Abstract
Introduction
Experimental Materials
Instruments
The following instruments were used: LC-MS (THERMO, UltiMate3000) (AB SCIEX, 5600QTOF) analytical balance (Mettler Toledo, XS205), rotary evaporator (ZhengZhou Creatwall, RE-52A A), circulating water multipurpose vacuum pump (ZhengZhou Creatwall, SHB-III), nebulizer (OMRON, C28P), embedding center (Huida, HD-310), rotary microtome (Huida, HD-330), optical microscope (OLYMPUS, IX73), high-speed centrifuge (Thermo, Pico 17), inverted microscope (Leica, DMI3000B), CO2 incubator (Thermo, 370), enzyme calibrator (Thermo, MULTISKAN FC), cell counter (Nexelom, Cellometer mini), and autoclave (Thermo, Hirayama HVE-50).
Cells
HL-60 leukemia cells, A549 lung cancer cells, SMMC-7721 liver cancer cells, MCF-7 breast cancer cells and SW480 colon cancer cells were purchased from the China General Microbiological Culture Collection Center.
Crude Drugs
Drugs and Reagents
Cisplatin (Meilunbio, N1001A), and Taxol (Meilunbio, D1106A) were used.
Fetal calf serum (lot 1706126), RMPI-1640 medium (lot 0023019), DMEM (lot 0024719), pancreatin (lot 0023518), double resistance (lot 1936917), PBS (lot 0044818), and DMSO (lot B821BA0018) were purchased from BI, and an MTS kit (Promega, 0000219904) was purchased from HKM.
The reagents included 95% alcohol (FUCHEN, AR), acetic ether (Huada CHEM, AR), alcohol (FUCHEN, AR), etc.
Experimental Methods
Preparation of the EAF
Anti-NSCLC Activity
Solution Preparation
For preparation of the positive control solution, cisplatin was added to 0.064, 0.32, 1.60, 8.00, and 40.00 μg/mL concentrations, and Taxol was added to concentrations of 0.008, 0.04, 0.20, 1.00, and 5.00 μg/mL.
Methods
Inoculated cells in the logarithmic growth phase were observed under an inverted microscope, and 10% fetal bovine serum culture medium was added to a single-cell suspension. A total of 3000 to 15 000 cells per well with a volume of 100 μL were inoculated into a 96-well plate. After 12 to 24 h, the cells were divided into 4 groups: the blank group, the EAF group, the cisplatin control group, and the paclitaxel control group.
The solutions to be measured were created with final concentrations of 100, 20, 4, 0.8, or 0.16 μg/ml EAF. Three wells were set up, with a final volume of 200 μL in each, and an equal volume of culture medium was added to the blank group. A positive control was used to determine the concentrations, and 3 additional wells were used.
After culturing at 37 °C for 48 h, 20 μL of MTS solution and 100 μL of culture medium were added to each well, and the culture medium in the wells was discarded. Then, 20 μL of MTS solution was added to each well, after which 100 μL of culture supernatant was discarded; 3 blank wells (20 μL of MTS solution and 100 μL of medium) were included and the plates were incubated for 2 to 4 h to determine the optical absorption after the reaction was complete.
For colorimetric analyses, a wavelength of 492 nm was selected, the optical absorption value of each well was read under a multifunctional microplate reader, and the results were recorded. The growth curve of cells was drawn with concentration as the abscissa and inhibition of cell proliferation as the ordinate after data processing. The IC50 values of the compounds were calculated via the Reed and Muench method.
Analysis of the Active Components
Sample Pretreatment
First, 30 mg of EAF was added, and 500 μl of 75% methanol in water was added; then, the mixture was vortexed for 60 s, ultrasonicated for 30 min, and fully dissolved. The mixture was centrifuged at 17 000×g for 15 min, after which the supernatant was vacuum-dried at room temperature. Then, 100 μl of 50% methanol aqueous solution was added, the sample was vortexed at 60 °C and filtered through a 0.22 µm organic filter.
