Abstract
Objective
Osteoarthritis (OA) is a common degenerative joint disease. The occurrence of OA slowly destroys the soft tissue structure of the patient’s joint. Severe cases could lead to disability. Current studies had shown that inhibition of chondrocytes pyroptosis could slow down the progression of OA. Our work aimed to explore the specific mechanisms and ways of regulating this process.
Design
In this work, the level of N6-methyladenosine (m6A) in clinical tissues was detected by ribonucleic acid (RNA) m6A dot blot. qRT-PCR (quantitative real-time polymerase chain reaction) was used to detect the messenger RNA (mRNA) expression level of m6A modified enzyme in clinical tissues. MTT (3-(4,5)-dimethylthiahiazo(-z-y1)-3,5-di-phenytetrazoliumromid) and flow cytometry were used to detect the effect of sh-METTL3 (methyltransferase like 3) and NIMA-related kinase 7 (NEK7) transfection on chondrocytes pyroptosis in OA. Western blot was used to detect the protein expression levels of pyroptosis-related proteins. ELISA (enzyme-linked immunosorbent assay) was used to measure the protein concentration of inflammatory cytokines. The SRAMP online database was used to predict the m6A site of NEK7. HE staining was used to assess the progression of OA in mice.
Results
The level of m6A in clinical samples of OA patients was higher, and METTL3 was significantly higher expressed in clinical samples of OA patients. We provided evidence that low expression of METTL3 inhibited chondrocytes pyroptosis. In addition, Rescue experiments and in vivo experiments had shown that METTL3 in combination with NEK7 inhibited the progression of OA by promoting chondrocytes pyroptosis.
Conclusions
METTL3 regulates m6A modification of NEK7 and inhibits OA progression.
Introduction
Osteoarthritis (OA) is a common bone and joint disease. The clinical manifestations of OA are joint redness, dysfunction, and joint deformity. Severe cases can even become disabled. 1 Statistics from the Incidence and Prevalence Database suggests that 355 million people suffer from arthritis worldwide, and predicts that the number will double by 2040. 2 The progression of OA is the result of a combination of factors, such as joint trauma, population aging, large base weight, and sedentary lifestyle. 3 At present, most of the diagnosis and treatment methods of OA are mainly aimed at relieving pain, reducing swelling, and improving joint function. 4 There are no effective drugs for the clinical treatment of OA. However, recent studies have found that targeted therapy based on the pathogenesis of OA to design personalized treatment processes was an effective clinical OA treatment scheme. 5 Therefore, we explored the pathogenesis of OA to find suitable therapeutic targets.
Pyroptosis is a kind of programmed cell death caused by inflammasome. 6 Pyroptosis is widely involved in the occurrence and development of cancer, cardiovascular disease, brain injury, and other diseases.7-10 Pyroptosis is caused by inflammasome. Pyroptosis of cells is mainly dependent on inflammatory caspase and GSDMs protein family. 11 The classic process of pyroptosis could be summarized as the activation of caspase1 by inflammasome (e.g., NLRP3) binding to ASC. Activated caspase1 spliced GSDMD (Gasdermin D). The release of GSDMD-N punctured the cell membrane. As the osmotic pressure of the cell changes, the cell swelled until the cell membrane bursts.12-14 When the cell membrane is broken, activated inflammatory cytokines, such as IL-1β (interleukin-1β), IL-18 (interleukin-18), IL-6 (interleukin-6), and TNF-α (tumor necrosis factor-α) are released. 15 Therefore, the pyroptosis process is often accompanied by a strong inflammatory response. 16 Further study of pyroptosis is helpful to understand its role in the occurrence and development of related diseases, which may provide innovative ideas for clinical treatment.
The NEK kinase family is a conserved serine/threonine kinase family that plays a crucial role in regulating signaling pathways in metabolic cells. 17 NIMA-related kinase 7 (NEK7) was the shortest member of the family (302 bp). 18 NEK7 participated in regulating various cellular biological behaviors. 19 Abnormal expression of NEK7 could lead to the increased number of mitotic cells, chromosome abnormalities, and programmed cell death. Studies have shown that NEK7 is involved in the formation and activation of NLRP3 inflammasome.20,21 In addition, NEK7 is the key regulatory protein of NLRP3 inflammasome involved in pyroptosis. 22 However, the specific regulatory mechanism and the complete molecular route of the pyroptosis process involved in NEK7 in OA have not been fully investigated.
N6-methyladenosine (m6A) is considered to be one of the main ways of RNA posttranscriptional modification. 23 NGS reveals that a third of RNA in mammals has 4 or so m6A modifications. 24 M6A modification also plays an important role in regulating RNA splicing, translation, stability, translocation, and higher structure. 25 The m6A modification process mainly relied on 3 types of m6A methylation modification enzymes: “writers” (methyltransferases: METTL3, METTL14 [methyltransferase like 14] and WTAP [Wilms tumor 1-associating protein]) are responsible for the catalysis of m6A modification. 26 “Erasers” are demethylase (ALKBH5 [alkylation repair homolog 5] and FTO [fat mass and obesity associated]). 27 “Readers” identify the targeted RNA modified by m6A. 28 Furthermore, m6A methylation has been reported to be involved in and influence the pyroptosis process. 29 However, the specific modification mode of m6A methylation on NLRP3 in OA has not been fully explained.
