallel predictions. These predictions are then processed by a weight-aware decision module, which generates the final output by accounting for the weighted contributions of each model's prediction.
Weight computation of employed model
In this study, a novel weight-aware decision approach is employed for the final classification. Unique weights are assigned to each deep model, with the weight of each model being proportional to its average validation accuracy during the training phase. Specifically, the weight 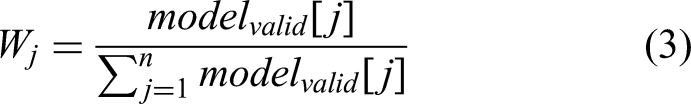
In this formula, the numerator
This approach is influenced by the class weight formula commonly applied to tackle class imbalances. In such cases, greater weights are allocated to underrepresented classes, preventing the model from favouring classes with a larger sample size. Similarly, the weight-aware decision method ensures a balanced contribution from models by assigning weights proportional to their validation performance, thereby improving overall prediction accuracy.
Weight-aware decision approach
In this module, a novel weight-aware decision mechanism is applied for final of image objects. After model-wise predictions, model-weights are iteratively added into the buckets of respective predicted classes. The weighted-aware decision approach is inspired by Khan et al.,
17
the mechanism involves aggregating the weighted predictions of each of the model and returning the output class with highest aggregate score. The work-flow of the weight-aware-decision approach is illustrated in Figure 9. Moreover, the detailed working principle of this module is outlined below:
Let, The unique weights of the employed deep models are denoted as Each possible output class for an input image maintains its own bucket. If the model generated predicted class matches the possible output class, the unique weight of that specific model is added to the corresponding bucket. This process continues for all other deep models. Finally, the bucket of the possible class with the highest aggregated scores is selected as the final output class for the input image.
The weight-aware decision approach is mathematically illustrate using equations (4) to (6).
Here,
The possible class correlation factor (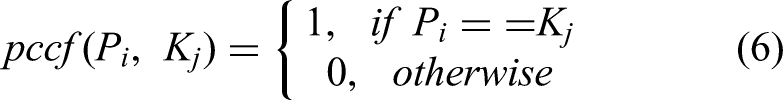
Here,
