Abstract
Objectives
Osteoarthritis (OA) is the most common joint disease. The occurrence and progression of OA are regulated by epigenetics. A large number of studies have shown the important regulatory role of noncoding RNAs in joint diseases. As the largest class of noncoding small RNAs, the importance of piRNAs in many diseases, especially cancer, has been increasingly recognized. However, few studies have explored the role of piRNAs in OA. Our study showed that hsa_piR_019914 decreased significantly in OA. This study aimed to demonstrate the role of hsa_piR_019914 as a potential biological target of OA in chondrocytes.
Design
The GEO database and bioinformatics analysis were used for a series of screenings, and the OA model using human articular chondrocytes (C28/I2 cells), SW1353 cells under inflammatory factor stimulation was used to determine that hsa_piR_019914 was significantly downregulated in OA. Overexpression or inhibition of hsa_piR_019914 in C28/I2 cells was achieved by transfecting mimics or inhibitors. The effect of hsa_piR_019914 on the biological function of chondrocytes was verified by qPCR, flow cytometry, and colony formation assays in vitro. The target gene of hsa_piR_019914, lactate dehydrogenase A (LDHA), was screened by small RNA sequencing and quantitative polymerase chain reaction (qPCR), LDHA was knocked out in C28/I2 cells by the transfection of siRNA LDHA, and the relationship between hsa_piR_019914, LDHA, and reactive oxygen species (ROS) production was verified by flow cytometry.
Results
The piRNA hsa-piR-019914 was significantly downregulated in osteoarthritis (OA). Hsa-piR-019914 reduced inflammation-mediated chondrocyte apoptosis and maintained cell proliferation and clone formation in vitro. Hsa-piR-019914 reduced the production of LDHA-dependent ROS through targeted regulation of LDHA expression, maintained chondrocyte-specific gene expression of ACAN and COL2, and inhibited the gene expression of MMP3 and MMP13.
Conclusions
Collectively, this study showed that hsa_piR_019914 was negatively correlated with the expression of LDHA, which mediates ROS production. Under the stimulation of inflammatory factors, overexpression of hsa_piR_019914 had a protective effect on chondrocytes in vitro, and the absence of hsa_piR_019914 exacerbated the negative effect of inflammation on chondrocytes. Studies on piRNAs provide new therapeutic interventions for OA.
Introduction
Osteoarthritis (OA) is one of the most common joint diseases and the main reason for chronic pain and disability in the elderly population. 1 With the aging of the population, the incidence rate of this disease is on the rise worldwide and is characterized by cartilage degeneration, osteophyte formation, subchondral osteosclerosis, chronic synovial inflammation, and meniscus injury. At present, studies have shown that the pathogenesis of osteoarthritis is mainly affected by age, sex, obesity, abnormal activity of osteoclasts and chondrocytes, dysfunction of the extracellular matrix, and other factors. 2 At present, early diagnosis of OA is very difficult, and it is usually diagnosed during the clinical manifestation or late stage. At this time, the effect of conservative treatment is very limited, and the quality of life of patients can only be improved by joint replacement surgery. 3 However, not all patients can undergo surgery, and postoperative complications and mortality can occur. 4 At present, there is no method for the early diagnosis and etiological treatment of osteoarthritis, and there is no particularly effective intervention or treatment for patients with advanced osteoarthritis except surgery. Therefore, it is critical to explore the pathophysiology and molecular regulatory mechanism of osteoarthritis to identify biomarkers for the early diagnosis of osteoarthritis and develop new targets for etiology and treatment.
Epigenetics has become a research hotspot in the field of life science in recent years, and noncoding RNA research is an important part of this field. Studies have confirmed that ncRNAs (miRNAs, lncRNAs, and circRNAs) play important roles in the biological processes of transcription and translation. NcRNAs are involved in the regulation of inflammation, chondrocyte apoptosis, and extracellular matrix metabolism and play an important role in the occurrence and development of OA. 5 With further research on microRNAs (miRNAs) and long noncoding RNAs (lncRNAs) among noncoding RNAs, research on piwi-interacting RNAs (piRNAs), which is a new research field, has become a new hotspot in life science research.6,7 PiRNA is a kind of small noncoding RNA composed of 24 to 31 nucleotides found in bacteria and somatic cells. PiRNA can bind to the piwi protein to form the piRNA/piwi complex, thus affecting transposon silencing, spermatogenesis, genome rearrangement, epigenetic changes, and protein regulation. 8 A large portion of piRNAs come from 2 types of large genome loci called piRNA clusters. 9 Similar to coding genes, single-stranded clusters are transcribed by RNA polymerase II, and double-stranded clusters are transcribed from 2 genomic chains, which rely on nearby coding genes to promote transcriptional initiation. 10 Two types of piRNA are transcribed in the nucleus. These primitive piRNAs mainly rely on the intermediate carrying 5’ U to bind to piwi and then are methylated in the cytoplasm to form a mature piRNA-piwi complex. Piwi protein is composed of the PAZ and piwi structure. After PAZ binds to the piwi protein, the 3’ end is constructed by the Zuc ribozyme. 11 PiRNA plays an important role in physiological and pathological processes. PiRNA functions by forming piRNA-RNA complementarity through a mechanism similar to miRNA, and the targets include mRNAs, 12 lncRNAs, 13 and pseudogenes. 14
