Abstract
Objective
Methods
Active components and potential targets of Yadanzi were retrieved from the Traditional Chinese Medicine Systems Pharmacology Database (TCMSP). BC-related genes were collected from public databases. A compound-target network was constructed using Cytoscape. Furthermore, protein-protein interaction (PPI) network construction, along with Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses, were performed to identify hub targets and signaling pathways. Finally, molecular docking,
Results
Fifteen active components of Yadanzi were screened from the TCMSP database. After excluding compounds without target information, two key bioactive compounds (beta-sitosterol and luteolin) and 73 potential targets were retained, among which 71 were associated with BC. Network analysis revealed AKT1, CASP3, and TP53 as hub targets. Enrichment analysis indicated these genes are primarily involved in apoptosis regulation, PI3K-Akt signaling, and other cancer-related pathways. Molecular docking demonstrated strong binding affinity between luteolin and AKT1/CASP3/TP53, as well as between beta-sitosterol and CASP3, with binding energies ranging from −6.4 to −7.8 kcal/mol.
Conclusion
This study provides a novel network pharmacology-based framework for elucidating the anti-BC mechanisms of BJOE. It was suggested that the key active component luteolin can inhibit BC cell proliferation, induce apoptosis and regulate the expression of related proteins associated with apoptosis. By integrating computational predictions with experimental validation, our work offers deeper insights into the pharmacological basis of BJOE and supports its potential as a complementary therapeutic agent for breast cancer.
Introduction
Since the mid-2000s, breast cancer incidence rates in females have increased gradually at an annual rate of approximately 0.6%. 1 According to Global cancer statistics 2022, breast cancer remains the most prevalent malignancy among women globally, accounting for 11.6% of new cancer cases. 2 Current treatments include surgery, chemotherapy, radiotherapy, and targeted therapies,3,4 but challenges such as toxicity and resistance persist. Traditional Chinese medicine, as a natural anti-cancer agent, has emerged as a promising alternative treatment option due to its efficacy, reduced side effects, and low toxicity.
TCM has been widely used in all kinds of malignant tumors for many years. It's reported that TCM can possibly regulate oncogenes and tumor suppressor genes, epigenetic modification, the microenvironment and cancer stem cells thus achieving antitumor effects.5–7 Unexceptionally, TCM plays an irreplaceable role in the field of breast disease. Li et al reported that Chinese patent medicines such as Rupi Sanjie Capsules and Xiakucao Oral Liquid could significantly improve the clinical efficacy in resolving hard lump for cyclomastopathy compared with the simple western medicine. 8 Aiduqing formula inhibits breast cancer metastasis by suppressing TAM/CXCL1/Treg pathway. 9 Icariin as a kind of plant-derived natural drug could effectively trigger apoptosis and inhibit the migration of breast cancer cells via the SIRT6/NF-κB signaling pathway. 10 It is worth noting, however, that the clinical application of TCM is often accompanied by certain challenges, such as non-uniform quality standards, variations in bioavailability, and a relative scarcity of large-scale clinical validation. Further research remains necessary to establish standardized protocols and clarify pharmacological mechanisms.
Yadanzi, derived from the dried mature fruit of
Due to the diverse active substances and complex action targets of TCM, previous studies have often struggled to pinpoint specific bioactive compounds and their precise mechanisms of action. This lack of mechanistic clarity has limited the translation of TCM findings into clinically applicable therapies, particularly under the evidence-based standards of modern oncology. Network pharmacology, an emerging field integrating pharmacology, computer science, and bioinformatics, has been increasingly recognized as a robust approach for TCM research. 21 To our knowledge, no prior studies have investigated the molecular mechanisms of Yadanzi oil in breast cancer using network pharmacology. This study firstly employs an integrated network pharmacology and experimental approach to systematically identify the active components of Yadanzi oil and their interactive targets, aiming to bridge the gap between traditional use and mechanistic validation. Graphical abstract of this research was shown in Figure 1.

Graphical abstract of this research.
