Abstract
Objective
Osteoarthritis (OA) is characterized by the chronic and progressive deterioration of articular cartilage. Chondrocyte senescence could lead to a shift in the balance between extracellular matrix (ECM) component synthesis and degradation. Small noncoding RNAs (sncRNAs), including microRNAs (miRNAs), P-element-induced wimpy testis-(PIWI-) interacting RNAs (piRNAs), small nucleolar RNAs (snoRNAs), small nuclear RNAs (snRNAs), and repeat-associated siRNAs (rasiRNAs), are a class of important epigenetic molecules. We aimed to gain insights into the changes and roles of sncRNA in chondrocyte senescence.
Design
Healthy mouse postnatal chondrocytes were isolated, and a replicative aging model was constructed. We used small RNA sequencing (small RNA-seq) to generate extensive small RNA data. We identified differentially expressed sncRNAs and performed tissue-specific analysis using real-time quantitative polymerase chain reaction (qRT-PCR). β-galactosidase staining was used to detect chondrocyte senescence. The results showed that the expression profiles of sncRNA in passage 5 chondrocytes were significantly different from those in passage 0 chondrocytes. The expression of sncRNA was tissue specific. We found that 40 miRNAs were upregulated and 70 miRNAs were downregulated during chondrocyte senescence, and that miR-132-5p expression inhibition prevented chondrocyte senescence. We found that 8 piRNAs were upregulated and 17 piRNAs were downregulated during chondrocyte senescence, and that piRNA piR_025576 overexpression delayed chondrocyte senescence. We found that 24 snoRNAs were upregulated and 28 snoRNAs were downregulated during chondrocyte senescence, and that snoRNA ENSMUSG00000087935 overexpression delayed chondrocyte senescence. We found that 5 snRNAs were upregulated and 6 snRNAs were downregulated during chondrocyte senescence, and that snRNA ENSMUSG00000064682 overexpression delayed chondrocyte senescence. We found that 1 rasiRNA was upregulated and 4 rasiRNAs were downregulated during chondrocyte senescence.
Conclusions
These findings might provide novel insights into OA pathogenesis and contribute to the development of candidates for targeted therapeutics in OA.
Introduction
Osteoarthritis (OA) is a prevalent multiple factor-associated disorder that is characterized by chronic and progressive deterioration of articular cartilage. Several risk factors for the development of OA exist, including prior joint injury, obesity, diabetes mellitus, hyperglycemia, and anatomical factors related to joint shape and alignment; however, the most prevalent risk factor is increasing age.1-3 Cellular senescence, one of the hallmarks of aging, is a form of irreversible cellular arrest characterized by senescence-associated beta-galactosidase (SA-β-Gal) activity and the release of harmful proinflammatory molecules, growth factors, and matrix metalloproteinases as part of the senescence-associated secretory phenotype (SASP). 4 As the accumulation of senescent cells reduces cellular proliferation and disordered tissue function and metabolism, senescence has been implicated in the pathogenesis and/or progression of a variety of aging-associated diseases, including OA.3,5
Although OA has recently been regarded as a whole joint disease rather than merely dysfunctional cartilage, chondrocytes are primarily known to have a major role in the pathology of OA.5-8 Chondrocytes, the unique resident cell type in articular cartilage, regulate anabolic and catabolic pathways to maintain cartilage homeostasis. The senescence of chondrocytes is expected to lead to a shift in the balance between extracellular matrix (ECM) component synthesis and degradation through metalloproteinase components (i.e., MMP1, MMP2, MMP3, MMP10, MMP13, and more) of the SASP response.5,9 There was a causal relationship between the occurrence and development of OA, which was significantly correlated with age and chondrocyte aging, but the more specific pathological mechanism has not been fully clarified. Studies have also confirmed that the expression of SA-β-Gal in articular cartilage is positively correlated with the severity of OA. 10 The increase in the positive rate of aging chondrocytes reduced the repair ability of cartilage, aggravated cartilage degeneration, and led to an increase in the incidence rate of OA. Understanding the molecular mechanism of chondrocyte aging is of great significance for the treatment of OA and the development of new therapies.
The exact mechanisms through which senescence can affect cartilage health remain elusive, although it is believed to be caused by multiple molecular mechanisms rather than a single etiology. Recent studies have shown that small noncoding RNAs (sncRNAs), a class of important epigenetic regulatory molecules, play an important role in regulating cellular senescence in OA.11-14 SncRNAs are short RNA species, typically fewer than 200 nucleotides in length, that are not translated into proteins but have other structural or regulatory biological roles. 15 These include microRNAs (miRNAs), small nuclear RNAs (snRNAs), small nucleolar RNAs (snoRNAs), P-element-induced wimpy testis (PIWI-)-interacting RNAs (piRNAs), small interfering RNAs (siRNAs), transfer RNAs (tRNAs), and repeat-associated siRNAs (rasiRNAs). SncRNAs are promising candidates for targeted therapeutics in OA due to their small size, and their activity can be modulated via small molecules and biological delivery systems.15,16
In the present study, primary mouse chondrocytes were isolated and subjected to 5 additional passages to simulate chondrocyte replicative senescence in vitro. To gain insights into sncRNA changes in chondrocyte senescence, we used small RNA sequencing (small RNA-seq) to generate extensive small RNA data. We identified differentially expressed sncRNAs and performed tissue-specific analysis. These findings might provide novel insight into OA pathogenesis and contribute to the development of candidates for targeted therapeutics in OA.