Therefore, our work aimed to explore the specific mechanism and regulation of m6A methylation modification of NEK7 during the progression of OA through in vitro and in vivo experiments. We hypothesized that METTL3 might act as the key enzyme in m6A methylation modification to promote chondrocytes pyroptosis and inhibit the formation of OA.
Materials and Methods
Cartilage Specimens Collection
Cartilage specimens were taken from patients in our hospital, who underwent Total Knee Arthroplasty (TKA) surgery. Normal cartilage tissue in the control group was taken from the knee joint of patients undergoing total hip replacement for femoral neck fractures. 30 The selected patients were all clinically diagnosed with OA. Patients with traumatic knee OA, rheumatoid arthritis, and infectious arthritis were excluded. All the enrolled patients were aware of the research purpose and specific procedure of the experiment before surgery. All participants signed informed consent. Our study has been approved by the ethics committee of our hospital.
Chondrocytes Culture
The cultivation method was as described above. 31 The collected cartilage tissue was cut into approximately 3 × 3 mm2 pieces with a sterile scalpel. The fragments were digested in 2% trypsin (Gibco, Darmstadt, Germany) for 30 minutes and co-incubated with collagenase for 16 hours. The product was filtered through a cellular sieve. The filtered cells were digested by DNA enzyme (Gibco) for 15 minutes. Primary chondrocytes were obtained after cells were centrifuged 3 times. The cells were grown in 20% Fetal Bovine Serum and 1% P/S Dulbecco’s Modified Eagle Medium/Nutrient Mixture F-12 (DMEM/F12; Gibco). The culture conditions were 37°C and 5% CO2. Lipopolysaccharide (LPS; 5 µg/ml; Sigma-Aldrich, Merck, Darmstadt, Germany) was used to induce primary chondrocytes at 37°C for 12 hours to construct cell models of OA. The OA cell lines used in this study are the second to fifth generation.
Chondrocytes Transfection
Short hairpin RNA METTL3 (sh-METTL3), over expression NEK7 (NEK7), and their negative control (sh-NC, vector) were purchased from GenePharma (Shanghai, China). When the integration degree of chondrocytes was about 80%, Lipofectamine 2000 (Thermo Fisher Scientific, CA) was used to perform cell transfection. The mRNA expression levels of METTL3 and NEK7 were determined by qRT-PCR.
Quantitative Real-time Polymerase Chain Reaction
The Total RNA Extraction Kit (Solarbio, Beijing, China) was used to extract total RNA from cells and tissues. A Universal RT-PCR Kit (M-MLV; Solarbio) was used to perform reverse transcription. 2×SYBR Green PCR Mastermix (Solarbio) was used to perform qRT-PCR. Primers involved in this article were listed in Supplemental Table S1. The relative expression levels of related genes were calculated by 2−ΔΔCT.
Stabilization Assay
Actinomycin D (2 mg/ml; Merck, Darmstadt, Germany) was treated with chondrocytes at 0, 6, 12, 18, and 24 hours. The mRNA expression levels of NEK7 were measured by qRT-PCR.
Western Blot
ProteoPrep Total Extraction Sample kit (Merck) was used to extract total proteins from chondrocytes. The target protein of the total protein was isolated by SDS-PAGE (polyacrylamide gel electrophoresis) and PVDF (polyvinylidenefluoride; Merck), and finally detected by an ECL kit (Beyotime, Beijing, China). After adding luminescent solution, the E-Gel Imager (Thermo Fisher Scientific) was used for exposure photography, image processing, and analysis. The main antibodies used in this study include NLRP3 (ab263899; Abcam, Cambridge, UK), ASC (ab307560; Abcam), caspase1 (ab179515; Abcam), GSDMD (ab215203; Abcam), and GADPH (ab9485; Abcam).
Dual-luciferase Reporter Assay
Site-directed Mutagenesis Kit (Sangon Biotech, Shanghai, China) was used to perform site-directed mutagenesis kit. We used the online software SRAMP (http://www.cuilab.cn/sramp) to predict the m6A sites of NEK7. MUT (Mutation) and WT (Wild type) fluorescent expression vectors for site1, site2, site3, and site4 of NEK7 were obtained from Genepharma. MUT or WT fluorescence expression vectors were co-transfected with sh-NC or sh-METTL3 into chondrocytes. The luciferase Reporter Assay System (Promega, Beijing, China) measured luciferase activity 48 hours later.
