Abstract
Objectives:
Long noncoding RNAs plays the important part in tumor biology. SNHG19 was found to be a new oncogenic lncRNA in some malignant tumors. However, the effect of SNHG19 in hepatocellular carcinoma has not been reported.
Methods:
The expression of SNHG19 in hepatocellular carcinoma tissues were detected by using quantitative Real-Time Reverse Transcription Polymerase Chain Reaction (qRT-PCR). Melanoma cases from The Cancer Genome Atlas were included in this study. cell counting kit-8 assay, Transwell, and scratch wound assay were used to explore the role of SNHG19 in melanoma cells. Luciferase reporter assays and RNA pull-down assay were used to explore the molecular mechanism of SNHG19 in hepatocellular carcinoma.
Results:
Here, we found that SNHG19 level was upregulated in hepatocellular carcinoma. Hepatocellular carcinoma patients with high levels of SNHG19 have shorter Disease-Free Survival (RFS). SNHG19 promotes the Protein Tyrosine Phosphatase 4A3 expression by sponging miR-137 to liberate Protein Tyrosine Phosphatase 4A3 mRNA transcripts. SNHG19 enhances the development of hepatocellular carcinoma by affecting miR-137/Protein Tyrosine Phosphatase 4A3 axis.
Conclusion:
These results demonstrated the effect of SNHG19 in the occurrence and progression of hepatocellular carcinoma. SNHG19 may be used as the specific molecular target in patients with hepatocellular carcinoma.
Introduction
Hepatocellular carcinoma (HCC) is one of the most common malignancies worldwide.1–3 HCC occupy nearly 80% of primary liver cancer and is the third most common cause of cancer-related death.2,4 The global incidence rate of HCC is increasing year by year, it ranks sixth in the world’s cancer incidence rate. 5 The treatment of HCC depends on the stage of HCC. 6 At present, comprehensive treatment strategies are mostly used, including ablation, resection, liver transplantation, transarterial embolization, and systemic treatment. 6 However, the current 5-year survival of HCC patients is very poor.7,8 Therefore, early detection and intervention are key to increase the survival rate of HCC patients. 9 The pathogenesis and development of HCC is still unknown, thus, further study of the mechanism of hepatocarcinogenesis is of great benefit to the diagnosis and treatment of HCC patients.
Most human DNA can be transcribed into RNA, but very few RNA are translated into proteins. 10 This sort of RNA without coding ability is termed noncoding RNA (ncRNA), lncRNA refer to a major subgroup of ncRNA with a length over 200 nucleotides.11,12 lncRNA participate in multiple biological processes of cancer occurrence and development by binding to DNA, RNA, and protein.13,14 Many studies have also shown that some specific lncRNA are involved in the malignant progression of HCC through regulating gene expression.15,16 Small Nucleolar RNA Host Gene 19 (SNHG19) was first detected in the cerebrum tissue of patients with Alzheimer’s disease. 17 Some scholars have confirmed that SNHG19 is highly expressed in breast cancer. 18 In addition, SNHG19 can enhance the ability of proliferation and metastasis in nonsmall cell lung cancer. 19 Some studies have analyzed the level of SNHG19 in HCC and found that it is overexpressed in tumor tissues. 20 Nevertheless, the exact effect and mechanism of SNHG19 in HCC are not very clear.
At present, the most in-depth research on the mechanism of lncRNA is that lncRNA participates in tumor progression as competitive endogenous RNAs (ceRNA). 21 Specifically, lncRNA upregulate the expression of target genes by competitively binding to target miRNA. 22 SNHG19 has also been proved to have the same mechanism. 19 In this report, we explored in detail the role and mechanism of SNHG19 in HCC. We found that SNHG19 expression level was upregulated in HCC. HCC patients with high levels of SNHG19 have shorter disease-free survival (RFS). SNHG19 can promote the expression of Protein Tyrosine Phosphatase 4A3 (PTP4A3) by removing the effect of miR-137 on its target gene PTP4A3 through sponging binding to miR-137. PTP4A3, a newly recognized oncogene, has been determined to be involved in progression, invasion, and metastasis of HCC. 23 We confirmed that SNHG19 enhances the proliferation and metastasis of HCC through the miR-137/PTP4A3 axis. Therefore, SNHG19 may be served as the specific molecular target in patients with HCC.