Materials and Methods
Collection of Active Components and Related Targets of Yadanzi
Effective components of Yadanzi were identified from the Traditional Chinese Medicine Systems Pharmacology Database (TCMSP; https://tcmsp-e.com/tcmsp.php). Based on ADME principles (absorption, distribution, metabolism, and excretion), active components were selected using widely adopted pharmacokinetic criteria 22 : oral bioavailability (OB) ≥ 30% and drug-likeness (DL) ≥ .18. Associated targets of Yadanzi were also obtained from the TCMSP database. A Perl package was employed to integrate these components and targets of Yadanzi for downstream analyses. To standardize nomenclature, target proteins were processed through the UniProt database (https://www.uniprot.org/) to obtain official gene symbols.
Identification of BC-Related Targets
With “breast cancer” applied as search term, the target genes related to BC were retrieved from GeneCards (https://www.genecards.org/), Online Mendelian Inheritance in Man (OMIM; https://omim.org/), PharmGkb (https://www.pharmgkb.org/), Therapeutic Target Database (TTD; http://db.idrblab.net/ttd/) and DrugBank (https://www.drugbank.ca/). After removing duplicate target genes, a Venn diagram was generated via Venny 2.1.0 to integrate genes from all databases. To identify overlapping targets between Yadanzi and BC-related genes, a corresponding Venn diagram was subsequently generated. The overlapping genes designated as drug-disease targets were identified as potential anti-BC targets of Yadanzi.
Construction of the Active Components-Target Network
Cytoscape 3.8.0 was used to construct a composite components-target network of Yadanzi against BC. Relationships between active components and potential targets were analyzed using Cytoscape's built-in network analysis tool. Nodes and edges were generated by the software. The “degree” value, defined as the number of edges per node, was used to quantify node importance. Core active components were identified by calculating their degree values. As such, this network provided a more visual and quantitative framework to map Yadanzi's anti-BC targets.
Construction of the Protein-Protein Interaction (PPI) Network
Overlapping genes (designated as drug-disease targets) were imported into the STRING database (https://string-db.org/) with an interaction score set to > .9 and species restricted to “
GO and KEGG Analysis
Entrez IDs of overlapping genes were obtained through the Bioconductor package org.Hs.egdb in R software (version 4.4.1). The org.Hs.egdb and clusterProfiler packages were used in R for Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses of overlapping genes. GO functional enrichment includes biological process (BP), cellular component (CC), and molecular function (MF) of potential target genes. The Pathview package was applied to generate signaling pathway maps, visually illustrating target-pathway interactions. Significantly enriched GO and KEGG terms were screened using thresholds of
Molecular Docking
Molecular docking was conducted to investigate interactions between molecules and their protein receptors. The active components of Yadanzi served as small molecule ligands, while target genes acted as receptors. The 2D structures of these ligands were available from the PubChem website (http://pubchem.ncbi.nlm.nih.gov/). Their structures were converted to 3D format using Chem3D Ultra 14.0, and saved in mol2 format. Subsequently, 3D structures of the screened core genes were downloaded from the PDB database (https://www.rcsb.org/) and imported into PyMOL 2.4.0 for protein dehydration and ligand removal. Using AutoDockTools 1.5.6, hydrogen atoms were added to the proteins with Gasteiger charges assigned, and non-polar hydrogens were merged. The Kollman united atom charge model was applied for protein charge calculation. The grid box dimensions were set to 40 × 40 × 40 Å and the center of the grid box was positioned around the active site of each protein. Both ligands and receptors were converted to “pdbqt” format for active pockets identification. Finally, Vina 1.5.6 was run to evaluate molecular docking and the visualization of docking results was performed using PyMOL 2.4.0.
Cell Lines and Reagents
The human BC cell lines MDA-MB-453 (HTB-131) and MDA-MB-231 (CRM-HTB-26) were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). These cells were cultured in Dulbecco's modified Eagle medium (DMEM; Gibco, China) supplemented with 10% fetal calf serum (Lonsera, China) and 1% penicillin-streptomycin (Servicebio, China). The cells were incubated at 37 °C in a humidified incubator containing 5% CO2. Luteolin was purchased from MedChemExpress Co., Ltd (China; Cat. No. HY-N0162, purity: 99.51%).