Materials and Methods
Cell Isolation and Culture
Mouse primary articular chondrocytes were isolated from the femoral heads, femoral condyles, and tibial plateaus of mice at postnatal day 5 as previously described. 17 The animal study was reviewed and approved by the Ethics Committee of Xinhua Hospital affiliated with Shanghai Jiao Tong University School of Medicine. Cartilage was minced into small pieces. After digestion with 0.25% trypsin and collagenase D solution at 0.5 mg/ml, chondrocytes were collected and cultured in Dulbecco’s modified Eagle medium DMEM (1 g glucose per ml; Sigma, cat. no. D5546) supplemented with 10% fetal bovine serum (FBS) (Gibco, Thermo Fisher Scientific, USA) and 1% penicillin-streptomycin. Chondrocytes at passages 0 to 5 were used in all experiments.
SA-β-Gal Staining
The Senescence Detection Kit (BioVision, San Francisco, CA, USA) was used to stain SA-β-Gal. Chondrocytes at passages 1 and 5 were rinsed in phosphate-buffered saline (PBS). After fixation with fixative solution at room temperature for 15 minutes, the cells were washed twice with PBS. Next, the cells were added to a staining solution mixture and incubated overnight at 37°C. Chondrocytes were then washed twice with PBS, and the senescent cells stained blue were analyzed under a light microscope.
qRT-PCR Analysis
Total RNA was extracted from cells using an RNeasy Mini kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions, and cDNA was synthesized from total RNA (1 µg) using reverse transcriptase (TaKaRa Biotechnology, Otsu, Japan). The complementary DNA (cDNA; 10 ng) products were used as the template for amplification using SYBR Green PCR Master Mix (TaKaRa Biotechnology). Furthermore, mRNA expression was detected by an ABI 7500 sequencing detection system (Applied Biosystems, Foster City, CA, USA). The primer lists are shown in Supplementary Material S1.
Small RNA-Seq
The original image data file obtained by high-throughput sequencing was transformed into the original sequenced reads by base calling analysis. The results are stored in FASTQ (FQ) file format, which contains the sequence information of sequenced reads and their corresponding sequencing quality information. First, the original data were preprocessed, joint sequences were removed, low-quality sequences were filtered, and then the FASTQ format was converted to a FASTA format. After data preprocessing, we used Bowtie software to compare the sequences to the reference genome, and only the sequences aligned to the reference genome were selected for subsequent analysis. To comprehensively predict noncoding RNAs of different lengths, we first extracted the sequencing sequences aligned to the reference genome in each sample