Abstract
Introduction
Acute lung injury (ALI) and its severe form, acute respiratory distress syndrome (ARDS), are key global determinants of morbidity and mortality in critically ill patients. This destructive disorder stems from diverse causes like pneumonia, sepsis, trauma, and COVID-19. Clinically, anti-inflammatory agents, particularly glucocorticoids, are often employed as the first-line therapy for ALI. Despite the limited efficacy of glucocorticoids in improving ALI survival, their prolonged use incurs severe complications and persistent sequelae, underscoring the urgent need for safe, effective, and cost-efficient therapies. 1
Acute lung inflammation featuring inflammatory cell infiltration and high-permeability pulmonary edema defines ALI, a condition where pro-inflammatory mediator accumulation is central to its pathogenesis. 2 Acute lung injury is the consequence of an inflammatory cascade initiated by over-secretion of pro-inflammatory cytokines, including tumor necrosis factor-α (TNF-α), interleukin (IL)-6, IL-1β, and IL-18.3,4 Research validates the IL-17/NF-κB signaling axis as central to lung inflammation regulation, whereby IL-17 drives inflammatory chemokine/cytokine expression through NF-κB—a pivotal transcription factor orchestrating the inflammatory cytokine cascade.5,6 NF-κB hyperactivation was identified in multiple inflammatory cell subtypes of sepsis and sepsis-induced ALI patients. 7 Therefore, precise regulation of inflammatory cascades represents a pivotal pharmacological approach for ALI management. 8
Oxalis corniculata L. belongs to the Oxalidaceae family, which is distributed in Asia, Europe, Africa, Oceania, South America and North America. Preclinical investigations show O. corniculata mitigates CCl4-induced acute hepatotoxicity by attenuating oxidative stress and inflammatory cascades, while concurrently protecting lung tissue via inhibiting blood cell degeneration, lipid peroxidation, and restoring antioxidant enzymes. 9 Considering excessive lung inflammation represents the key pathological hallmark of ALI, we propose O. corniculata could mitigate ALI.
This study's objective to validate the hypothesis drove its investigation into O. corniculata's therapeutic efficacy and molecular mechanisms in ALI. First, the LPS-induced ALI model in Sprague-Dawley (SD) rats was used to evaluate the alleviating effects of 70% alcohol extract from O. corniculate (EOC) on ALI. Next, LC-MS was employed to identify potential active compounds in EOC. Finally, To analyze EOC's therapeutic mechanisms in ALI, network pharmacology and western blotting were employed, with structural analysis verifying how its active constituents associate with target proteins (Figure S1). These findings may provide a foundation for developing novel therapeutic agents against ALI.
Experimental Materials
Instruments
The following instruments were used: LC-MS (UltiMate3000 / Q-Exactive Plus, ThermoFisher, USA), analytical balance (EL204, Mettler Toledo, Switzerland), rotary evaporator (R-300, BUCHI, Switzerland), embedding center (BMJ-A, Changzhou Zhongwei, China), rotary microtome (Leica-2016, Leica, Germany), optical microscope (BA210Digital, MOTIC, China), high-speed centrifuge (Multifuge X3R ThermoFisher, USA), CO2 incubator (Tianjing Taisote, MJX-150BIII, China), microplate reader (Varioskan LUX, ThermoFisher, USA).
Animal
In accordance with China's National Institutes of Health Care and Use of Laboratory Animals (1996 revision; approval no. 2001363), all procedures approved by Guizhou Medical University's Animal Ethics Committee after a 7-day acclimatization involved 8-week-old male SPF-grade SD rats (200-250 g) from Beijing HFK Bioscience Co., Ltd, housed at the university's Guizhou Provincial Key Laboratory of Pharmaceutics Experimental Animal Center. Animals were housed under standardized conditions during the study: 12-h light/dark cycle with free access to autoclaved rodent feed and water. This study adheres to the ARRIVE 2.0 guidelines for reporting. 10
Cells
RAW264.7 (18K29C022), a murine macrophage cell line, was procured from Wuhan Boster Biological Technology, Ltd
Crude Drugs
O. corniculata was harvested from the fields of Gaopo Township, Huaxi District, Guiyang City, Guizhou Province, and identified by Associate Professor Chunhua Liu of Guizhou Medical University as the whole herb of Oxalis corniculata L., which belongs to the family Oxalidaceae.
Drugs and Reagents
Lipopolysaccharide (LPS, from Escherichia coli O55:B5 (0000153963, Merck KGaA, Germany), dexamethasone (DEX, 5305022, Solarbio, China), nitric oxide (NO) assay kit (20230225, Nanjing Jiancheng, China), hematoxylin staining solution (G1004, Wuhan Servicebio, China), eosin Y solution (YE2080, Hefei Bomei, China), ELISA kits (TNF-α, ZC-37624; IL-6, ZC-36404; IL-18, ZC-36389; IL-1β, ZC-36391, ZCIBIO, China), and enhanced chemiluminescence (ECL, 34580, ThermoFisher, USA).
The reagents included alcohol (AR, Tianjin Kemio, China), acetonitrile (AR, Merck, Germany), formic acid (AR, ThermoFisher, USA), etc.
Experimental Methods
Preparation of the EOC
O. corniculata (1.2 kg) was cut into segments and extracted three times by reflux extraction with 70% ethanol (1:10, 1:8, 1:8, w/v), each time for 1.5 h. The EOC was acquired by reducing pressure to concentrate combined extracts, then drying (341 g, extraction yield: 28.4% w/w).