Materials and methods
Human tissue samples
Our study involved the HCC database of the Cancer Genome Atlas (TCGA, 374 HCC tissues vs 50 tissues of normal liver), and the data were obtained and analyzed on StarBase (http://starbase.sysu.edu.cn/). Twenty-two tumor tissues from HCC patient, matched normal hepatic tissue were gained in the Changshu No. 2 People’s Hospital. Two pathologists completed pathological diagnosis of related tissues independently. The study was sanctioned by the ethics committee of the Changshu No. 2 People’s Hospital.
Cell lines and cell culture
We obtained the cell line of HCC (HepG2, SMMC7721, and Huh7) in the American Type Culture Collection (ATCC, USA). LO2, normal human hepatic cell line, was derived from the Culture Collection of Chinese Academy of Sciences (Shanghai, China). We used Dulbecco’s modified Eagle’s medium (DMEM; Gibco, USA) to culture related cell lines, and added 10% fetal bovine serum (Invitrogen, USA), and culture environment was 37°C humid condition with 5% CO 2 .
Oligonucleotides and plasmids
We obtained small-interfering-RNA (siRNA) of SNHG19 and the mimic and inhibitor of miR-137 from GenePharma (Shanghai, China). In order to construct PTP4A3 plasmid, we used pcDNA3.1 vector (Invitrogen, USA). To transfect oligonucleotides and plasmids into HCC cell, the reagent of lipofectamine 3000 (Invitrogen, USA) was applied.
Quantitative RT-PCR
In order to extract RNA, TRIzol (Invitrogen, USA) was used. We applied reverse transcription Kit (Applied Biosystems, CA, USA) to perform reverse transcription. For amplification reaction, ABI StepOnePlus System (Applied Biosystems, CA, USA) was carried out. The experimental steps are carried out according to the instructions, and the reaction conditions are set. The sequences of primer: SNHG19 forward 5′-CGTCCAGGCCTGGCCTAC-3′ and SNHG19 reverse 5′-GCTCGCGACGAAACCTGC-3′; PTP4A3 forward 5′-CAGCCAGTCTTCCACTACCTT-3′ and PTP4A3 reverse 5′-GCTTCCTCATCACCCACAACC-3′; GAPDH forward 5′- GTCAGCCGCATCTTCTTTTG-3′ and GAPDH reverse 5′- GCGCCCAATACGACCAAATC-3′. The relative expression was expressed using 2–∆∆Ct method.
Western blot
We first extracted the protein, and quantified it by using bicinchoninic acid method. After separation, the protein was transferred to polyvinylidene fluoride membrane. We blocked the membrane with 5% skimmed milk, subsequently incubated with the primary antibody overnight. We detected the signal on the membrane after incubating the secondary antibody for 1 h. Antibodies against PTP4A3 (Affinity Biosciences, 1:1000, USA) was used.
Cell counting kit-8 assay
To evaluate the proliferation level of HCC cell, cell counting kit-8 (CCK-8) assay was used. We seeded the transfected cells into 96-well plates with 5000 cells/well. CCK-8 reagent (Beyotime, Shanghai, China) was added to cells at different times. An additional 2 h of 37°C incubation is required before being measured. Microplate reader from the Thermo Scientific was used to detect absorbance, and the optical density is 450 nm.
Transwell assay
To detect the invasion level of HCC cell, Transwell assay was used. The cell was placed in medium that does not contain serum, then stored in the superstratum of chamber Matrigel-coated chambers (BD Biosciences, USA). DMEM added 10% fetal bovine serum was placed to the down-layer zone as an inducer. We then continued to culture the cells for 24 h. Invading HCC cells were dyed with crystal violet.
Scratch wound assay
To detect the migration level of HCC cells, scratch wound assay was used. Cells were placed in the 6-well plates. We use tip of 200 μL pipette to scratch the cell layer. We then continued to culture the cells for 24 h. The wound width was detected at 0 and 26 h.