Methods
The mobile phase conditions were as follows: (1) column temperature, 40 °C; sample size, 3 μL. (2) In positive ion mode, solution A was 0.1% formic acid in water and solution B was 0.1% formic acid in acetonitrile. In negative ion mode, solution A was water (2 mM ammonium acetate) and solution B was acetonitrile. The elution methods for the mobile phase were as follows: flow rate of 400 μL/min, 0 to 1.5 min (A:B = 95:5), 1.5 to 2.5 min (A:B = 90:10), 2.5 to 14 min (A:B = 60:40), 14 to 22 min (A:B = 5:95), 22 to 25 min (A:B = 5:95), 25 to 26 min (A:B = 95:5), and 26 to 30 min (A:B = 95:5).
The mass spectrometer conditions were as follows: ion source, ESI; nebulizer pressure, 60 psi; assisted gas pressure, 60 psi; air curtain pressure, 35 psi; temperature, 650 °C; spray voltage, 5000 V (positive ion mode) and 4000 V (negative ion mode); primary acquisition range, 50-1200; impact energy, 30 eV; and 10 secondary spectra per 50 ms.
Compound Structure Analysis
The literature on the chemical constituents of
Active Component Analysis
All compounds obtained by LC‒MS were input into the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, https://old. tcmsp-e.com/index.php) and the Database of Constituents Absorbed into Blood and Metabolites of Traditional Chinese Medicine (DCABM-TCM, http://bionet.ncpsb. org.cn/dcabm-tcm/) to identify the active components of
Mechanistic Research
Target Prediction and Network Construction
TCMSP and the Bioinformatics Analysis Tool for Molecular Mechanics of Traditional Chinese Medicine (Batman-TCM, http://bionet.ncpsb.org.cn/batman-tcm/) were used to collect and predict the targets of chemical components. Disease-related target genes were obtained from the GeneCard database (https://www.genecards.org/), OMIM database (https://omim.org/), and DisGeNET database (https://www. disgenet.org/), with the keyword “non-small cell lung cancer,” and shared targets were identified by Venny2.1.0 software. Cytoscape 3.7.1 software was used to visualize the common drug-disease targets, and a drug–disease–target network diagram was constructed. The shared target genes were input into String (http://stringdb.org/). The conditions were set as medium confidence > 0.9, and the free protein targets were eliminated to construct a protein‒protein interaction (PPI) network.
GO Function Analysis and KEGG Pathway Enrichment Analysis
Using Bioconductor software, with the help of the clusterProfiler, pathview, and Dose packages, the gene ontology and pathway enrichment data were obtained and visualized by R4.4.1 software, and GO function enrichment and KEGG pathway analyses were performed. A
Molecular Docking
First, the names of key targets were searched in the RCSB protein database (https://www.rcsb.org). The screening conditions were Homo sapiens, and the sequences were reviewed. The crystal structure data of the targets were obtained, and the 3D structure was saved in PDB format. The residues and water were removed by PyMOL2.5, and the ligands were structured by Chem3D 19.0 and stored in mol2 format. Finally, the key targets were identified with MOE 2019 software for protein preparation and molecular docking verification. The docking was performed with the Discovery Studio software (Version 4.5). The binding sites and all compound conformations were identified, and the LibDock was selected for docking. The site and molecule conformation with the highest LibDockScore were determined for final interaction. The binding pocket 3D view and the intermolecular forces distance 2D view were displayed. The verification results were visually analyzed by PyMOL 2.5.
Statistical Analysis
Origin 2022 software (OriginLab Corporation, USA) was used for data analysis. The data are expressed as the mean ± SD for a minimum of 3 independent experiments. One-way analysis of variance and the least significant difference test were used to analyze significant differences in multiple and single comparisons, respectively.
Results
Anti-NSCLC Activity
The results of the antitumor activity screening showed that EAF significantly inhibited the proliferation of 5 tumor cell lines, and the intensity was weaker than that of cisplatin and paclitaxel; however, the concentrations that inhibited cell proliferation were at the microgram level. As shown in Table 1, the IC50 of EAF inhibition of the proliferation of the HL-60 cell line was 56.72 μg/mL, that of A549 was 41.13 μg/mL, that of SMMC-7721 was 36.69 μg/mL, that of MDA-MB-231 was 48.63 μg/mL, and that of SW480 was 44.48 μg/mL. As shown in Figure 1, an EAF concentration of 100 μg/mL showed the strongest inhibition of the lung cancer cell lines.
