Abstract
Background
Hepatocellular carcinoma (HCC) was one of the most prevalent and aggressive cancers worldwide, marked by a high mortality rate. Previous clinical treatments predominantly relied on chemotherapeutic agents, which are often associated with severe side effects and a high risk of drug resistance. Therefore, the development of natural compounds for HCC treatment has emerged as a promising therapeutic strategy. Acacetin, a natural flavonoid found in various plants, possesses potent anti-inflammatory and anticancer properties. However, its mechanisms of action remain poorly understood.
Objective
An integrated strategy combining network pharmacology, molecular docking, molecular dynamics simulation, and
Methods
Firstly, we identified acacetin-related targets and HCC-related targets from public databases and determined their overlapping genes. Secondly, network pharmacology and enrichment analyses were performed to predict the potential biological functions and mechanisms underlying the anti-HCC effects of acacetin. Subsequently, molecular docking and molecular dynamics simulation were conducted to evaluate the stability and reliability of the interactions between candidate targets and acacetin. Finally, a series of
Results
Network pharmacology analysis identified 125 targets of acacetin, 2228 HCC-related targets, and 77 overlapping targets. The protein-protein interaction (PPI) network construction dentified six core targets: TP53, AKT1, EGFR, CASP3, ESR1, and MMP9. GO and Kyoto Encyclopedia of Genes and Genomes enrichment analyses revealed significant involvement of negative regulation of apoptosis, the PI3K-Akt signaling pathway, and the p53 signaling pathway. Molecular docking and molecular dynamics simulation confirmed stable binding between acacetin and the core targets.
Conclusion
This study demonstrated the multi-target and multi-pathway nature of acacetin in the treatment of HCC. The proposed mechanism involves inhibition of key genes MMP9, suppression of the PI3K-AKT signaling pathway, reduced cancer cell proliferation, and induction of apoptosis. These findings offered a comprehensive theoretical basis for the therapeutic potential of acacetin in HCC treatment.
This is a visual representation of the abstract.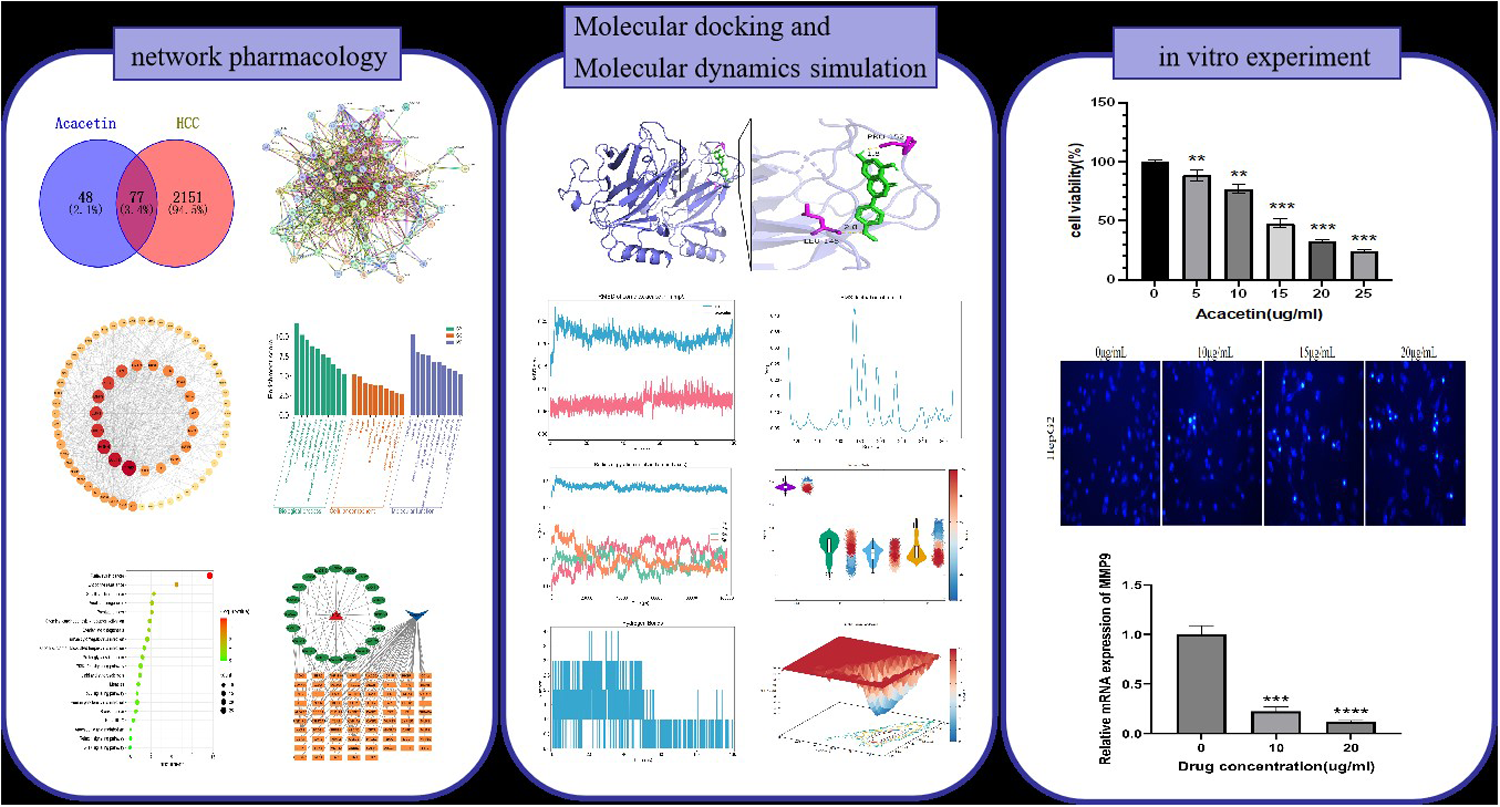
Introduction