Cell Pyroptosis Assays
FLICA 660 Caspase-1 Assay Kit (ImmunoChemistry Technologies, CA, USA) was used for cell pyroptosis assays. Active Caspase-1 was detected in MCF-7 and MDA-megabyte (MB)-453 cell lines as recommended by the manufacturer. At the same time, propyl iodide (PI; Life Technologies, CA, USA) was added. IP was used to label cells with membrane pores. The fluorescence intensity of the cells was analyzed by flow cytometry (CytoFLEX, CA). Average fluorescence intensity was quantified using FlowJo v10.8.1 (OR).
Cell Viability Assay
The MTT cell viability and proliferation assay kit (ScienCell, CA) was used to assess the cell viability of chondrocytes. The transfected cells were added with 20 µl MTT solution and incubated for 4 hours. Then 150 µl of DMSO was added to the cells. Finally, cell viability was measured at 490 nm by MultiskanFC microplate reader (Thermo Fisher Scientific).
RNA m6A Dot Blot Assays
Total RNA is heated and cooled rapidly. The cooled total RNA was dripped onto Immobilon-Ny+ nylon coil film (Yiqi, Shanghai, China) and blocked by purple diplomatic membrane. The membrane was co-incubated with m6A antibody (1:1000; Abcam) at 4°C for 12 hours. The membrane was then coupled with Immunoglobulin G (IgG; Beyotime Biotechnology, Shanghai, China) for 1 hour. Finally, the membrane was visualized by the Imaging System (Bio-Rad, Universal Hood II, CA).
RNA-Binding Protein Immunoprecipitation
RIP (RNA-binding protein immunoprecipitation) lysis buffer (Millipore, MA) was used to lyse chondrocytes, which were then left overnight at 4°C. RNA protein immune complexes were precipitated by protein A/G magnetic beads. RNA was purified and verified by qRT-PCR.
MeRIP-qPCR
The extraction steps for Total RNA are shown above. 3 µg Synaptic Systems (RIP) anti-m6A was mixed with 200 µg RNA for 2 hours and then A/G magnetic beads were added. 50 µl of immunoprecipitation reagent (Thermo Fisher Scientific) was used to elute RNA. The enrichment of immunoprecipitated RNA was detected by qRT-PCR.
Enzyme-linked Immunosorbent Assay
The IL-1β ELISA kit, IL-18 ELISA kit, IL-6 ELISA kit, TNF-α ELISA kit, and IL-10 ELISA kit (Abcam) were used to detect the protein concentration of inflammatory cytokines. Cells were collected and centrifuged at 1,000 g for 20 minutes. ELISA kits were used to detect inflammatory components in cell supernatant and mouse serum.
Construction of OA Mice Models
Twenty male healthy C57BL/6J mice (10 weeks) were purchased from the National Rodent Laboratory Animal Resources Center (Shanghai, China). All mice were raised and treated in accordance with the procedures set by the Animal Resource Center of our School of Medicine. We referred to the experiment of Xu and Xu. 32 Fifteen mice were randomly selected for medial meniscus (DMM) resection to establish mice models of OA knee joint. Sham operation was performed on mice in sham group. Two weeks after DMM operation, 10 mice were randomly selected and divided into 2 groups. Mice in both groups were injected weekly with 5 µg sh-METTL3 or sh-NC (GenePharma). Six weeks after surgery, knee joint specimens were collected for analysis.
Hematoxylin-Eosin Staining
The knee joint of mice was fixed, paraffin-embedded, and sliced. Mice knee sections were stained with hematoxylin and eosin. Cartilage tissues were further stained with safranine O/sodium acetate buffer. Olympus BX51 optical microscope was used to observe and stain the slices. The Olympus DP72 digital camera (Tokyo, Japan) was used to obtain the image.
Statistical Analysis
All data were presented as the means ± standard deviation. The data statistical significance analyses and statistical mapping were performed using Prism GraphPad 8.0 software. Data were analyzed using 1-way analysis of variance (ANOVA) and Student t test. P < 0.05 was considered statistically significant.