and then clustered the sequences of all samples using cd-hit to remove the same or similar sequences. Then, we used infernal software to search the RFAM database. After searching, we matched the small RNA sequence in the RFAM database and then compared the sequencing data of each sample to count the read count of small RNA of each sample. The read count was additionally subjected to TPM standardization to calculate the proportion of each million reads in the comparison. For the experiments with biological repetition, we used DESeq for the analysis. For experiments without biological duplication, we used edger for the analysis. Finally, the genes with Padj less than 0.05 and a difference multiple greater than or equal to 2 were selected as small RNAs with significant differential expression.
Statistical Analysis
Statistical analysis was performed using IBM SPSS Statistics 24, and graphs were generated by GraphPad Prism 9. Independent sample t tests were used to analyze the mRNA expression results from the qPCR assays. P values less than 0.05 were considered significant.
Results
Elevation of Cell Senescence with Increasing Passages
First, the morphological changes in the chondrocytes were recorded. Mouse cartilage chondrocytes were isolated and continuously subcultured to passage 5. At passage 1, chondrocytes had a characteristic tetris-shaped, smooth cell surface, and grew regularly. Unlike the cells at passage 1, almost all chondrocytes at passage 5 exhibited irregular shapes with pseudopods and became larger and flatter (
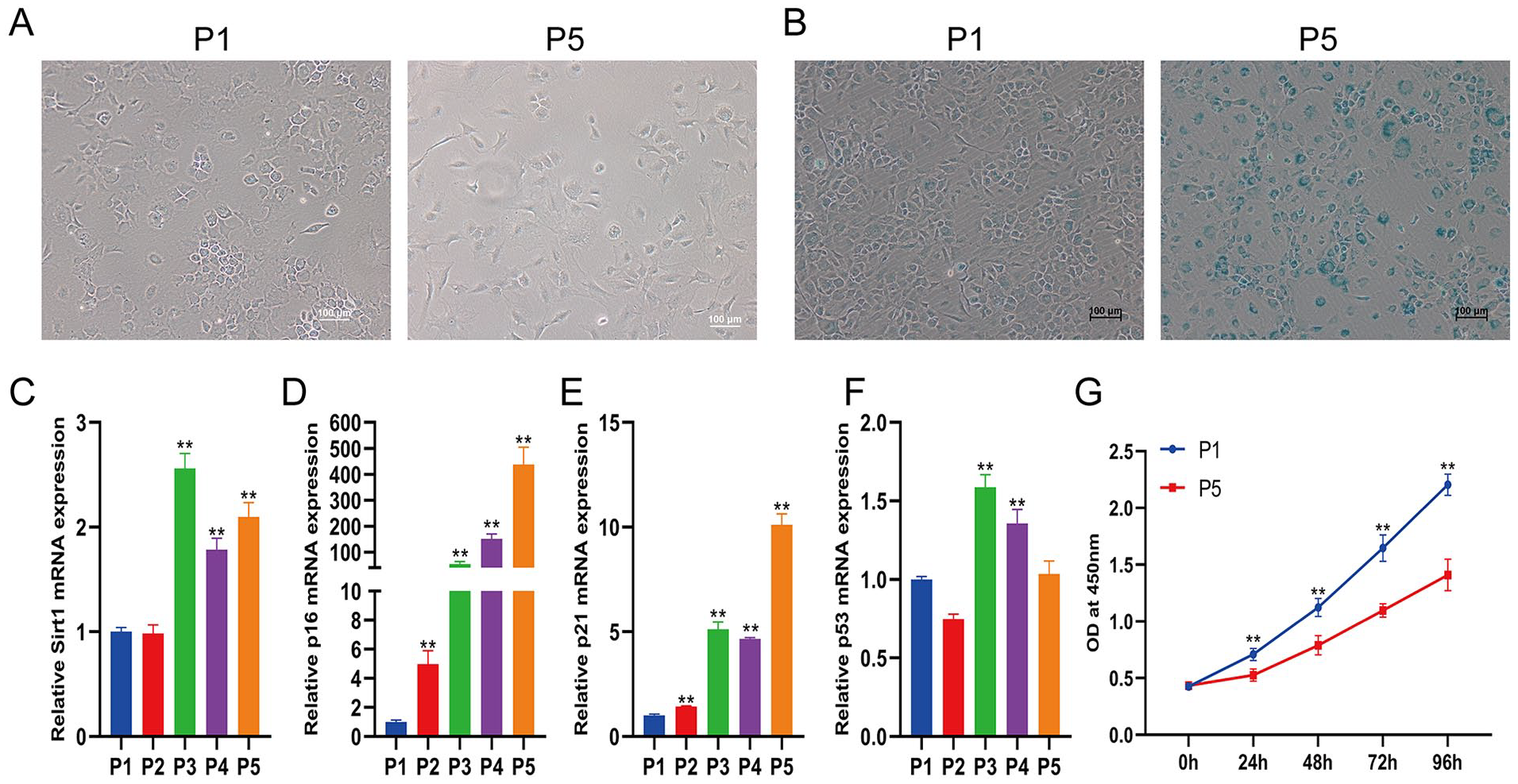
Elevation of cell senescence with increasing passages. (
Identification of Differentially Expressed miRNAs and Tissue-Specific Analysis
As important epigenetic regulators, miRNAs bind to specific regions of target mRNAs, resulting in reduced protein expression and thus participating in many cellular processes. We performed small RNA-seq to analyze the miRNA expression profiling change in replicative senescence chondrocytes. The small RNA-seq results showed that 40 miRNAs were upregulated and 70 miRNAs were downregulated in chondrocytes at passage 5 compared with those at passage 0 (
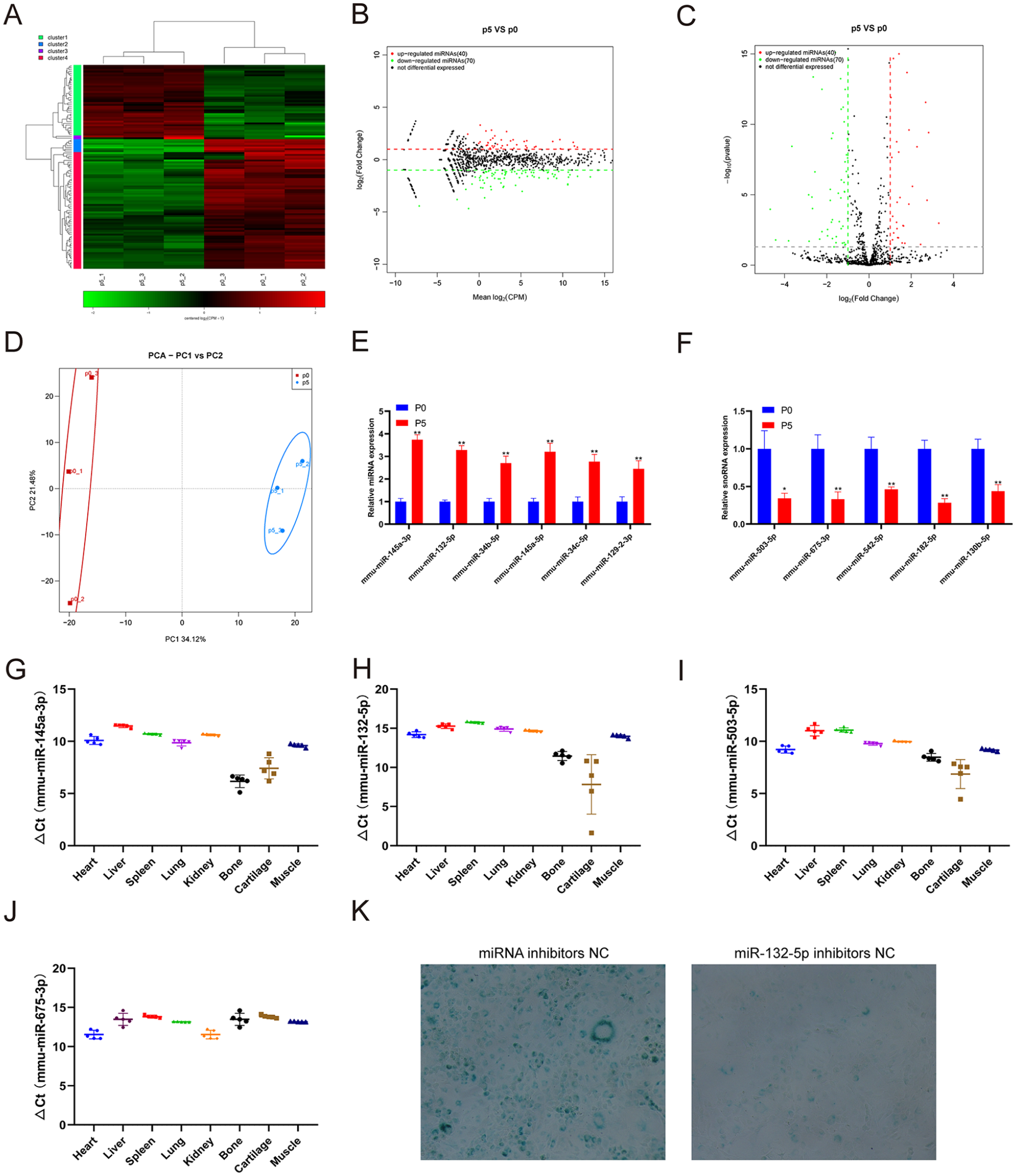
Identification of differentially expressed miRNAs and tissue-specific analysis. (
Identification of Differentially Expressed piRNAs and Tissue-Specific Analysis
Compared with miRNAs, piRNAs are a relatively new type of sncRNA and have attracted increasing attention in recent years. piRNAs, which have a length of 24-31 nucleotides, play regulatory roles, such as silencing the transcriptional gene process, regulating mRNA stability, interacting with multiple proteins, and maintaining stem cell functions.