Cell Viability Assay
BC cells were plated in 96-well plates at a density of 3000 cells per well. To determine the IC50 of luteolin against BC cells, cells were treated with fresh culture medium containing luteolin at concentrations of 1, 10, 20, 50, 100, 200, or 400 μM, or with vehicle control (0.1% DMSO in medium) for 48 h. To measure cell viability, the cells were treated with luteolin (20, 40, or 80 μM) and incubated at 37 °C for further detection. The absorbance of each well at 450 nm was measured by Pro-11 Multiskan FC (Thermo Fisher, USA) after incubation with Cell Counting Kit-8 (CCK-8, Yeasen, China) solution following the manufacturer's instructions. Each assay was performed in triplicate.
Wound Healing Assay
Cells were seeded in 6-well plates at a density of 8 × 104 cells/well and allowed to attain confluent monolayers. The cells were then treated with luteolin at various concentrations and incubated. A line or a shape of “≡”was drawn at the center of each well using a pipette tip to create the wound area. Images were captured using an Olympus CKX53 microscope (Olympus, China) at 0, 24 and 48 h after scratching. The wound healing rates were quantitatively analyzed by measuring the changes in the scratch area over time using ImageJ software.
Cell Apoptosis Assay
Cell apoptosis analysis was performed using a FITC-Annexin V Apoptosis Detection Kit (Yeasen Biotech, China). Cells were harvested by mild trypsin digestion and dual-stained with FITC-Annexin V and propidium iodide according to the manufacturer's instructions. After dissociation from 6-well plates, cells were resuspended in flow cytometry staining buffer (Thermo Fisher, USA). Afterwards, the cells were labelled with conjugated antihuman CD44-Phycoerythrin (PE), CD24-Fluorescein Isothiocyanate (FITC), or Ep CAMPE (Thermo Fisher, USA), incubated for at least 30 min on ice in darkness, and then washed twice with flow cytometry washing buffer. Finally, apoptosis was assessed using a CytoFlex LX flow cytometer (Beckman Coulter, USA).
Western Blot Analysis
MDA-MB-453 and MDA-MB-231 cells were treated with luteolin (20 μg/mL) for 48 h and total protein was extracted using RIPA lysis buffer (Beyotime, China). BCA Protein Assay Kit (Beyotime, China) was utilized to detect the concentration of protein. The total protein (20 μg) was separated by 7.5% or 10% SDS–PAGE (Epizyme, China) and then transferred onto polyvinylidene fluoride (PVDF) membranes (Millipore). QuickBlock™ Blocking Buffer for Western Blot (Beyotime, China) was applied in blocking PVDF membranes. Antibodies specific for the following proteins were applied: BAX (1:20000, 50599-2-Ig, Proteintech), BCL2 (1:5000, 12789-1-AP, Proteintech), STAT3 (1:5000, 80149-1-RR, Proteintech), p-STAT3 (1:2000, 28945-1-AP, Proteintech), GAPDH (1:50000, 60004-1-Ig, Proteintech), β-actin (1:20000, 66009-1-Ig, Proteintech).β-actin or GAPDH served as the internal control.
In Vivo Tumor Growth Model
Five-week-old female BALB/c nude mice (purchased from GemPharmatech LLC) were used to establish a xenograft model. Briefly, MDA-MB-231 cells (5 × 106) suspended in 200 µL of phosphate-buffered saline (PBS) were inoculated subcutaneously into the right flank of each mouse. When the tumor volume reached an approximate diameter of 0.5-0.6 cm, the tumor-bearing mice were randomly assigned into two groups (n = 5 per group):The control group received an equal volume of saline via oral gavage once daily. The luteolin treatment group received luteolin (20 mg/kg) via oral gavage once daily. The treatment regimen consisted of five doses, administered every other day. After the final treatment, all mice were euthanized by cervical dislocation. Tumors were carefully excised and harvested. Tumor volume was measured using a digital caliper and calculated according to the formula: V = 1/2 × length × width². Tumor weight was measured using an electronic balance.