Results
High Levels of m6A Methylation in OA Patients
In order to explore the effect of m6A methylation on OA patients, we tested the level of m6A methylation and the mRNA level expression of m6A modification enzyme in clinical samples of OA patients. As shown in
Figure 1A
, the level of m6A methylation in OA patients was higher than that in the control group. Meanwhile, the expression of m6A modification enzyme METTL3 was significantly increased in OA patients (1.00 ± 0.04 vs. 2.47 ± 0.19, P < 0.001,
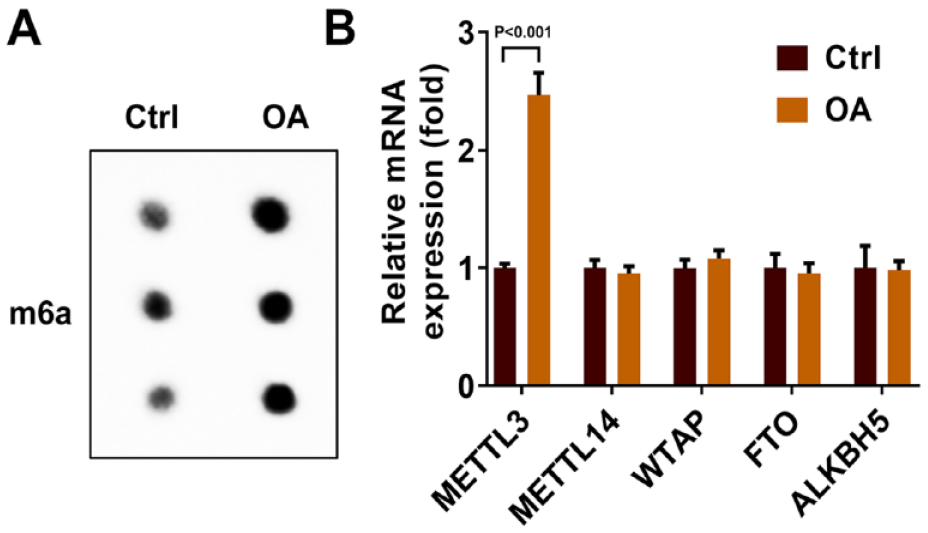
M6A levels in OA patients. (
METTL3 Knock-down Inhibited Pyroptosis in OA Cell Models
To confirm the influence of METTL3 on the progression of OA, we reduced the expression of METTL3 in chondrocytes and established the OA cell model by LPS induction. As shown in
Figure 2A
