Abstract
Objective
Osteoarthritis (OA) is the most common arthritic disease in humans. Nevertheless, the pathogenic mechanism of OA remains unclear. This study aimed to explore that heat-shock transcription factor 1 (HSF1) facilitated interleukin-1 beta (IL-1β) chondrocyte injury by increasing Notch1 O-linked N-acetylglucosamine (O-GlcNAc) modification level.
Design
Human chondrocytes were incubated with 5 ng/ml interleukin-1 beta (IL-1β) for 24 h to establish OA cell model. The messenger RNA (mRNA) or protein expressions were assessed using reverse transcription-quantitative polymerase chain reaction, western blot, or immunofluorescence. Chondrocyte viability was examined by Cell Counting Kit-8 assay. Enzyme-linked immunosorbent assay was employed to detect the secretion levels of interleukin-6 (IL-6) and interleukin-8 (IL-8). Immunoprecipitation was adopted to detect Notch1 O-GlcNAc modification level. The interaction between HSF1 and epidermal growth factor–like (EGF) domain–specific O-GlcNAc transferase (EOGT) promoter was analyzed by dual-luciferase reporter gene and chromatin immunoprecipitation assays.
Results
Herein, our results demonstrated that HSF1, EOGT, Notch1, and Notch1 intracellular domain (NICD1) expressions in chondrocytes were markedly increased by IL-1β stimulation. EOGT elevated Notch1 expression in IL-1β-treated chondrocytes by increasing Notch1 O-GlcNAc modification level. EOGT silencing reduced IL-1β-induced chondrocyte inflammatory injury. In addition, HSF1 knockdown relieved IL-1β-induced chondrocyte inflammatory injury. Molecular interaction experiment proved that HSF1 transcriptionally activated EOGT expression in IL-1β-treated chondrocytes.
Conclusions
HSF1 promoted IL-1β-induced inflammatory injury in chondrocytes by increasing EOGT-mediated glycosylation of Notch1.
Introduction
Osteoarthritis (OA) is a common degenerative disease defined primarily by articular cartilage destruction and encompassing the whole articular tissue, resulting in articular cartilage degeneration, fracture, defect, and damage to the entire articular surface. 1 The elderly have a relatively high incidence of OA, with approximately 80% of people above the age of 75 having OA. 2 OA is the leading cause of disability in the elderly, severely affecting patients’ quality of life. 3 Traditional therapies, including drug therapy and acupuncture therapy, cannot cure OA completely. 4 It is suggested that understanding the pathogenic mechanisms of OA can greatly aid in the development of innovative OA therapy techniques.
Notch is a highly conserved cell surface receptor that plays an important role in cell fate determination by regulating embryogenesis and differentiation and apoptosis during postnatal development. 5 As widely reported, Notch signaling activation in chondrocytes contributes to OA progression.6,7 During cartilage degradation, Notch is highly expressed in articular chondrocytes and can translocate from the cell surface to the nucleus. 6 In addition, excessive Notch signaling exacerbated OA by inhibiting chondrocyte proliferation and terminal differentiation. 8 It is suggested that understanding the mechanism of Notch activation contributes to the development of novel therapeutic strategies for OA. O-glycosylation is a post-translational modification of proteins, which has important functional significance in both physiological and disease contexts. 9 O-glycosylation plays important role in mediating protein quality control, molecular recognition, interaction, and protease protection by regulating the chemical and physical properties of proteins, and O-glycosylation dysregulation can lead to various defects in the organism. 10 Many studies have found that protein O-glycosylation is linked to the development of OA. A recent study 11 found that protein glycosylation was dysregulated in osteoarthritic equine carpal synovial fluid. In addition, it was previously described that accumulation of O-linked N-acetylglucosamine (O-GlcNAc) modified proteins aggravated OA by triggering hypertrophic alterations in chondrocytes. 12 Notably, glycation changes affect epidermal growth factor–like (EGF) repeat sequences of the extracellular domain of Notch receptors. 13 EGF domain–specific O-GlcNAc transferase (EOGT) can add O-GlcNAc to EGF repeats of Notch protein. EOGT-induced aberrant O-glycosylation and dysregulation of Notch occur in various diseases.14,15 However, it is uncertain whether Notch1 overexpression in OA is connected to EOGT-mediated O-glycosylation alteration, which warrants additional investigation.