Statistical Analysis
The experimental data were shown as the mean ± standard deviation. GraphPad Prism 6.0.1 was used for the statistical analysis. The differences between groups were determined by one-way ANOVA or Student's t test. A significant difference was assumed if *
Results
Screening of Active Components and Targets of Yadanzi Against BC
With oral bioavailability (OB) ≥ 30% and drug-likeness (DL) ≥ 0.18 as screening criteria, we identified 15 active components of Yadanzi from the TCMSP database (Table 1). Associated targets for Yadanzi's active components were also obtained from the TCMSP database. By applying a Perl package, we integrated the components with the targets, ultimately identifying 2 active components (beta-sitosterol and luteolin) and 73 potential targets of them (Table S1) after excluding 13 components lacking target information. Official symbols of all potential genes were obtained through the UniProt database.
Active Components of Yadanzi Identified from the TCMSP Database.

Using “breast cancer” as the search term, we retrieved 15692 (GeneCards), 414 (OMIM), 296 (PharmGKB), 97 (TTD), and 60 (DrugBank) BC-related genes. After deduplication, we obtained 15787 unique BC-related genes and plotted a Venn diagram to reveal the overlaps between databases (Figure 2A). As is shown in Figure 2B, intersection of the 73 potential targets with 15787 BC-related genes yielded 71 overlapping genes, identifying Yadanzi's critical targets against BC.

Network pharmacological research on Yadanzi against breast cancer. (A) Venn diagram of the overlaps of BC-related genes from different databases (GeneCards, OMIM, TTD, PharmGkb and DrugBank). (B) Venn diagram of Yadanzi-BC interaction targets. (C) “Yadanzi-active component-targets” network: the green nodes represent the active components and the blue nodes represent the target genes, with gray edges indicating that nodes can interact. (D-F) PPI network and core target genes.
“Yadanzi-active components-targets” network
The “Yadanzi-active components-targets” network was built using Cytoscape 3.8.0 to demonstrate how Yadanzi counteracts BC. The outer circle graphs represent the targets, the inner graphs display active components (Figure 2C). Node sizes were proportional to their degree values. The network contained 73 nodes: 2 active components and 71 targets, connected by 77 edges. Network topology analysis revealed luteolin (MOL000006; DC = 52) demonstrated more interconnection relationships with the targets than beta-sitosterol (MOL000358; DC = 25), suggesting its predominant role in Yadanzi's anticancer activity.
PPI Network Construction and Analysis
Using the 71 overlapping targets, we constructed the protein-protein interaction (PPI) network through the STRING database with confidence score ≥ 0.9 and disconnected nodes hidden (Figure 2D). The PPI network included 57 nodes and 160 edges, with an average node degree of 5.61 and a local clustering coefficient of 0.66. Based on CytoNCA function, we conducted topological analysis measuring six parameters: Betweenness Centrality (BC); Closeness Centrality (CC); Degree Centrality (DC); Eigenvector Centrality (EC); Local Average Connectivity (LAC); and Neighborhood Connectivity (NC). Nodes with all 6 parameter values exceeding the mean were filtered for the next screening round. Through two screening rounds, three core target genes were filtered: AKT1, CASP3 and TP53 (Figure 2E-F).
GO and KEGG Pathway Enrichment Analysis
GO and KEGG pathway enrichment analyses were performed on 71 overlapping targets using the Bioconductor package and clusterProfiler package in R. GO analysis included biological process (BP), molecular function (MF) and cellular component (CC) analysis with significance thresholds of

GO and KEGG analysis. (A) Bar graph of GO enrichment analysis results of biological process (BP), molecular function (MF) and cellular component (CC). (B) Bubble chart of the top 10 terms extracted according to the p.adjust value based on GO enrichment analysis. (C) Bar graph of KEGG enrichment analysis of the overlapping targets. (D) Bubble chart of the top 30 pathways extracted according to the p.adjust value based on KEGG enrichment analysis.