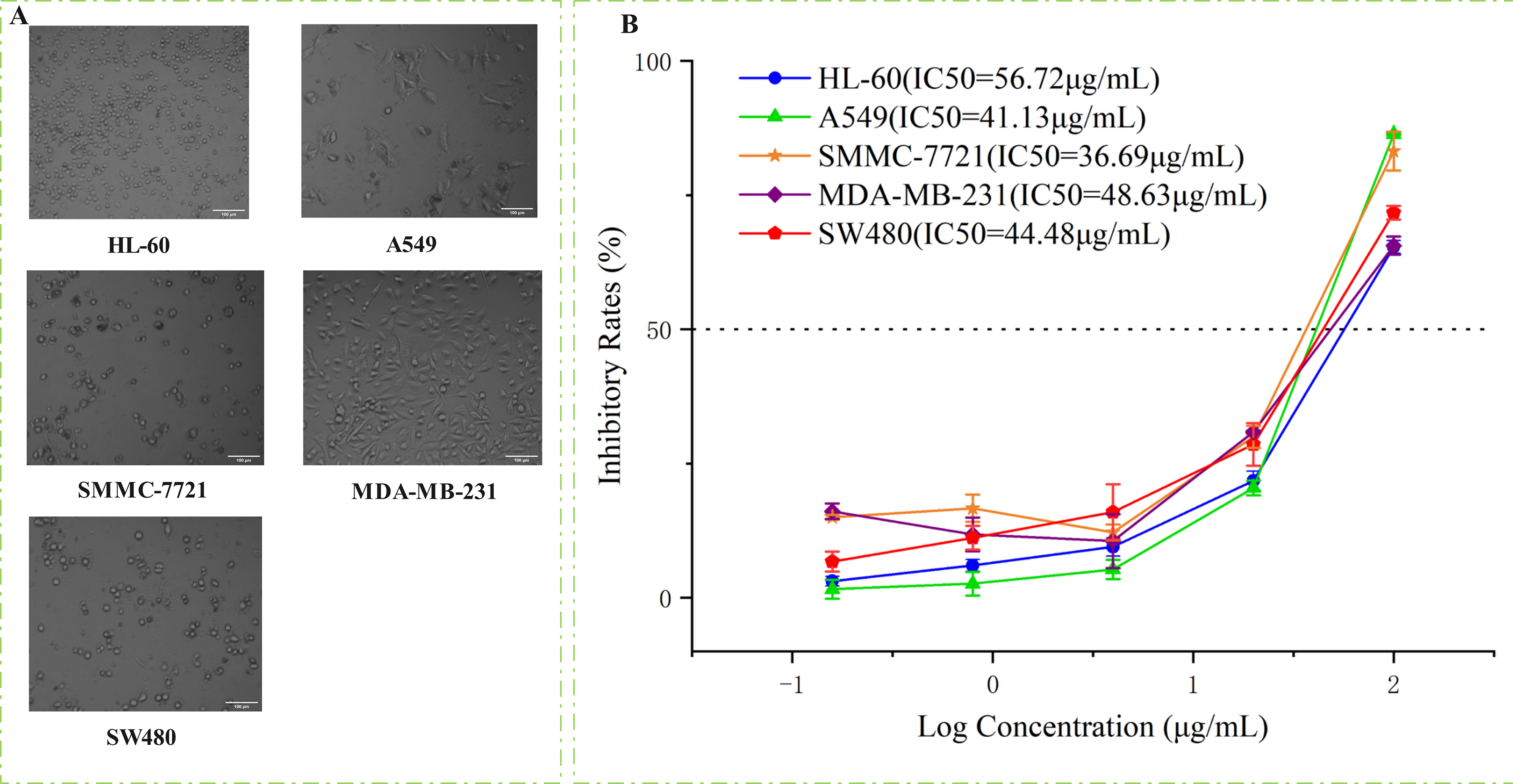
Cell state diagram after 48 h of administration (A) and inhibition of tumor cell proliferation by different concentrations of EAF solution (B).
Screening of Antitumor Activity.
