Abstract
Objective
Osteoarthritis (OA) is an incurable joint disease characterized by pronounced pain. MicroRNAs constitute epigenetic mechanisms that may affect OA progression by contributing to changes in chondrocyte phenotype. This study investigates for the first time whether there is a link between miRNA-1 (miR-1) and OA pathogenesis, and the molecular mechanisms involved.
Design
OA-associated gene expression, including MMP-13, ADAMTS5, and COL2A1 was compared in chondrocytes from non-OA and OA cartilage, and in SW1353 cells over- and underexpressing miR-1. Bioinformatics and luciferase reporter assay were conducted to confirm whether FZD7 was a target of miR-1. The effects of miR-1 on FZD7 expression and downstream Wnt/β-catenin signalling were investigated.
Results
Non-OA and OA chondrocytes differed significantly in the expression of miR-1 and OA-associated genes. MiR-1 over- and underexpression in SW1353 cells, respectively, reduced and enhanced gene expression associated with cartilage catabolism. FZD7, which has an important role in the Wnt/β-catenin signaling pathway, was shown to be a potential target of miR-1. MiR-1 binding to FZD7 increased the levels of phosphorylated (inactivated) β-catenin, thereby preventing downstream β-catenin signaling.
Conclusions
Inhibition of Wnt/β-catenin signaling by miR-1 in chondrocytes may attenuate the expression of genes that regulate the activity of catabolic enzymes. This finding may be useful for future investigations of molecular targets for OA treatment.
Introduction
Osteoarthritis (OA) is a chronic degenerative joint disease characterized by progressive destruction of articular cartilage that is often associated with significant pain. 1 Due to the irreversible nature of the disease and the lack of an effective cure, OA remains a leading cause of global disability for which chronic management creates a major burden for health care systems worldwide. 2 Although some common risk factors have been identified for OA, including age, gender, prior joint injury, obesity, genetic predisposition, and abnormal mechanical loading, 3 the disease is idiopathic in the majority of OA patients. The pathogenesis of OA is complex and not fully understood, and the severity of disease symptoms (such as pain) can have almost no association with structural changes in the joint (such as cartilage and bone changes) or the phenotype of cells (such as chondrocytes). These factors make OA a difficult disease to treat, and the effects of current clinical therapies are typically limited to relieving symptoms rather than interfering with disease progression.
Degenerative changes in OA are correlated with alterations in chondrocyte gene expression involved in maintaining the integrity and function of articular cartilage.4,5 Over the past decade, growing evidence has pointed to the important role of epigenetic mechanisms in modulating cell phenotype in OA.6-8 Interactions with the environment can produce epigenetic alterations in gene expression that contribute to disease etiology and are transmitted in a sustained manner to daughter cells throughout many doublings. Of these epigenetic mechanisms, microRNAs (miRNAs) are of particular interest due to their roles in maintaining a normal chondrocyte phenotype and controlling inflammatory response pathways, 7 which may affect OA disease progression. miRNAs are a class of small, noncoding single-stranded RNAs of 20 to 25 nucleotides that regulate gene expression at the posttranscriptional level, affecting cell fate during proliferation and differentiation. 9 This regulatory mechanism involves sequence-specific interactions with the 3′-untranslated regions (3′-UTRs) of specific mRNA targets, leading to mRNA degradation and/or translational repression. 10
Almost one-third of mammalian mRNA expression is regulated by miRNAs, 11 which have diverse but critical functions in tissue development and disease.12,13 Alterations in the expression of miRNAs are found to be closely related to chondrogenesis and OA.14,15 A range of miRNAs have been explored in this regard and some of their specific roles in regulating chondrocyte activity and OA pathogenesis have been elucidated.16,17 For instance, miR-140 is a cartilage-specific miRNA that is expressed during embryogenesis and adult articular cartilage, with important roles in OA pathogenesis through the regulation of ADAMTS-5 and IGFBP-5.18,19 Other relatively well-investigated miRNAs which are differentially expressed in OA and/or have important roles in OA pathogenesis include miR-27,19,20 miR-34a, 21 miR-146,14,22 and miR-675. 23
Proteins of the Wnt/β-catenin pathway are known to be overexpressed in the joints of OA patients and in animal models of disease, and pathway activation is thought to contribute to chondrocyte and matrix degradation in joint pathologies.24,25 Despite there being a range of studies investigating the roles of various miRNAs in OA, there is insufficient evidence to suggest that these affect Wnt/β-catenin signaling as one of the major pathways modulating OA progression. In this context, we have identified miR-1 as a possible epigenetic link between Wnt/β-catenin signaling and OA. MiR-1 has been shown to modulate the activity of the Wnt/β-catenin pathway, which in one study promoted cardiomyocyte differentiation in pluripotent stem cells and suppressed their commitment to becoming endothelial cells. 26 The same study also identified FZD7 as a new miR-1 target. However, the only known association between miR-1 and joint pathology is that miR-1 plays an important role in regulating the chondrocyte phenotype and the late stage of chondrogenic differentiation, thereby contributing to cartilage integrity. 27 Interestingly, there has been more evidence suggesting that changes in miR-1 are correlated with other diseases that often occur as comorbidities of OA, 28 including cardiovascular disease 29 and Parkinson’s disease. 30 Based on this information, we hypothesized that non-OA and OA chondrocytes may have different levels of miR-1 expression, and that overexpression of miR-1 may inhibit OA pathogenesis by targeting FZD7 of the Wnt/β-catenin signaling pathway. This is the first study to define a link between miR-1 and OA, where miR-1 overexpression was found to inhibit the expression of catabolic genes related to OA development, and suppress the Wnt/β-catenin pathway through FZD7 in chondrocytic cells.