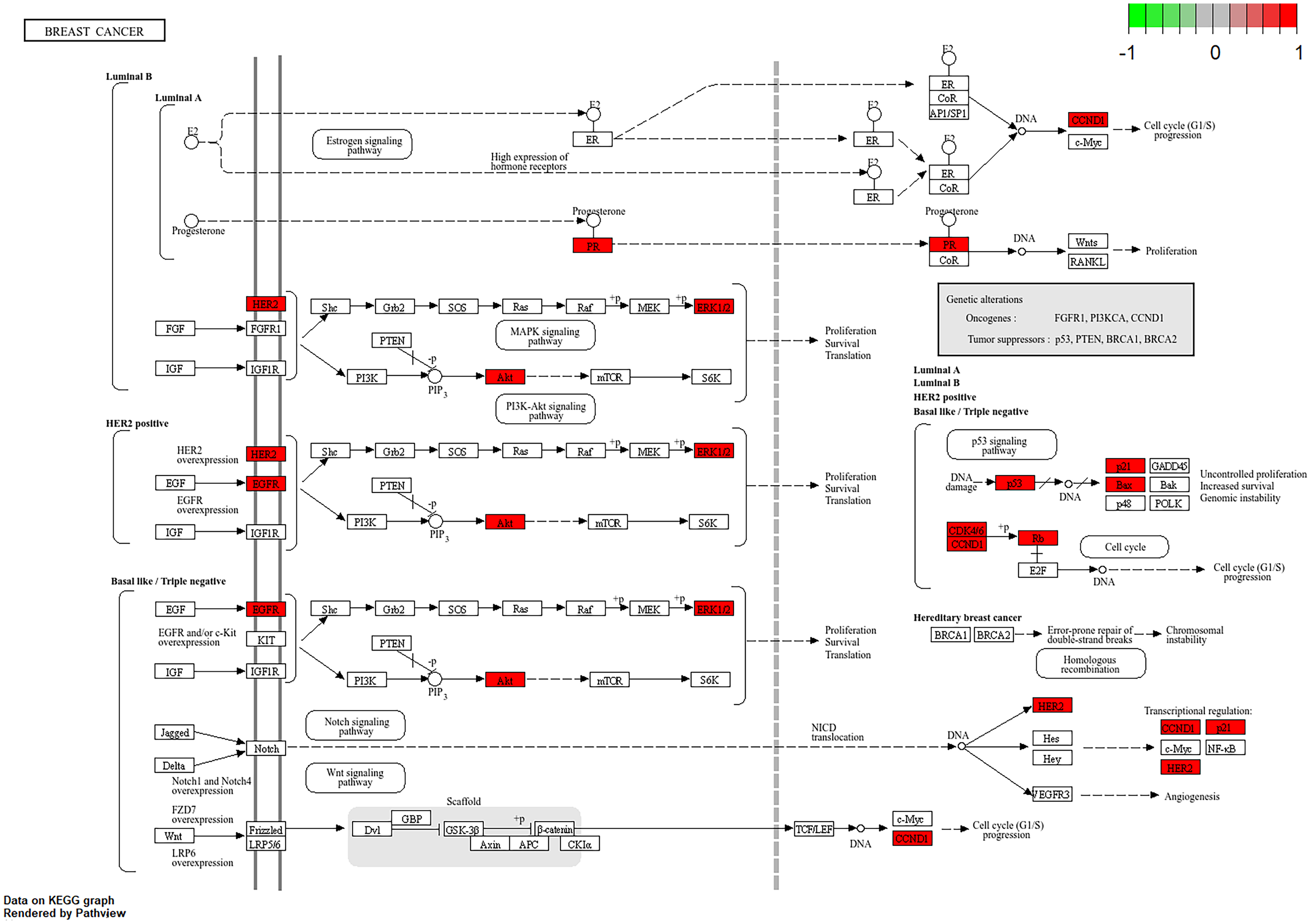
Signaling pathway map of BC targets treated with Yadanzi. The pathway diagram is classified by the type of breast cancer. The red nodes represent the most potential targets, arrows represent the activation effect, T arrows represent the inhibition effect, and segments show the activation effect or inhibition effect.