Drug Administration
SD rats were randomly allocated into six groups (n = 6 per group): control, model (LPS only), dexamethasone (DEX, 3.0 mg/kg), low-dose (EOC, 0.8 g/kg), middle-dose (EOC, 1.6 g/kg), and high-dose (EOC, 3.2 g/kg). The EOC was dosed based on human equivalent dose (HED) calculations using human-rat body surface area conversion. The reference human dose was 0.43 g crude herb/kg/day, which was derived from classical Chinese herbal pharmacopoeias the commonly used therapeutic dosage range for Oxalis corniculata in traditional practices for respiratory conditions. 11 This human dose was converted to yield a pharmacologically equivalent rat dose of 0.8 g EOC/kg/day (designated as the low dose). Middle and high doses were set at 2-fold (1.6 g/kg/day) and 4-fold (3.2 g/kg/day) of this rat dose, respectively. For 7 days, all treatments were intragastrically administered bidaily, the EOC suspension prepared in 0.5% carboxymethyl cellulose sodium (CMC-Na) (0.20 g/ml), control/model groups given equal volumes of 0.5% CMC-Na, and on day 7, LPS (10 mg/kg) was i.p. injected 30 min after final gavage, samples collected 6 h later.
Collection of Blood Samples and Lung Tissues
Post-drug administration, rats were anesthetized, whole blood obtained via abdominal aorta, euthanized under deep anesthesia, and lungs harvested for pathology.
Hematoxylin-Eosin (HE) Staining
Lungs were excised and immersed in 10% formaldehyde for fixation, then subjected to gradient dehydration. Small sections measuring 1 × 1 × 0.3 cm³ were cut from the lung tissues for paraffin embedding. Sections underwent slicing, bleaching, HE staining, and light microscope observation of lung tissue pathology.
Enzyme-Linked Immunosorbent Assay (ELISA)
Blood samples from each group were subjected to ELISA analysis for inflammatory factors (IL-6, TNF-α, IL-18, IL-1β), following the manufacturer's standard procedures.
Analysis of the Blood Components
Take 1 mL of the serum from both the EOC treated and control groups, add 500 μL 2% formic acid aqueous solution to each, vortex mix thoroughly, then add 4 times the volume of acetonitrile, vortex mix again. Samples underwent sonication (40 kHz, 10 min) and centrifugation (12,000 r/min), after which the supernatant was transferred to fresh tubes, nitrogen-dried at 37°C, reconstituted in 200 μL methanol, vortexed, and re-sonicated (40 kHz, 10 min). Finally, centrifuge at 12 000 r/min for 10 min to obtain the deproteinized serum samples.
The deproteinized serum samples were loaded onto a Hypersil GOLD C18 column (100 mm × 2.1 mm, 1.9 μm) at 40°C for the qualitative analysis of EOC components in rat serum after EOC administration. Using mobile phase A (water/0.1% formic acid) and B (acetonitrile) at 0.3 mL/min, gradient elution was optimized as: 5% B (0-1 min), linear 5%–30% B (1-15 min), 30%–60% B (15-25 min), 60%–95% B (25-28 min), with 1-µL injection. Fitted with an high-voltage electrospray ion source (H-ESI) source (3500 V spray voltage), the MS system had sheath gas at 30 Arb, Aux gas at 10 Arb, capillary temperature 350°C, Aux gas heater 320°C, and conducted full m/z scans (100-1500). The impact energy gradients are 20, 40, and 60 eV. The scanning mode is full scanning + positive and negative ion switching mode with data-dependent secondary scanning. The primary and secondary resolutions are 70000 and 17500, respectively.
Screening of Potential EOC Targets for Treating ALI
Canonical SMILES of identified blood components, after being searched in PubChem (https://pubchem.ncbi.nlm.nih.gov/), were imported into Swiss Target Prediction (https://www.swisstargetprediction.ch/). Following the removal of duplicate and zero-probability targets, Cytoscape 3.9.0 was used to build the compound-target network. ALI-related targets were collected fromthe herapeutic target (TTD) database (https://db.idrblab.net/ttd/), online mendelian inheritance in man (OMIM) database (https://omim.org/), GeneCards database (https://www.genecards.org/), tand disease gene network (DisGeNET) database (https://www.disgenet.org/home/). Duplicate targets were removed, and component-disease overlapping targets were identified via Venn diagram.
Construction and Analysis of the Protein–Protein Interaction (PPI) Network
Overlapping targets and candidate genes were defined as EOC's potential ALI-treatment targets, with PPI data retrieved from STRING (https://string-db.org/). The PPI network was constructed with Cytoscape 3.9.1. Topological indices, including “degree centrality (DC)”, “closeness centrality (CC1)”, “betweenness centrality (BC)”, and “eigenvector centrality (EC)” were calculated using CytoNCA, and the core targets of the PPI network were obtained.
Enrichment Analysis of Gene Ontology (GO)and Kyoto Encyclopedia of Genes and Genomes (KEGG)Pathways
GO functional and KEGG pathway enrichment analyses were performed using DAVID (https://david.ncifcrf.gov/summary.jsp). GO functions included biological process (BP), molecular function (MF), and cellular component (CC2). Using active components, potential targets, and KEGG's top 20 pathways, component-target-pathway networks were constructed.
Cell Culture
To prepare for subsequent experiments, researchers thawed RAW264.7 cells, grew them in high-glucose DMEM at 37°C with 5% CO2 for 24 h, examined them microscopically, subcultured at 80–90% confluence, and harvested logarithmic-phase cells.
Cell Viability Assay
RAW264.7 cells (100 μL, 3 × 105 cells/mL) were seeded in 96-well plates and incubated with EOC (0-200 μg/mL) for 24 h; CCK-8 was added, and optical density (OD) was read at 450 nm using a microplate reader.
Western Blotting
Seeded in six-well plates at 3 × 105 cells/mL, RAW264.7 cells were allocated to six experimental groups: normal, LPS-induced model (0.25 μg/mL), three EOC groups (10, 20, 50 μg/mL), and DEX (50 μg/mL). Then, EOC solutions were added to each group at designated final concentrations, followed by 3 h pretreatment, LPS (0.25 μg/mL final) stimulation for 24 h (except normal group), and cell lysis in protein extractant for BCA-based protein quantification. Proteins were transferred to PVDF, incubated with primary antibodies at 4°C, washed, and probed with HRP-secondary antibodies for 1 h at RT. Visualization used an enhanced chemiluminescence (ECL) detection system, and images were captured via chemiluminescence imaging.