It may be stated that, the proposed weight-aware decision approach demonstrates improved accuracy and reliability compared to traditional majority voting for multi-class problem. Unlike majority voting, where bucket values are incremented by a fixed value of 1, this approach aggregates unique weights into the corresponding buckets. This effectively mitigates the issue of tie scenarios, a notable drawback of conventional majority-based methods. There is high possibility that, weights in this mechanism are a distinct real number. The final prediction is determined based on the total aggregated model weights, significantly reducing the likelihood of ties during classification. This ensures a more precise and effective decision-making process.
For example, consider a scenario where model-1, model-3, and model-4 predict the input as meningioma, while model-2 predicts it as glioma. In this case, the if prediction of the model matches the possible output class, then weights assigned to particular model is aggregated for the respective bucket of that possible classes. The class with the highest aggregated value is selected as the final output. In this instance, meningioma is returned as the final predicted class for the input image. Figure 9 clearly illustrates the working principle of the weight-aware decision approach for a general audience.
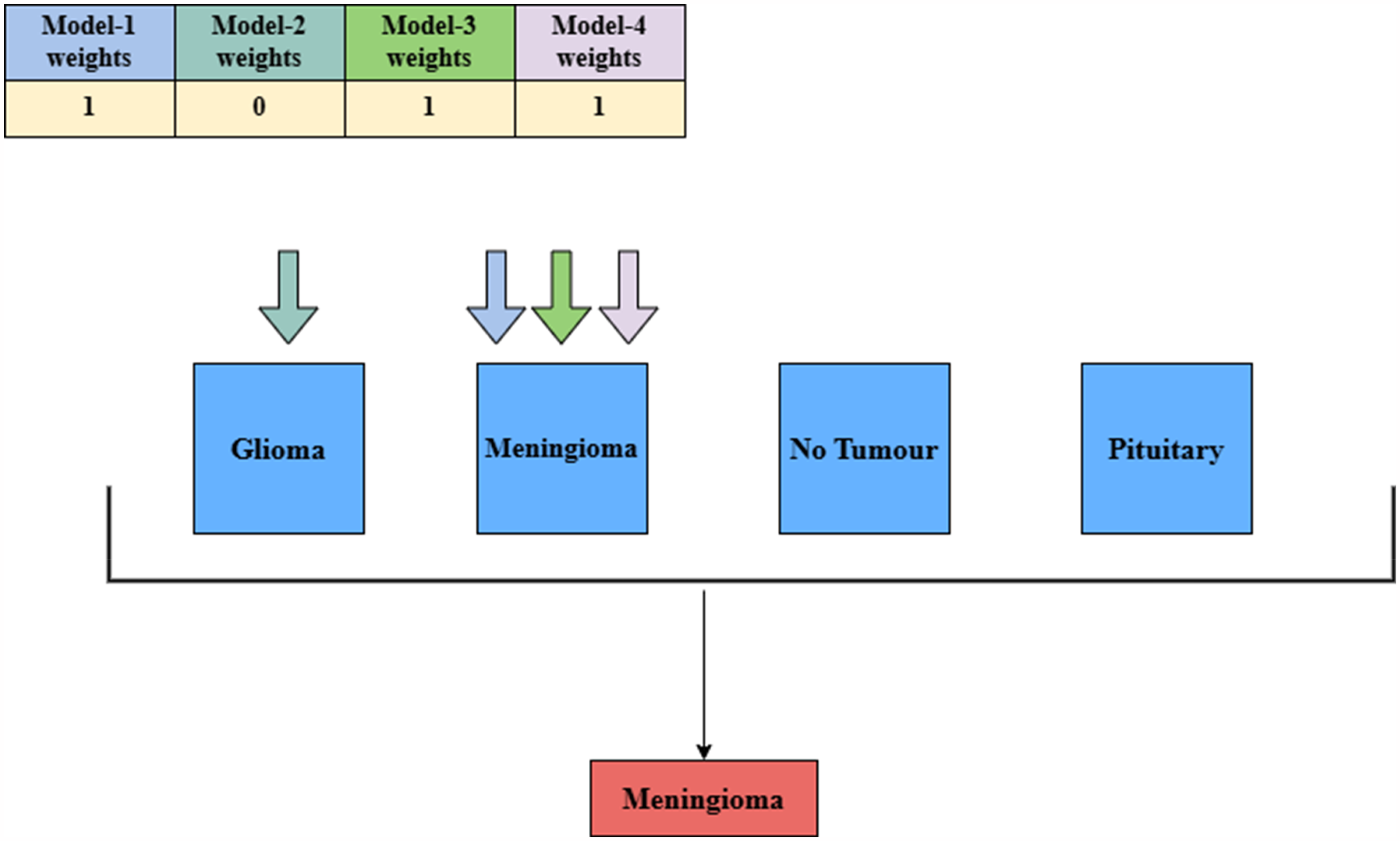
Working principle of weight-aware decision approach of the developed framework. This figure illustrates the weighted mechanism employed in the proposed framework. It provides a detailed explanation of the model-weighting process, using a scenario to demonstrate its application and functionality within the system.
Development of web application
The proposed framework is extremely dependable in detecting MRI brain scan images. To make it applicable in real-world situations, the model was deployed using Stream lit, an open-source python framework that enables users to develop and deploy web applications and visualizations. The ‘NeuroVision’ application also includes data visualizations and provides prediction probabilities and the final output class. The frameworks and libraries utilized for the development of the web application are reported in Table 3. Besides, the user interface of the designed web application is depicted in Figure 10.