The heat-shock transcription factors (HSFs), regulated by thermal stress, are a family of DNA-binding proteins that regulate gene expression at the transcriptional level. Heat-shock transcription factor 1 (HSF1) is one of the most studied members of this family. HSF1 is modulated by a multi-molecular chaperone complex composed of different heat-shock proteins. 16 Phosphorylated HSF1 assembles into homologous trimers and accumulates in the nucleus after activation to control gene expression. 17 A previous study showed that HSF1 was activated in synovial tissues of rheumatoid arthritis patients, and pro-inflammatory cytokines could induce HSF1-DNA binding activation in cultivated synovial fibroblast-like cells (SFCs). 18 The evidence presented above implies that inflammation can induce HSF1 activation and transcriptional activation of downstream factor expression under pathological conditions, exacerbating inflammatory disease. However, the role of HSF1 in OA has not been thoroughly clarified, which is valuable for us to investigate. Herein, we predicted that HSF1 has potential binding sites to EOGT promoter by using JASPAR (http://jaspar.genereg.net/), suggesting that HSF1 may regulate the transcription of EOGT.
In general, it is speculated that HSF1 transcriptionally activated EOGT expression during OA progression, and then EOGT promotes O-glycosylation of Notch1 and activates Notch1 signal, thus exacerbating chondrocyte inflammatory injury. Our research suggests that HSF1 may be a potential target for the diagnosis and therapy of OA.
Materials and Methods
Isolation and Culture of Primary Human Chondrocytes
The cartilage tissues were collected from patients (4 females and 1 male) with femoral neck fractures undergoing total hip replacement surgery. The study was authorized by the Ethics Committee of Honghui Hospital, Xi’an Jiaotong University before enrollment of patients. All participants signed informed consent. Primary chondrocytes were isolated from cartilage tissues as previously described. 19 Chondrocytes were cultured in DMEM (Sangon, Shanghai, China) supplemented with 10% fetal bovine serum (FBS; Sangon) at 37°C with 5% CO2. The second generation of chondrocytes were used for follow-up experiments.
Cell Treatment
When human chondrocytes were in logarithmic growth phase, the OA cell model was established according to the previous report. 20 In brief, we established an inflammatory and a senescence model, which consisted of stimulating human chondrocytes with IL-1β (PeproTech, NJ) at 5 ng/ml for 24 h.
Cell Transfection
The small-interfering RNA of EOGT (si-EOGT), si-HSF1, and their negative controls, obtained from GenePharma (Shanghai, China), were transfected into cells with Lipofectamine™ 3000 (Invitrogen, Carlsbad, CA). In brief, cells were inoculated into a 6-well plate the day before transfection (18-24 h) to allow the cell density to reach about 50% to 60% on the second day. Before performing the transfection procedure, each well was replaced with 2 ml of fresh medium (serum and antibiotic-free). For cells in each well to be transfected, Opti-MEM (Gibco, Baltimore, MD) was added into 2 clean sterile centrifuge tubes (125 μl/tube), then 100 pmol siRNA or si-NC was added to one of the tubes and 5 μl of Lipofectamine 3000 was mixed in another tube gently. After standing at room temperature for 5 min, the culture medium containing the siRNA was gently added into the culture medium containing Lipofectamine 3000 transfection reagent. After incubating at room temperature for 20 min, the Lipofectamine 3000-siRNA mixture was added dropwise to the well. To achieve the highest transfection efficiency, cells were replaced with fresh complete cultures after 5 h of transfection. After 24 h, the transfection efficiency was detected by reverse transcription-quantitative polymerase chain reaction (RT-qPCR) and western blot.
RT-qPCR
The total RNA from chondrocytes was extracted by TRIzol® reagent (Thermo Fisher Scientific, Waltham, MA). The concentration and purity of all samples were measured via a NanoDrop 2000 (Thermo Fisher Scientific). The extracted RNA (200 ng) was reverse transcribed into cDNA using the Reverse Transcription Kit (Toyobo, Tokyo, Japan). Quantitative PCR was carried out using SYBR (Thermo Fisher Scientific) and a LightCycler 480 Real-Time PCR system (Roche, Shanghai, China). GAPDH was employed as the reference gene. The data were analyzed using the 2−ΔΔCT method. The primers were listed as follows (5′-3′):
HSF1 (F): ACCCATGCTTCCTGCGTGGC
HSF1 (R): TGCTTCTGCCGAAGGCTGGC
EOGT (F): AGAGGCGTTCACTACATCACT
EOGT (R): GCAGCCTGAAGGACAAGATAC
GAPDH (F): GACTCATGACCACAGTCCATGC
GAPDH (R): AGAGGCAGGGATGATGTTCTG
Western Blot
The proteins were isolated with RIPA (radioimmunoprecipitation assay). Then, the proteins were quantified by a BCA kit (Beyotime). Subsequently, total protein (20 μg) was isolated by 10% SDS-PAGE (sodium dodecyl sulfate-polyacrylamide gel electrophoresis) and transferred onto the PVDF (polyvinylidene difluoride) membrane (Millipore, MA). Then, the membranes were sealed with 5% skim milk for 1 h and incubated overnight with antibodies against EOGT (Proteintech, Wuhan, China; 1:1000, 27595-1-AP), Notch1 (Santa Cruz Biotechnology, Dallas, TX; 0.1 µg/ml, sc-6014R), cleaved Notch1 (NICD1; Cell Signaling Technology, Danvers, MA; 1:1000, #4147), HSF1 (Abcam; 1:50000, ab52757), O-GlcNAc-modified proteins (Thermo Fisher Scientific; 1:1000, MA1-072) and GAPDH (Abcam; 1:10000, ab8245, ab9485) at 4°C, then hybridized with secondary antibody (Abcam; 1:10000, ab7090, ab124055) for 60 min. Blots were visualized by an ECL detection kit (Beyotime).