Methods
Tissue Samples and Chondrocyte Isolation
This research was approved by the Ethics Committee of Tianjin Hospital, China (2014-002). All methods were carried out in accordance with relevant guidelines and regulations. Discarded cartilage tissues were obtained from 10 non-OA patients undergoing traumatic above-knee amputation and 10 OA patients undergoing total knee replacement surgery. The OA patients were clinically diagnosed to be Kellgren-Lawrence (K-L) grades 3 to 4 based on radiographic examination. All clinical specimens were obtained after patients gave informed written consent. The 2 groups were paired and cartilage samples were matched by age, sex, and body mass index (Supplementary Table 1).
Chondrocytes were isolated from the articular cartilage of clinical specimens obtained from non-OA and OA patients. Cartilage samples were enzymatically digested using 0.15% (w/v) collagenase (CLS-2, Worthington, USA) for 16 hours at 37 °C in maintenance medium. Isolated chondrocytes were washed in phosphate buffered saline and filtered using 100 μm cell strainers (BD Biosciences, USA). The cells were seeded at high density (1 × 104 cells/cm2) and kept in maintenance medium for 2 days prior to gene expression analysis. Maintenance medium consisted of Dulbecco’s modified Eagle Medium (DMEM, Gibco, UK) supplemented with 10% fetal bovine serum (FBS, HyClone, USA), 100 U/mL penicillin (Gibco), and 100 mg/mL streptomycin (Gibco).
SW1353 Cell Culture and Transfection
Human chondrosarcoma cells (SW1353) were obtained from the American Type Culture Collection (ATTC). Cells were grown in DMEM supplemented with 10% FBS at 37 °C with 5% CO2. The cells were transfected with miR-1 agomir (miR-1 agonist; 5′-UGGAAUGUAAAGAAGUAUGGAG-3′), miR-1 antagomir (miR-1 antagonist; 5′-CUCCAUACUUCUUUACAUUCCA-3′), and agomir and antagomir negative controls (NC; 5′-UUCUCCGAACGUGUCACGUTT-3′, and 5′-UUGUACUACACAAAAGUACUG-3′), which were scrambled miRs (RiboBio, China), using Lipofectamin 2000 (Invitrogen, USA) according to the manufacturer’s instructions. Transfected cells were kept in maintenance medium and gene expression analysis was conducted 48 hours posttransfection.
Plasmid Construction
For the FZD7 3′UTR reporter assay, the psiCHE-FZD7 3′UTR-wt plasmid was constructed by annealing the sense and antisense strands of the putative binding site sequence for miR-1, and inserting the annealed sequence downstream of the stop codon of the Renilla luciferase gene in the psiCHECK-2 dual-luciferase reporter plasmid (Promega). The psiCHE-FZD7 3′UTR-mt plasmid was constructed using a mutated sequence (randomly disorganized seed region) in the putative binding site for miR-1. To construct the FZD7 overexpression vector (3.1-FZD7), the FZD7 coding sequence was amplified by polymerase chain reaction (PCR) and cloned into the Nhel and Xhol sites in the pcDNA3.1 vector (Invitrogen).
Bioinformatics and Luciferase Reporter Assay
FZD7 was predicted to be a prime target of miR-1 using the TargetScanHuman search program (www.targetscan.org). The complementary sequences in miR-1 and FZD7 mRNA were confirmed by microRNA.org.
SW1353 cells were grown at a density of 5.5 × 104 cells/cm2 in 48-well plates. The cells were transfected with psiCHECK-2 recombination vector (50 ng), together with miR-1 agomir (miR-1) or agomir negative control (NC). At 48 hours after transfection, Firefly luciferase and Renilla luciferase activities were measured using the Dual-Glo Luciferase Assay System (Promega) with a Synergy 2 Multi-Mode microplate reader (Biotek, USA).
Gene Expression Analysis
Total RNA was isolated from cells using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. Briefly, after homogenizing samples using the TRIzol reagent, chloroform was added and RNA was precipitated using isopropanol. The RNA was resuspended in 20 μL purified water.