Molecular Docking
Plasma-absorbed prototype compounds were selected as docking ligands to corroborate network pharmacology predictions and experimental findings. Following download from PubChem, 3D SDF conformer files of components were converted to MOL2 using Open Babel GUI; PDB files of the top four target proteins (TNF, IL-1β, Caspase3 (CASP3), Nuclear factor NF-κB p105 subunit (NFκB1)) were retrieved from RCSB (https://www.rcsb.org). PyMOL removed water/redundant ligands, while AutoDock4 v1.5.7 hydrogenated acceptor proteins, outputting PDBQT files. Upon hydrogenation, small molecules were processed to auto-generate torsion trees in the software, which were saved as a PDBQT ligand file for minimal binding energy calculation. The top-affinity conformations were selected as the final docking model and visualized using PyMOL, serving as the definitive docking result.
Molecular Dynamics (MD) Simulation
Following selection of the optimized protein–ligand complex from docking outputs, GROMACS 2022 was employed to perform MD simulations. For small molecules, GAFF force field was used; protein molecules underwent processing with AMBER14SB force field combined with TIP3P water model. Simulation of the constructed protein–ligand complex system was performed under periodic boundary conditions and constant temperature/pressure, with hydrogen bonds constrained via the LINCS algorithm at a 2 fs time step in MD. PME (Particle Mesh Ewald) was utilized to compute electrostatic interactions at 1.2 nm cutoff.
Non-bonding interactions were cutoff at 10 Å (10-step updates), with the system subjected to 100 ns MD simulation at 298 K/1 bar. Following simulation, VMD and PyMOL analyzed the trajectory, while the “gmx_MMPBSA.py” script calculated MMPBSA binding free energy for protein–ligand complexes.
Statistical Analysis
Normality of distribution was evaluated by the Shapiro–Wilk test, and variance homogeneity was confirmed via Levene's test, with data analysis conducted using SPSS Statistics (17.0, IBM, USA). For datasets violating assumptions, logarithmic transformation was applied prior to parametric analysis.
Independent-measures one-way analysis of variance (ANOVA) assessed intergroup dynamic changes, followed by Tukey's post hoc test (α=0.05), with partial η² effect sizes (0.01/0.06/0.14) and G*Power 3.1 power analysis (0.82 for medium effects, two-tailed, α = 0.05).
As mean ± standard error of the mean (SEM), continuous data were reported, all tests two-tailed and significance set at P ≤ .05.
Results
Activity of EOC in Alleviating ALI
EOC attenuates pulmonary edema and inflammatory cell infiltration. To verify EOC can reduce the pulmonary edema, the lung tissues were observed. Lung tissues in the control group were dry/non-hyperemic, contrasting with the model group's hardened, congested tissues with red exudate (ALI phenotype), and EOC/DEX groups showed better morphology and less bleeding (Figure 1A).

EOC Attenuates LPS-Induced ALI. (A) Gross Pathological Changes in Lung Tissue (Representative Images). (B) HE Staining Showing Neutrophilic Infiltration: Green Arrows Indicate Sites of Neutrophil Accumulation. Magnification: 400×; Scale Bars: 100 μm.
HE staining was used to assess lung pathology, validating EOC-mediated reduction of inflammatory cell infiltration in ALI by revealing significantly increased infiltration in LPS-administered versus control lungs. EOC at different concentrations and DEX both alleviated LPS-induced ALI to varying degrees, as demonstrated by results from the EOC and DEX groups (Figure 1B).
EOC suppresses IL-6, TNF-α, IL-18 and IL-1β secretion. ELISA-measured plasma cytokines (IL-6, TNF-α, IL-18 and IL-1β) showed significantly higher secretion in LPS versus control groups, validating EOC's anti-inflammatory effect in ALI. In contrast, EOC administration inhibited the secretion of these cytokines: IL-6 (F5, 30 = 14.607, P < .001, Figure 2A), TNF-α (F5, 30 = 21.939, P < .001, Figure 2B), IL-18 (F5, 30 = 21.551, P < .001, Figure 2C), and IL-1β content (F5, 30 = 25.491, P < .001, Figure 2D). Data were expressed as the mean ± SD (n = 6).

EOC Suppresses the Secretion of (A) IL-6, (B) TNF-α, (C) IL-18 and (D) IL-1β in Plasma of LPS-Induced ALI Rats. Data Were Expressed as the Mean ± SD (n = 6), IL-6 (F5, 30 = 14.607, P < .001); TNF-α (F5, 30 = 21.939, P < .001); IL-18 (F5, 30 = 21.551, P < .001); IL-1β (F5, 30 = 25.491, P < .001). Groups are Annotated (a–c) for ANOVA/Tukey HSD with Shared Letters Being Insignificantly Different.
Potential Active Components in Serum Collected After EOC Administration
LC-MS was used for chemical profiling of EOC-administered serum to identify candidate therapeutic substances of EOC. Figure S1 shows the total ion chromatograms in both positive and negative ion modes. Serum after EOC oral administration showed 19 detected compounds (Table 1), nine confirmed by standards (Table 1). Among these 19 prototype compounds, 10 were flavonoids, 7 were organic acids, 1 was coumarin, and 1 was another type (Figure S2-S3). Serum pharmacochemistry indicates that the compounds in the serum are potentially active traditional Chinese medicine (TCM) components. Therefore, 19 authentic compounds were identified in the serum as active components for further study.
The MS Information of Prototype Compounds After EOC Administration.

Represents the results were confirmed by comparison with the standard.