User interface of the developed web application. (a) Interface before uploading the input image, (b) corresponding user interface after classification of brain tumour from input image.
Tools and library packages utilized to build and deploy the NeuroVision web application.

Salient features of NeuroVision
The salient features of the developed framework are mentioned as follows:
User-friendly interface: The application is designed with a responsive, intuitive interface that provides a smooth experience and makes navigation easier for users of all experience levels. Weighted-aware decision approach: At the core of the framework is a weighted decision mechanism that integrates predictions from multiple models by assigning weights based on the verification accuracy of each model. This approach allows models with higher performance to significantly contribute to the final prediction, increasing accuracy and robustness. Classification Class Information: An extensible section provides detailed information about each classification class, including tumor type, symptoms, and potential treatments. This feature educates users about health risks and encourages informed decisions. Prediction probability indicator: Above the prediction results, the probability extender displays the confidence of the model for each prediction, so that the user can effectively interpret the reliability of the output. Practical Utility for Medical Professionals: The application's integration of accurate predictive algorithms with informative displays makes it a valuable tool for medical professionals, supporting real-world clinical assessments and decision-making in healthcare settings.
Experiment results and analysis
This section provides a comprehensive performance assessment of the developed framework using different evaluation metrics. To evaluate the stability and convergence of the model learning capability, training accuracy and loss curves are shown and obtained findings are examined. The generated confusion matrices for different deep models are presented to illustrate their effectiveness in the classification task. Furthermore, the classification accuracy of various deep models, along with the final performance of the developed framework (after applying the weight-aware decision approach), is reported. Finally, a performance comparison between the developed framework and existing models is presented, followed by an insightful discussion based on the findings.
Empirical findings
To demonstrate optimal training, the training vs. validation accuracy and corresponding loss graphs for different deep models over 25 epochs are presented in Figure 11. It provides insightful observation into the model's convergence and learning stability. The models assessed include DenseNet-169, EfficientNetV2B2, VGG19, and Xception, each showcasing distinct learning patterns. DenseNet-169 and EfficientNetV2B2 demonstrate steady loss reduction, while VGG19 and Xception show fluctuating trends, reflecting variations in model architecture and optimization behaviour. This comparison allows for a clear evaluation of each model's effectiveness in minimizing error over time, highlighting their relative training efficiencies. The number of epochs required is the point at which the curve starts to plateau. Identifying this is necessary since training the model for an excessive number of epochs may cause the model to identify the noise in the data, leading to overfitting.

Performance assessment of deep models using training vs. validation accuracy and loss graphs. (a-b) Training vs. Validation accuracy and loss curves for DenseNet169, (c-d) corresponding graphs for EfficientNetV2B2, (e-f) graphs for VGG19, and (g-h) graphs for Xception, respectively.
The training and validation accuracy, along with the corresponding loss values, are illustrated in Figure 11 and are detailed in Tables 4 and 5. It has been observed that the developed framework performs with good efficiency and high accuracy. Combination of VGG19, Xception, and EfficientNetV2 were equivalent to the proposed framework. However, whereas accuracy gives an overview of how the model performs generally, precision, recall, and f1-score must be looked at in addition for a better evaluation, especially on the accuracy of differentiating the individual classes. High precision shows fewer false positives, while high recall ensures that most true cases are detected. Both are very important in the diagnosis of brain tumours. The performance comparisons of individual fine-tuned models and the proposed framework are reported in Table 6.
Training and validation accuracies of the model during fine-tuning.

Training and validation losses of the model during fine-tuning.

Obtained classification accuracies of all employed deep models and final accuracy of developed framework after weight-aware decision approach.

Figure 12 shows the confusion matrices for each deep learning model used. These matrices help to clarify the recall, accuracy and F1 scores and provide insight into the performance of each model in correctly identifying each class. These matrices can reveal a model's strengths in class identification and highlight areas of misclassification, leading to targeted hyperparameter tuning to improve performance in specific categories.