Reactive oxygen species (ROS) are the products of normal mitochondrial metabolism. 15 Under physiological conditions, the redox system in cells is balanced by ROS and reactive nitrogen (RNS). 16 Oxidative stress is the result of the loss of balance between the production of reactive oxygen species and intracellular antioxidant defense and scavenging, which may lead to cell death and promote the occurrence and development of diseases. 17 Oxidative stress is associated with many inflammatory responses. Oxidative stress not only causes the destruction of articular cartilage but also leads to inflammatory changes in articular cartilage. For example, in acute inflammation, neutrophils rapidly produce and release large amounts of ROS, which is known as oxidative eruption, and the synovial fluid of joints and the synovium of patients with OA often accumulates a large number of neutrophils, which can release a large number of free radicals. 18 During chronic inflammation, the production and clearance of ROS are imbalanced, and pathogens and host cells are in an environment of excessive ROS for a long time. 19 With further research, the relationship between oxidative stress and many diseases has been confirmed. Chronic inflammatory diseases such as diabetes, chronic obstructive pulmonary disease, tumor diseases such as colon and breast cancer, ischemia–reperfusion injury, rheumatoid arthritis and other autoimmune diseases, neurodegenerative diseases such as Alzheimer’s disease, atherosclerosis, and other cardiovascular diseases are all closely related to oxidative stress. 20 Oxidative stress is one of the main causes of cartilage tissue injury in osteoarthritis. Osteoarthritis chondrocytes can produce a large amount of ROS,21,22 and excessive ROS can lead to cartilage matrix degradation, 23 prevent the synthesis of hyaluronic acid and proteoglycan, 24 destroy the cell membrane, 25 and induce chondrocyte apoptosis. 26
LDH is a tetramer enzyme formed by the arrangement and binding of 2 subunits: LDHA and LDHB. As a key enzyme, LDHA participates in the last step of cell glycolysis, mainly catalyzing pyruvate to lactic acid and oxidizing NADH to NAD+, to provide energy for cells, which plays an important role in anaerobic glycolysis. Lactate dehydrogenase A has a higher affinity for pyruvate and a faster catalytic rate than LDHB. 27 LDHA is involved in mediating oxidative stress, 28 extracellular matrix degradation, 29 cell proliferation and migration, 30 and cell metabolism, 31 and it has been widely and deeply studied as an important target for drug development and disease treatment. LDHA is considered to be a sensor of ROS. 32 Many studies have reported that cells can be protected from oxidative stress by inhibiting LDHA-mediated ROS production.33-35 Chondrocytes under inflammatory conditions undergo a metabolic transition regulated by NF-κB activation, which leads to the reprogramming of cell metabolism to glycolysis and LDHA. Inflammation and metabolism can regulate each other to regulate the degradation of cartilage. LDHA binds with NADH to stabilize NF-κB, a key proinflammatory mediator in chondrocytes, and promotes catabolic changes induced by ROS. LDHA-mediated ROS generation in chondrocytes is a potential therapeutic target for osteoarthritis. 36
In this study, we screened the abnormal expression of piRNAs in OA, verified the biological function of hsa-piR-019914 in chondrocytes, and revealed that hsa-piR-019914 reduced the production of ROS through targeted inhibition of LDHA.
Methods
Bioinformatics Analysis
Bioinformatics analysis was carried out at https://www.omicstudio.cn/tool using OmicStudio software. Volcano maps (or other graphics) were drawn based on R on the OmicStudio platform (https://www.r-project.org/https://www.omicstudio.cn/tool). A small RNA sequencing dataset (GSE143514) including 3 OA patients and 5 normal controls was downloaded from the GEO datasets for piRNA expression analysis. The Cutadapt program was used to remove the connector sequence in the original offline data. The Trimmatic program was used to remove low-quality sequences to obtain clean data. The Fastqc program was used to count the data volume of the clean data and retain fragments larger than 15 nt for subsequent analysis. A bowtie program was used to compare clean data to the piRNA database and genome, an edger was used to analyze the difference in piRNA expression, and piRNAs with significant differences between the 2 comparison groups were selected according to P value <0.05 and FC>2 or FC<0.5.
Cell Culture
The human chondrocyte lines C28/I2 and SW1353 were purchased from the Cell Bank of the Chinese Academy of Sciences. The cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM)/F-12 (Thermo Fisher, California, USA) with 10% fetal bovine serum (FBS, BIOEXPLORER, California, USA) at 37°C in a humidified chamber with 5% CO2.
RNA Extraction and Quantitative Real-Time Polymerase Chain Reaction
TRIzol reagent (Invitrogen, California, USA) was used to extract the total RNA from cells. For piRNA analysis, an EZB-miRT2-plus-L kit (EZBioscience, Roseville, USA) was used to reverse transcribe the piRNA into cDNA, and the relative expression level of piRNA was normalized to U6 controls. For mRNA analysis, a HiScript III 1st Strand cDNA Synthesis Kit (Vazyme, Nanjing, China) was used to reverse transcribe the mRNA into cDNA, and the relative expression level of genes was normalized to the internal control GAPDH. Real-time quantitative polymerase chain reaction (q-PCR) was conducted in a LightCycler 480 instrument and a ChamQ Universal SYBR qPCR Master Mix kit (Vazyme, Nanjing, China) according to the manufacturer’s instructions. Relative piRNA and mRNA expression was calculated using the 2-ΔΔCt method. All q-PCR was performed in triplicate on a LightCycler 480 System (Roche, Basel, Switzerland). The primers were obtained from Sangon Biotech (Shanghai, China) Co., Ltd. ( Table 1 ).
The Sequence of Primers.