18
We performed small RNA-seq to analyze piRNA expression profiling changes in chondrocytes undergoing replicative senescence. The small RNA-seq results showed that 8 piRNAs were upregulated and 17 piRNAs were downregulated in chondrocytes at passage 5 compared with those at passage 0 (
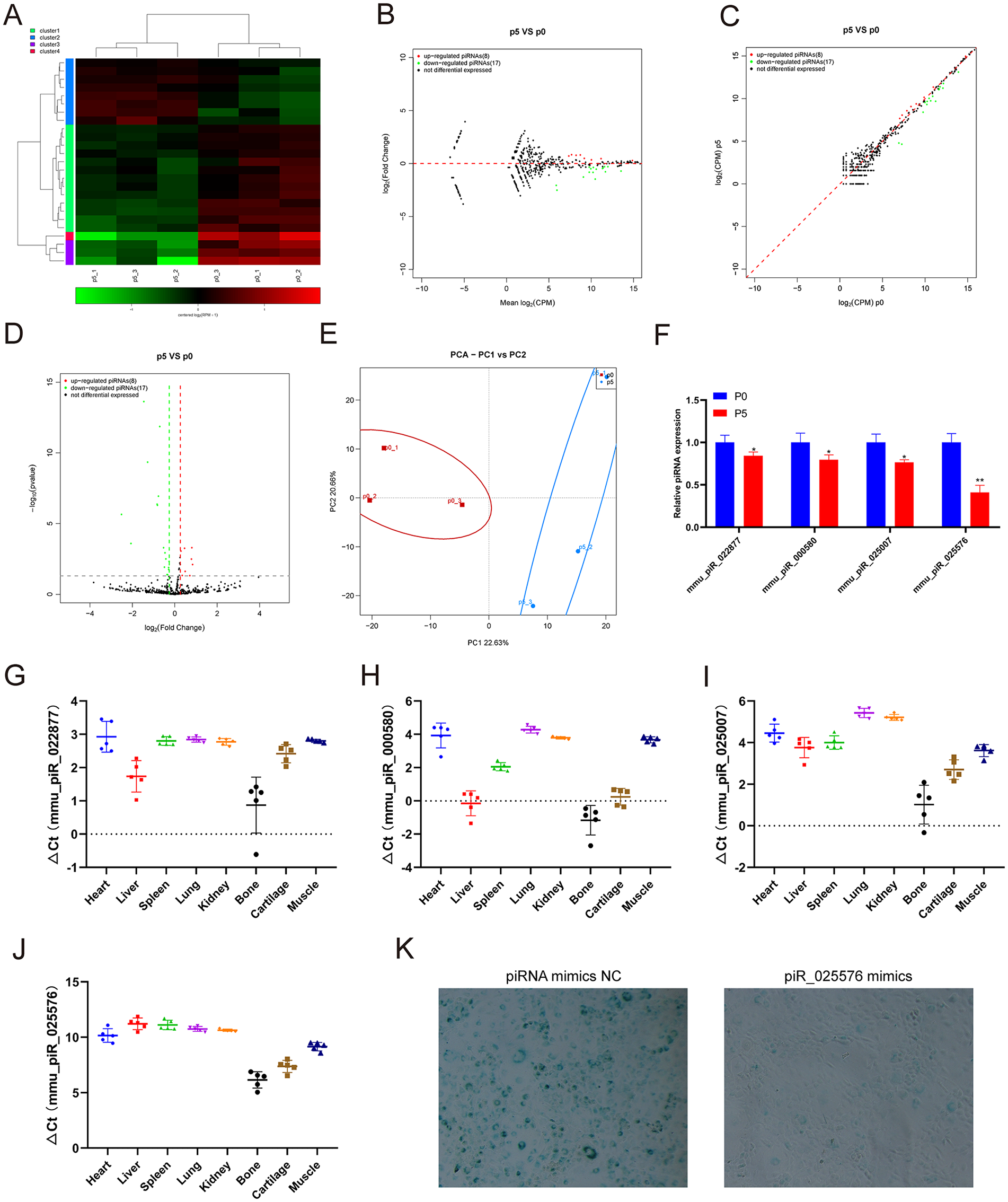
Identification of differentially expressed piRNAs and tissue-specific analysis. (
Identification of Differentially Expressed snoRNAs and Tissue-Specific Analysis
SnoRNAs direct chemical modification of other RNA substrates to fine-tune spliceosomal splicing and are mainly involved in endoribonucleolytic pre-rRNA processing.