Immunoprecipitation
Human chondrocytes were incubated with 5 ng/ml IL-1β for 24 h combined with small interfering RNA negtive cotrol (si-NC) or si-EOGT transfection. The proteins were extracted from cells using radioimmunoprecipitation buffer (Transgene, Beijing, China). Equal amounts of protein (200 µg) were then incubated with anti-Notch1 (Abcam; 1:200, 10062-2-AP) antibody overnight and subsequently incubated with Protein A/G-agarose (Santa Cruz) for 4 h. The beads were washed with lysis buffer. The immunoprecipitated protein was then eluted from the magnetic bead and subjected to western blot analysis by using anti-O-GlcNAc (Abcam; 1:200, ab2739).
Cell Counting Kit-8 Assay
Cells were cultured in 24-well plates (2 × 104 cells/well) for 24 h and incubated with Cell Counting Kit-8 (CCK-8) solution (10 μl; Sangon) at 37°C for 3 h. Absorbance at 450 nm was subsequently analyzed.
Enzyme-Linked Immunosorbent Assay
The secretion levels of interleukin (IL)-8 and IL-6 were examined by human IL-8 enzyme-linked immunosorbent assay (ELISA) kit (Solarbio, Beijing, China; SEKH-0016) and human IL-6 ELISA kit (Solarbio; SEKH-0013), respectively. All operations were strictly carried out in accordance with the instructions.
Immunofluorescence
Cells were fixed with 4% paraformaldehyde for 10 min, permeabilized with 0.3% Triton X-100 and sealed with goat serum for 30 min. Cells were subsequently incubated overnight with antibodies against MMP13 (Proteintech, Wuhan, China; 1:500, 18165-1-AP) and AdAMTS5 (ABclonal, Wuhan, China; 1:200, A2836) for 2 h. Then, cells were incubated with the secondary antibody (Abcam; 1:3000, ab7090) for 1 h, and the nucleus was stained with DAPI (Sangon). Cells were subsequently sealed and observed under a fluorescence microscope (Olympus, Tokyo, Japan).
Dual-Luciferase Reporter Gene Assay
EOGT promoter fragments containing HSF1 wild-type (WT)/mutant (MUT) binding sites were amplified by PCR and inserted into the pGL3 reporter plasmid (GenePharma) expressing firefly luciferase (F-luc). Cells were co-transfected with si-HSF1 or sh-NC, and the luciferase activity was subsequently examined. The Renilla luciferase functioned as the internal control.
Chromatin Immunoprecipitation Assay
Chromatin immunoprecipitation (ChIP) assay was used to detect the binding relationship between HSF1 and EOGT promoter. Cells were fixed with 1 % formaldehyde solution for 10 min and quenched with 125 mM glycine for 5 min, and DNAs were fragmented into 200 to 500 bp length fragments by sonication. The cell lysates were then incubated with anti-HSF1 (Abcam; 1:50, ab52757) or anti-IgG (Abcam; 1:1000, ab171870) at 4°C overnight. DNA was enriched and subjected to gel electrophoresis analysis.
Statistical Analysis
All cells were randomly grouped according to experimental requirements. All the data from 3 independent experiments were analyzed by SPSS 19.0 and expressed as means ± SD. The normality of data was assessed by the Shapiro-Wilk test. Considering a significance level of 5%, there were no significant deviations from the normality of all data (P > 0.05). The differences among 2 groups were analyzed by Student’s t tests. One-way analysis of variance (ANOVA) was performed to assess the differences among multiple groups. The P values less than 0.05 were considered significant.
Results
The High Expression of Notch1 in IL-1β-Treated Chondrocytes was Associated With EOGT-Mediated O-GlcNAc Modification
As reported, Notch signaling activation in chondrocytes contributes to OA progression.