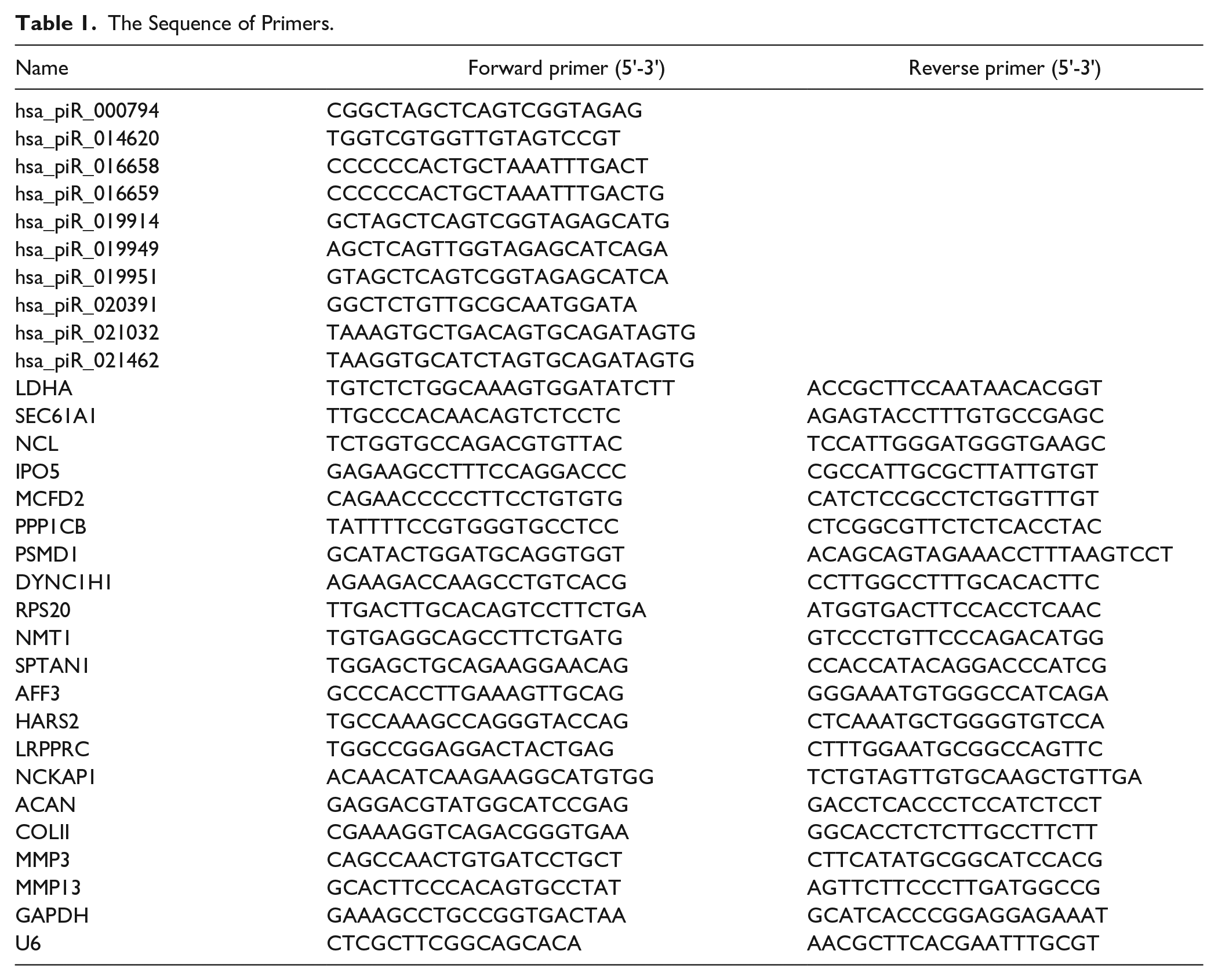
Cell Transfection
The hsa-piR-019914 mimics, hsa-piR-019914 inhibitors, LDHA siRNA, and the negative control siRNA were purchased from General Bio-company (General Biol, Anhui, China). According to the manufacturer’s instructions, all mimics, inhibitors, and siRNA plasmids were transfected into C28/I2 cells using Lipofectamine™ RNAiMAX Transfection Reagent (Thermo Fisher, California, America).
Model of OA
In this study, 10 ng/ml interleukin (IL)-1β, 10 ng/ml tumor necrosis factor (TNF-α), or 1 µg/ml lipopolysaccharide (LPS) was used to stimulate C28/I2 cells or SW1353 cells to establish the model of OA in vitro for 24 h.
CCK-8 Analysis
The cells were inoculated into each well of a 96-well plate at a density of 2000 cells per well, and the cells were detected after they adhered to the well. This point was considered 0 h. Ten microliters of CCK-8 detection reagent was added at 0, 12, 24, 48, 72, and 96 h and incubated for 2 h. The absorbance at 450 nm was measured by a microplate instrument.
Clone Formation Analysis
The cells were inoculated into a 6-well cell culture plate at a density of 300 cells per well. The cells were cultured for 2 weeks, and the culture medium was changed every 3 days. After culture, the cells were fixed with 4% paraformaldehyde solution and stained with 0.1% crystal violet. After staining, the cells were washed twice with phosphate-buffered saline (PBS), and the number of clones was counted after drying.
Apoptosis Detection By Annexin V-FITC/PI and Flow Cytometry
C28/I2 cell apoptosis was determined by flow cytometry using an Annexin V-FITC Apoptosis Detection kit (Beyotime, Shanghai, China). C28/I2 cells were digested and collected with trypsin without ethylenediaminetetraacetic acid (EDTA), rinsed twice with cold PBS, and 500 μl binding buffer was added to suspend the cells according to the manufacturer’s instructions. Five microliters of Annexin V-FITC was added and mixed well, and 5 μl of propidium iodide was added, mixed well, and incubated in the dark at room temperature for 15 min. Flow cytometry was conducted on a BD Aria flow III cytometer (BD, New Jersey, USA). The results were analyzed by FlowJo software v10.
ROS Detection By H2DCFDA and Flow Cytometry
H2DCFDA staining was performed to detect the levels of ROS in C28/I2 cells. According to the manufacturer’s instructions, H2DCFDA was diluted with serum-free culture medium at 1:1000 to a final concentration of 10 μM. The cells were centrifuged, and the diluted probe was added to 1 × 106 cells/ml and incubated at 37°C in the dark for 30 min. After incubation, the analysis was performed on a BD Aria flow III flow cytometer (BD, New Jersey, USA). FlowJo software v10 was used to analyze the results.
mRNA Sequencing
The data were uploaded to the gene expression database (Gene Expression Omnibus, GEO) (numbered GSE227945). Total RNA was extracted using TRIzol reagent according to the manufacturer’s instructions. Furthermore, mRNA sequencing was performed by LianChuan Biological Company (Hangzhou, China). The samples were 3 cases of mimic NC-transfected C28/I2 cells and 3 cases of mimic hsa_piR_019914-transfected C28/I2 cells. After RNA was extracted by the TRIzol method, total RNA was purified by an RNeasy Kit (QIAGEN, Dusseldorf, Germany), and then cDNA libraries were constructed. RNA-seq was performed on an Illumina NovaSeq 6000 platform according to the manufacturer’s instructions, and reads were generated.