Clone formation assay
We seeded cells with different treatment factors into the 6-well plates at a density of 400 cells/well. The seeded cells were cultured in the incubator and fixed with 4% paraformaldehyde after 14 days. The cells were then stained with crystal violet and photographed.
Fluorescence in situ hybridization
We used RiboTM Fluorescent In Situ Hybridization (FISH) Kit (RiboBio, Guangzhou, China) to perform the FISH assay, and strictly follow the experimental steps. We synthesized SNHG19 probe (red fluorescence) from RiboBio (Guangzhou, China), nuclei were stained with Hoechst. Confocal microscopy is used to take representative photos.
Luciferase reporter assay
Binding sites were predicted. PTP4A3 3ʹ-UTR and SNHG19 fragments containing binding sites of miR-137 were inserted into reporter plasmid. We used the mutant as the control after constructing the wild-type reporter plasmid. The activity of luciferase was measured through Luciferase Reporter Assay System (Promega, USA) when transfection of related oligonucleotides and luciferase reporter plasmids.
RNA pull-down assay
We entrusted Guangzhou Ruibo to synthesize biotinylated miR-137. After transfection, cells were cultured with M-280 streptavidin magnetic beads (Invitrogen, USA). We used biotinylated mutants and NC as the control. In order to analyze the SNHG19 level in the bound RNA, qRT-PCR was used.
Xenograft tumor and immunohistochemistry staining assay
The nude mice were obtained from the Beijing Experimental Animal Center (Beijing, China). These mice were subcutaneously injected with HepG2 cell to form xenograft tumors. SNHG19 siRNA were locally injected into xenograft tumors every 5 days. The volume of xenograft tumor was calculated every 5 days. After 30 days, we killed them and dissected tumor tissue. We euthanized mice using a carbon dioxide chamber, and this study did not involve animal anesthesia. We set the humane endpoint at 30 days after tumor formation and ensured that the tumor weight did not exceed 10% of the mice weight. The tumor was not located in a position where the mice is functioning normally. The mice weight loss did not exceed 20%, and the tumor did not develop ulcers and the mice did not engage in self-harm behavior. The study was sanctioned by the ethics committee of the Changshu No. 2 People’s Hospital. Immunohistochemistry was performed using antibody PTP4A3 (Affinity Biosciences, USA).
Statistical analysis
The data were assessed through SPSS13.0 (mean ± SD). The statistically significant was analyzed by one-way ANOVA or t-test. The experiment was independently repeated three times. The disease-free survival was plotted, and the Spearman’s correlation analysis was performed. When p < 0.05, it was considered that this group of data was statistically significant.
Results
SNHG19 was overexpressed in HCC, it served as the risk factor for the survival rate of HCC patients
We first detected SNHG19 level in 374 HCC tissues and 50 normal liver tissue in the public HCC datasets (TCGA, StarBase (http://starbase.sysu.edu.cn/), and discovered that SNHG19 was overexpressed in HCC tissues (Figure 1(a)). We confirmed the result by analyzing the SNHG19 level in 22 HCC tissues and adjacent normal tissue (Figure 1(b)). Next, we also found that the level of SNHG19 was upregulated in HCC cell (HepG2, SMMC7721, Huh7) compared to human hepatic epithelial cell (LO2) (Figure 1(c)). The disease-free survival of HCC patients with high expression of SNHG19 was shorter by analyzing the data of TCGA-HCC prognostic database through GEPIA (http://gepia.cancer-pku.cn/index.html) (Figure 1(d)). These results indicated that SNHG19 is critic in the progression of HCC.

SNHG19 was overexpressed in HCC and served as a risk factor for survival rate of patients with HCC. (a) StarBase (http://starbase.sysu.edu.cn/) was used to detect the expression of SNHG19 in TCGA HCC dataset. (b) The level of SNHG19 was analyzed in 22 HCC tissues and adjacent normal tissue. (c) The expression profile of SNHG19 in HCC cell lines (HepG2, Huh7, SMMC7721) and normal human hepatic epithelial cell (LO2). (d) The disease-free survival of HCC patients in TCGA was analyzed by using GEPIA (http://gepia.cancer-pku.cn/index.html).