, we detected successful transfection of sh-METTL3 in chondrocytes by qRT-PCR (1.00 ± 0.07 vs 0.25 ± 0.04, P < 0.001). Then we examined the cell function. The MTT experiment showed a significant decrease in cell viability in the OA group compared with the control group (100.00 ± 7.04 vs. 27.08 ± 3.19, P < 0.001). However, this was reversed by METTL3 silencing (25.92 ± 3.58 vs. 70.31 ± 4.05, P < 0.001,
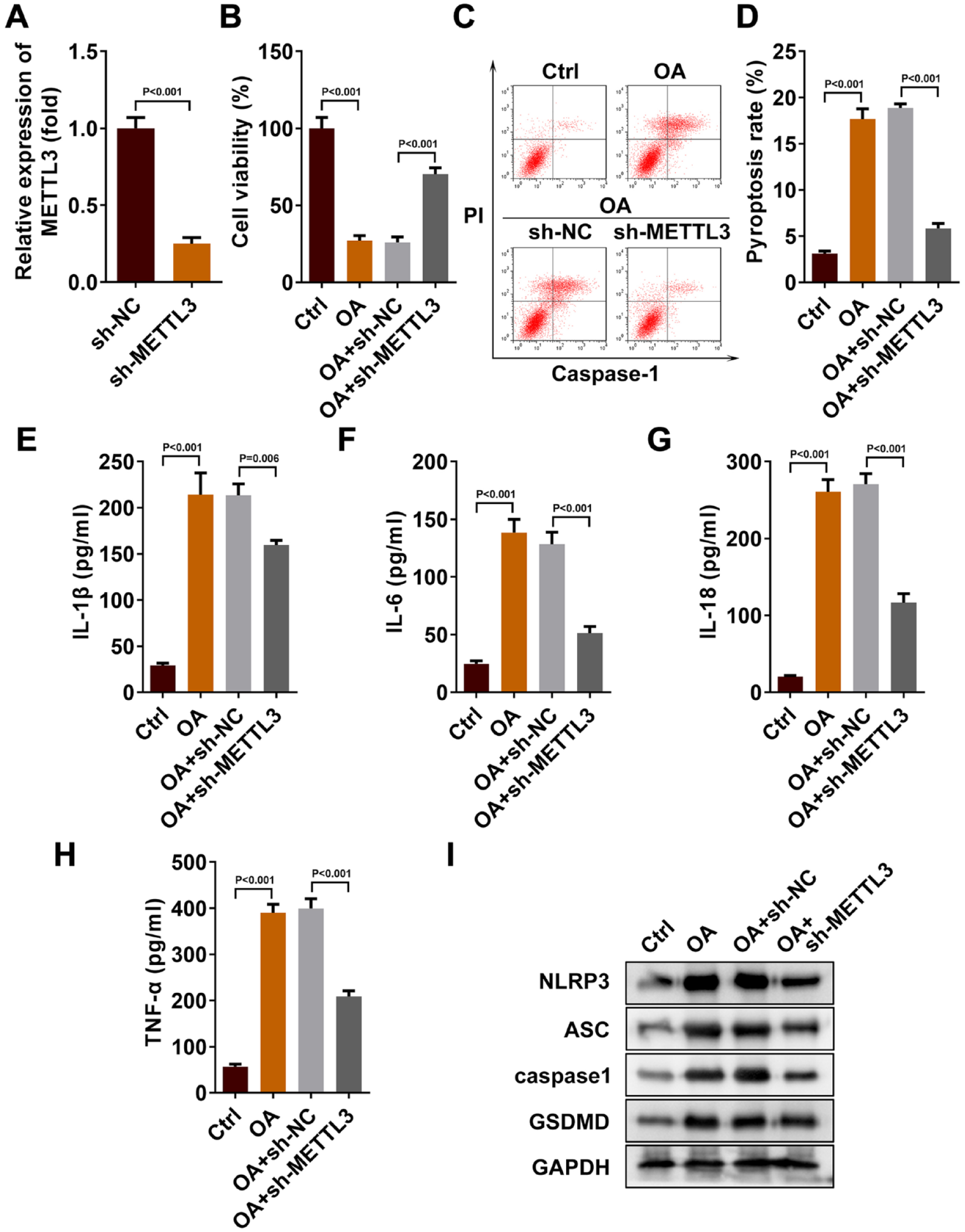
METTL3 was the key enzyme for m6A methylation modification of OA. (
METTL3 Promoted m6A Methylation of NEK7
The activation of NLRP3 inflammasome by NEK7 has been repeatedly reported in Nature.20,33 This effect in turn affected the pyroptosis processes in which NLRP3 was involved. To further elucidate the completed pathway of pyroptosis in chondrocytes during the progression of m6A methylated OA, we analyzed the changes in NEK7 when METTL3 expression decreased in chondrocytes. According to MeRIP-qPCR, we found that m6A levels in NEK7 were significantly reduced when METTL3 was knockdown (1.00 ± 0.11 vs. 0.25 ± 0.03, P < 0.001,
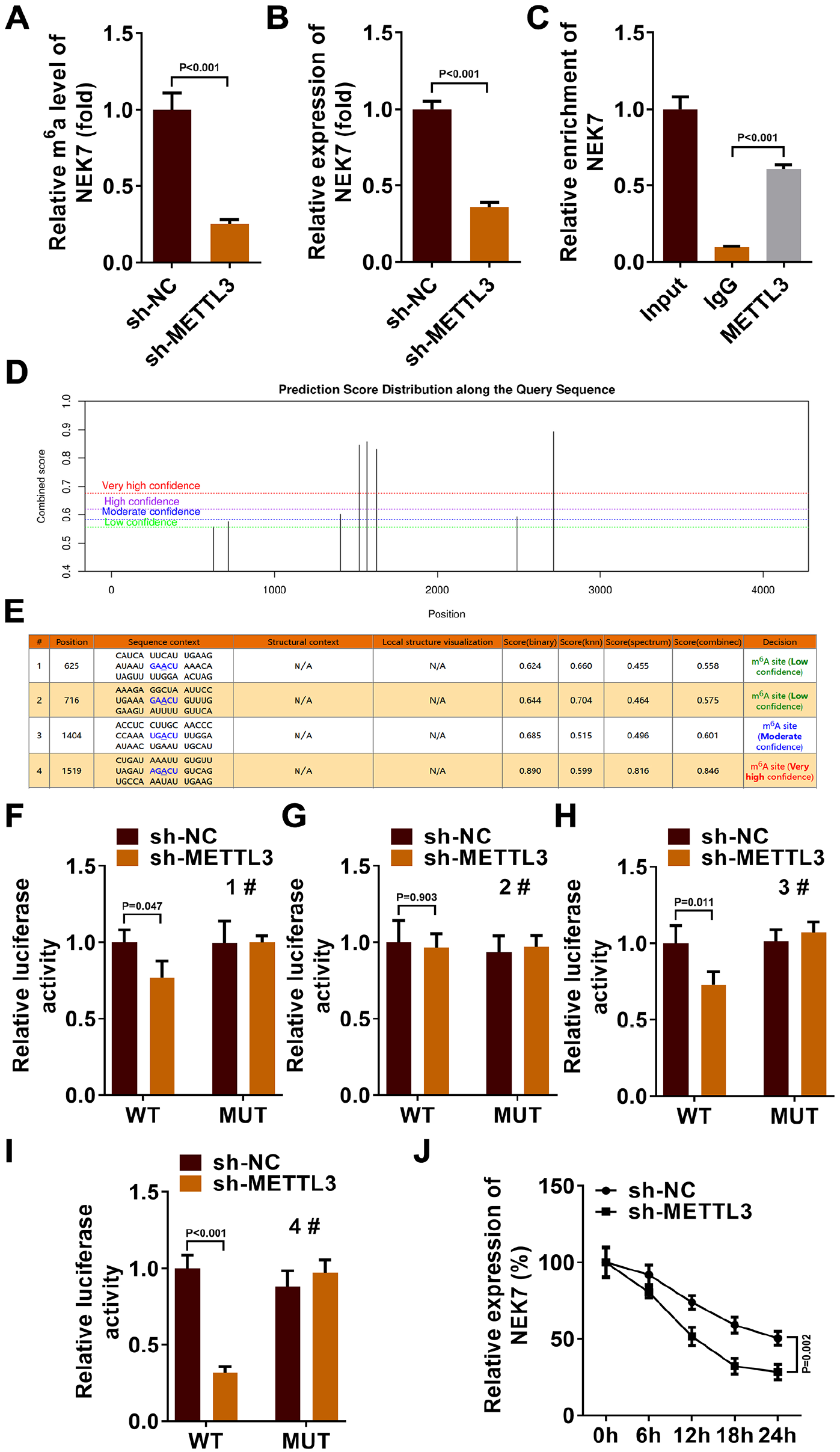