After sequencing, raw data (raw data) were obtained. After filtering, joints were removed, pollution was removed, and the reference genome was compared. High-quality mapped reads (MAPQ ≥ 30) were used for subsequent analysis. The Cutadapt program was used to remove the joint sequence in the original offline data. The Trimmatic program was used to remove low-quality sequences to obtain clean data. The FastQC program was used to count the data volume of clean data and the proportions of q20 and q30. The HISAT2 program was used to compare clean data to the reference genome. StringTie was used to splice reads into transcripts and analyze gene expression levels. Edger or DEGseq was used for differential gene expression analysis. Cluster Profiler was used to analyze the GO/KEGG pathways of differentially expressed genes.
Statistical Analysis
All tests in this study were repeated at least 3 times with similar results. Data were expressed as the mean ± standard deviation (SD). Unpaired 2-tailed Student’s t-test was carried out for the comparison between 2 groups and one-way analysis of variance (ANOVA) or 2-way ANOVA was used for multiple group comparisons with Tukey post hoc test. P < 0.05 was considered as statistically significant. GraphPad Prism software (Version 6.01) was used for statistical analysis. 37 According to the results of bioinformatics analysis and mRNA sequencing, differentially expressed genes were screened by fold change ≥2 and P ≤ 0.05 as the standard.
Result
Decreased Expression of piRNA-019914 in Chondrocytes Stimulated By Inflammation
To screen piRNAs that play an important role in OA, we analyzed the miRNA expression profiles of OA patients and normal controls in the GEO database (GSE143514). Bioinformatics analysis showed that there were 11 significantly downregulated piRNAs (P < 0.05 and fold change > 2) in OA compared with normal controls (
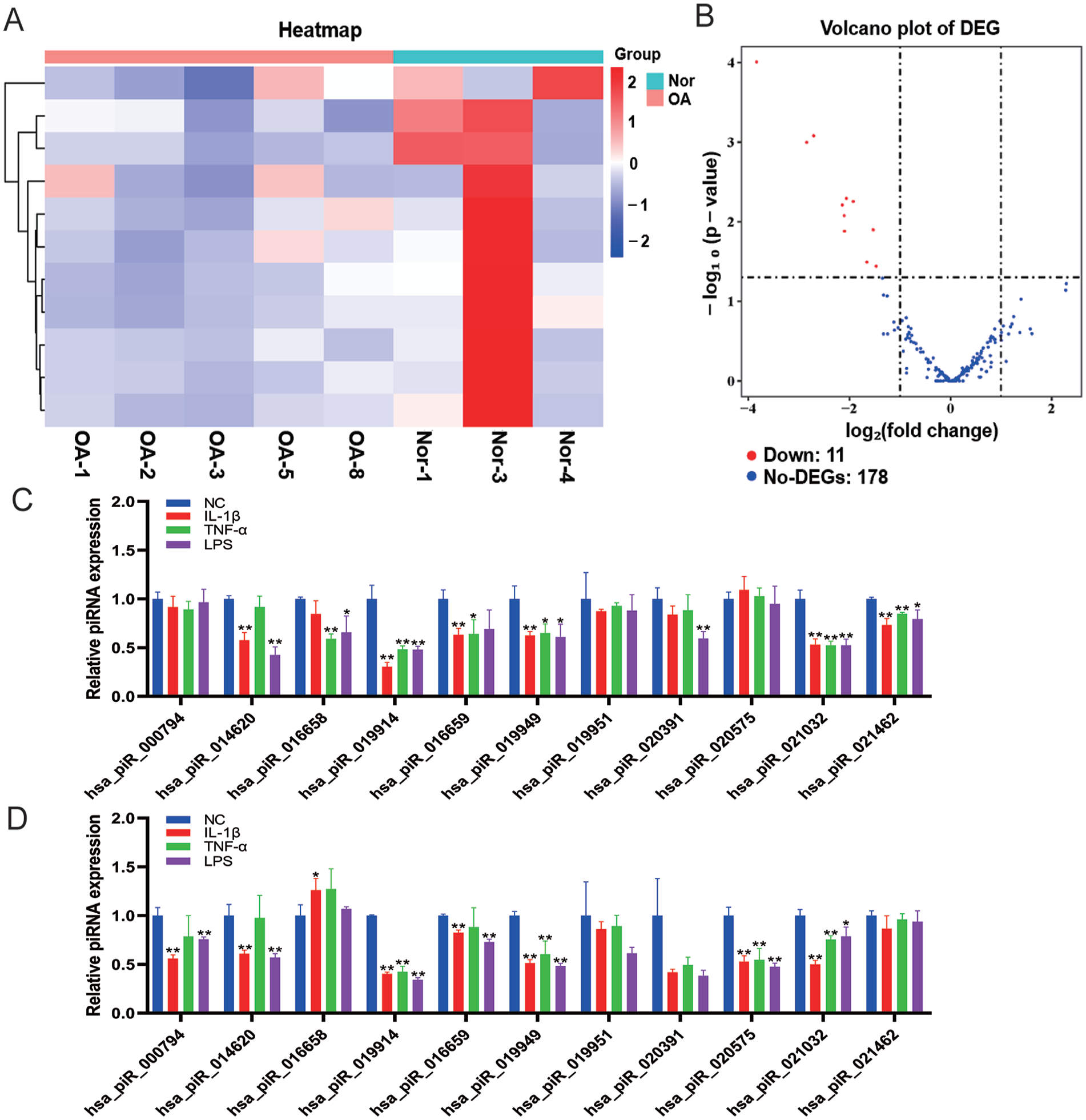
Decreased expression of piRNA-019914 in chondrocytes stimulated by inflammation. (
piRNA hsa_piR_019914 Overexpression Promotes the Anabolic Metabolism of Chondrocyte
To study the role of hsa-piR-019914 in chondrocyte function, C28/I2 cells were transfected with hsa-piR-019914 mimics, and the expression of hsa-piR-019914 in C28/I2 cells was significantly increased after transfection (