6
In the current research, it was observed that O-GlcNAc modification of Notch1 protein in chondrocytes was significantly increased by IL-1β treatment (
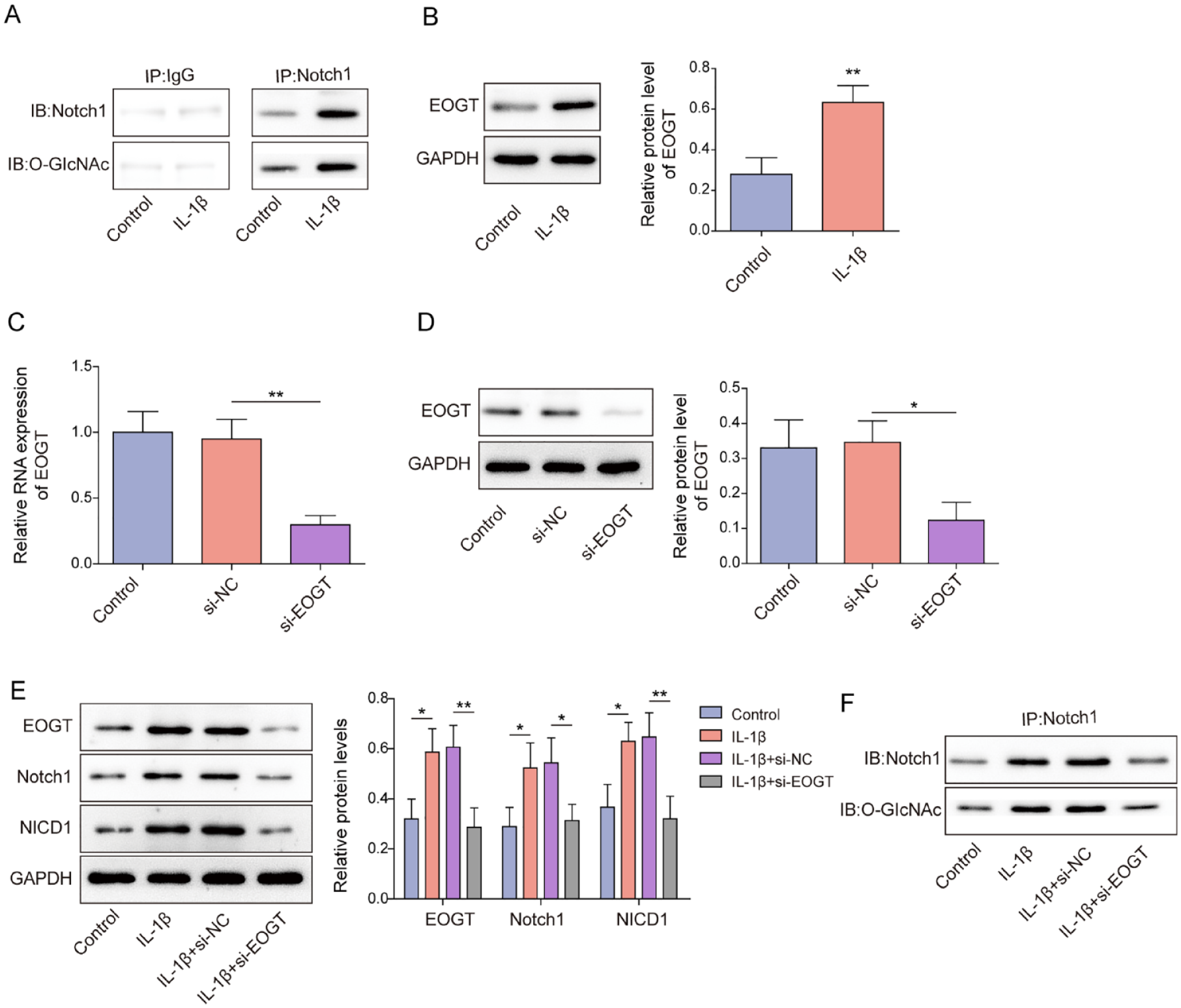
The high expression of Notch1 in IL-1β-treated chondrocytes was associated with EOGT-mediated O-GlcNAc modification. Human chondrocytes were incubated with 5 ng/ml IL-1β for 24 h. (A) IP was adopted to detect Notch1 O-GlcNAc modification level in cells. (B) EOGT protein level in cells was measured by western blot. (C-D) EOGT expression in chondrocytes following si-NC or si-EOGT transfection was measured by RT-qPCR and western blot. IL-1β-treated chondrocytes were transfected with si-NC or si-EOGT. (E) Western blot was adopted to examine EOGT, Notch1, and NICD1 protein levels in cells. (F) Notch1 O-GlcNAc modification level was measured using IP. Data were expressed as mean ± SD. All our data were obtained from 3 independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001. EOGT = epidermal growth factor–like domain–specific O-GlcNAc transferase; IL-1β = interleukin-1 beta; RT-qPCR = reverse transcription-quantitative polymerase chain reaction.