For each sample, reverse transcription into cDNA was performed using 1 μg total RNA using M-MLV reverse transcriptase (Promega, USA) according to the manufacturer’s instructions. Gene expression levels were quantified using a 7900HT Fast Real-Time PCR System (Applied Biosystems, USA) and normalized to GAPDH. Primer sequences were purchased from Sigma-Aldrich. For miR-1, reverse transcription and PCR reactions were performed using the All-in-One miRNA qRT-PCR Detection Kit (GeneCopoeia, USA) with primers for mature miRNA hsa-miR-1-3p (No. HmiRQP0044). MiR-1 expression levels were normalized to U6. Relative expression for all genes were calculated using the comparative Ct (2−ΔΔCT) method.
Western Blotting
Cells were lysed using a cell lysis buffer (Beyotime, China) with a protease inhibitor cocktail (Sigma) and phosphatase inhibitors (5 mM Na4P2O7, 50 mM NaF, 1 mM vanadate). The protein content of the lysate was quantified using a BCA assay. Total protein (20 μg) from each sample was electrophoresed on 12% (w/v) sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a 0.45 μm polyvinylidene fluoride (PVDF) membrane (Sigma), which were blocked for 2 hours with 5% skim milk. The membranes were incubated at 4 °C overnight with rabbit monoclonal antibodies (CST, China) anti-FZD7 (No. 3975), anti-Cyclin (No. 4138), anti-c-Myc (No. 9402), and anti-GAPDH (No. 5174). Monoclonal anti-β-catenin (No. 8480) and monoclonal anti-phospho β-catenin (No. 4176) antibodies were, respectively, used to detect total β-catenin and nonactive (phosphorylated) β-catenin. Secondary antibodies conjugated to horseradish peroxidase (HRP) were added for 1 hour. The blots were visualized using enhanced chemiluminescence (ECL) reagent (Thermo Fisher Scientific, USA). The blot intensity was quantified using molecular imaging software (Carestream Health, USA) after normalizing to the corresponding loading control.
Statistical Analysis