Knockdown of SNHG19 levels suppressed the proliferation and metastasis of HCC cell
In order to study the biological role of SNHG19 in HCC cells, the siRNA of SNHG19 was transfected into HepG2 and Huh7 cell (Figure 2(a)). The proliferation ability of HCC cell was analyzed through CCK8 assays, and found that the proliferation of HepG2 and Huh7 cell was inhibited after down-regulating of SNHG19 levels (Figure 2(b)). Meanwhile, SNHG19 siRNA transfected HCC cells showed significant inhibition of invasion and metastasis (Figure 2(c) and (d)). Clone formation assay also confirmed that the growth ability of HCC cells was significantly inhibited by SNHG19 siRNA (Figure 2(e)). These data suggested that SNHG19 enhanced the proliferation and metastasis of HCC cells.

Knockdown of SNHG19 levels inhibited the proliferation and metastasis of HCC cells. (a) Transfection efficiency of SNHG19 siRNA was determined by using qRT-PCR. (b) The proliferative ability of HepG2 and Huh7 cells was determined by CCK8 assay. (c) The effect of SNHG19 siRNA on the migratory ability of HepG2 and Huh7 cells was assessed by the scratch wound assay. Scale bar, 200 μm. (d) The invasive capacity of HepG2 and Huh7 cells was assessed by the Transwell assay. Scale bar, 50 μm. (e) The proliferative capacity of HepG2 and Huh7 cells was assessed by Clone formation assay.
SNHG19 functioned as a sponge of miR-137 in HCC
We further explored the mechanism of SNHG19 in HCC. lncRNA proved to act as a role of ceRNA that can mediate miRNA in many tumors.20,21 We first identified the cell localization of SNHG19 by using FISH, and found that SNHG19 was located in both nucleus and cytoplasm (Figure 3(a)). Bioinformatics software discovered that miR-137 have a binding site in SNHG19 (Figure 3(b)). We synthesized SNHG19 luciferase reporter plasmids, and mutant vectors was also constructed (Figure 3(b)). The luciferase activity was repressed by miR-137 mimic in wild-type plasmid (Figure 3C). To confirm the direct binding of SNHG19 and miR-137 in HCC cells, we perform the RNA pull-down assay. SNHG19 was proved to be pulled down by biotinylated-miR-137 (Figure 3(d)). Moreover, the miR-137 levels were promoted after transfection of SNHG19 siRNA (Figure 3(e)). These showed that miR-137 and SNHG19 bind directly in HCC.

SNHG19 acted as a miR-137 sponge in HCC. (a) FISH showed that SNHG19 was mainly distributed in both cytoplasm and nucleus in PC cells. Scale bar, 25 μm. (b) The binding sites of miR-137 on the SNHG19. (c) Luciferase activity of PC cells transfected with SNHG19-WT or SNHG19-MUT reporter together with miR-137 or NC. (d) HCC cells were transfected with biotin-labeled miR-137, and assayed by biotin-pull-down. SNHG19 levels were analyzed by qRT-PCR. (e) The expression of miR-137 in HCC cells following transfection with SNHG19 siRNA.
SNHG19 promote PTP4A3 expression by competitive binding miR-137 in HCC cells
Bioinformatics software (StarBase, miRDB, TargetScan) found that PTP4A3 maybe the potential target. The 3ʹ-UTR of PTP4A3 shares the binding sites with SNHG19 (Figure 4(a)). The wild-type PTP4A3 3ʹUTR luciferase reporter plasmids were constructed, and mutant plasmid as control. The luciferase activity of PTP4A3 wild-type plasmid was depressed by miR-137 mimic (Figure 4(b)). In HCC cells, miR-137 also suppressed the mRNA and protein expression levels of PTP4A3 (Figure 4(c) and (d)). These results confirmed that PTP4A3 is the target gene of miR-137. The SNHG19 siRNA suppressed the luciferase activity in PTP4A3 wild-type plasmid, and this inhibition effects can be abolished through co-transfected miR-137 inhibitor (Figure 4(e)). The protein and mRNA expression levels of PTP4A3 were decreased in SNHG19 siRNA transfected group, these effects can also be reversed by miR-137 inhibitor (Figure 4(c) and (d)). The TCGA-HCC database showed that SNHG19 has a positive correlation with PTP4A3 mRNA in HCC (Figure 4(f)). These indicated SNHG19 promotes the expression of PTP4A3 through sponging miR-137.