EOGT Knockdown Alleviated IL-1β-Induced Chondrocyte Inflammatory Injury
Then EOGT knockdown was induced to investigate the role of EOGT in IL-1β-treated chondrocytes. ELISA results demonstrated that IL-1β stimulation significantly increased IL-6 and IL-8 secretion levels in chondrocytes, but EOGT knockdown attenuated this effect (
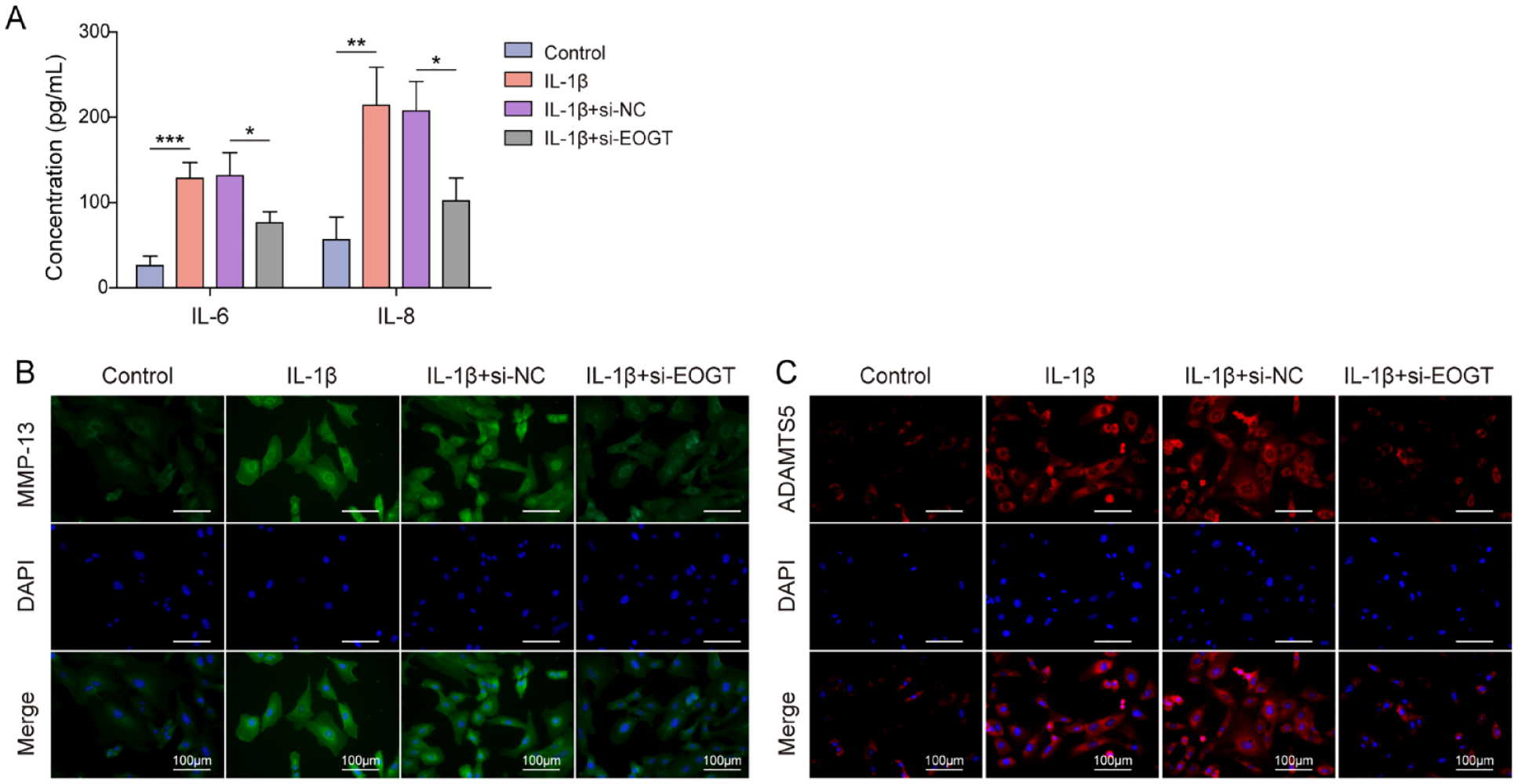
EOGT knockdown alleviated IL-1β-induced chondrocyte inflammatory injury. EOGT knockdown was induced in IL-1β-treated chondrocytes. (A) IL-6 and IL-8 secretion levels in chondrocytes were assessed by ELISA. (B-C) IF was adopted to detect MMP13 and ADAMTS5 fluorescence intensity in chondrocytes. Data were expressed as mean ± SD. All our data were obtained from 3 independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001. EOGT = epidermal growth factor–like domain–specific O-GlcNAc transferase; IL-1β = interleukin-1 beta; ELISA = enzyme-linked immunosorbent assay.