Hepatocellular carcinoma (HCC), a malignant tumor originating from hepatocytes, accounted for approximately 85% to 90% of primary liver cancers and was recognized as the second leading cause of cancer-related deaths worldwide. 1 In our country, it was identified as the fourth most common malignancy, posing a significant threat to public health. Although the combination of chemotherapy, surgery, and radiotherapy remains a cornerstone of HCC treatment, challenges such as drug resistance and severe adverse effects are frequently encountered. 2 The pathogenesis of HCC involves complex interactions among multiple genes, molecular pathways, and environmental factors, which aligns with the multi-component, multi-target, and multi-pathway characteristics of traditional Chinese medicine (TCM). In recent years, TCM has garnered increasing attention in medical research due to its low toxicity and ability to exert multi-target regulatory effects in the treatment of malignant tumors. 3 With ongoing advancements in biochemical and pharmacological research, TCM is increasingly acknowledged as a promising source of alternative and complementary anti-tumor therapies.4,5
Acacetin, a natural flavonoid compound, was widely distributed in nature and could be found in various plants such as the acacia tree, thistles, herbs, and chrysanthemum. 6 It was extensively studied and was shown to possess a broad spectrum of biological activities, including antimicrobial, anti-inflammatory, anticancer, antioxidant, anti-obesity, anti-malarial, and vascular relaxant effects. 7 In recent years, the potential anti-tumor value of acacetin was increasingly explored, particularly in cancers such as lung cancer, HCC, and gastric cancer. The anti-tumor mechanisms of acacetin were primarily attributed to several key actions: inhibition of the expression of relevant receptors and transcription factors; suppression of carcinogenic metabolite secretion; inhibition of tumor cell proliferation; promotion of tumor cell apoptosis; regulation of key signaling pathways; prevention of tumor invasion and migration; and inhibition of VEGF production.8–13 Unlike conventional anti-tumor agents, acacetin was notable for its multi-target activity, enabling it to simultaneously modulate multiple molecular pathways—a key advantage that positioned it as a promising candidate for cancer therapy. Further research into the mechanisms and clinical applications of acacetin was expected to offer new insights and opportunities for the development of more effective anti-tumor therapies.