Molecular Docking Analysis
After excluding 13 components lacking target information, beta-sitosterol and luteolin as small-molecule ligands were docked with three core target proteins (AKT1, CASP3, TP53) to simulate component-target interactions. The results suggested that luteolin can form a stable docking model with AKT1, CASP3 and TP53 (Figure 5A-C); beta-sitosterol can dock with CASP3 protein ligands (Figure 5D). The docking data were analyzed by AutoDockTools 1.5.6, which included the center position and the dimensions (length × width × height) of the grid box according to the interaction site between the small molecule and the target. Furthermore, the binding energy was calculated.

3D Molecular docking model of the two key active components with the three hub-targets. (A) Luteolin and AKT1. (B) Luteolin and CASP3. (C) Luteolin and TP53. (D) Beta-sitosterol and CASP3.
Binding energies < −5 kcal/mol indicate strong interactions, with lower values correlating to greater stability. The binding energy of luteolin with AKT1, CASP3 and TP53 was −6.4 kcal/mol, −7.8 kcal/mol and −6.4 kcal/mol, respectively. The binding energy of beta-sitosterol with CASP3 was −7.3 kcal/mol.
Validation of the Effect of Luteolin in BC Cells
To validate Yadanzi's anti-BC effects, we selected luteolin for experimental validation, which is high ranking in our network pharmacology analysis and commercially available. As shown in Figure 6A, luteolin demonstrated concentration-dependent cytotoxicity with IC50 values of 76.47 μM (MDA-MB-231) and 85.25 μM (MDA-MB-453). CCK-8 assay showed that luteolin inhibited the proliferation of MDA-MB-231 and MDA-MB-453 cells, in a time- and dose-dependent manner (Figure 6B). To determine the effect of luteolin on metastatic capacity of BC cells, we performed a wound healing assay. The results indicated that luteolin significantly reduced the migration ability of MDA-MB-231 and MDA-MB-453 cells (Figure 6C and Figure S1). Flow cytometry was also conducted to measure the effects of luteolin on BC cells. Luteolin potently induced apoptosis in both cell lines (Figure 6D).

Validation of the effect of luteolin in BC cells. (A) IC50 of luteolin. (B) Luteolin shows a dose- and time-dependent effect on the viability of BC cells. (C) Luteolin shows the ability to inhibit the migration of BC cells. (D) Luteolin treatment significantly induced apoptosis of BC cells. The data are shown as the mean ± SD of three experiments. *P <0.05, **P < 0.01, ***P < 0.001 compared with the control group.
Subsequently, the expression levels of apoptosis-related proteins in MDA-MB-231 and MDA-MB-453 cells were detected by Western blot. The results indicated that the expression of BAX was upregulated, but the expression of BCL2 was downregulated by treatment with luteolin (Figure 7A). It's reported that STAT3 has an oncogenic role and may inhibits apoptosis by regulating the expression of its downstream target genes like BCL2 and is also regulated by the PI3K-Akt signaling pathway. 23 As an intermediate key factor, the level of STAT3 and phosphorylated STAT3 (pSTAT3) were also measured. As shown in Figure 7B, after treated with luteolin, the level of STAT3 was downregulated, whereas the levels of the pSTAT3 was increased. Therefore, we hypothesis that luteolin may depressed the progression and induced the apoptosis of BC cells by inhibiting STAT3 and promoting its phosphorylation.

Verification of the effect of luteolin at the molecular level and animal level. (A) The expression levels of BAX and BCL2 were detected by Western blot. (B) The expression levels of STAT3 and pSTAT3 were measured by Western blot. (C) MDA-MB-231 cells were subcutaneously injected into nude mice (n = 5 mice per group for different treatment) and tumors were excised and photographed. (D) Comparison of the tumor weight between the two groups. (E) Comparison of the tumor volume between the two groups.