Data for all experiments were obtained from at least 3 independent samples, and all results were expressed as mean ± standard deviation (SD). Statistical analysis was performed using SPSS 13.0 (SPSS, USA). One-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test was used for statistical comparisons. A P value of 0.05 was considered statistically significant.
Results
Gene Expression Profile of Chondrocytes from Non-OA and OA Patients
Chondrocytes isolated from knee cartilage of non-OA and OA patients were tested for differences in OA-related gene expression. Compared with non-OA chondrocytes, the expression level of COL2A1 was decreased in OA chondrocytes, while the expression levels of matrix metalloproteinase–9 (MMP-9), MMP-13, BMP-2, and ADAMTS5 were increased (
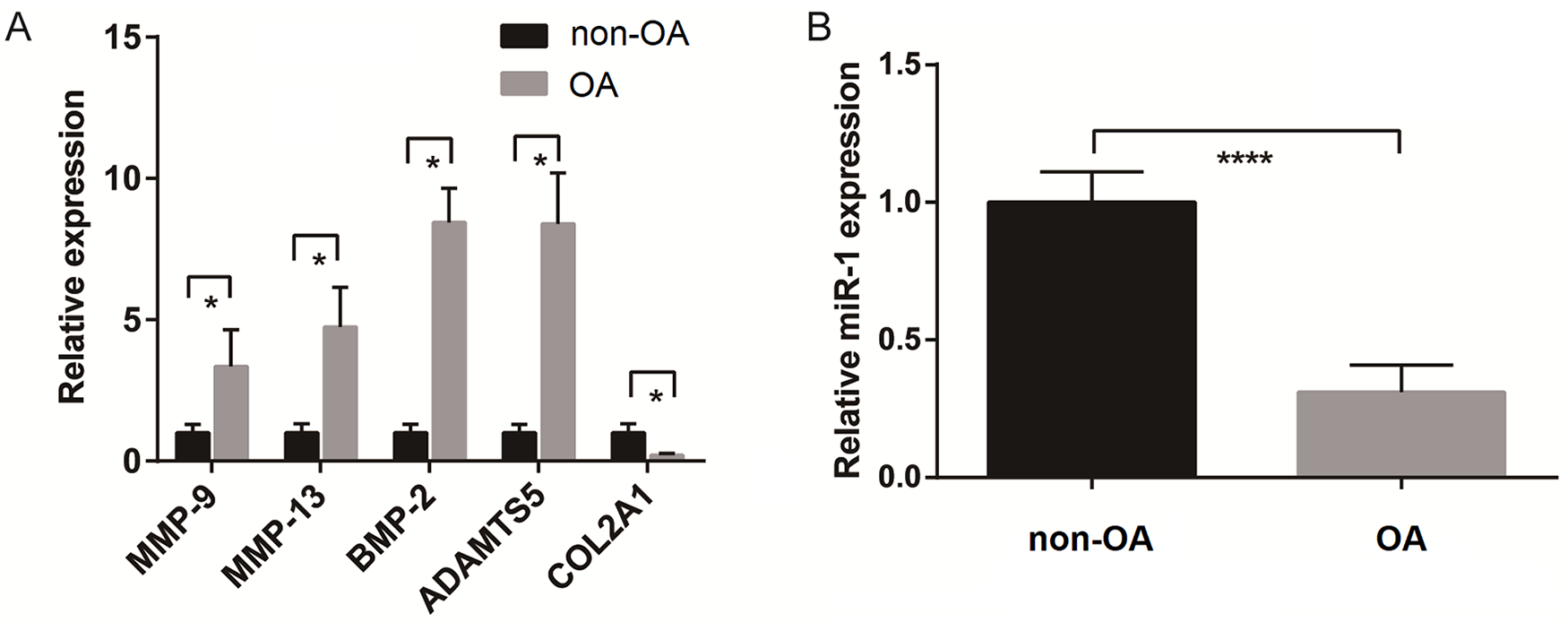
Gene expression profile of chondrocytes from nonosteoarthritis (non-OA) and OA patients. Expression levels of (
Effect of miR-1 Overexpression on the Gene Expression Profile of SW1353 Cells
SW1353 cells transfected with miR-1 antagomir, miR-1 agomir, and negative controls (NC) were tested for the expression of miR-1 and OA-related genes at 48 hours posttransfection. Cells transfected with miR-1 antagomir showed lower miR-1 expression compared with the NC group (P < 0.05), while miR-1 expression in cells transfected with miR-1 agomir were significantly higher than the NC group (P < 0.0001) (
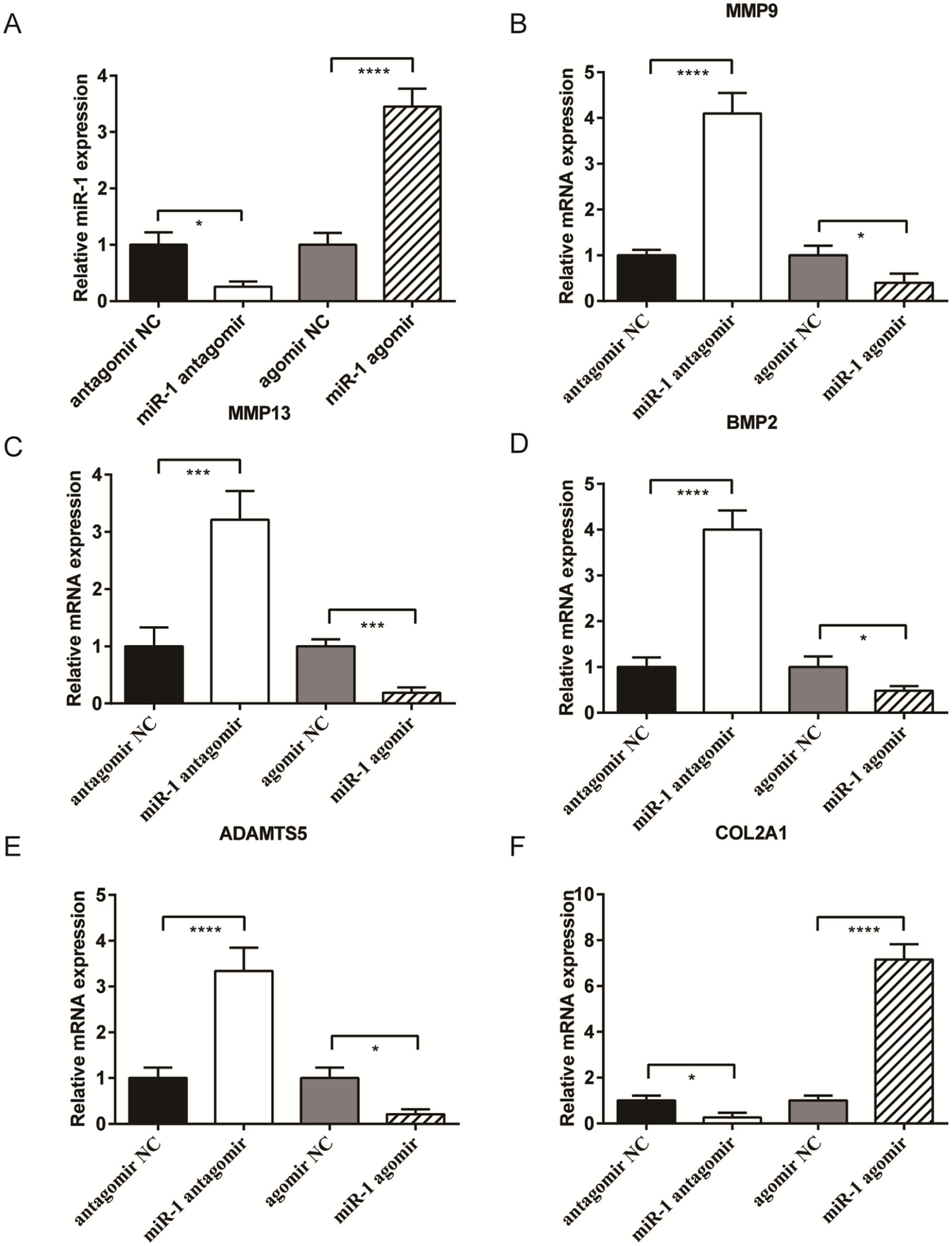
Effect of miR-1 over-expression on the gene expression profile of SW1353 cells. Expression levels of (
The inhibition of miR-1 expression in SW1353 cells caused significant increases in the expression of genes associated with cartilage catabolism, as well as greatly reduced COL2A1 expression (
miR-1 Regulation of FZD7 Expression
To determine the mechanism by which miR-1 inhibits the expression of genes associated with cartilage catabolism, we investigated a potential target of miR-1. FZD7, as a key component of the Wnt/β-catenin signaling pathway, was predicted by the TargetScan database as a prime target of miR-1, with a highly conserved complementary miR-1 binding site in its 3′UTR across vertebrates from humans to cows (Fig. 3A and B).