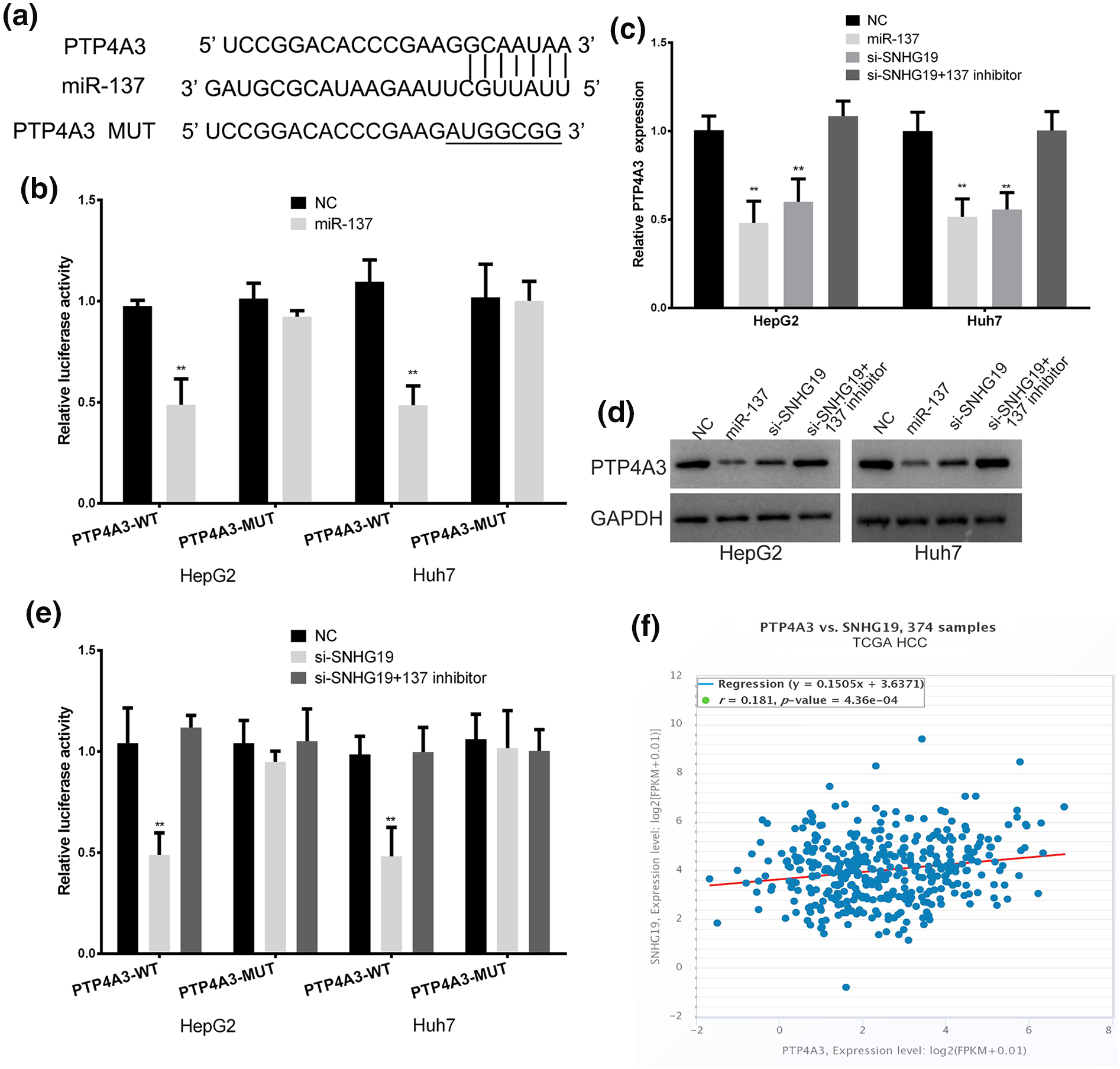
SNHG19 promote PTP4A3 expression by competitive binding miR-137 in HCC cells. (a) The binding sites of miR-137 within the 3ʹ-UTR of PTP4A3. (b) Luciferase activity of HCC cells transfected with PTP4A3-WT or PTP4A3-MUT reporter. (c) The expression of PTP4A3 mRNA in HCC cells transfected with miR-137 mimic, SNHG19 siRNA, or SNHG19 siRNA plus miR-137 inhibitor. (d) Western blots identified PTP4A3 protein expression changes, GAPDH was used as a control. (e) Luciferase assay of HCC cells transfected with PTP4A3-WT or PTP4A3-MUT reporter together with SNHG19 siRNA or SNHG19 siRNA plus miR-137 inhibitor. (f) TCGA dataset revealed a significant positive correlation between PTP4A3 mRNA and SNHG19 in HCC tissues.