HSF1 Was Highly Expressed in IL-1β-Treated Chondrocytes and Was Involved in Notch1-Mediated Inflammatory Injury
HSF1 is a major transcription factor that regulates the heat-shock response. As shown in
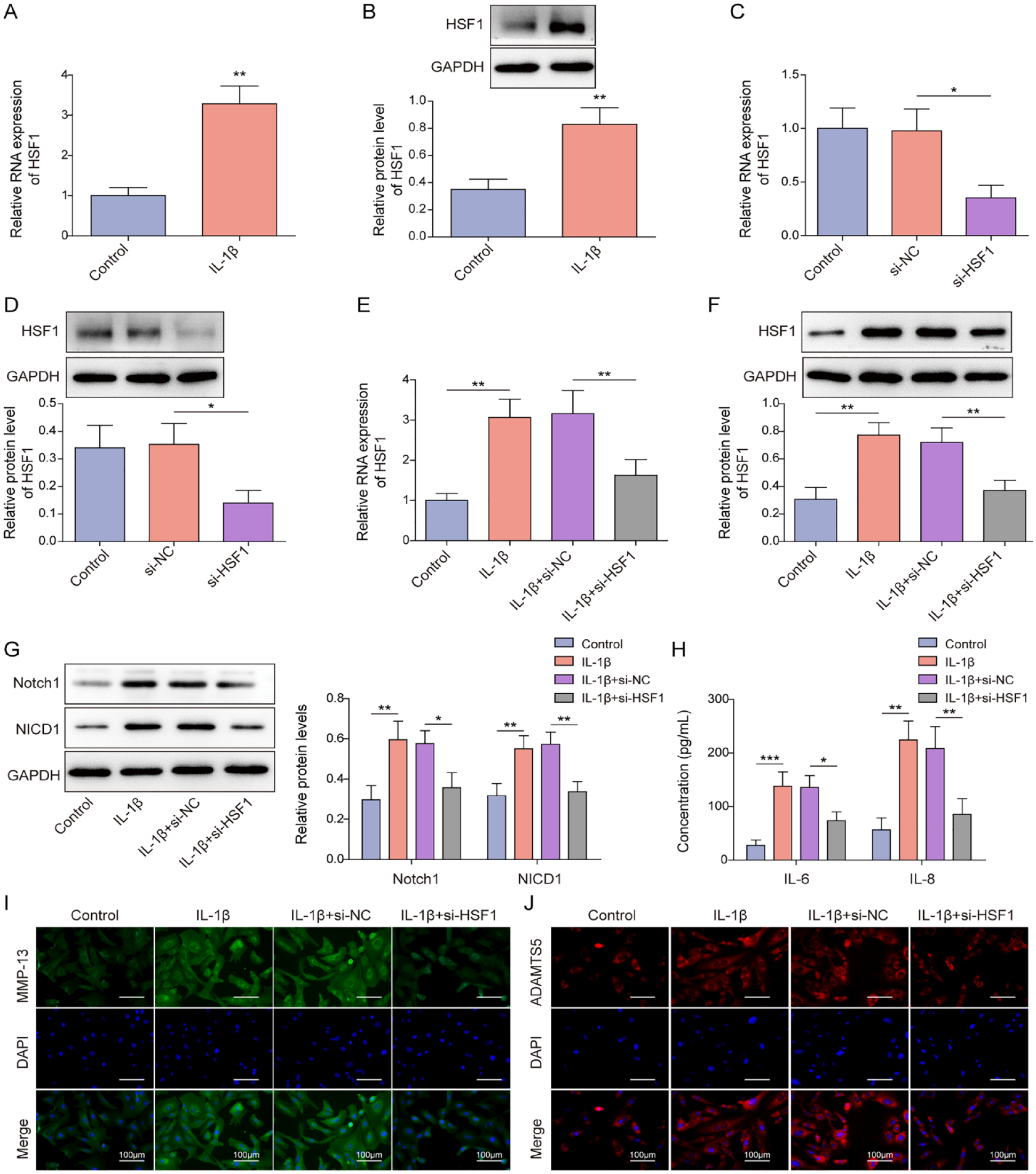
HSF1 was highly expressed in IL-1β-treated chondrocytes and was involved in Notch1-mediated inflammatory injury. (A-B) HSF1 expression level in human chondrocytes after IL-1β stimulation was measured by RT-qPCR and western blot. (C-D) RT-qPCR and western blot were adopted to examine HSF1 expression level in human chondrocytes following si-NC or si-HSF1 transfection. HSF1 knockdown was induced in IL-1β-treated chondrocytes. (E-F) HSF1 expression in chondrocytes was measured by RT-qPCR and western blot. (G) Notch1 and NICD1 protein levels in cells were examined using western blot. (H) ELISA was adopted to detect IL-6 and IL-8 secretion levels in chondrocytes. (I-J) IF was adopted to detect MMP13 and ADAMTS5 fluorescence intensity in cells. Data were expressed as mean ± SD. All our data were obtained from 3 independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001. HSF1 = heat-shock transcription factor 1; EOGT = epidermal growth factor–like domain–specific O-GlcNAc transferase; IL-1β = interleukin-1 beta; RT-qPCR = reverse transcription-quantitative polymerase chain reaction; ELISA = enzyme-linked immunosorbent assay; IF = immunofluorescence.