Binding effect of METTL3 and NEK7. (
In addition, when the expression level of METTL3 decreased, the stability of NEK7 also enhanced (
METTL3 Regulated the m6A Modification of NEK7 Gene to Promote OA Cell Model Pyroptosis
To further elucidate the mode of action of pyroptosis affected by METTL3/NEK7 axis, we analyzed the cases of pyroptosis after NEK7 overexpression. As shown in
Figure 4A
, we detected successful transfection of NEK7 in chondrocytes by qRT-PCR (1.00 ± 0.08 vs. 2.66 ± 0.14, P < 0.001). Then we examined the cell function. MTT assay results showed that overexpression of NEK7 inhibited the increase of OA cell viability induced by silencing METTL3 (240.40 ± 11.18 vs. 165.00 ± 7.27, P < 0.001,
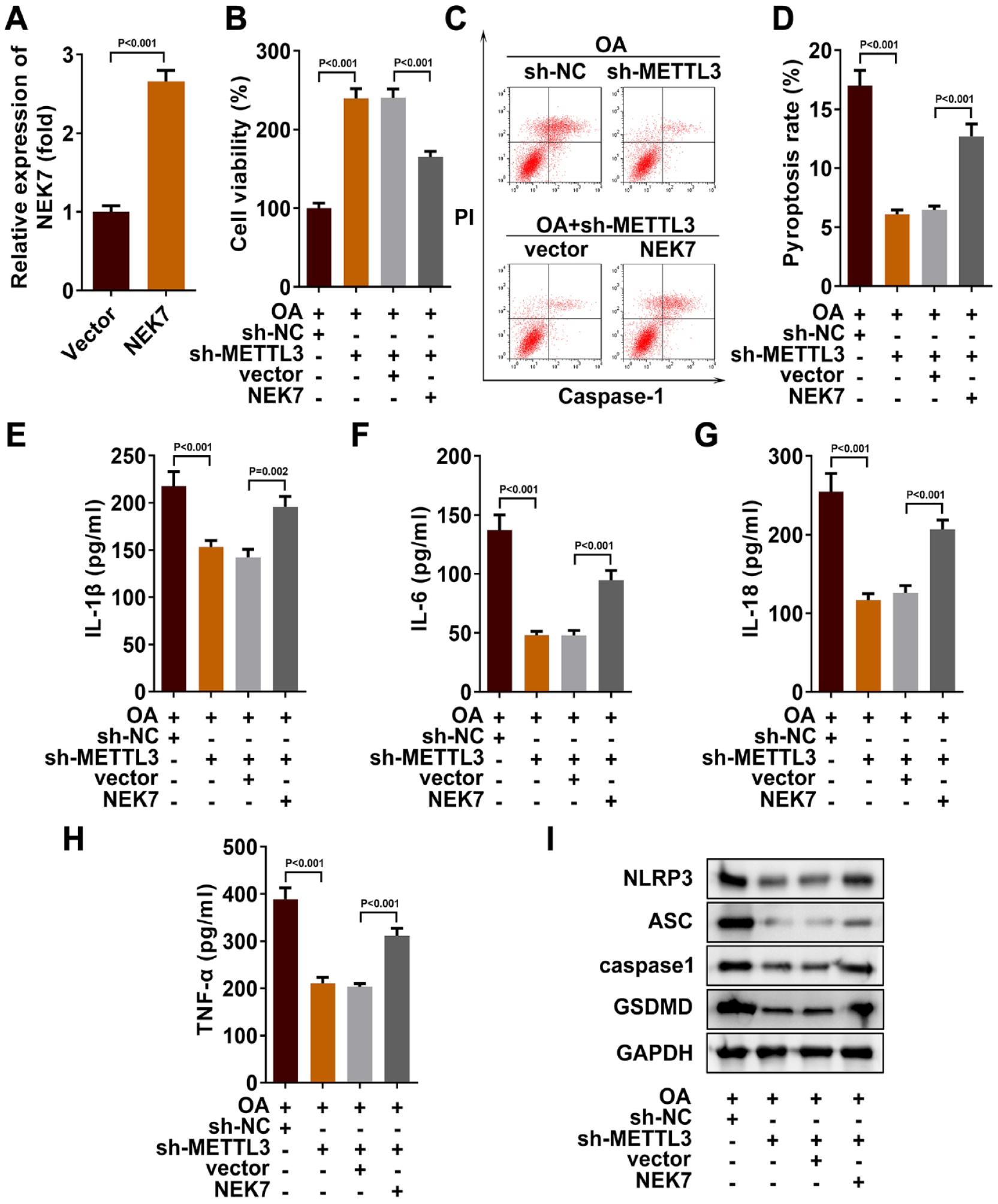
METTL3 regulated m6A modification of NEK7 gene to promote pyroptosis. (
Knockdown METTL3 Expression Inhibited the Progression of OA in Mice