Confusion matrix of employed models and developed framework. (a-e) Confusion Matrix of Densenet169, EfficientNetV2, VGG19, Xception and finally the developed framework.
Figure 13 provides a comparative analysis of the proposed framework on 2 external datasets. The Kaggle_1 dataset represents the dataset utilised for model training and the accuracy indicates the model performance on the testing dataset. The figshare_1 dataset and the Kaggle_2 datasets are the external dataset used to evaluate the proposed framework efficiency and performance on real-time data. The proposed framework achieved an accuracy of 98.7 on the Kaggle_1 dataset, 97.52 on the Figshare_1 dataset and an accuracy of 94.94 on the Kaggle_2 dataset. These values clearly indicate that the proposed model is performing remarkably well and achieving high performance on real-time data.
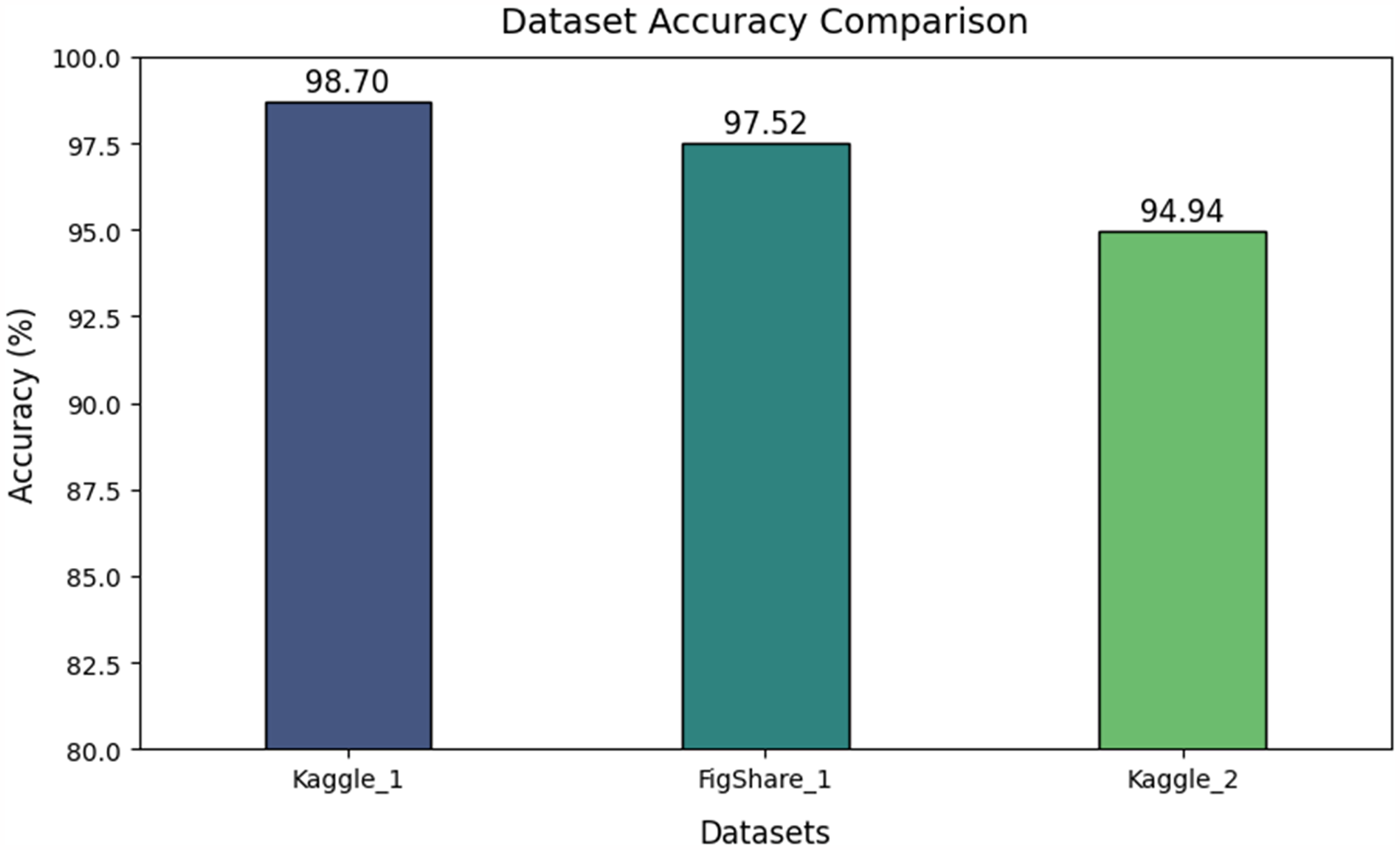
Comparative analysis of proposed framework on external datasets.
Low precision or recall in medical diagnosis can lead to misdiagnosis, thereby resulting in inappropriate or wrong treatments or delayed diagnosis. Hence, it becomes very essential to have a deep analysis of performance across various metrics. Thus, it is essential to comprehensively understand the strengths and limitations of a model, particularly in real-time medical applications, where performance deficiencies can lead to life-threatening scenarios. This analysis points out the necessity of using robust metrics other than accuracy for reliable and clinically relevant results in practical settings.
To conduct an in-depth analysis of the performance of various models, Table 7 presents the results in three stages: first, the performance of individual models; second, the performance of model combinations; and finally, the performance of the developed framework using a weight-centric decision approach with all employed models. These results can offer general users’ valuable insights to select the most suitable model combination based on their specific requirements.
Individual performance of different deep model, combinations of models and final framework using weight-aware decision approach.