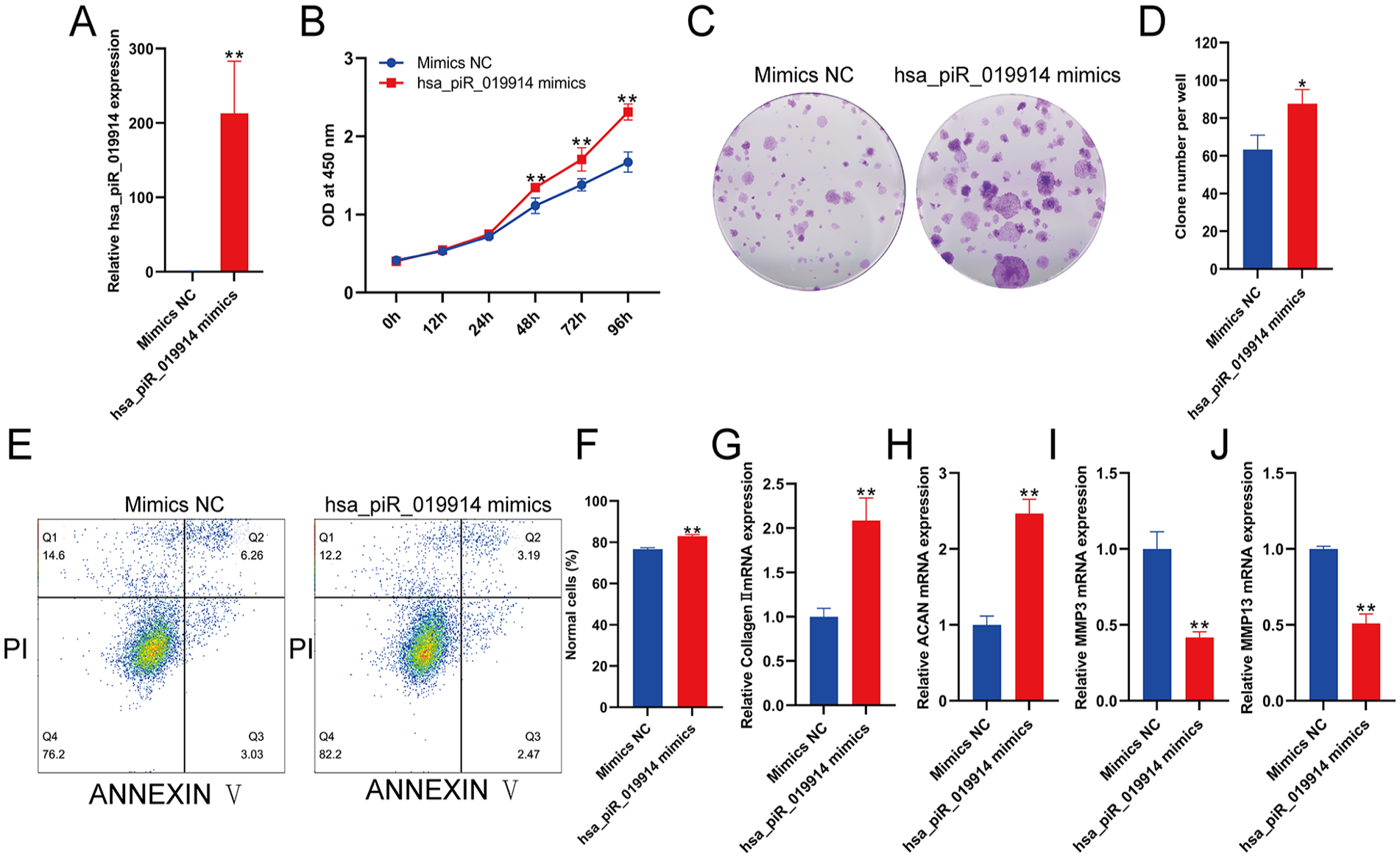
piRNA hsa_piR_019914 overexpression promotes the anabolic metabolism of chondrocytes. (
piRNA hsa_piR_019914 Suppression Promoted the Catabolic Metabolism of Chondrocyte
To study the role of hsa-piR-019914 in chondrocyte function, C28/I2 cells were transfected with hsa-piR-019914 inhibitors, and the expression of hsa-piR-019914 in C28/I2 cells was significantly inhibited after transfection (
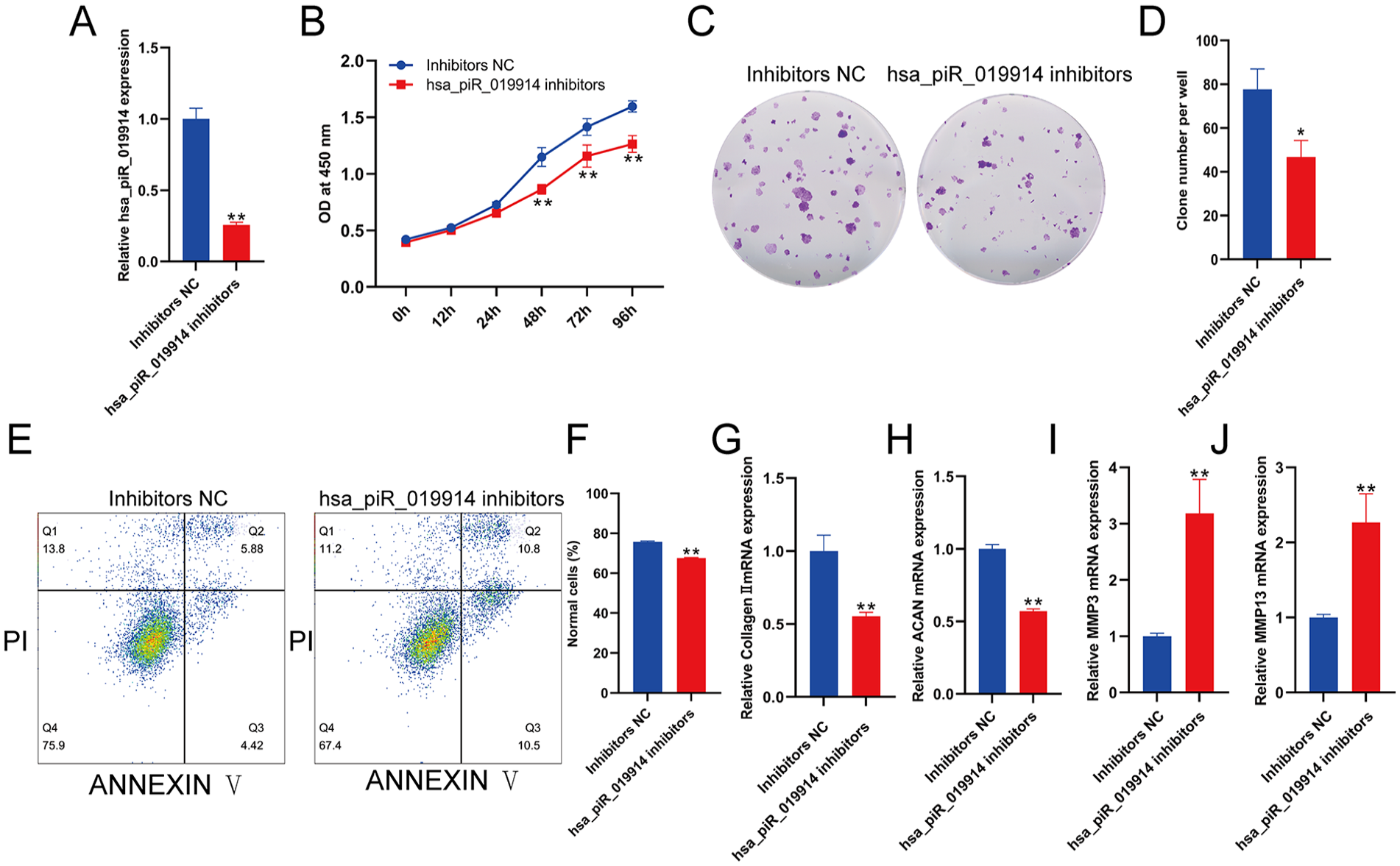
piRNA hsa_piR_019914 suppression promotes the catabolic metabolism of chondrocytes. (
piRNA hsa_piR_019914 Regulated the Expression of LDHA
To identify the target gene of hsa-piR-019914, we performed mRNA sequencing. The samples were 3 groups of C28/I2 cells transfected with mimics NC and 3 groups of C28/I2 cells transfected with hsa-piR-019914 mimics. The sequencing data were uploaded to the GEO database [GSE227945]. The heatmap (
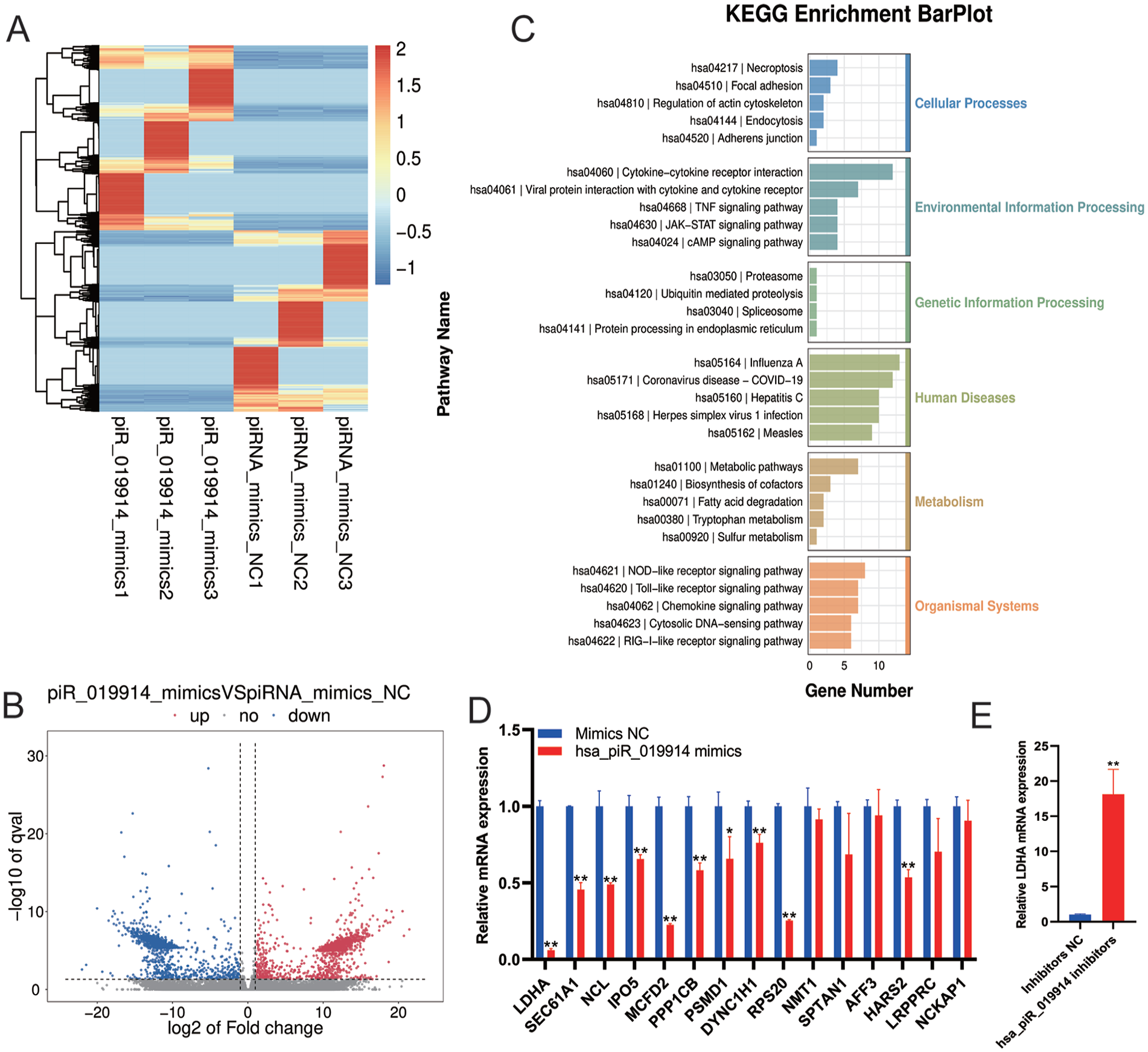
The piRNA hsa_piR_019914 regulated LDHA expression. (
piRNA hsa_piR_019914 Regulated the Oxidative Stress in Chondrocyte
Lactate dehydrogenase A–mediated ROS production plays an important role in OA.