Screening of Potential EOC Targets for Treating ALI
Based on the 19 authentic compounds identified earlier, 183 putative targets were predicted using the Swiss Target Prediction platform. Following duplicate removal, 2774 ALI targets from GeneCards, DisGeNET, OMIM, TTD were analyzed by Venn online tool to identify overlapping targets across the two sets (Figure 3A). This analysis revealed 76 potential ALI-related targets that could be modulated by EOC. The EOC-authentic compounds-ALI-potential target network (97 nodes, 208 edges) was constructed via Cytoscape 3.9.1, with circles denoting ALI targets and arrows representing EOC-derived compounds (Figure 3B).
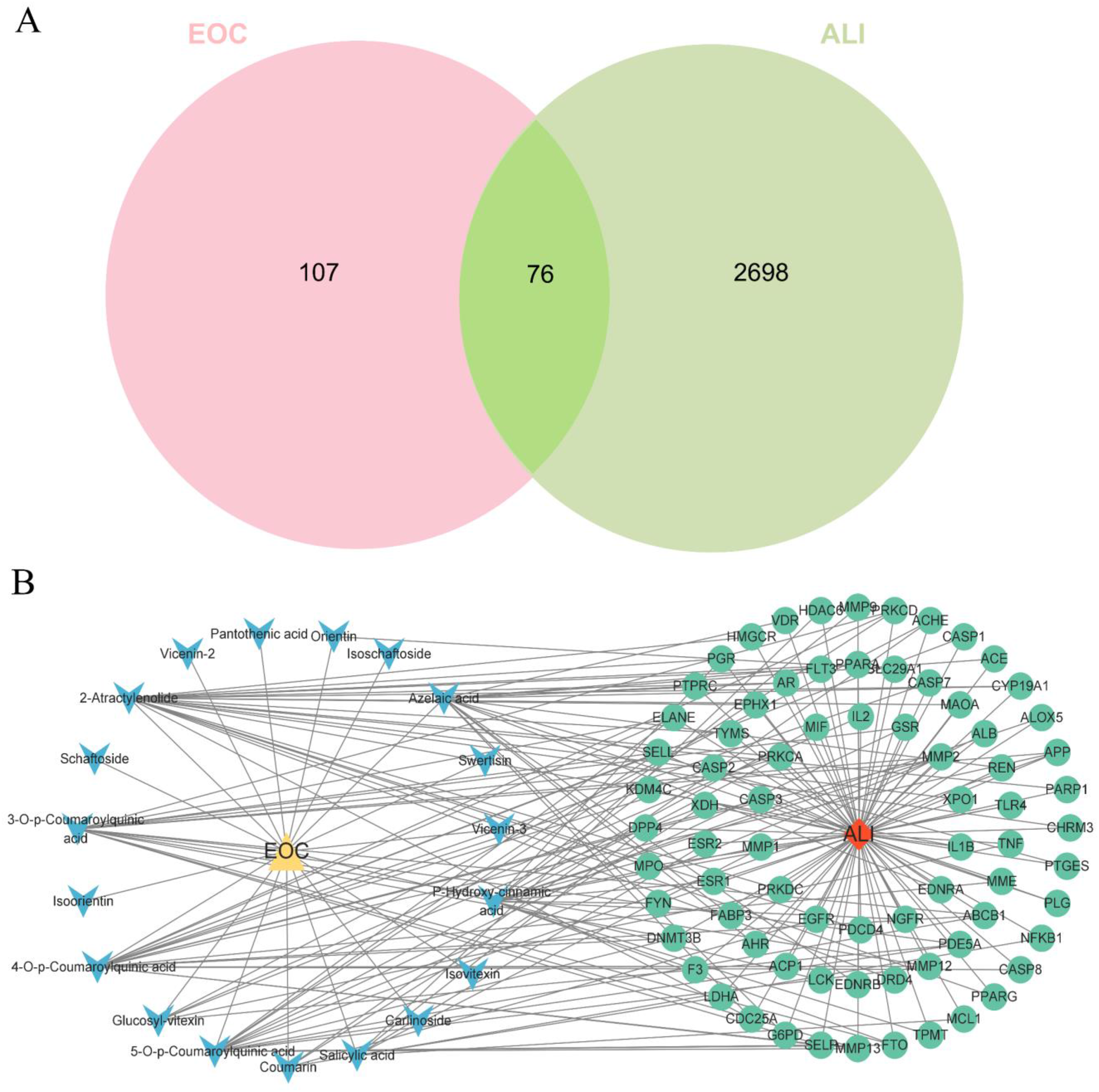
The Potential Targets for EOC Treating ALI. (A) Veen Plots of the Cross-Targets Between ALI and 19 Authentic Compounds in EOC; (B) Drug-Authentic Compounds-Disease-Target Network Diagram by Cytoscape 3.9.1. the Circles Represented Targets for ALI, Arrows Represented 19 Authentic Compounds.
Construction of the PPI Network
STRING analysis of 76 overlapping targets (Figure 4A) generated PPIs, with node size and color reflecting degree centrality. Based on DC, CC1, and BC values of the component nodes (which were ≥18.58, ≥ 0.54, and ≥65.54, respectively) (Figure 4B), 15 core targets were identified: TNF, IL1B, ALB, CASP3, NFKB1, EGFR, MMP9, ESR1, PPARG, TLR4, ACE, REN, PARP1, AR, and XDH (Figure 4C, Table S1).