In Vivo Tumor Growth Model
The antitumor effects of luteolin were further investigated
Discussion
Previous studies have explored the potential anticancer effects of the Yadanzi oil and its components in cancer treatment such as lung cancer and gastric cancer. Yadanzi oil was reported to have significant inhibitory effects on cell growth and induced apoptosis.12,24 Compounds of Yadanzi also presented excellent anticancer activity.
For example, luteolin (3,4,5,7-tetrahydroxy flavone), a flavonoid found in vegetables, medicinal herbs and fruits, has antioxidant and pro-oxidant, as well as antimicrobial, anti-inflammatory, and cancer-preventive properties. 25 Luteolin has also been reported to repress androgen receptor-positive TNBC by inactivating the AKT/mTOR signaling pathway and reversing the epithelial-mesenchymal transition (EMT). 26 Wu et al put forward that luteolin may inhibit TNBC by inducing apoptosis and autophagy through SGK1-FOXO3a-BNIP3 signaling. 27 Additionally, beta-sitosterol, another component of Yadanzi, has also been studied for its potential antitumor effects.28,29 Although the anticancer activity of Yadanzi and its compounds has been extensively investigated, the effects and potential mechanisms of Yadanzi in breast cancer have not been researched.
In this study, we identified 15 bioactive components and 158 associated targets in Yadanzi through the TCMSP database. The component-target integration identified 2 active components (beta-sitosterol and luteolin) and 73 potential targets. From public databases, we collected 15787 BC-related targets, and 71 candidate targets of Yadanzi for BC treatment were identified by using Venn diagram analysis. We further constructed the PPI network to visualize the interaction of the proteins and identified AKT1, CASP3 and TP53 as core therapeutic targets in Yadanzi's anti-BC effects. Molecular docking analysis revealed that the key active components (beta-sitosterol and luteolin) can bind stably with targets (AKT1, CASP3 and TP53).
AKT1, one of the isoforms of the AKT family, has been the most widely studied effector of PI3 K signaling and the focus of numerous studies in diverse fields of biology and medicine. 30 In the CAPItello-291 trial, the selective AKT inhibitor capivasertib combined with fulvestrant therapy improved progression-free survival (PFS) in patients with hormone receptor-positive, HER2-negative advanced breast cancer. 31 Wu et al proposed that the receptor tyrosine kinase HER2 recruits AKT1 to disrupt STING signaling and suppress the surveillance of cellular damage and antitumor immunity. 32 Additionally, previous studies indicate that AKT1 plays an indispensable role in tumors such as gastric cancer, 33 colorectal cancer 34 and ovarian cancer. 35 Caspases are crucial mediators of programmed cell death (apoptosis). Among them, caspase-3 (CASP3) is a frequently activated death protease that catalyzes the specific cleavage of many cellular proteins and participates in physiological and pathological processes.36–39 Apoptosis is typically considered an anti-oncogenic process as caspase activation promotes the elimination of genetically unstable or damaged cells. 40 Due to its critical role in apoptosis, CASP3 has become a primary target for cancer treatment, with numerous components reported and patented for their caspase-3-mediated cytotoxicity. 41 The TP53 gene is a critical tumor suppressor that plays a fundamental and multifaceted role cancer development and cancer therapy. 42 Its importance in tumor prevention is demonstrated by the frequent detection of TP53 genetic alterations through immunohistochemical testing in human cancers, including breast cancer. As a transcription factor, TP53 regulates diverse pathways such as cell cycle arrest, DNA repair, apoptosis, autophagy, and metabolism. 43 Apoptosis is among the most extensively studied cellular processes induced by the TP53 protein and has been the focus of much of the investigation into its tumor-suppressor effect. 44 Therefore, the aforementioned targets may critically contribute to BC pathogenesis and represent potential therapeutic targets for Yadanzi in BC treatment.
To explore the underlying molecular mechanisms and signaling pathways of Yadanzi in BC treatment, we performed GO and KEGG enrichment analyses. The results indicated that epithelial cell proliferation, regulation of apoptotic signaling pathway, cyclin-dependent protein serine/threonine kinase activity, PI3K-Akt signaling pathway, p53 signaling pathway, and TNF signaling pathway may represent potential mechanisms of Yadanzi in BC treatment. The PI3K-Akt signaling pathway, the most frequently mentioned pathway in this study, was both associated with the active component luteolin and the key target AKT1. It is reported that the PI3K-Akt signaling pathway plays an essential role in diverse cellular processes and is aberrantly activated in cancers, which promotes tumor development and progression. 45 Consistently, both the pathways and the three core target genes relate to the pathophysiological process of cancer.