19
The role of snoRNAs in cartilage aging has attracted increasing attention.
12
We performed small RNA-seq to analyze the snoRNA expression profiling change in replicative senescent chondrocytes. The small RNA-seq results showed that 24 snoRNAs were upregulated and 28 snoRNAs were downregulated in chondrocytes at passage 5 compared with those at passage 0 (
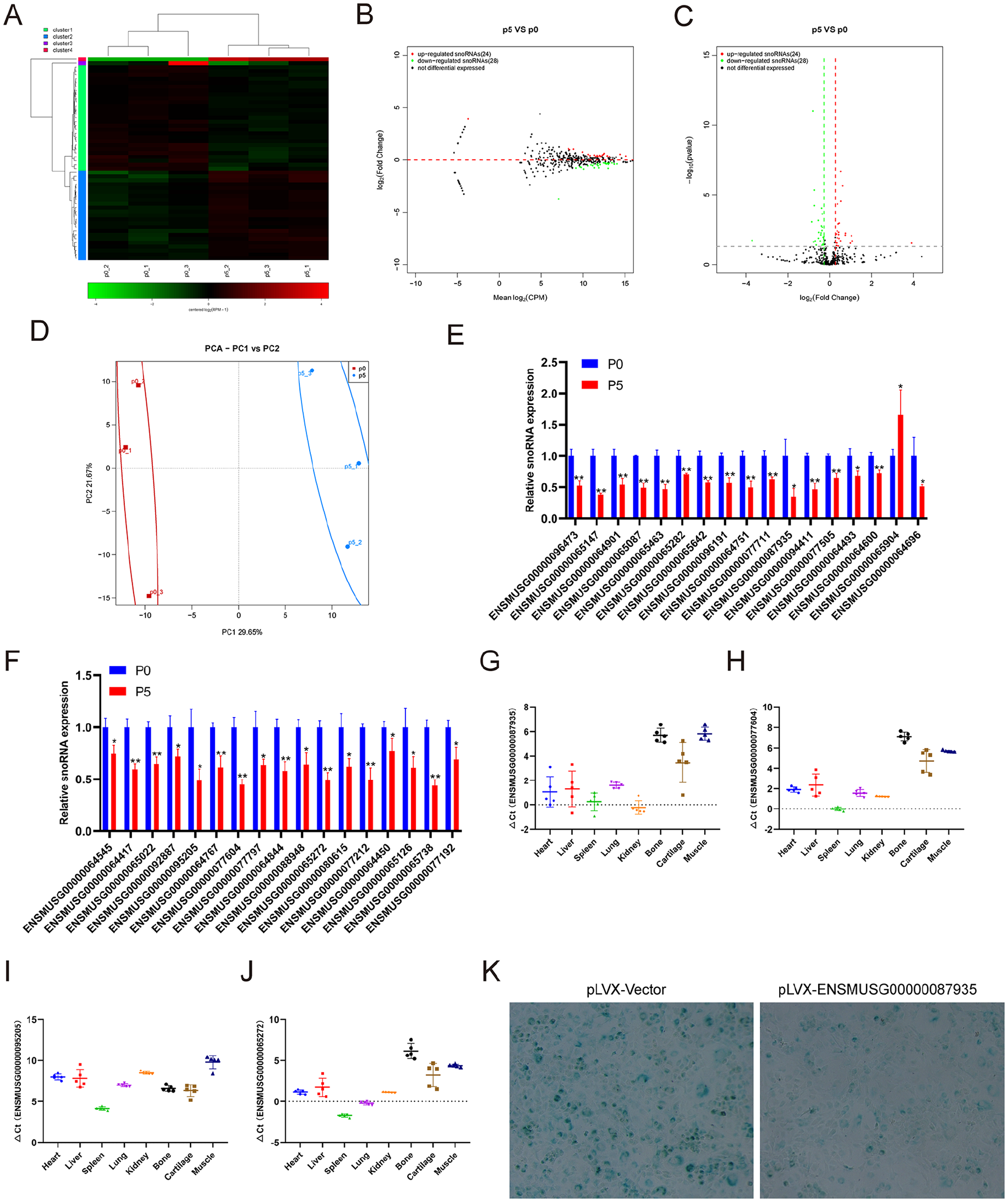
Identification of differentially expressed snoRNAs and tissue-specific analysis. (
Identification of Differentially Expressed snRNAs and Tissue-Specific Analysis
SnRNA is the main component of RNA spliceosomes in the posttranscriptional processing of eukaryotes, with a length