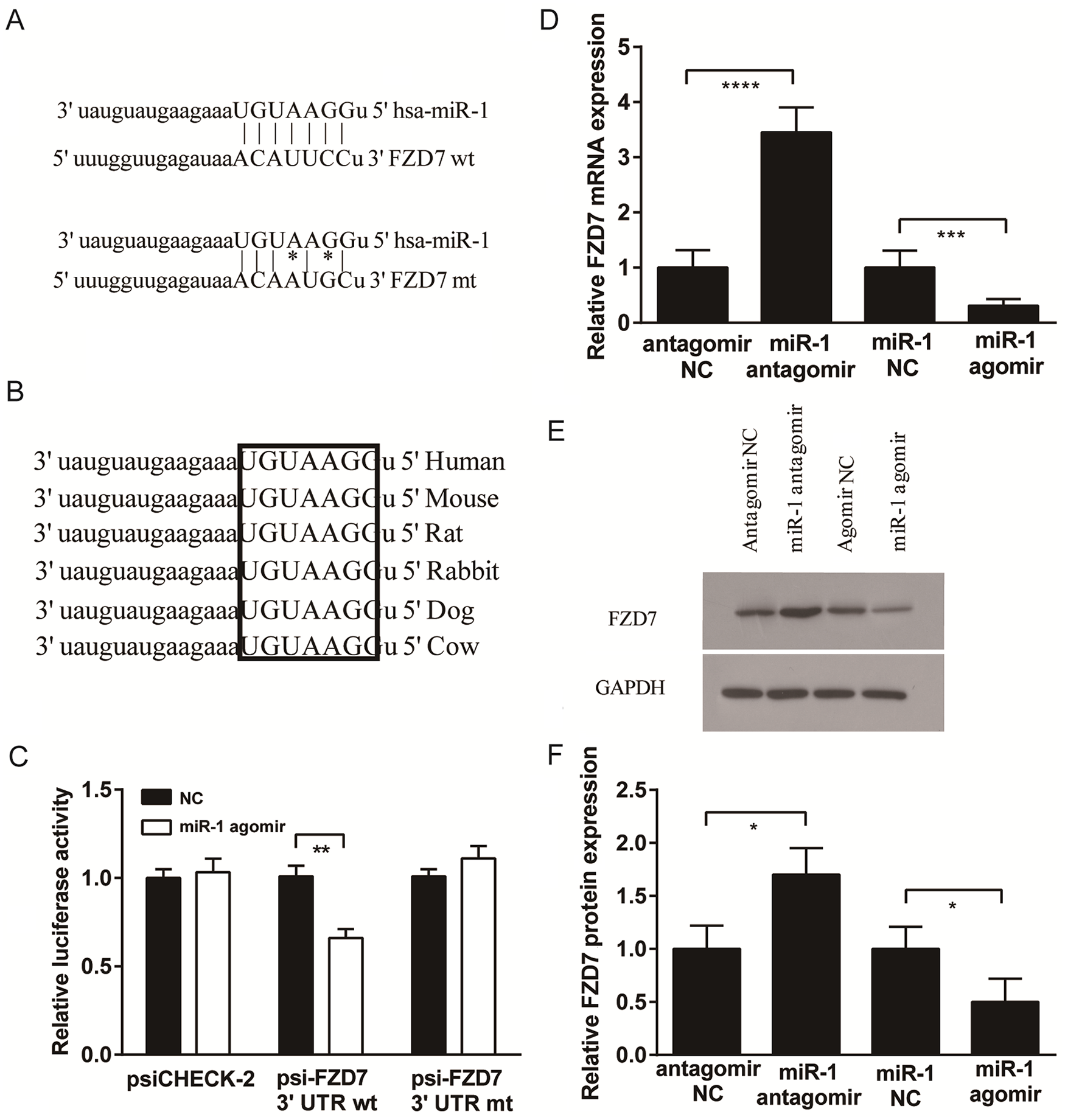
MiR-1 regulation of FZD7 expression. (
To confirm that FZD7 is a target for miR-1, sequences of wild-type 3′UTR of FZD7 containing the putative binding site for miR-1 were synthesized and inserted downstream of the luciferase gene in SW1353 cells. Renilla luciferase expression was inhibited in SW1353 cells when the construct (psiCHE-FZD7 3′UTR-wt) was co-transfected with miR-1 agomir compared with the NC (P < 0.01) (
To examine whether miR-1 can regulate FZD7 expression, the levels of FZD7 mRNA and protein expression were measured in SW1353 cells transfected with miR-1 agomir, miR-1 antagomir, and negative controls (NC). At 48 hours posttransfection, FZD7 mRNA expression was significantly reduced in the miR-1 agomir group but significantly increased in the miR-1 antagomir group (
Effect of miR-1 on Wnt/β-Catenin Signaling Pathway
FZD7 is a transmembrane receptor for Wnt/β-catenin signalling. After being transfected to induce or inhibit miR-1 expression, the total β-catenin protein levels in SW1353 cells remained similar among groups, but the levels of nonactive (phosphorylated) β-catenin were significantly elevated in the miR-1 agomir group and significantly reduced in the miR-1 antagomir group compared with NCs (
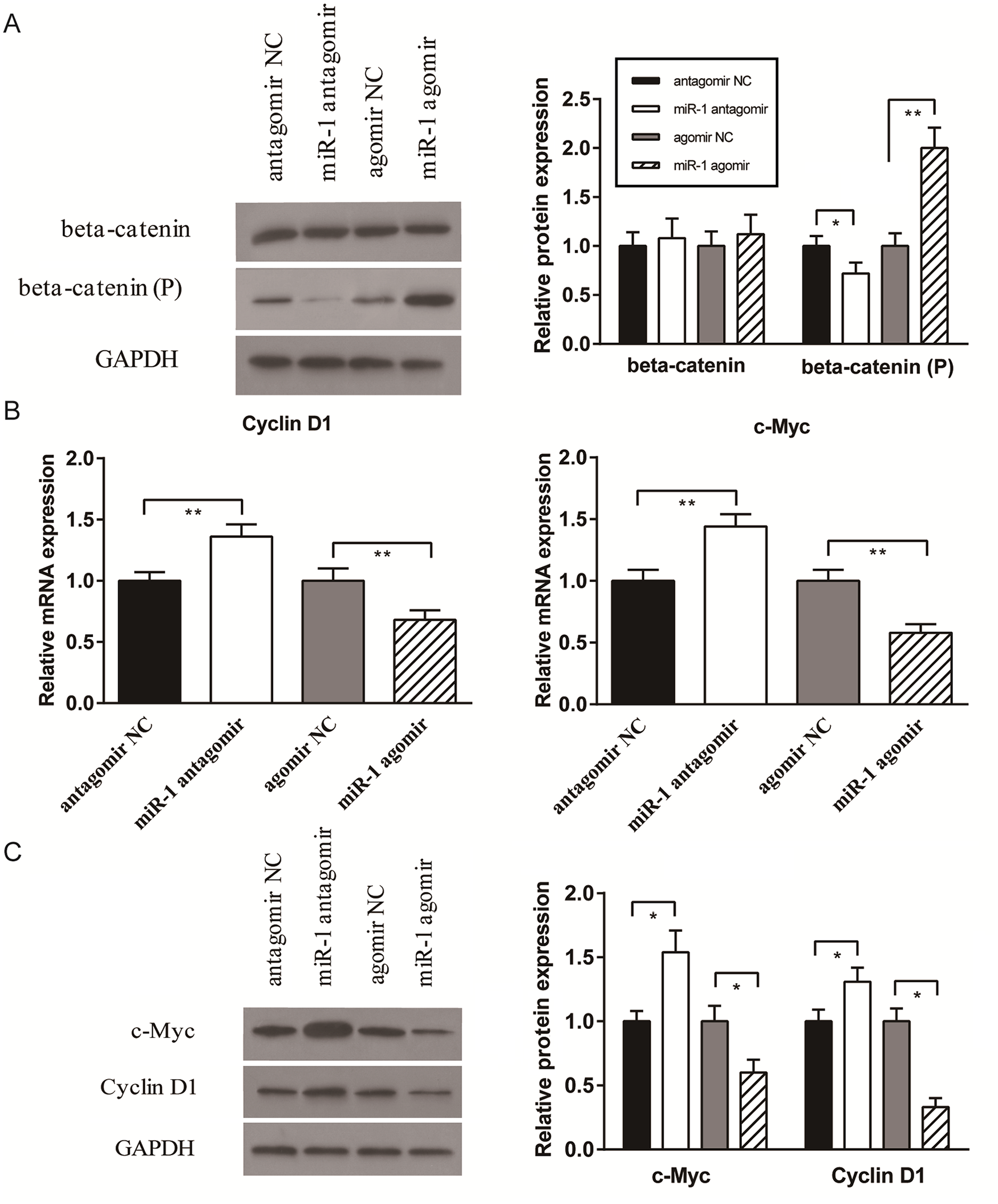
Effect of miR-1 on Wnt/β-catenin signaling pathway. The expression levels of (
In addition, we examined the mRNA and protein expression of Cyclin D1 and c-Myc, which are well-known downstream targets of Wnt/β-catenin signaling. For both Cyclin D1 and c-Myc, mRNA (
Discussion
The pathogenesis of OA is characterized primarily by cartilage degradation accompanied by local inflammation. As one of the key types of epigenetic mechanisms, miRNAs are now understood to have essential roles in modulating and maintaining normal cartilage development and homeostasis. 7 Normal levels of miRNA expression are disrupted in pathological disorders such as OA, either as a cause or consequence of disease pathogenesis and progression. 