36
Flow cytometry showed that overexpression of hsa-piR-019914 decreased ROS production in C28/I2 cells, while inhibition of hsa-piR-019914 led to an increase in ROS production in C28/I2 cells (
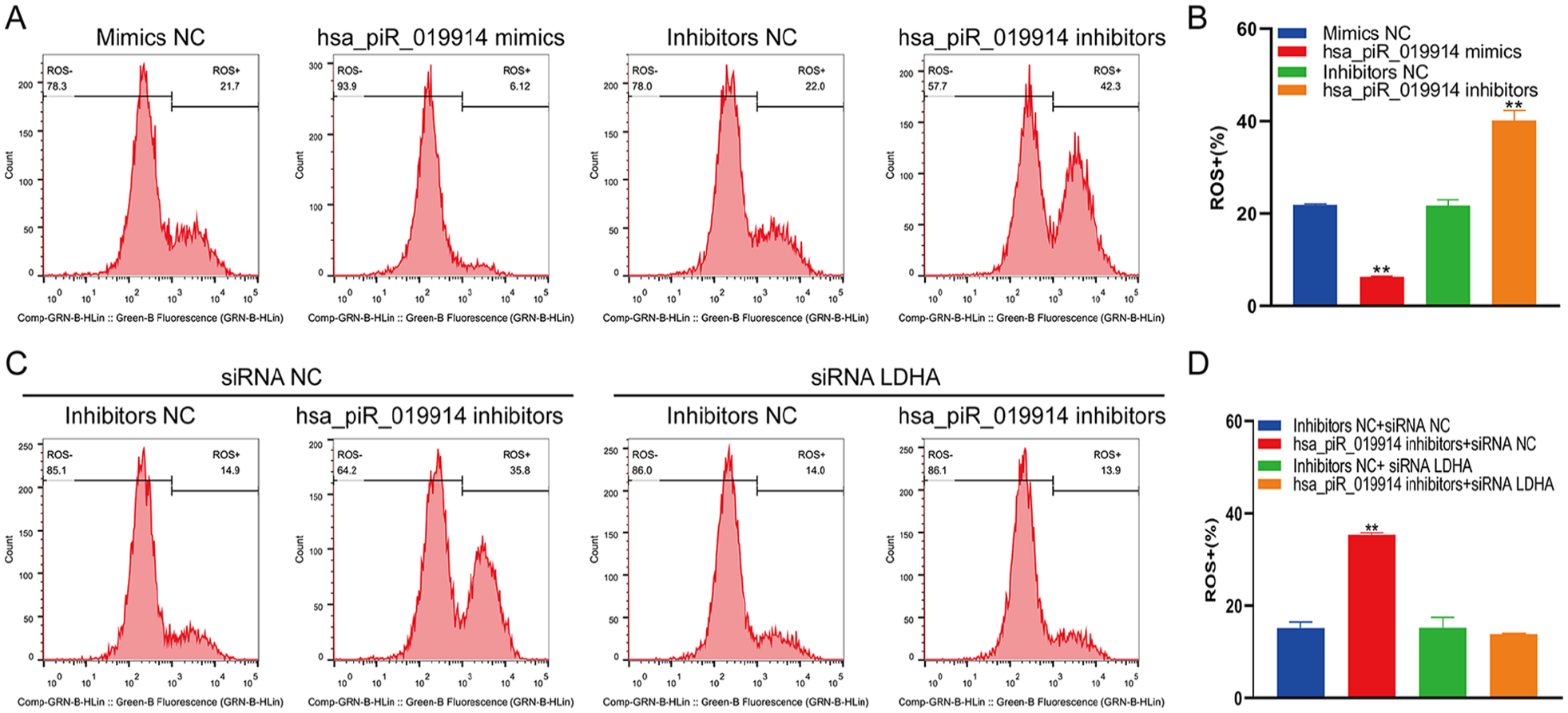
The piRNA hsa_piR_019914 regulated oxidative stress in chondrocytes. (
Discussion
The etiology and pathogenesis of osteoarthritis are complex, and the exact molecular mechanism is not clear. As the number of high-throughput profiling studies and mechanistic investigations of noncoding RNAs in OA has increased, it has been confirmed that noncoding RNAs play an important role in the pathogenesis and are potential therapeutic targets of OA. 38 As a recently discovered non-small coding RNA, piRNA has been widely studied in the field of cancer, and its great research value and potential have been verified. With the development of high-throughput sequencing and other advanced detection techniques, it is easy to detect the abnormal expression of piRNA/piwi protein in diseases. Several databases, such as piRbase and piRNA Bank, can be used for piRNA functional analysis, target RNA prediction and homologous piRNA searches. The mechanism of piRNA in osteogenic differentiation has been studied, 39 and it has been found that piRNA is a potential target in osteoporosis. 40 However, no research has explored the role of piRNAs in OA.
In this study, we screened the abnormal expression of piRNAs in OA through the GEO database and bioinformatics analysis. Next, to verify the correlation between candidate piRNAs and the pathogenesis of OA, we constructed an in vitro model of OA chondrocytes and detected the expression of candidate piRNAs in OA chondrocytes. According to the results, we eliminated the piRNAs with no significant difference. We found that hsa_piR_019914 was significantly downregulated in OA.
To study the biological function of hsa_piR_019914 in chondrocytes, overexpression or inhibition of hsa_piR_019914 in C28/I2 cells was performed by transfection of hsa_piR_019914 mimics and hsa_piR_019914 inhibitors, and the transfection efficiency was verified by qPCR. Many studies have confirmed the important role of piRNAs in regulating cell proliferation and apoptosis.41-45 The normal function and activity of chondrocytes is an important condition for maintaining cartilage homeostasis. 46 Degenerative lesions of articular cartilage are the main feature of OA, and there is obvious apoptosis in OA cartilage. 47 Many studies have blocked chondrocyte apoptosis and regulated chondrocyte activity to treat OA. In this study, the CCK-8 assay results showed that overexpression of hsa_piR_019914 increased the proliferation of C28/I2 cells and inhibition of hsa_piR_019914 inhibited the proliferation of C28/I2 cells, and similar results were observed in the colony formation assay. Then, to determine whether hsa_piR_019914 can regulate the apoptosis of C28/I2 cells, we detected apoptosis by flow cytometry. The apoptosis level of cells overexpressing hsa_piR_019914 decreased, and inhibition of hsa_piR_019914 increased apoptosis. These results showed that hsa_piR_019914 maintained the proliferation of OA chondrocytes and reduced the apoptosis level of OA chondrocytes.