To further explore the role of the METTL3/NEK7 axis in vivo, we constructed a mouse model of OA. HE staining was used to assess the progression of OA in mice. It was found that, compared with OA group and OA + sh-NC group, the synovial cells in sham group and OA+sh-METTL3 group were arranged in an orderly manner, with loose connective tissue and less inflammatory cell infiltration. This suggested that METTL3 knockdown effectively inhibited the progression of OA in mice (
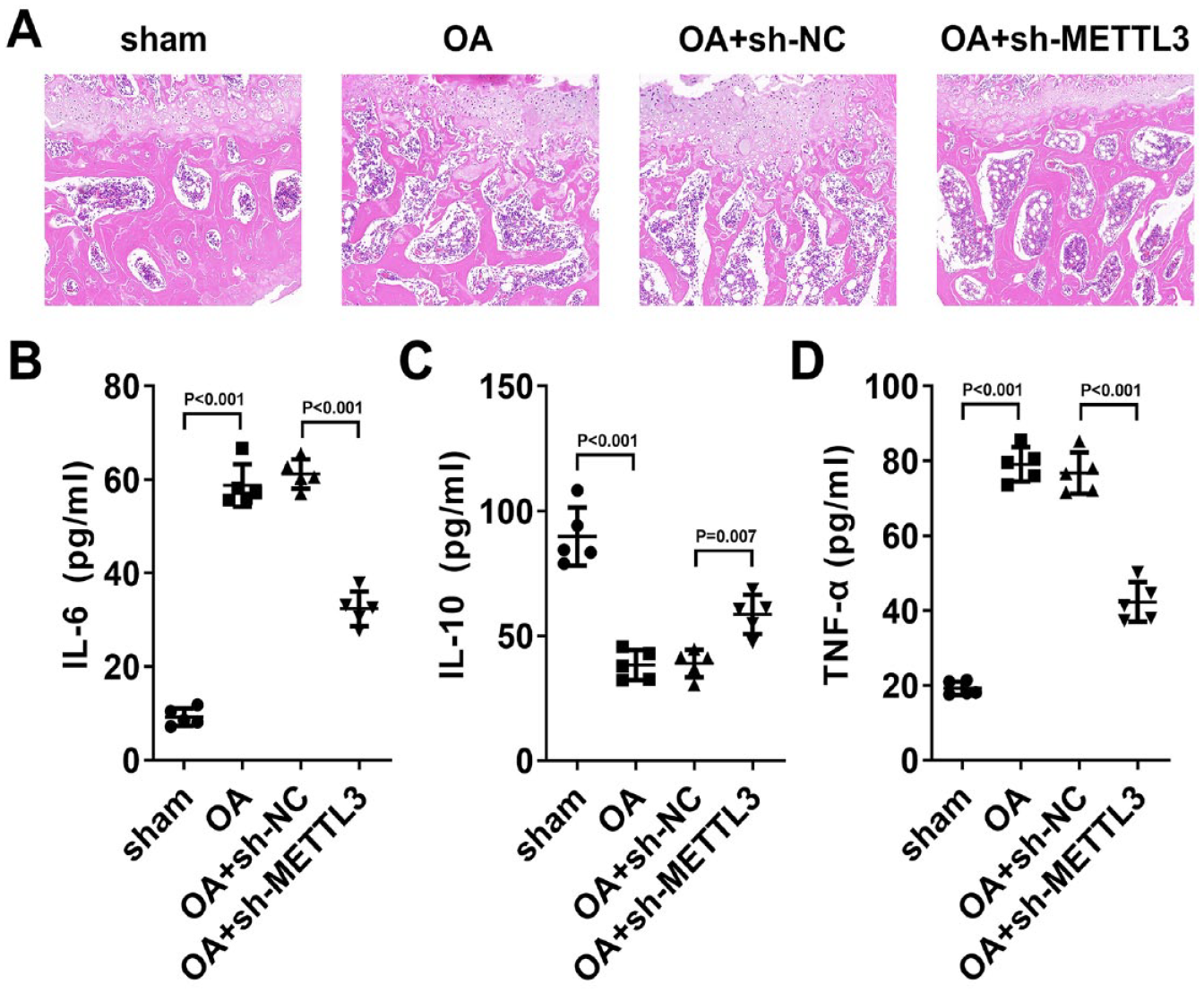
Knockdown METTL3 expression inhibited the progression of OA in mice. (
Discussions
Recent studies have shown that the vitality of chondrocytes played a key role in the progression of OA. 34 Chondrocytes pyroptosis mediated the development of OA. 35 We aimed to analyze the ways and mechanisms that regulate chondrocyte pyroptosis during the progression of OA. Our study showed that NEK7 promoted OA cell model pyroptosis. Furthermore, this process was mediated by m6A methylation of METTL3.