HSF1 Transcriptionally Activated EOGT Expression in IL-1β-Treated Chondrocytes
As revealed in
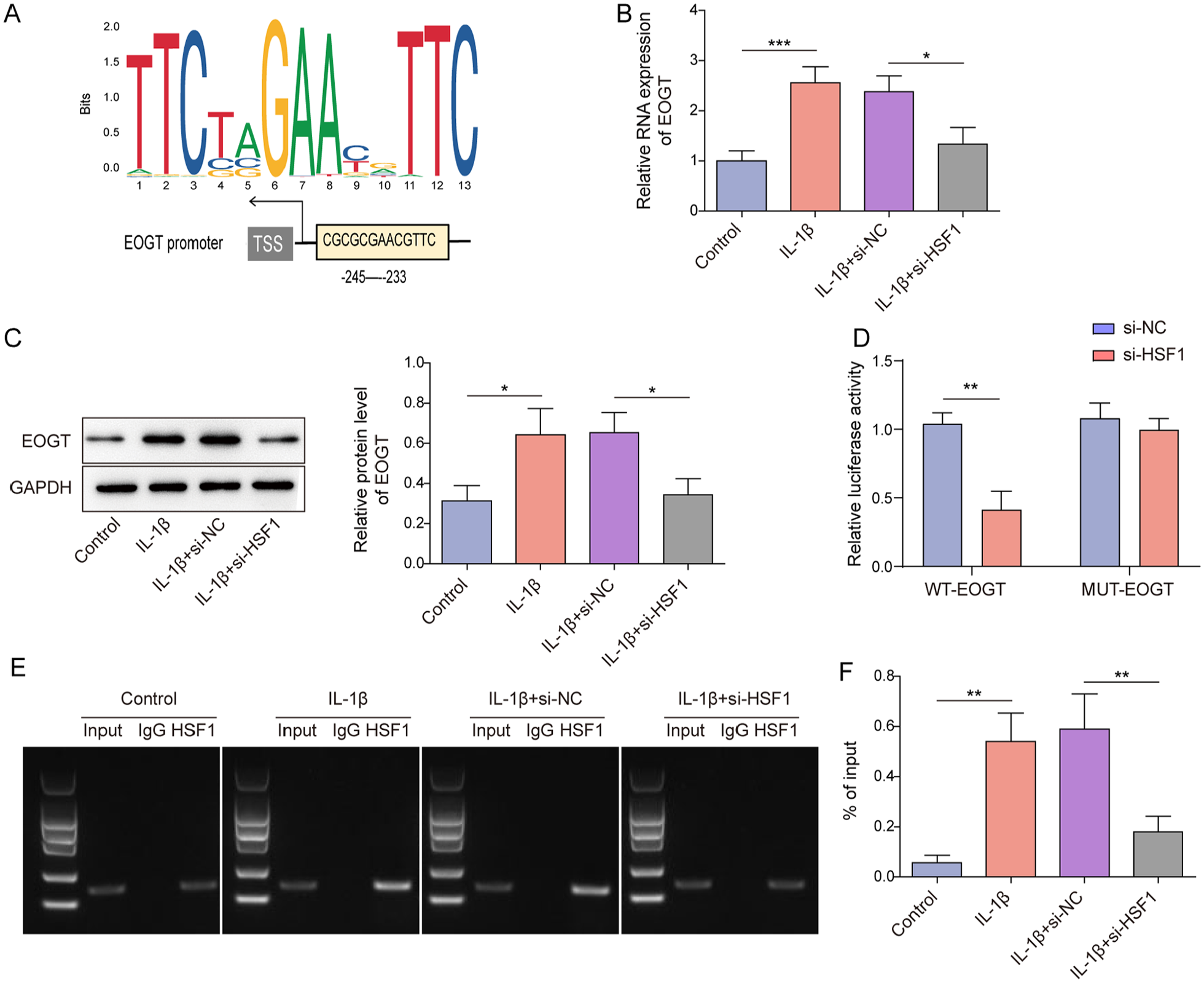
HSF1 transcriptionally activated EOGT expression in IL-1β-treated chondrocytes. (A) The potential binding sites between HSF1 and EOGT promoter was predicted using JASPAR. (B-C) IL-1β-treated chondrocytes were transfected with si-NC or si-HSF1, and EOGT expression level in cells was assessed by RT-qPCR and western blot. (D) The interaction between HSF1 and EOGT promoter was analyzed using dual-luciferase reporter gene assay. (E-F) The interaction between HSF1 and EOGT promoter was analyzed using ChIP assay. Data were expressed as mean ± SD. All our data were obtained from 3 independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001. HSF1 = heat-shock transcription factor 1; EOGT = epidermal growth factor–like domain–specific O-GlcNAc transferase; IL-1β = interleukin-1 beta; RT-qPCR = reverse transcription-quantitative polymerase chain reaction; ChIP = chromatin immunoprecipitation.