SNHG19 enhances the growth and metastasis of HCC cell through miR-137/PTP4A3 axis
We further explored whether SNHG19 exerts its effect in HCC through the miR-137/PTP4A3 pathway. Subsequently, we transfected miR-137 mimic, SNHG19 siRNA, SNHG19 siRNA + miR-137 inhibitor, and PTP4A3 plasmid + miR-137 mimic into HCC cells. The protein level of PTP4A3 in different intervention groups were detected (Figure 5(a)). The miR-137 mimic depressed the growth and metastasis of HCC cells, and PTP4A3 plasmid abolished the effect of miR-137 (Figure 5(b)–(e)). Therefore, we proved that miR-137 plays a role of antitumor gene in HCC by targeting PTP4A3. More importantly, the negative regulatory role of SNHG19 siRNA on the PTP4A3 level, metastasis, growth of HCC cell was abolished through co-transfected miR-137 inhibitor (Figure 5(b)–(e)). To sum up, these results indicated that SNHG19 acts its cancer promoting effect in HCC through decoying miR-137 to upregulate PTP4A3 expression.
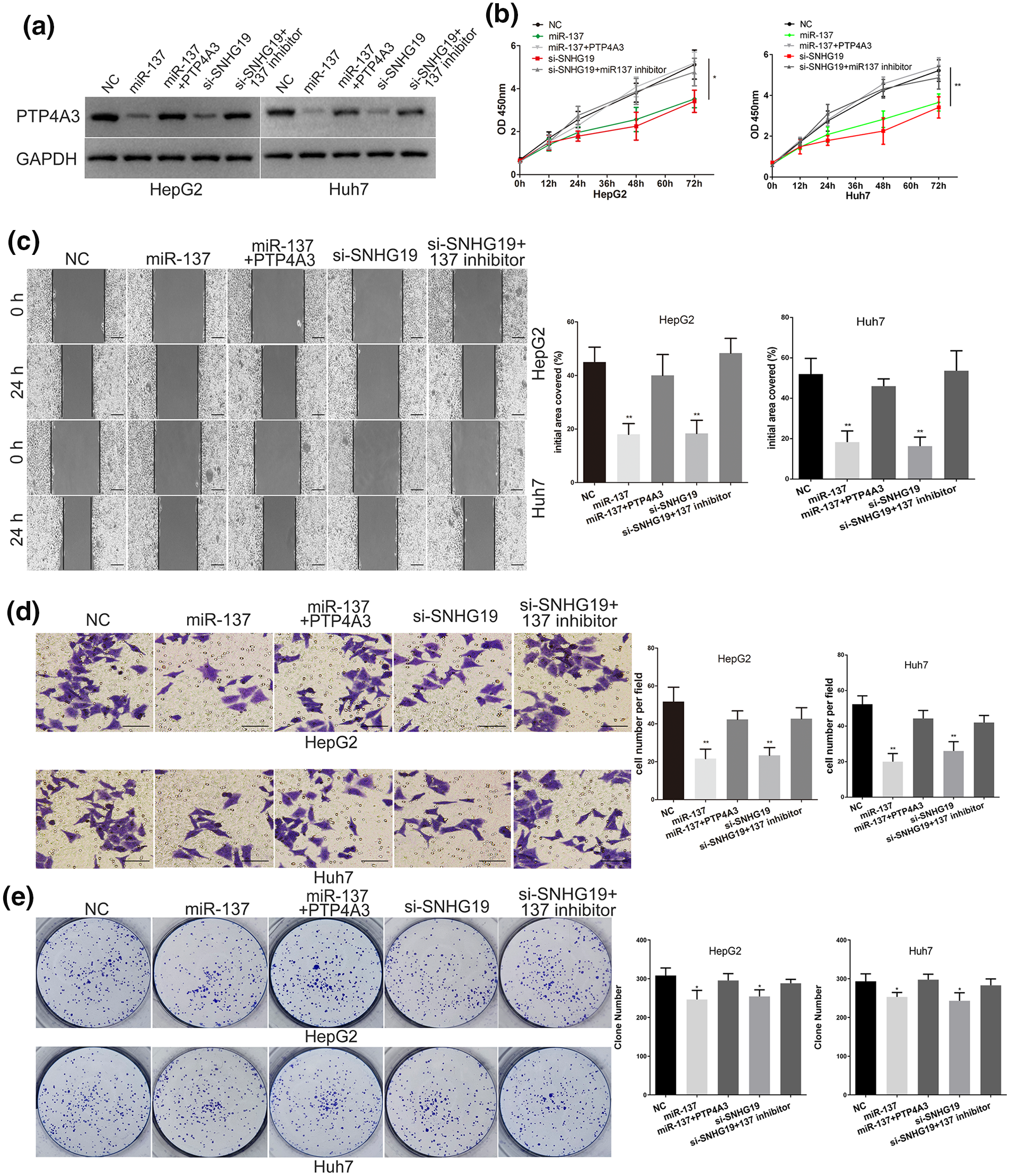
SNHG19 promotes the growth and metastasis of HCC cell via regulating miR-137/PTP4A3 axis. (a) Western blots identified PTP4A3 protein expression changes in transfected HCC cells, GAPDH was used as a control. (b) The proliferative ability of HCC cells was determined by CCK8 assay. (c) The migratory ability of HepG2 and Huh7 cells was assessed by the scratch wound assay. Scale bar, 200 μm. (d) The invasive capacity of HepG2 and Huh7 cells was assessed by the Transwell assay. Scale bar, 50 μm. (e) The proliferative capacity of HepG2 and Huh7 cells was assessed by Clone formation assay.
SNHG19 exerts its oncogenic activity in vivo