12 Over 25 different miRNAs have been identified as either having altered expression in OA or functional implications in OA pathogenesis.16,17 Interestingly, miR-1 has been identified as having a key role in the regulation of chondrocyte hypertrophy, 27 but its target in chondrocytes is unknown, and its effects on OA-related gene expression in chondrocytes have not been investigated. This is the first study to show that miR-1 is differentially expressed in OA compared with non-OA chondrocytes, and its underexpression in OA may lead to increased activation of the Wnt/β-catenin pathway that then modulates the expression of key genes implicated in OA pathogenesis.
We found that the transcription levels of several key proteins were significantly different between OA and non-OA chondrocytes, including cartilage degrading enzymes (MMP-9, MMP-13, and ADAMTS5), markers of cartilage repair (BMP-2) 31 and cartilage matrix proteins (COL2A1). These results corroborated with an increasing number of studies examining global gene expression profiles of OA compared with non-OA human cartilage samples, which reported differential expression of both anabolic and catabolic matrix genes.32,33 The OA chondrocytes in our study also showed significantly lower miR-1 expression compared to non-OA chondrocytes, suggesting that its underexpression in OA may be linked to disease pathogenesis.
To elucidate the link between miR-1 expression and OA-associated gene transcription, we studied the effects of miR-1 over- and underexpression in chondrocytes by transfecting SW1353 cells with a miR-1 agonist and antagonist, respectively. Although SW1353 is a chondrosarcoma cell line with limited similarities in gene expression profile compared with primary human chondrocytes, they have similar responses as primary chondrocytes to catabolic cytokines such as IL-1β, and have been used in many studies to investigate protease expression and regulation in chondrocytic cells, 34 or molecular mechanisms of pathogenesis in OA.35-37 In our study, we chose SW1353 cells for use as a reliable transfection host that replicated some of the key characteristics of