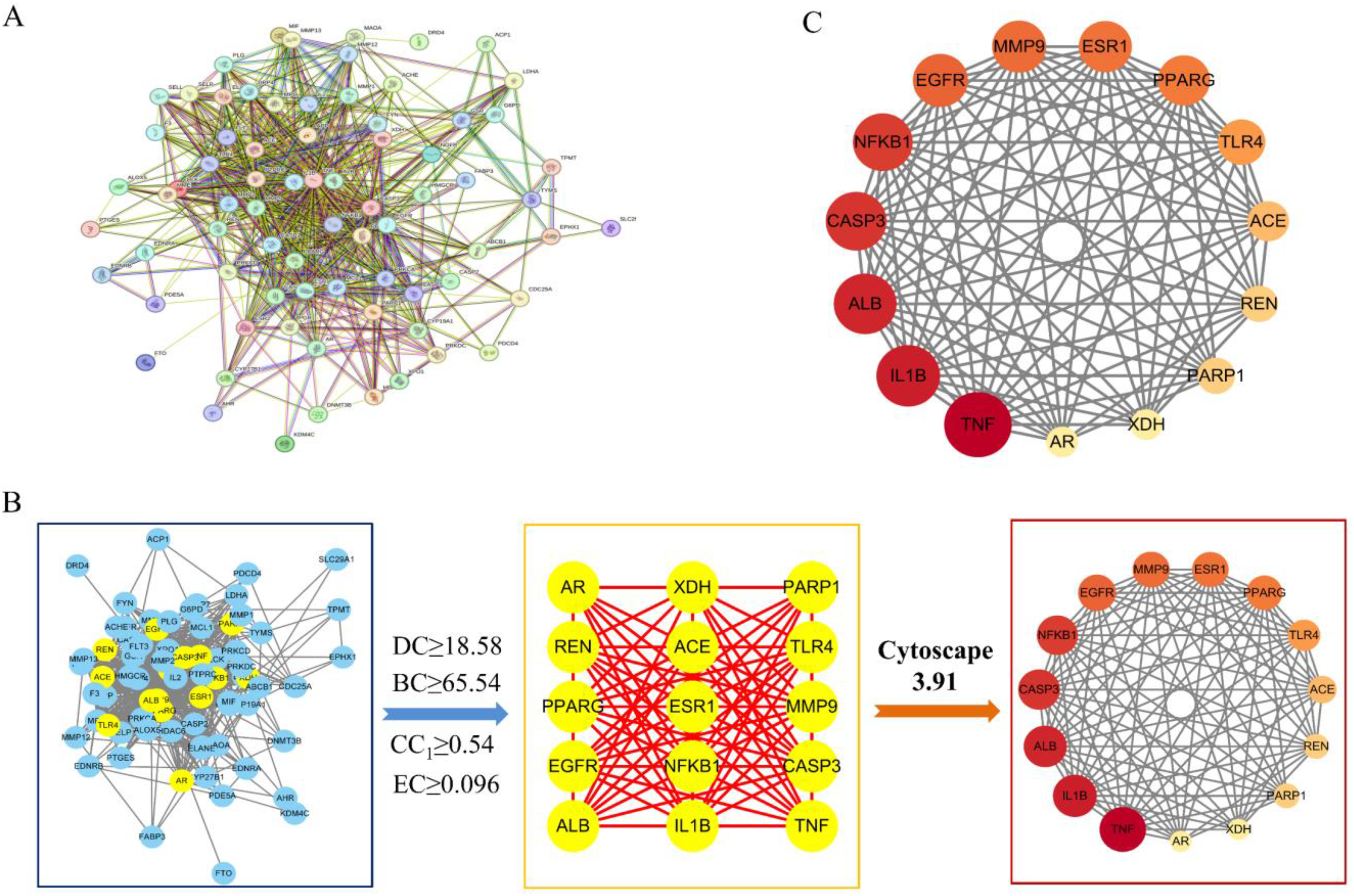
Core Targets and Their Screening Process Diagram. (A) Protein-Protein Network Constructed Using String Database; (B) The Screening Process of Core Targets; (C) PPI Network Diagram of Core Targets.
GO and KEGG Enrichment Analysis
To explore EOC's ALI treatment mechanisms, GO enrichment analysis showed that BP-associated processes most significantly included positive regulation of nitric oxide biosynthesis, lipopolysaccharide-mediated signaling, and MAP kinase activity. In terms of CC2, the targets were primarily enriched in macromolecular complexes, membrane rafts, receptor complexes, and chromatin. MF results revealed a strong association with identical protein binding, enzyme binding, estrogen receptor binding (Figure 5A). Enrichment analysis of KEGG pathways yielded 62 significant results (p < 0.05). Figure 5B shows the 20 top-ranking pathways based on their p-values. These results indicate that IL-17/NF-κB signaling pathways were the most important mechanisms by which EOC exerts its treatment effects on ALI.

GO and KEGG Enrichment Analysis. (A) Results of GO Enrichment Analysis of the Potential Targets; (B) Results of KEGG Enrichment Analysis of the Potential Targets.
EOC Inhibits IL-17/NF-κB Pathways
EOC's dose-dependent effects on IL-17/NF-κB pathways were tested by western blot to clarify its ALI-alleviating mechanisms. In LPS-induced RAW264.7 cells (Figure S4), LPS upregulated IL-17/p-NF-κBp65 (Fig. 6B), while EOC reduced IL-17/GAPDH (Fig. 6C; F = 44.790, P < .001) and p-NF-κBp65/NF-κBp65 (Fig. 6D; F = 165.074, P < .001) ratios significantly.

Expression of IL-17, NF-κBp65, and p-NF-κBp65 Proteins in RAW264.7 Cells. Representative Cell Viability (A) (F6, 18 = 72.729, P < .001), Western Blots (B), Quantitative Analysis for IL-17 (C), and p-NF-κBp65/ NF-κBp65 (D) Protein Expression. Data were Expressed as the Mean ± SD (n = 3), IL-17/GAPDH Protein Ratio (F5, 15 = 44.790, P < .001); p-NF-κBp65/ NF-κBp65 Protein Ratio (F5, 15 = 165.074, P < .001). Groups are Annotated (a–f) for ANOVA/Tukey HSD with Shared Letters Being Insignificantly Different.