The xenograft model was used to explore the role of SNHG19 in vivo. Knockdown of SNHG19 inhibited the growth of tumor between 20 and 28 days (Figure 6(a) and (b)). We killed the mice and stripped the tumor after 30 days. In the SNHG19 knockdown group, the level of miR-137 increased, while the expression of PTP4A3 decreased (Figure 6(c) and (d)). These results indicated that SNHG19 promoted the in vivo progression of HCC through regulating the miR-137/PTP4A3 pathway.

SNHG19 exerts its oncogenic activity in vivo. (a) The excision tumor in nude mice of HCC xenograft model. (b) Differences in tumor volume among groups. (c) PCR identified SNHG19 and miR-497-5p expression changes. (d) The expression of PTP4A3 was examined by immunohistochemical staining of sections from the xenograft model. Scale bar, 25 μm.
Discussion
LncRNA, a class of ncRNA with a length over 200 nucleotides, involves many biological processes, including gene transcription and protein translation.24,25 Some special lncRNAs were identified to be abnormally expressed in many tumor tissues, and act the significant role in genesis and metastasis of many tumor.26,27 SNHG19, a new recognized lncRNA, was first detected in brain tissues of Alzheimer’s disease. 17 Recently, SNHG19 has been discovered to be highly expressed in many tumors, including nonsmall cell lung cancer, breast cancer, and HCC.18–20 SNHG19 has been proved to be a carcinogenic lncRNA in some tumors. 19 But the specific role of SNHG19 in HCC was unclear. We first detected SNHG19 expression level by using TCGA-HCC database and our HCC samples. SNHG19 is upregulated in HCC tissues, and the high level of SNHG19 is a harmful factor for disease-free survival of HCC patients. SNHG19 also enhanced the metastasis and growth capability of HCC cell. We next research focuses on which mechanism does SNHG19 play its role.