Active Component Analysis
The total ion flow diagram of EAF in positive and negative ion modes is shown in Figure 2, and a total of 33 compounds were identified, as shown in Table 2, including 12 flavonoids, 9 coumarins, 7 organic acids, 1 naphthoquinone, 2 glycosides, and 3 other types of compounds. The active ingredients with antitussive effects were rutin and luteolin.14,15 The active ingredients with antitumor activity were calycosin, marmesin, 7-hydroxy-coumarin, 6-hydroxy-4-methylcoumarin, rutin, 6,7-dihydroxycoumarin, 2,3-dihydroxybenzoic acid, p-coumaric acid, caffeic acid, and so on.16–24 The active ingredients with antilung cancer effects were 4-hydroxybenzoic acid, pinocembrin, luteolin, 3′-O-methylluteolin, hesperetin, shikonin, 4-hydroxycoumarin, and so on.25–31

Positive ionization mode (A) and negative ionization mode (B).
Results of Chemical Composition Analysis.

Analysis of Cleavage of the Major Compounds
The results showed that 12 flavonoids were the most abundant components in the ethyl acetate fraction. The excimer ion peak of compound 3 was m/z 285.04[M-H]-. Removing 1 molecule of C7H2O4 or 1 molecule of C3O2 resulted in the fragment ion peaks at m/z 133.03 and m/z 217.05. Removing 1 molecule of O or 1 molecule of C2H2O resulted in the fragment ion peaks at m/z 107.01 and m/z 175.04. Removing 1 molecule of C2 resulted in a fragment ion peak at m/z 151.00, which was attributed to luteolin. The excimer ion peak of compound 2 was m/z 283.06 [M-H]-, the fragment ion peak at m/z 268.04 was a result of removing a molecule of CH3, and the fragment ion peak at m/z 148.01 was a result of removing a molecule of O; this peak was determined to be calycosin. The cleavage pathway is shown in Figure 3a.

Fragmentation pathway of calycosin (A). Fragmentation pathway of vitexin 2 “-O-β-D-glucoside (B). Fragmentation pathway of 5,7-dihydroxy-4-methylcoumarin (C). Fragmentation pathway of caffeic acid (D).
There were 9 coumarin compounds, and this group of compounds was the second most abundant among all groups of compounds. The excimer ion peak of compound 20 was at m/z 191.03 [M-H]-. Removing a molecule of CO or a molecule of CO2 resulted in fragment ion peaks at m/z 163.04 and m/z 147.05, which were attributed to 5,7-dihydroxy-4-methylcoumarin. The cleavage pathway is shown in Figure 3b.
There were 7 organic acid compounds, and this group of compounds was the third most abundant group. The excimer ion peak of compound 29 was at m/z 179.04 [M-H]-. Removing a molecule of C8H7O2, a molecule of C3H3O3 or a molecule of C3H2O2 resulted in fragment ion peaks at m/z 43.99, m/z 93.03 and m/z 109.03. Removing a molecule of CO or a molecule of H2O resulted in fragment ion peaks at m/z 65.04 and m/z 91.02, which were attributed to caffeic acid. The pyrolysis pathway is shown in Figure 3c.
Two compounds were identified as glycosides, which was the fourth most abundant group of compounds. The ion peak of compound 17 was at m/z 593.15 [M-H]-. A fragment ion peak at m/z 283.06 was formed after removing glucoside. Removing a molecule of O, a molecule of C9H8O2 or a molecule of C8H7O2 resulted in fragment ion peaks at m/z 267.07, m/z 135.01 and m/z 148.02. Removing a molecule of CH2 resulted in a fragment ion peak at m/z 253.05. A fragment ion peak at m/z 93.03 was found after removing a molecule of C9H5O3; this peak was identified as vitexin 2″-O-β-D-glucoside. The cleavage pathway is shown in Figure 3d.
Active Compounds
A total of 6 active components, including hesperetin, pinocembrin, shikonin, luteolin, marmesin, and calycosin, were detected; the details are shown in Table 3. Studies have shown that pinocembrin, luteolin, hesperetin, and shikonin have anti-NSCLC activity.26–29
Information About the Active Components.

Target Prediction and Network Construction
A total of 6833 NSCLC disease targets and 230 drug targets were evaluated, and 175 common targets were ultimately identified via Venny software, as shown in Figure 4a. Figure 4b shows the constructed drug–disease–target network, which included 182 nodes and 497 interconnected edges. An increase in connected edges indicates that the compound is more active. Drug–disease–target network analysis revealed that luteolin (106 interconnected edges), pinocembrin (80 interconnected edges) and hesperetin (65 interconnected edges) were the main active components. The PPI network diagram was constructed as shown in Figure 4c. Based on analysis of the PPI network, 6 targets with high degree composite values—AKT1, IL6, EGFR, BCL2, CASP3, and ESR1—were identified as key targets. Among them, EGFR is a target that has been thoroughly studied and has achieved remarkable results in recent years.