To validate these computational predictions,
Nevertheless, this study has several limitations. First, using different databases may introduce variations in target collection and data processing, potentially leading to bias. Second, the assessment of cytotoxicity in non-cancerous cell lines, comprehensive pharmacokinetic investigations and systematic toxicological assessments were required to enhance clinical relevance. Moreover, follow-up experiments using CRISPR-Cas9 knockdown or siRNA-mediated silencing of the targets (eg, AKT1, CASP3, and TP53) to explicitly test their necessity in mediating Yadanzi's effects will form the basis of our subsequent research efforts. What's more, while luteolin was subjected to a more detailed mechanistic validation, beta-sitosterol represents another interesting component worthy of investigation in future work.
Conclusion
In summary, our study is the first to systematically investigate the anti-BC effects and potential mechanisms of Yadanzi through network pharmacology, molecular docking,
Supplemental Material
sj-xlsx-1-npx-10.1177_1934578X251409306 - Supplemental material for Mechanistic Exploration and Experimental Validation of Yadanzi Oil in Treating Breast Cancer Based on Network Pharmacology and Molecular Docking Technology
Supplemental material, sj-xlsx-1-npx-10.1177_1934578X251409306 for Mechanistic Exploration and Experimental Validation of Yadanzi Oil in Treating Breast Cancer Based on Network Pharmacology and Molecular Docking Technology by Xiaoqing Shi, Mingyu Wang, Donglian Xu, Jinpeng Chen and Jue Wang in Natural Product Communications
Supplemental Material
sj-jpg-2-npx-10.1177_1934578X251409306 - Supplemental material for Mechanistic Exploration and Experimental Validation of Yadanzi Oil in Treating Breast Cancer Based on Network Pharmacology and Molecular Docking Technology
Supplemental material, sj-jpg-2-npx-10.1177_1934578X251409306 for Mechanistic Exploration and Experimental Validation of Yadanzi Oil in Treating Breast Cancer Based on Network Pharmacology and Molecular Docking Technology by Xiaoqing Shi, Mingyu Wang, Donglian Xu, Jinpeng Chen and Jue Wang in Natural Product Communications
Footnotes
Acknowledgements
Grateful acknowledgement is made to Haiyan Shen and Lexin Wang who gave us considerable help by means of continuous support, mentorship, and training on this project.
Ethical Approval and Consent to Participate
All experimental procedures involving animals were approved by the Institutional Animal Care and Use Committee of Nanjing Medical University (IACUC-2309011) and conducted in accordance with the guidelines of the Jiangsu Center for Medical and Pharmaceutical Animal Experiments.
Author Contributions
XS and MW designed the research; MW performed the experiments; XS completed the data analysis and drafted the manuscript. JC, DX and JW conducted quality control on it. All authors have read and approved the final manuscript. The authors declare that all the data were generated in-house and that no paper mill was used.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was partially supported by the Chinese Society of Clinical Oncology Foundation (Y-HR2020MS-0421), Jiangsu Province Elderly Health Research Project (LKM2022033, LKM2023023), Wu Jieping Medical Foundation Special Fund for Clinical Research (320.6750.2023-11-29), and China Breast Surgery Young Physician Research Award Advantage Support Program (2020-CHPASLP-01).
Chinese Society of Clinical Oncology Foundation, Wu Jieping Medical Foundation Special Fund for Clinical Research, China Breast Surgery Young Physician Research Award Advantage Support Program, Jiangsu Province Elderly Health Research Project, (grant number Y-HR2020MS-0421, 320.6750.2023-11-29, 2020-CHPASLP-01, LKM2022033, LKM2023023).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data used to support this study are available from the corresponding author upon reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