Molecular Docking
Docking 19 active compounds with IL-17/NF-κB pathway core targets (TNF, IL1B, CASP3, NFKB1) evaluated EOC's further effects on ALI (Table S2). Inversely correlated with binding stability, ligand-receptor binding energy <−5.0 kcal/mol indicated effective interaction and <−7.0 kcal/mol strong binding. Figure 7 revealed that NFKB1 had the strongest binding energy with the 19 blood entry components. The binding energy with flavonoids, including vicenin-3, isovitexin, isoschaftoside, orientin, isoorientin, schaftoside, glucosyl-vitexin, swertisin and carlinoside, were less than −7 kcal/mol, which showed that flavonoids are the main active ingredients of EOC. Figures 8 and 9 also showed that NFKB1 was the important core target for IL-17/NF-κB pathways. Representative molecular docking of vicenin-3, isovitexin, isoschaftoside with NFKB1 is illustrated in Figure 10.

Molecular Docking Heat Map.

IL-17 Signaling Pathway.

NF-κB Signaling Pathway.
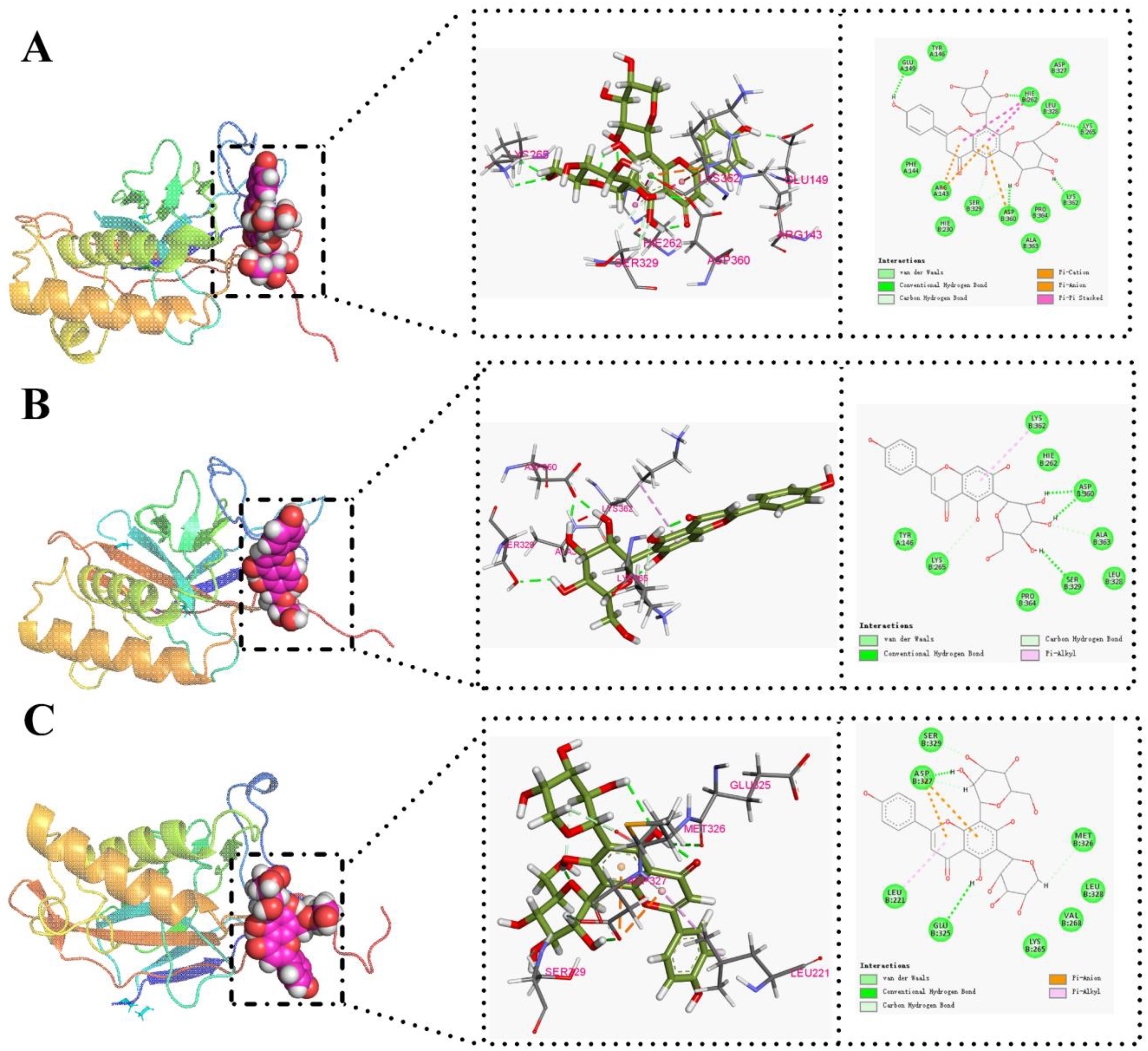
Molecular Docking of NFKB1 with (A) Vicenin-3, (B) Isovitexin, (C) Isoschaftoside.
Molecular Dynamics Analysis
As a metric for atomic displacement in MD simulations, RMSD assesses target protein–ligand complex stability via initial-state fluctuation analysis. RMSD stabilization with <0.2 nm fluctuations generally indicates system stability. During the 100 ns MD simulation, vicenin-3, isovitexin, and isoschaftoside showed stable conformational fluctuations (Fig. 11A). Stable NFKB1 binding was indicated by consistent RMSD fluctuations (0.2-0.4 nm) and similar trends in the three ligands.

The Fluctuation Plot of NFKB1–Ligand Complexes. (A) RMSD; (B) SASA.