Comparative analysis
The performance of the developed framework is compared with other existing methods, and the results are presented in Table 8, to highlight its effectiveness and superiority in the current domain. The analysis reveals that our framework achieves a classification accuracy of 98.7% by employing weight-aware ensemble of four advanced deep models. This finding underscores the strength of the weight-aware decision approach, particularly for multi-class classification problems.
Performance comparison of developed framework with existing state-of-the art models on Kaggle brain tumour dataset.

Ablation study
Ablation study is a research approach to machine learning and deep learning which helps in understanding the contributions of each components or feature within an architecture. 51 This section demonstrates the impact of various data augmentation techniques utilised and its impact on the proposed framework performance. We also demonstrated the trade-off between computational time for each model combination and its performance.
Impact of data augmentation
Data augmentation is a technique utilised to improve the dataset samples and introduce complex features into the dataset enhancing the number of features and the model's ability to grasp the features and helps in achieving better performance. 52 Our approach presents the impact of the data augmentation techniques used, horizontal flipping, vertical flipping, and rotation. The performance of our proposed framework for each augmentation combination is presented in Table 9.
Impact of data augmentation on model performance.

Computational time
Computational time refers to the duration required to generate an output from a given input. This factor plays a critical role in machine learning tasks, particularly in real-time applications. A comparative assessment on computational time per sample across different combinations of employed models and the final developed framework is reported in Table 10. It is also essential to acknowledge the inherent trade-off between computational time and performance.
Execution time of a sample for different architectural combinations.

The results of this research underscore both the strengths and weaknesses of the proposed model framework for brain tumor classification. Implementing a weighted-decision approach led to a significant boost in classification accuracy for various tumor types, particularly when tested on external datasets. This methodology illustrated how model weighting can improve decision consistency by utilizing the strengths of individual models based on their validation accuracy. However, there are several limitations that need to be considered. For example, the variation found in external datasets may influence performance due to inconsistencies in imaging quality, pre-processing methods, or class distribution, indicating the necessity for further assessment on a wider range of datasets. Moreover, the suggested model could enhance its performance by incorporating more advanced ensemble techniques or attention mechanisms that can selectively focus on important features within images.
Conclusion and future directions
The objective of the research was to create a strong deep learning model for classifying brain tumours. We adjusted each model to suit our data by fine-tuning them, assigning weights to each based on their validation accuracies. These model weights are incorporated into the weight-decision process to predict the final output, resulting in a remarkable accuracy of 98.7%. The model's effectiveness has been tested on external datasets to show its performance with real-time data. This study emphasizes the importance of adjusting and assigning model weights based on performance, enabling better-performing models to have greater influence in predictions. Despite the positive outcomes, there is considerable room for additional enhancements. In the future, there could be an emphasis on utilizing image segmentation methods as well as classification techniques to detect tumour regions and analyse a wide range of datasets for categorizing different types of main tumours.
Several avenues for future work can be explored to enhance the utility and accuracy of the proposed framework, as discussed below:
Advanced Image Segmentation: Incorporating advanced image segmentation techniques could precisely identify and isolate the tumour region, potentially improving classification outcomes and aiding in detailed diagnostic processes. Dataset Expansion: Expanding the dataset to include a wider variety of tumour types and imaging conditions would enhance the model's generalization capability. Real-time Optimization: Optimizing the model for faster inference while maintaining improved accuracy could make it suitable for clinical settings and real-time applications. Hybrid Models: Future research could explore hybrid models combining classification and detection approaches to simultaneously identify tumour types and localize the tumour area. Multimodal Data Integration: Incorporating multimodal data, such as patient medical history and genetic information, along with MRI images, could improve prediction accuracy and offer personalized diagnostic insights. Robust Evaluation Frameworks: Developing robust evaluation frameworks to ensure model performance across diverse populations would improve the framework's reliability and scalability for widespread adoption in medical applications.
Footnotes
Acknowledgements
The first four authors would like to express their gratitude to VIT-AP University, Amaravati, Andhra Pradesh, while the last author is thankful to King Abdulaziz University, Jeddah, Saudi Arabia, for providing the necessary support to carry out this work.
ORCID iDs
Ethical considerations
Our institution does not require ethical approval for reporting individual cases or case series.
Author contributions/CrediT
TRSS contributed to data curation, methodology, validation, visualization, writing original draft; SNM contributed to conceptualization, reviewing the draft and editing; NRP contributed to data visualization, reviewing the original draft and editing; TK contributed to conceptualization, formal analysis, data visualization, work administration, reviewing the original draft. MD contributed to writing review and editing.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