chondrocytic cells. Using these cells, we found that miR-1 overexpression resulted in the downregulation of gene expression for MMP-9, MMP-13, and ADAMTS5, and essentially reversed the trends observed in primary OA chondrocytes with regard to the relative expression levels of OA-associated genes. These findings suggested an essential role of miR-1 in regulating the phenotype of chondrocytes and their expression of catabolic genes that may contribute to OA pathogenesis.
The molecular mechanisms responsible for OA progression are highly complex and not fully understood. Nevertheless, several signaling pathways are known to be involved in the development of OA, including mitogen-activated protein kinase (MAPK), nuclear factor κB (NF-κB), and Wnt/β-catenin. 24 Among these, the Wnt/β-catenin pathway is particularly interesting as it is involved in embryonic development of cartilage and bone and has a key regulatory role in joint remodeling. 38 The activation of Wnt/β-catenin signaling is thought to contribute to excessive cartilage matrix remodeling and degradation in joint pathologies. 24 Overexpressing or preventing the degradation of β-catenin protein has been shown to induce an OA-like phenotype in articular chondrocytes and mouse models, and significantly affect the expression of chondrocyte marker genes, including MMPs, ADAMTSs, aggrecan, and collagen type II.39,40 Studies have shown that molecular 41 or pharmacological42,43 agents inhibiting Wnt/β-catenin signaling in chondrocytes reduced the expression of matrix-modulating enzymes including MMPs, which might contribute to dampening disease progression in OA. It is well established that the Wnt/β-catenin pathway involves the interactions of Wnt ligands with Frizzled receptors on the cell membrane. 44 In the absence of Wnt ligand-receptor interactions, β-catenin is phosphorylated and subsequently undergoes proteasomal destruction. On Wnt ligand-receptor interaction with Frizzled receptors, β-catenin is stabilized, accumulates, and can translocate to the nucleus to modulate the expression of specific target genes.