Solvent-Accessible Surface Area (SASA) captures the solvent-exposed molecular surface area, tracking conformational changes and ligand exposure during simulations to assess protein–ligand binding interactions. During MD simulation, the average SASA values (nm2) were 114.74 ± 2.09 (vicenin-3, black), 112.89 ± 2.70 (isovitexin, red), and 115.02 ± 2.67 (isoschaftoside, blue). SASA values of these compounds fluctuated minimally throughout the simulation, confirming stable interaction with the protein complexes (Figure 11B).
Molecular Mechanics–Poisson-Boltzmann Surface Area (MMPBSA) calculates protein–ligand binding energy by post-processing MD trajectories, evaluating interaction stability via bound/unbound free energy differences. Following selection of stable-RMSD segments from the three complexes and extraction of 10-nanosecond trajectories for MMPBSA energy analysis, the total binding energy (ΔG) was calculated as the sum of enthalpic (ΔH) and entropic (−TΔS) contributions. Table 2 summarizes the contributions from various components, including the Van der Waals force (VDW), Coulomb interaction energy (COU), molecular mechanics (MM), polar solvation energy (PB), and nonpolar solvation energy (SA). Reliability of computed free energy values was strengthened by the close consistency between gmx_mmpbsa.bsh ΔH and gmx_mmpbsa.py ΔG. As shown in Table 2, vicenin-3 exhibited significantly lower ΔG and ΔH values than other ligands, corroborating molecular docking results and reinforcing its promise as a target protein interaction candidate.
The △G of Each complex and the Contributions of Individual Energy Components (kJ/mol).

Note: In this table, MM is the sum of VDW and ELE, △H is the sum of MM, PB, and SA, △G is the sum of △H) and–T△S.
Discussion
Acute pulmonary inflammation with inflammatory cell infiltration and edema defines ALI, whose progression correlates with TNF-α, IL-6, IL-1β, and IL-18 release.2-4 TNF-α is a pivotal cytokine initiating innate immunity by stimulating macrophage secretion of other cytokines. IL-6 drives acute phase response amplification of the inflammatory cascade.12,13 Inflammatory cascades in ALI are triggered and exacerbated by pyroptosis-derived IL-1β and IL-18. 14 Furthermore, By enhancing alveolar/vascular endothelial permeability to induce pulmonary edema, excessive IL-1β release shows contrast with IL-18's inverse survival association in ARDS.15,16 In this study, EOC effectively attenuated pulmonary edema and inflammatory cell infiltration in LPS-induced ALI rats (histopathological/HE analysis), accompanied by marked decreases in lung TNF-α, IL-6, IL-1β, and IL-18. Collectively, these findings indicate that EOC exerts a significant suppressive effect on the dysregulated inflammatory response in ALI.
An effective in vivo strategy for direct-acting substance identification, serum pharmacochemistry is crucial for elucidating therapeutic mechanisms of herbal medicines via bioactive constituent discovery. 17 Potent anti-inflammatory/antioxidant flavonoids are vital for inflammatory disease (eg, ALI) prevention and therapy. 18 OC contains various bioactive compounds, such as flavonoids and organic acids, with flavonoids being established as major contributors to its anti-inflammatory effects.19–21 For instance, vicenin-2 modulates LPS-induced TNF-α and total nitrite production22,23; vicenin-3 significantly inhibits IL-1β-induced NO and PGE2 24 ; isovitexin exerts anti-inflammatory effects by suppressing the secretion of IL-1β, IL-6, IL-8, and TNF-α25–27; isoschaftoside-mediated inhibition of LPS-induced NO and pro-inflammatory cytokines (iNOS, TNF-α, IL-1β, and COX2) is notable; activation of the EPCR-dependent JAK2/STAT3 pathway mediates the alleviation of sepsis-induced ALI in mice by isoorientin. 28 Additionally, swertisin, orientin,29,30 salicylic acid, 3-O-p-coumaroylquinic acid, and protocatechuic acid have documented anti-inflammatory properties.31–33 Identified by LC-MS/MS in rat serum post-EOC administration, 19 prototype compounds (10 flavonoids, 7 organic acids, 1 coumarin, 1 other) were all found in the crude extract, indicating their active component potential. Notably, while the crude extract contains diverse constituents, serum analysis primarily detected flavonoids and organic acids, highlighting the discrepancy between traditional extract analysis and serum pharmacochemistry in identifying bioavailable active components.34–36 Leveraging NF-κB's regulatory role in ALI inflammation, molecular docking assessed NF-κB1 (p50) binding of 19 serum compounds, demonstrating universal affinity and flavonoid-dominant peak binding energies.37–39 Therefore, we conclude that flavonoids, including Vicenin-3, isovitexin, isoschaftoside, orientin, isoorientin, schaftoside, glucosyl-vitexin, swertisin, and carlinoside, constitute the primary bioactive basis for EOC's anti-ALI effects.
Tissue damage is mediated by IL-17, a Th17-secreted cytokine binding IL-17R to activate NF-κB and induce inflammatory mediator expression.5,40 Researchers have confirmed NF-κB pathway overactivation in sepsis-associated ALI, noting that the NF-κB dimer (eg, p50/p65) is sequestered in the cytoplasm by IκB proteins in quiescent cells. 7 Inflammatory stimuli (eg, LPS) activate the IKK complex (IKKα/IKKβ), leading to IκB phosphorylation and degradation. Induction of NF-κB p65 nuclear translocation promotes gene transcription for TNF-α, IL-6, and IL-1β inflammatory mediators. 37 Validated by Western blotting, EOC suppressed LPS-induced NF-κB activation, paralleling cytokine/pathological improvement, and although NF-κB's resolution role complicates targeting, it remains a primary anti-inflammatory intervention target.38,39 Molecular docking results identify NF-κB1 as a key target of EOC flavonoids, which suppress the IL-17/NF-κB axis to protect against ALI.