OA is a common degenerative joint disease. The occurrence of OA would slowly destroy the soft tissue structure of the patient’s joint and even lead to disability in severe cases.1,2,36 The incidence of OA also increases as people’s lifestyle changes. 3 There are no effective drugs for the clinical treatment of OA. However, inhibition of pyroptosis in chondrocytes could slow the progression of OA, which has attracted widespread attention.37,38 Li et al. 39 found that P2x7-induced chondrocytes pyroptosis played a key role in promoting inflammatory responses and the development of OA. Yu et al. 40 found that Corni Fructus treated OA by inhibiting the pyroptosis process of chondrocytes. M6A is one of the main ways to regulate and modify RNA after transcription. This modification process is widely involved in a variety of cellular activities, including pyroptosis. 41 Our study found that high methylation of m6A occurred in clinical tissues when OA occurred. This was consistent with the results of Ren et al. 30 At the same time, they also found that METTL3 was highly expressed in the development of OA. This aroused our interest and we further analyzed the role of m6A methylation modification enzyme METTL3. On one hand, Chen et al. 42 found that m6A methylation that inhibited METTL3 inhibited the progression of OA. On the other hand, Liu et al. 43 found that METTL3-mediated m6A methylation of NLRP3 inflammatory promoted podocyte pyroptosis in diabetic nephropathy, thereby ameliorating kidney injury. We found the same mechanism of action in OA through in vitro and in vivo experiments. METTL3 was overexpressed due to the inflammatory response of OA. Inhibition of METTL3 expression promoted cell viability in OA cell models and inhibited pyroptosis processes of chondrocytes, and attenuated OA in mice.
NEK7 is a protein kinase that regulates the cell cycle. It exists widely in all kinds of life activities. 44 Sun et al. 45 found that knockdown NEK7 inhibited the progression of OA in mice. Our study complemented this mechanism with cell function experiments. We found that overexpression of NEK7 significantly promoted chondrocyte pyroptosis and inhibited the progression of OA. In addition, the binding and activation between NEK7 and the inflammasome NLRP3 have been reported in Nature many times.20,33 This effect affects pyroptosis through the formation of inflammasome complexes through the recruitment of ASCs and shear of caspase1.21,22,46 Our study also demonstrated the effect of overexpression of NEK7 on pyroptosis-related proteins (NLRP3, ASC, caspase1, and GSDMD) and inflammatory cytokines (IL-1β, IL-18, IL-6, IL-10, and TNF-α). Interestingly, we found a complementary relationship between NEK7 and METTL3 by double luciferin reporting and rescue experiments. These results complemented the molecular regulatory mechanisms of pyroptosis promoted by NEK7. METTL3 acted as the m6A methylation modifier of NEK7. METTL3 combined with NEK7 positively regulated the m6A methylation level and expression level of NEK7.
Our study suggested that METTL3 regulated NEK7 gene m6A modification and regulation of chondrocyte pyroptosis in the development of OA. However, there were still some limitations in this study. We just analyzed the role of METTL3-mediated m6A methylation in OA progression. It is still unclear which “reader” played a recognition role in OA progression. In the future, we will conduct more in-depth research to improve the molecular mechanism of m6A modification in OA.
Conclusion
In conclusion, our work revealed for the first time the mechanism by which METTL3 regulated the m6A modification of NEK7 gene to inhibit the progression of OA, which might provide a new understanding of the roles of m6A and NEK7 in OA.
Supplemental Material
sj-docx-1-car-10.1177_19476035231200336 – Supplemental material for METTL3 Regulates the m6A Modification of NEK7 to Inhibit the Formation of Osteoarthritis
Supplemental material, sj-docx-1-car-10.1177_19476035231200336 for METTL3 Regulates the m6A Modification of NEK7 to Inhibit the Formation of Osteoarthritis by Xiaochuan Xiong, Hao Xiong, Jun Peng, Yingjie Liu and Yang Zong in CARTILAGE
Footnotes
Author Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by J.P., Y.L., and Y.Z. The first draft of the manuscript was written by X.X. and X.H. and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Acknowledgments and Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of Shanghai Jiao Tong University–affiliated Sixth People’s Hospital.
Consent to Participate
Informed consent was obtained from all individual participants included in the study.
Consent to Publish
The authors affirm that human research participants provided informed consent for publication of the images in Figures.
Data Availability
The data sets used and/or analyzed during this study are available from the corresponding author on reasonable request.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