Discussion
OA is the most prevalent degenerative joint disease affecting 70% of the global population. 24 Notably, existing therapies can only relieve and control the symptoms of OA. 25 Abnormal activation of inflammatory response can lead to inflammatory injury of chondrocytes, leading to cartilage degradation and accelerating the progression of OA. 26 Our results showed that HSF1 promoted IL-1β-induced inflammatory injury in chondrocytes by increasing EOGT-mediated glycosylation of Notch1.
Notch, which has 4 mammalian components (Notch 1-4), is activated by ligands on adjacent cells. 27 After ligand binding, the NICD is lysed and transferred to the nucleus, where it regulates downstream target gene transcription. 27 Pathological Notch activation leads to the development of early and progressive OA pathology. 28 Consistently, our findings showed that Notch and NICD1 levels in human chondrocytes were markedly increased by IL-1β treatment. The specific mechanism of Notch activation during OA progression was subsequently investigated. O-glycosylation modification is a form of protein post-translation modification, which functions in regulating various cellular processes. As widely illustrated, EGF repeats of Notch protein are modified by O-linked glycans, and the extracellular domain is also modified by glycosylation. 14 Our results demonstrated that IL-1β treatment increased O-GlcNAc modification of Notch1 protein in human chondrocytes. EOGT can add O-GlcNAc to EGF of Notch protein. 29 Notch glycosylation abnormality mediated by EOGT dysregulation can induce many human diseases. 14 However, it is uncertain whether Notch1 upregulation in OA is related to EOGT-mediated O-glycosylation modification. Herein, EOGT was highly expressed in IL-1β-treated chondrocytes, and EOGT knockdown reversed the promoting effect of IL-1β on Notch1 O-GlcNAc modification level in chondrocytes. In addition, EOGT knockdown could relieve IL-1β-induced chondrocyte inflammatory injury. Collectively, EOGT facilitated IL-1β-induced inflammatory injury in chondrocytes by mediating glycosylation of Notch1 protein.
HSF1, as a transcription factor, has received a lot of interest in recent years for its involvement in regulating stress-related genes and the onset and progression of diseases. HSF1 has been shown to respond to stress factors, such as pro-inflammatory factors, by transcriptionally regulating downstream targets. 30 Notably, HSF1 was previously shown to be significantly elevated in rheumatoid arthritis, and this upregulation was induced by pro-inflammatory cytokines. 18 Our findings showed that HSF1 expression in human chondrocytes were markedly increased by IL-1β treatment; this is the first study to examine the role of HSF1 in OA. In addition, HSF1 knockdown inhibited IL-1β-induced inflammatory injury of chondrocytes. Notably, our following investigations showed that HSF1 transcriptionally activated EOGT expression in IL-1β-treated chondrocytes by directly targeting EOGT promoter. Collectively, HSF1 transcriptionally activated EOGT expression to increase Notch1 O-GlcNAc modification level and expression, thereby promoting IL-1β-induced inflammatory injury of chondrocytes.
Taken together, our research provided evidence that HSF1 upregulation promoted EOGT-mediated Notch1 O-GlcNAc modification, thereby promoting IL-1β-induced chondrocyte inflammatory injury, revealing that targeting HSF1 might be a potential therapeutic strategy for OA. The regulation of HSF1 by specific small molecules or related drugs might have important implications for OA treatment. Of course, there are some limitations to our article. For instance, we did not investigate the role of HSF1 in OA at the animal level, and we will make up for this deficiency in future studies.
Footnotes
Acknowledgments and Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Shaanxi Provincial Natural Science Foundation (2023-JC-YB-810) and General Research Projects of Xi’an Municipal Health Commission (2023yb28).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Consent to Participate
The study was authorized by the Ethics Committee of Honghui Hospital, Xi’an Jiaotong University before enrollment of patients. All participants signed informed consent.