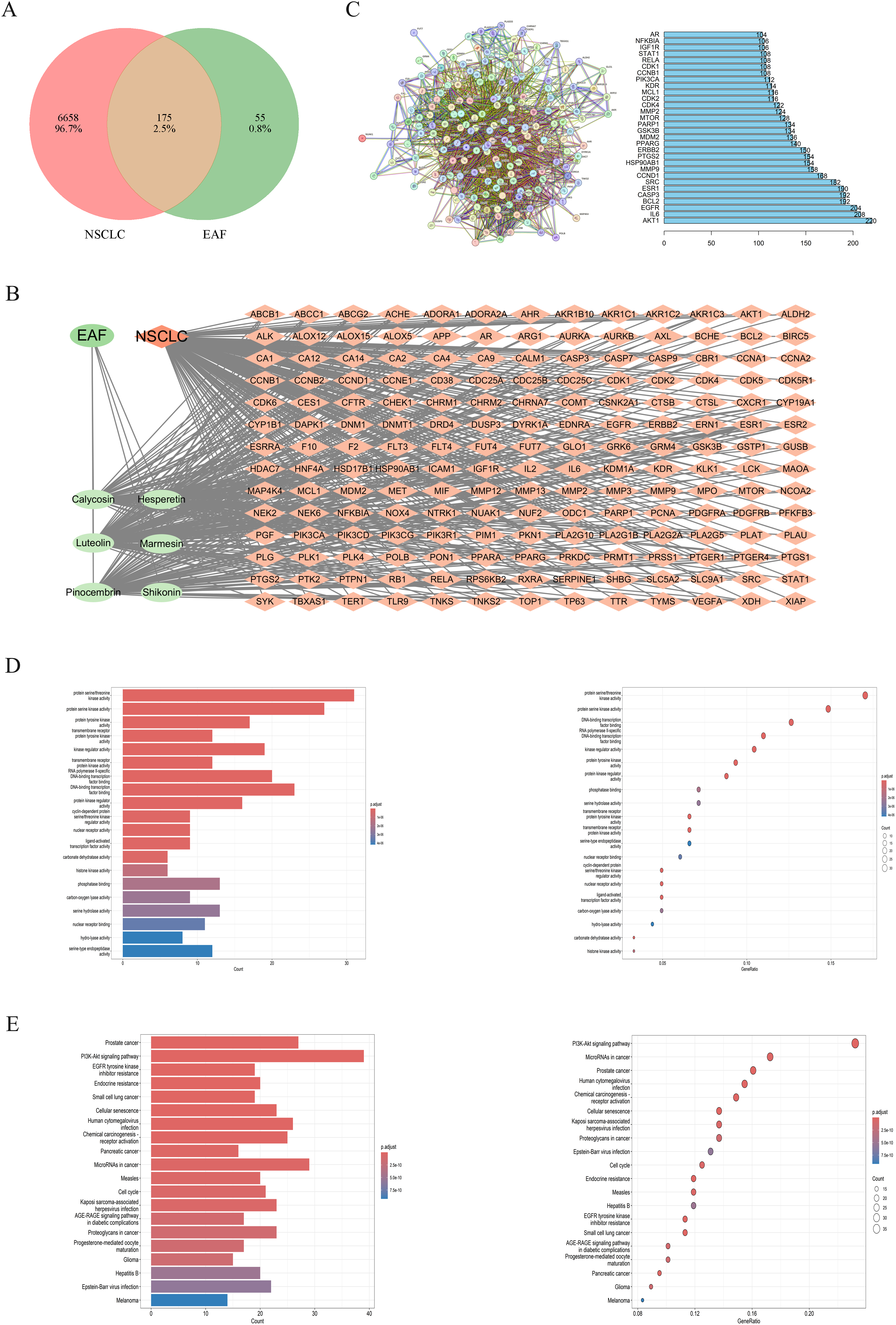
Venn diagram of drug–disease interactions (A). Drug–disease–target network (B). PPI network (C). GO analysis diagram (D). KEGG analysis diagram (E).