At present, the well-established theory is that lncRNA can exert its biological functions as the ceRNA to regulate gene expression in some tumors.28–31 ceRNA is a kind of RNA with miRNA binding sites, which can competitively bind miRNA to reduce miRNA binding to target genes. 32 In this way, ceRNA plays its role by regulating the expression of tumor-related gene. Similarly, SNHG19 has been proved to promote the malignant progression of nonsmall-cell lung carcinoma by regulating the miR-137/E2F7 pathway. 19 MiR-137 has been proven to suppress the growth and progression of HCC. 33 Here, we further studied whether SNHG19 function as ceRNA in HCC. SNHG19, miR-137, and PTP4A3 were found to have the potential ceRNA correlation in HCC based on bioinformatics software and TCGA database. We further proved that PTP4A3 is a target gene of miR-137, and SNHG19 can bind to miR-137. The inhibitory effect of SNHG19 knockdown on the luciferase activity of wild-type PTP4A3 plasmids and PTP4A3 level can be reversed by miR-137 inhibitor. These proved that SNHG19 promotes the PTP4A3 expression by decoying miR-137 in HCC.
PTP4A3 is a member of the phosphatase of regenerating liver (PRL) family, commonly known as PRL-3. 34 PTP4A3 is involved in regulating the signal cascade that closely related to cell growth and viability.35,36 PTP4A3 was confirmed to be highly expressed in HCC, and participated in progression, invasion, and metastasis of HCC. 23 It was also found that overexpression of PTP4A3 was associated with decreased survival rate of HCC patients. 23 Here, we demonstrated that miR-137 exerts its anti-cancer effect by targeting PTP4A3. In addition, we indicated that miR-137 inhibitor can neutralize the role of si-SNHG19 on growth and metastasis of HCC cells. To sum up, this study demonstrated the ceRNA network of SNHG19/miR-137/PTP4A3 in HCC. However, there are still some limitations in this article. First, this study did not conduct a formal sample size calculation, which may introduce several limitations to the validity and reliability of the findings, including increasing the risk of Type II errors and reducing the generalizability of the findings due to potential underrepresentation of population variability. Second, the clinical sample size of this study is still relatively small. Third, there is still a long way to go in the clinical translational application of this study. In the future, we will gather larger samples to draw more reliable conclusions. Scholars have confirmed that repurposing drugs and giving various vitamins as D as prophylactic with a modulatory effect with a positive impact on HCC. 37 Therefore, the development of antitumor drugs is essential. It is worthwhile to develop new targeted drugs for HCC based on this research in the future.
Conclusion
Our results proved that SNHG19 is key to the malignant progression of HCC. SNHG19 inhibits the binding between miR-137 and PTP4A3 through competitive binding to miR-137, resulting in the up-regulation of PTP4A3 by liberating PTP4A3 mRNA. Further study on the internal mechanism of SNHG19/miR-137/PTP4A3 axis is helpful for us to find novel biological targets for the HCC patients.
Footnotes
Ethical considerations
The study was conducted in accordance with the Declaration of Helsinki principles. The study was sanctioned by the ethics committee of the Changshu No. 2 People’s Hospital. Significant efforts were made to minimize both the number of animals and their respective suffering. Ethical approval for this study was obtained from *Ethics committee of the Changshu No. 2 People’s Hospital (2020-KY-046DY)*.
Consent to participate
The informed consent was obtained from each participant. Written informed consent was obtained from all subjects before the study.
Author contributions
XS and WL conceived and designed the experiments; YD, WL, FL, and ZW performed the experiments. WL and XS provided the technical support. WL, YD, and FL analyzed and interpreted the data. YD, XS, and FL wrote the manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by Joint Open Project of Jiangsu Key Laboratory for Tumor biotherapy (XZSYSKF2020006), Science and technology project of Changshu Health Commission (CSWS202010), Science and technology project of Changshu No. 2 People’s Hospital (CSEY2021010), Open Project of Jiangsu Key Laboratory for Encephalopathy bioinformatics 2024 (34), Kuncheng talent project 2024.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