of 100-215 nucleotides. At present, there is no literature on snRNAs directly related to chondrocytes or cartilage. We performed small RNA-seq to analyze the snRNA expression profiling change in chondrocytes undergoing replicative senescence. The small RNA-seq results showed that 5 snRNAs were upregulated and 6 snRNAs were downregulated in chondrocytes at passage 5 compared with those at passage 0 (
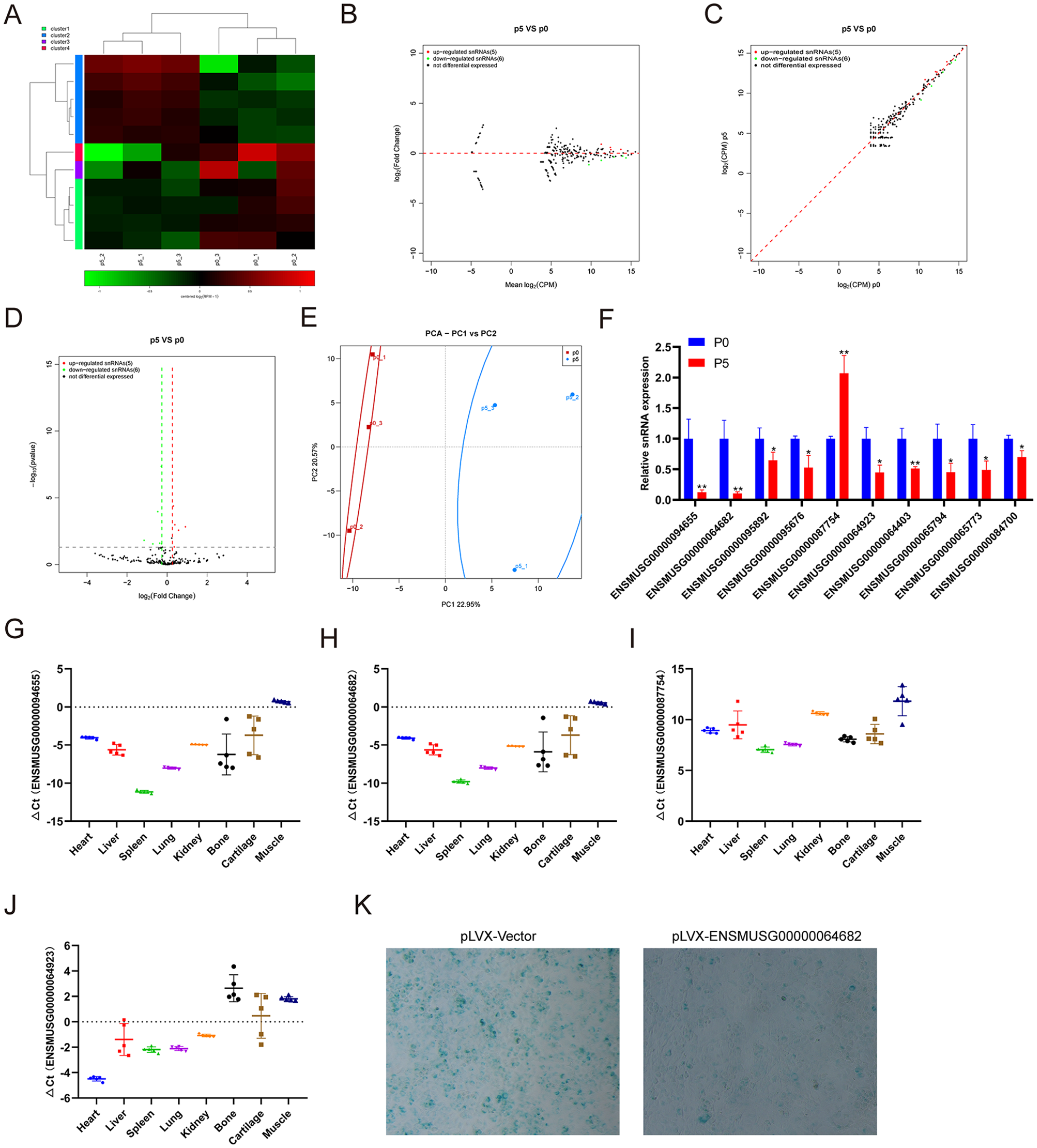
Identification of differentially expressed snRNAs and tissue-specific analysis. (
Identification of Differentially Expressed rasiRNAs
RasiRNAs of 24-28 nucleotides in length may assume a role in sperm confrontation and consolidation.