The findings reveal that EOC potently reduces LPS-induced ALI by inhibiting the IL-17/NF-κB signaling cascade and decreasing levels of TNF-α, IL-6, IL-1β, IL-18. For the first time, flavonoids (vicenin-3, isovitexin, isoschaftoside, orientin) were systematically identified as EOC's primary anti-ALI constituents by serum pharmacochemistry, with their mechanism shown to involve NF-κB1 binding and NF-κB pathway modulation. This provides crucial evidence for the pharmacodynamic material basis and molecular mechanism underlying OC's potential in ALI treatment.
However, this study has limitations that warrant attention in future research. Firstly, the investigation was confined to a single LPS-induced, sepsis-related ALI model; extending evaluations to models reflecting other etiologies (eg, viral or trauma-induced ALI) is essential to assess the broader applicability of EOC. Secondly, serum pharmacochemistry identified 19 bioavailable components, and molecular docking predicted their core target proteins within the IL-17/NF-κB signaling pathways (TNF, IL-1β, CASP3, and NF-κB1). Nonetheless, the specific mechanistic roles of individual constituents (especially those identified as core flavonoids) on this pathway remain to be fully elucidated. Determining whether their effects are synergistic, additive, or independent will require dedicated experimental approaches, such as orthogonal compound combination screening, isobolographic analysis, and dose-response matrix assays. Future studies investigating the interaction dynamics among these components (as preliminarily explored in Supplemental Materials Table S3) are essential and will significantly accelerate clinical translation. Thirdly, although the IL-17/NF-κB pathway was implicated as central, potential crosstalk with other critical inflammatory pathways (eg, NLRP3 inflammasome, MAPK cascades) was not explored; investigating these interconnections would provide a more comprehensive view of EOC's multi-target anti-inflammatory mechanism. Fourthly, the computational predictions from molecular docking regarding ligand-target interactions require experimental validation (eg, via surface plasmon resonance (SPR), cellular thermal shift assay (CETSA), or target knockdown/inhibition studies) to confirm binding affinity and functional relevance. Finally, the current assessment focused on acute efficacy; longer-term studies evaluating chronic toxicity, sustained efficacy, and pharmacokinetics are needed to provide comprehensive safety data for potential clinical translation. These limitations outline important directions for future work to deepen our understanding; they do not diminish the value of the core findings presented here but rather establish a foundation for further elucidating the complex mechanisms of OC in ALI and advancing its therapeutic application.
These limitations outline directions for future research to deepen our understanding; they do not diminish the value of the core findings presented here but rather establish a foundation for further elucidating the complex mechanisms of OC in ALI and advancing its therapeutic application.
Conclusions
This study demonstrates OC ameliorates ALI via dual targeting of NF-κB1 and the IL-17/NF-κB signaling axis, leading to reduced pulmonary edema, inflammatory cell infiltration, and lung TNF-α, IL-6, IL-1β and IL-18 levels, with flavonoids identified as the primary therapeutic constituents. This study demonstrates OC's capacity to rescue ALI, highlighting its therapeutic potential for clinical translation in lung disease management.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X251361195 - Supplemental material for The Activity, Composition, and Molecular Mechanism of Oxalis corniculata in Alleviating Acute Lung Injury
Supplemental material, sj-docx-1-npx-10.1177_1934578X251361195 for The Activity, Composition, and Molecular Mechanism of Oxalis corniculata in Alleviating Acute Lung Injury by Xue Ma, Bao Zhang, Li Jiang, Weimi Kuang, Meiqin Wang, Yang Wang, Jia Sun and Yongjun Li in Natural Product Communications
Supplemental Material
sj-docx-2-npx-10.1177_1934578X251361195 - Supplemental material for The Activity, Composition, and Molecular Mechanism of Oxalis corniculata in Alleviating Acute Lung Injury
Supplemental material, sj-docx-2-npx-10.1177_1934578X251361195 for The Activity, Composition, and Molecular Mechanism of Oxalis corniculata in Alleviating Acute Lung Injury by Xue Ma, Bao Zhang, Li Jiang, Weimi Kuang, Meiqin Wang, Yang Wang, Jia Sun and Yongjun Li in Natural Product Communications
Supplemental Material
sj-png-3-npx-10.1177_1934578X251361195 - Supplemental material for The Activity, Composition, and Molecular Mechanism of Oxalis corniculata in Alleviating Acute Lung Injury
Supplemental material, sj-png-3-npx-10.1177_1934578X251361195 for The Activity, Composition, and Molecular Mechanism of Oxalis corniculata in Alleviating Acute Lung Injury by Xue Ma, Bao Zhang, Li Jiang, Weimi Kuang, Meiqin Wang, Yang Wang, Jia Sun and Yongjun Li in Natural Product Communications
Footnotes
Abbreviations
Acknowledgements
This study was funded by the Guizhou Provincial Department of Science and Technology, Science and Technology Bureau, and Guiyang Municipal Government.
Ethical Approval
Guizhou Medical University Animal Ethics Committee approved all animal experiments herein (approval no. 2001363).
Authors’ Contributions
Xue Ma, Bao Zhang, Li Jiang and Wei-Mi Kuang performed all of the experiments; Xue Ma, Bao Zhang, Mei-Qin Wang and Yang Wang analyzed the data; Xue Ma and Li Jiang wrote the manuscript. Bao Zhang, Jia Sun and Yong-Jun Li were the project leaders and designed the experiment.
Funding
This work was supported by the Science and Technology Program of Guizhou Province (ZK [2023] general project 008, ZK [2023] general project 329, QKHZ [2023] 006, ZK [2021] general project 529), Science and Technology Bureau, Guiyang Municipal Government (2024235).
Declaration of Conflicting Interests
The authors have no conflicts of interest to declare in this article.
Statement of Human and Animal Rights
No human subject studies are reported here. All animal experiments were approved by the Guizhou Medical University Animal Ethics Committee (approval no. 2001363) and conducted in compliance with relevant guidelines.
Statement of Informed Consent
This study involved no human subjects, and informed consent was not required.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