Further studies are needed to determine the potential target genes of hsa_piR_019914. We examined the hsa_piR_019914 overexpression group and negative control group of C28/I2 cells by small RNA sequencing. We uploaded the sequencing results to the GEO database [GSE227945]. KEGG pathway analysis showed that the differentially expressed miRNAs were mainly related to apoptosis, oxidative stress and the inflammatory signaling pathway. Here, our results demonstrated that the expression of hsa_piR_019914 was negatively correlated with the expression of LDHA. Lactate dehydrogenase A has been thoroughly studied as an important therapeutic target in many diseases, such as cancer. Arra et al. 36 showed that LDHA-mediated ROS generation in chondrocytes was a potential therapeutic target for osteoarthritis.
To further investigate whether the biological function of hsa_piR_019914 was mediated by LDHA, we transfected C28/I2 cells with the hsa_piR_019914 inhibitor and siLDHA and then used flow cytometry to detect ROS(+) cells. Our results showed that overexpression of hsa_piR_019914 reduced the production of ROS, and inhibition of hsa_piR_019914 increased the production of ROS. When the expression of hsa_piR_019914 was inhibited and the expression of LDHA was knocked down, the production of ROS decreased significantly. These results indicate that hsa_piR_019914 is involved in regulating the production of ROS, and hsa_piR_019914 inhibits the expression of LDHA and LDHA can mediate the production of ROS, 36 suggesting that hsa_piR_019914 is a potential target for the treatment of OA that can reduce the production of ROS in chondrocytes by targeting hsa_piR_019914 to inhibit the expression of LDHA.
Chondrocytes can produce a large amount of ROS under the pathological conditions of OA.20,21 Excessive ROS induce oxidative stress, increase the expression of matrix metalloproteinases, lead to degradation of the cartilage matrix, and block the synthesis of ACAN and COL2.24,48 Acan and Col2a1 are specific genes in chondrocytes, and MMP3 and MMP13 are matrix metalloproteinases that mediate the degradation of cartilage matrix. Our results show that the overexpression of hsa_piR_019914 can protect the expression of ACAN and COL2 in chondrocytes and reduce the expression of MMP3 and MMP13, and inhibiting hsa_piR_019914 can increase the expression of MMP3 and MMP13 in OA and decrease the expression of ACAN and COL2.
In this work, the following findings were highlighted: (1) piRNA hsa-piR-019914 was downregulated in OA, (2) the piRNA hsa_PiR_019914 can promote the proliferation of chondrocytes and reduce apoptosis, and (3) the piRNA hsa_piR_019914 can reduce the production of LDHA-dependent ROS by inhibiting the expression of LDHA, which has anti-inflammatory and anticatabolic effects.
Supplemental Material
sj-tif-1-car-10.1177_19476035231181094 – Supplemental material for PiRNA hsa_piR_019914 Promoted Chondrocyte Anabolic Metabolism By Inhibiting LDHA-Dependent ROS Production
Supplemental material, sj-tif-1-car-10.1177_19476035231181094 for PiRNA hsa_piR_019914 Promoted Chondrocyte Anabolic Metabolism By Inhibiting LDHA-Dependent ROS Production by YuXuan Gao, Wen Yan, Liangye Sun and XiaoLing Zhang in CARTILAGE
Supplemental Material
sj-tif-2-car-10.1177_19476035231181094 – Supplemental material for PiRNA hsa_piR_019914 Promoted Chondrocyte Anabolic Metabolism By Inhibiting LDHA-Dependent ROS Production
Supplemental material, sj-tif-2-car-10.1177_19476035231181094 for PiRNA hsa_piR_019914 Promoted Chondrocyte Anabolic Metabolism By Inhibiting LDHA-Dependent ROS Production by YuXuan Gao, Wen Yan, Liangye Sun and XiaoLing Zhang in CARTILAGE
Supplemental Material
sj-tif-3-car-10.1177_19476035231181094 – Supplemental material for PiRNA hsa_piR_019914 Promoted Chondrocyte Anabolic Metabolism By Inhibiting LDHA-Dependent ROS Production
Supplemental material, sj-tif-3-car-10.1177_19476035231181094 for PiRNA hsa_piR_019914 Promoted Chondrocyte Anabolic Metabolism By Inhibiting LDHA-Dependent ROS Production by YuXuan Gao, Wen Yan, Liangye Sun and XiaoLing Zhang in CARTILAGE
Supplemental Material
sj-xlsx-4-car-10.1177_19476035231181094 – Supplemental material for PiRNA hsa_piR_019914 Promoted Chondrocyte Anabolic Metabolism By Inhibiting LDHA-Dependent ROS Production
Supplemental material, sj-xlsx-4-car-10.1177_19476035231181094 for PiRNA hsa_piR_019914 Promoted Chondrocyte Anabolic Metabolism By Inhibiting LDHA-Dependent ROS Production by YuXuan Gao, Wen Yan, Liangye Sun and XiaoLing Zhang in CARTILAGE
Footnotes
Acknowledgments and Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was supported by a Grant for Key Research and Development Plan of Anhui Province, a Special Project on population health (202004j07020003).
Author Contributions
Conceptualization: Y.X. Gao, W Yan, L.Y. Sun, and X.L. Zhang; methodology and formal analysis: Y.X. Gao, W. Yan; drafting of the article: Y.X. Gao, W. Yan; review and editing: all authors; supervision: X.L. Zhang; acquisition of funding: Y.X. Gao, L.Y. Sun, and X.L. Zhang. All authors have read and agreed to the published version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The Medical Research Ethics Committee of the Second Hospital of Shanxi Medical University.
Availability of Data and Materials
The datasets used and/or analyzed during the present study are available from the corresponding author upon reasonable request.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