20
Unfortunately, there are no studies in the literature focusing on repeat RNAs that are directly related to chondrocytes or cartilage. We performed small RNA-seq to analyze the repeat RNA expression profiling change in replicative senescent chondrocytes. The small RNA-seq results showed that only one repeat RNA was upregulated, and 4 repeat RNAs were downregulated in chondrocytes at passage 5 compared with passage 0 (
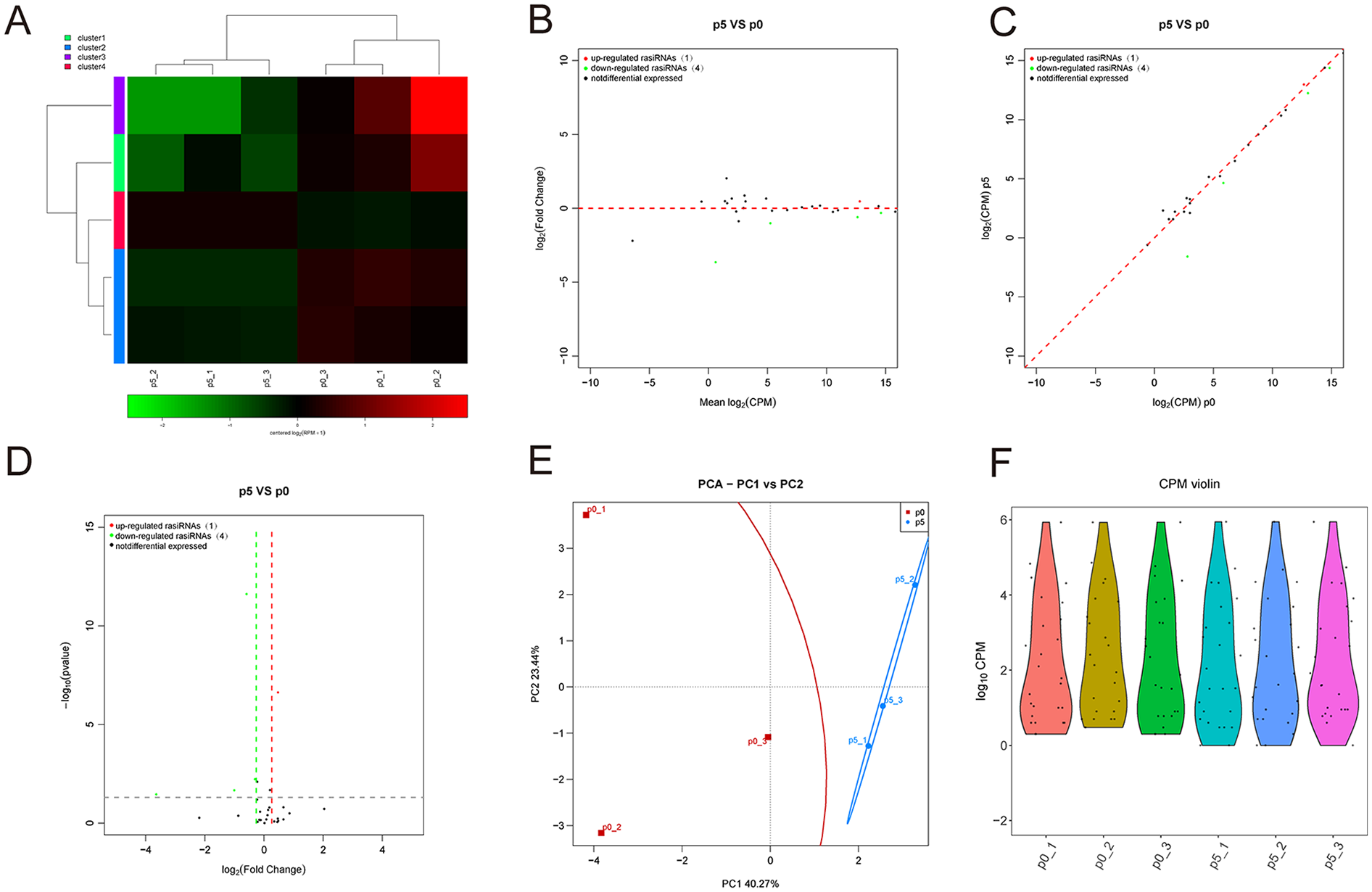
Identification of differentially expressed rasiRNAs and tissue-specific analysis. (
Discussion
Cellular senescence is a driver of various aging-associated disorders, including OA. OA chondrocytes exhibit features that are similar to senescent cells, such as specific senescent markers, telomere shortening, and increased SA-β Gal activity.21,22 Exploring anti-cellular senescence strategies may be a promising approach to prevent or cure OA. The exact mechanism of chondrocyte senescence remains elusive, although two different mechanisms of senescence have been suggested in chondrocytes, including stress-induced premature senescence (SIPS) and replicative senescence.23,24 It was concluded that replicative senescence contributes to either the development or the progression of OA. 21
SncRNAs, a large class of regulatory molecules, are involved in organism development and coordination of biological processes, including metabolism, maintaining genome integrity, and immune and stress responses. 25 Recent studies have suggested that sncRNAs play an important role in regulating cellular senescence in OA.11-14 The aim of this study was to systematically investigate the sncRNA profile changes in senescent chondrocytes after long-term expansion in vitro. We found that mouse chondrocytes at passage 5 showed obvious senescence characteristics, including increased SA-β Gal activity and senescence-related gene expression. Moreover, the expression profiles of miRNAs, snRNAs, snoRNAs, piRNAs, and rasiRNAs in passage 5 chondrocytes were significantly different from those in passage 0 chondrocytes.
MiRNAs, the most studied sncRNAs, exert their negative regulatory functions by partially or fully binding complementary sequences in the 3ʹ untranslated region of their messenger RNA (mRNA) targets, thus inhibiting mRNA translation. Our results showed that 6 miRNAs were upregulated and 5 miRNAs were downregulated (
PiRNAs are a type of small RNA with a length of approximately 24-13 nucleotides. The precursors of piRNAs are derived from tandem repeat sequences called piRNA clusters and form a mature piRNA/PIWI complex via two route-dependent or independent “Ping-Pong” amplification pathways.
18
In the past, piRNAs were thought to exist only in the reproductive system to regulate the growth and development of germ cells. Recently, it was reported that piRNAs are also expressed in several other human tissues with tissue specificity.
13
PiRNAs play roles in transcriptional or posttranscriptional gene silencing pathways by combining Piwi subfamily proteins to form piRNA complexes. At present, piRNA research mainly focuses on its regulation of the occurrence and development of cancer, including colorectal cancer, breast cancer and lung cancer.