In our study, FZD7 (Frizzled-7) was predicted to be a prime target of miR-1. Since our findings suggested that miR-1 had an important role in regulating OA-related gene expression in chondrocytic cells, we proceeded to investigate whether the effects of miR-1 were mediated by its binding to FZD7. This would inhibit Wnt binding to FZD7 and prevent β-catenin activation, 45 thereby reducing the downstream expression of catabolic genes. Although blocking FZD7 to reduce Wnt/β-catenin signaling has been performed in other studies, which resulted in growth inhibition in a range of tumor types,46,47 modulation of this same pathway has never been attempted in OA research. For the first time, our study demonstrated an important functional link between miR-1-FZD7 binding and Wnt/β-catenin signaling in chondrocytes. The regulation of catabolic gene expression through this pathway might contribute to changes in chondrocyte phenotype and play a role in the development of OA. We confirmed that miR-1 has a specific binding site in FZD7, and miR-1 binding reduces the mRNA and protein expression of FZD7 in chondrocytic cells. This results in increased levels of phosphorylated β-catenin that suppresses downstream β-catenin signaling. Other studies have shown that reduced activation of the Wnt/β-catenin pathway is associated with the downregulation of catabolic enzymes implicated in cartilage degradation and OA progression.39,40
There were some limitations in our study. First, although the findings from the luciferase reporter assay and gene expression analysis suggested potential interactions between miR-1 and the 3′UTR of FZD7, the results were relative and did not indicate direct binding. The precise mechanism for miR-1 regulation of Wnt/β-catenin signaling through FZD7 needs to be confirmed in future studies. Second, chondrocyte hypertrophy plays a key role in the development and progression of OA, which is partly associated with the activation of Wnt/β-catenin signaling.48,49 Although all of the gene markers we have examined in this study are relevant to chondrocyte hypertrophy, namely MMP-9, MMP-13, BMP-2, ADAMTS5, and COL2A1, there are other markers that are closely associated with hypertrophic differentiation, such as COL10A1 and RUNX2, as well as a range of markers associated with other pathways, including inflammatory signaling and mineralization. 48 Given the known association between miR-1 and late-stage chondrocyte differentiation, 27 it would be highly relevant to investigate the possible pathways by which miR-1 may regulate chondrocyte hypertrophy through Wnt/β-catenin signaling in future studies. Third, our study was performed using a chondrocytic cell model that does not fully replicate the characteristics of native tissues. This model does not allow the demonstration of a causal connection between miR-1 induced alterations in Wnt/β-catenin signaling and changes in cartilage degradation at the tissue level. The in vitro effects of miR-1 overexpression observed in this study will need to be verified in human OA cartilage explants, as well as in pathologically relevant in vivo models of OA.
To our knowledge, this study provides the first evidence of a functionally important role for miR-1 regulation of the Wnt/β-catenin pathway in chondrocytes. By inhibiting Wnt/β-catenin signaling through FZD7, miR-1 overexpression in chondrocytes can lead to changes in gene expression that potentially contribute to attenuating OA pathogenesis. It should be noted that a range of miRNAs implicated in cancer have been shown to regulate Wnt/β-catenin signaling through FZD7 and other Frizzled receptors, 50 some of which may be associated with pathways relevant to OA development and warrant further investigation. Future studies will be necessary to explore the functional and clinical implications of miR-1 and other relevant miRNAs in OA treatment. Although several clinical trials are currently ongoing for miRNA-based treatment of several diseases, including hepatitis C, liver cancer, and heart failure,51,52 miRNA-based treatment of OA will present unique challenges due to the relative inaccessibility of chondrocytes within the joint structure. Nevertheless, our work provides new evidence for the role of miR-1 in the regulation of chondrocyte phenotype and gene expression, which may be useful for future studies exploring epigenetic mechanisms of OA pathogenesis and developing related treatments.
Supplemental Material
sj-pdf-1-car-10.1177_1947603520973255 – Supplemental material for MicroRNA-1 Modulates Chondrocyte Phenotype by Regulating FZD7 of Wnt/β-Catenin Signaling Pathway
Supplemental material, sj-pdf-1-car-10.1177_1947603520973255 for MicroRNA-1 Modulates Chondrocyte Phenotype by Regulating FZD7 of Wnt/β-Catenin Signaling Pathway by Yang Yang, Yawei Wang, Haobo Jia, Bing Li, Dan Xing and Jiao Jiao Li in CARTILAGE
Footnotes
Acknowledgments and Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants from the National Natural Science Foundation of China (81973606, 81501919 and 81702208), the Australian National Health and Medical Research Council (GNT1120249), and Peking University People’s Hospital Scientific Research Development Funds (RDY2019-11).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This research was approved by the Ethics Committee of Tianjin Hospital, China (2014-002). All methods were carried out in accordance with relevant guidelines and regulations.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