GO Function and KEGG Pathway Enrichment Analysis
Through GO enrichment analysis, 176 molecular functions were identified (
A total of 146 signaling pathways were identified by KEGG pathway enrichment analysis, including prostate cancer, the PI3K-Akt signaling pathway, EGFR tyrosine kinase inhibitor resistance, endocrine resistance, small cell lung cancer, cellular senescence, human cytomegalovirus infection, chemical carcinogenesis-receptor activation, pancreatic cancer, and microRNAs in cancer. Many of these pathways have been reported to be closely related to NSCLC.36–39 The 20 key signaling pathways are shown in Figure 4e.
Molecular Docking
Through the above data analysis, 3 active components (pinecembrin, luteolin, and hesperetin) were selected for molecular docking with 5 key anti-NSCLC target proteins (EGFR, AKT1, IL6, BCL2, and CASP3). As shown in Figure 5, the lower the binding energy, the better the binding; additional details are shown in Table 4. The 3 active ingredients bound well to the anti-NSCLC target proteins, and hesperetin bound best.

Molecular docking.
Affinity Values.

Discussion
The technical flowchart of this study is shown in Figure 6. The antitumor activity of EAF was evaluated by the MTS assay, the chemical constituents of the ethyl acetate fraction were assessed by LC-MS, the active ingredients were identified by TCMSP and DCABM-TCM, and the molecular mechanism EAF alleviation of NSCLC was explored by network pharmacology. Finally, molecular docking technology was used for verification. This research has scientific integrity but also has certain limitations.
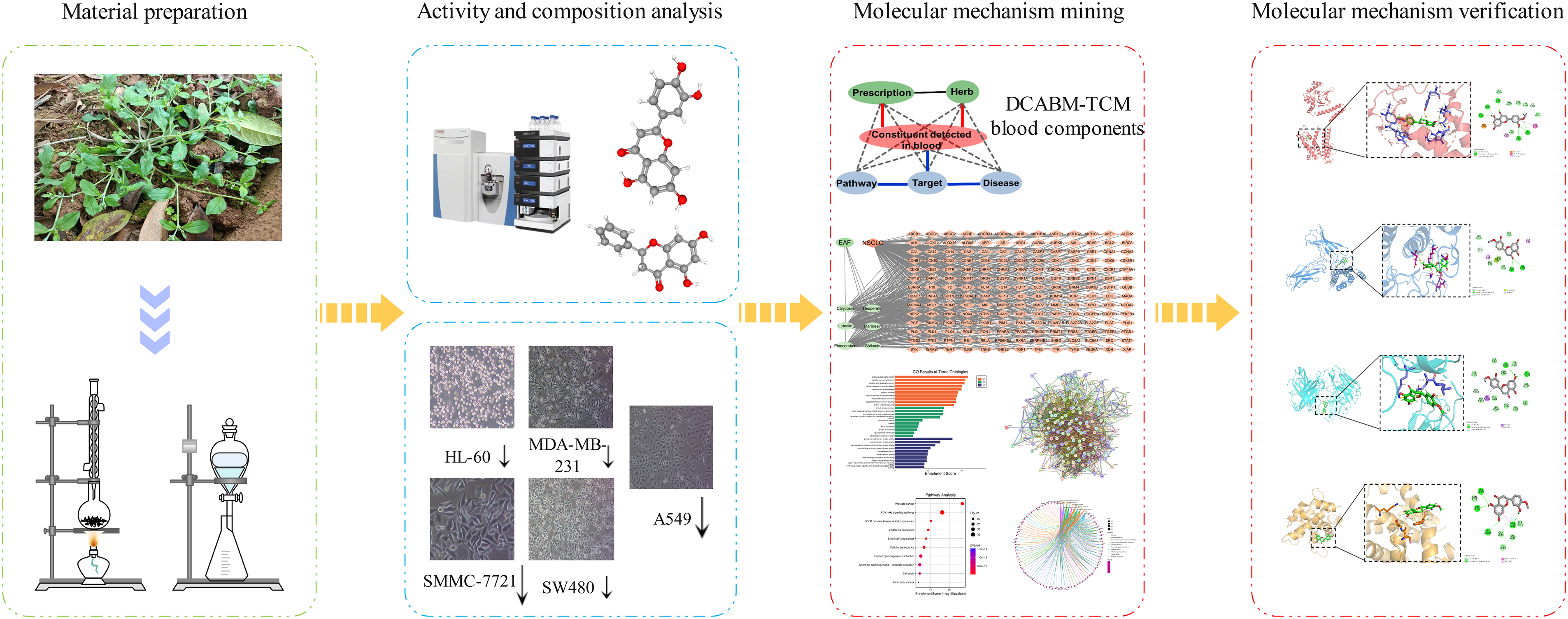
Technical workflow.