13
To our knowledge, there are no reports on the relationship between piRNAs and OA or cartilage, and the role of piRNAs in chondrocytes is unknown. Our results showed that the expression of piR_022877, piR_000580, piR_025007, and piR_025576 was decreased in chondrocytes at passage 5 compared with passage 0, while no piRNA was verified to upregulate piR_022877, piR_000580, piR_025007, and piR_025576 (
SnoRNAs, a class of evolutionarily conserved noncoding small guide RNAs with lengths of 60-300 nucleotides, are extensively studied noncoding RNAs that primarily accumulate in the nucleoli.
42
SnoRNAs are involved in the direct chemical modification of other RNA substrates and the regulation of alternative splicing and posttranscriptional modification of snRNA, tRNA, and mRNA, while others exhibit miR-like activity.
43
Recently, snoRNAs have been increasingly studied in aging and OA. A previous mouse study demonstrated alterations in the snoRNA profile of young compared with old and OA compared with healthy controls in joints and serum, highlighting the potential of snoRNAs to be used as novel markers for OA and aging.
44
In human cartilage, snoRNAs are differentially expressed due to aging (including SNORD96A and SNORD44) and OA (including SNORD26 and SNORD116).
12
Altering SNORD26 or SNORD96A expression resulted in changes in chondrogenic-, hypertrophic-, rRNA- and OA-related gene expression.
12
U3, one of the most abundant snoRNAs playing a key role in ribosome biogenesis, was found to have significantly lower expression in human OA cartilage and chondrocytes. The protein translational capacity of chondrocyte cultures with diminished U3 snoRNA expression was significantly reduced, while ectopic induction of U3 snoRNA expression resulted in increased translational capacity.
45
In the present study, we detected 17 downregulated snoRNAs in chondrocytes at passage 5 compared with passage 0 (
SnRNA is the main component of the RNA spliceosome in the posttranscriptional processing of eukaryotes and participates in splicing pre-mRNA and noncoding transcripts in cells. In humans, its length is approximately 60-330 nucleotides, and it can be divided into 14 categories. U1-U7 are rich in uridylate. U3 snRNA is related to the maturation of 28S rRNA in the nucleolus, while U1 is related to the splicing and processing of precursor mRNA in the nucleus. Rn7SK is a highly conserved snRNA transcribed by RNA polymerase III and is a regulator of Pol II activity through inhibiting the function of positive transcriptional elongation factor b. A recent study demonstrated that Rn7SK participates in the regulation of cellular senescence, with the transient knockdown of Rn7SK in mesenchymal stem cells (MSCs) leading to delayed senescence, while its overexpression shows the opposite effects.
46
However, the role of snRNA in chondrocytes and its relationship with OA have not yet been reported. We found that the expression of snRNA ENSMUSG00000087754 increased and the other 9 snRNAs decreased at passage 5 (
At present, the treatment of OA is still a clinical challenge, and the understanding of the molecular mechanism is still unclear. SncRNAs are promising candidates for targeted therapeutics in OA due to their small size, and their activity can be modulated via small molecules and biological delivery systems.15,16 However, the study of sncRNAs in the aging process of chondrocytes is not sufficient. Anti-senescence therapy is a frequently discussed clinical issue, and research into this topic involves new drug research, drug-target protein detection, and disease mechanism exploration. Through screening, we will be able to verify the target genes of the above differentially expressed sncRNAs in subsequent in vivo and in vitro studies. The sequencing results showed that rasiRNA was differentially expressed in senescent chondrocytes. However, due to the scarcity of research on rasiRNA and its short length, it is difficult to verify the sequencing results by conventional real-time PCR. More studies are needed to explore the role of rasiRNA in chondrocyte aging.
Conclusion
In this study, we detected the expression changes in miRNAs, piRNAs, snoRNAs, snRNAs and repeat RNAs during chondrocyte aging by high-throughput small RNA sequencing and detected the expression of miRNAs, piRNAs, snoRNAs, and snRNAs during chondrocyte aging by qRT-PCR. We found that the expression of miRNA, piRNA, snoRNA, snRNA, and repeat RNA changes significantly during chondrocyte aging and is tissue specific.
Supplemental Material
sj-xlsx-1-car-10.1177_19476035221118165 – Supplemental material for Changes in Small Noncoding RNA Expression during Chondrocyte Senescence
Supplemental material, sj-xlsx-1-car-10.1177_19476035221118165 for Changes in Small Noncoding RNA Expression during Chondrocyte Senescence by Fei Xiao, Chenglong Wang, Jianping Peng, Xing Zhou, Ding Ma, Yu Wang, Yanpeng Li, Xiaodong Chen and Chuandong Wang in CARTILAGE
Footnotes
Author Contributions
Conceptualization: Xiaodong Chen and Chuandong Wang; methodology: Fei Xiao, Chenglong Wang and Jianping Peng; software: Yanpeng Li, Xing Zhou, Ding Ma and Yu Wang; data curation: Chuandong Wang and Fei Xiao; writing, review, and editing: Fei Xiao and Chao Shen, Junfeng Zhu, Chuandong Wang.
Acknowledgments and Funding
This work is supported by Shanghai Sailing Program (20YF1429100), National Natural Science Foundation of China (82172473, 81802191, and 82072462), Natural Science Foundation of Shanghai (19ZR1433100), Interdisciplinary of Medicine and Engineering Foundation of Shanghai Jiao Tong University (YG2019ZDA22), and Biomedical Technology Support Program of Shanghai (20S31900200). Natural Science Foundation of Shandong Province (ZR2019PH068)
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This study was conducted under the standard approved by the Xinhua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