The MTS method was selected for in vitro antitumor activity experiments. EAF had certain antitumor effects and inhibited NSCLC growth. The IC50 was 41.13 μg/mL. This method is simple and has high sensitivity, so the experimental results are accurate. The experiment did not include an ethanol control group, mainly because the extraction and concentration processes removed ethanol and had no effect on the results of activity testing.
The effective components were determined by LC-MS, TCMSP, and the DCABM-TCM database. The screening criteria were an OB ≥ 30, a DL ≥ 0.18, and blood component information. There are many methods for the detection of blood components. Yang S-C et al 40 used LC-MS to detect the blood components of the Nourishing Blood Diuretic Formula in rat serum. With the help of advanced big data analysis technology, we used the DCABM-TCM database to identify blood components, which reduced the cost of research and development. However, there are some limitations also. If the database data are not updated in a timely manner and the coverage is not complete, there will be information missing from the results. Therefore, further research and analysis are needed.
GO enrichment analysis revealed 175 GO entries, and the top 20
KEGG enrichment analysis revealed that 146 pathways were significantly enriched. Many of the top 20 signaling pathways were closely related to the treatment of NSCLC. Activation of the PI3K/Akt pathway can promote resistance to cisplatin-induced apoptosis in NSCLC. 42 Inactivating the PI3K/AKT pathway can promote the development of NSCLC. 43 Astragali Radix exerts anti-NSCLC effects through the PI3K/AKT pathway. 44 The role of the PI3K/AKT pathway in NSCLC has been widely studied, and the PI3K/AKT pathway is a signaling pathway that is important in research. CD70 gene expression is upregulated in NSCLC cells with acquired EGFR tyrosine kinase inhibitor resistance, and CD70 inhibits drug resistance in NSCLC by upregulating the expression of the therapeutic target of EGFR, tyrosine kinase inhibitor-resistant CD70. 34 Cell senescence-related genes also affect the treatment of NSCLC. 45 The above analysis showed that the signaling pathways we identified have research value and provide evidence about the molecular mechanism of EAF against NSCLC.
Five key target proteins with high scores were selected based on molecular docking, and many studies have shown that these proteins are related to NSCLC. Experimental validation revealed that MFXD treatment inhibited the proliferation of NSCLC cells by downregulating the expression of EGFR. 46 Inhibition of Akt1 expression increases the sensitivity of NSCLC to drugs. 47 ROS-dependent apoptosis triggered by BCL2 and CASP3 can inhibit the proliferation of NSCLC cells. 48 Based on the above research and analysis, the efficacy of new anti-NSCLC agents has great potential. Modern scientific and technological exploration based on the theory of traditional Chinese medicine is worthy of further study and exploration.
Conclusion
In the present study, we found that EAF had anti-NSCLC activity and analyzed its chemical composition and components in the blood. The molecular mechanism underlying the anti-NSCLC effect was preliminarily predicted to be related to the PI3K-Akt signaling pathway, EGFR tyrosine kinase inhibitor resistance and other signaling pathways. Molecular docking technology verified the accuracy of the results. This study provides a reference for the efficacy and clinical development of
Footnotes
Acknowledgments
This study was supported by the Suqian Talent Officea, Suqian Science and Technology Bureau, and Jinhua Science and Technology Bureau.
Authors’ Note
Cheng-pu Liao and Yan-ping Zhou made equal contributions to the work.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This study was approved by the Ethics Committee of Suqian Affiliated Hospital of Nanjing University of Chinese Medicine (No. 2020SE-1-20-001).
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the 2022 “Suqian Elite” Youth Project, Jinhua Science and Technology Bureau Youth Talent Fund Project, Suqian Science and Technology Bureau Natural Science Foundation Youth Science and Technology Talents Project (Grant Nos. 2023KR03 and K202202).
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There were no human subjects included in this study, and informed consent was not available.