To gain a deeper understanding of the pharmacological mechanisms by which drugs treat diseases, various computational methods were increasingly being employed, including network pharmacology, molecular docking, and molecular dynamics simulation. Network pharmacology was a novel approach that integrated systems biology, network analysis, and pharmacology to analyze the potential mechanisms of action of single or compound drugs. It was frequently used to investigate the multi-target effects of complex natural compounds and served as a powerful tool for exploring the pharmacological effects, mechanisms, and toxicity of TCM. 14 Molecular docking was a computational simulation technique used to predict the binding modes and affinities between small molecules (drugs) and protein receptors (targets). 15 Molecular dynamics simulation further verified and refined these binding interactions, offering a more detailed, molecular-level understanding of the drug-target relationship. 16 The development and application of network pharmacology, molecular docking, and molecular dynamics simulation had enhanced research on the safety, efficacy, and mechanisms of TCM, thereby strengthening its credibility and increasing its acceptance in modern medicine.
In this study, a combination of network pharmacology, molecular docking, and molecular dynamics simulation was employed to comprehensively elucidate the anti-HCC potential of acacetin, identify its molecular targets, and explore its mechanisms of action.
Materials and Methods
Network Pharmacology
Acacetin Target Screening
The structure of acacetin was obtained from the TCMSP (http://tcmspw.com/tcmsp.php) database and imported into the PharmMapper database (https://lilab-ecust.cn/pharmmapper/index.html). Targets with normalized fit scores greater than 0.9 were considered as potential targets of acacetin. The canonical SMILES information of acacetin was retrieved from PubChem (https://pubchem.ncbi.nlm.nih.gov/) and then uploaded to the Swiss Target Prediction database (https://SwissTargetPrediction.cn) to identify additional potential targets. Using “acacetin” as a keyword, relevant targets were also identified in the TCMSP database. After compiling all target information related to acacetin from the aforementioned sources, duplicate targets were removed to generate a non-redundant list of acacetin-associated targets. All target names were standardized using the UniProt database (https://sparql.uniprot.org/).
Acquisition of Disease Targets
“Hepatocellular carcinoma” was used as the keyword to search for HCC-related targets in the OMIM (https://www.OMIM.org), TTD (https://db.idrblab.net/ttd/), and GeneCards database (https://www.GeneCards.org). The search results from these three databases were integrated, and gene names were standardized using the UniProt database (https://www.uniprot.org/).
Screening of Common Targets of Acacetin-HCC
The Venn diagram of common drug-disease targets was generated by inputting the acacetin targets and HCC disease targets into the online tool Venny 2.1.0 (https://bioinfogp.cnb.csic.es/tools/venny/index.html).
PPI Network Construction and Acquisition of Core Targets
The common drug-disease targets were input into the STRING database (http://string-db.org/) to construct a protein-protein interaction (PPI) network, with the species set to Homo sapiens and a confidence score threshold above 0.400. 17 The resulting PPI network was then imported into Cytoscape version 3.10.2 for visualization and further analysis. Core targets were identified through cluster analysis using the CytoNCA plugin within Cytoscape.
GO Functional Analysis and KEGG Pathway Analysis
To identify intersecting targets, the DAVID database (https://david.ncifcrf.gov) was used to perform Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses.
18
The species was set to Homo sapiens, and a significance threshold of
Construction of “Drug-Target-Pathway-Disease” Network
The “drug-target-pathway-disease” network was constructed using Cytoscape version 3.10.2. To clearly illustrate the relationships between key targets and pathways, only targets associated with the top 20 KEGG pathways ranked by smallest
Molecular Docking
Molecular docking between acacetin and the core targets was performed to verify their interactions. First, the molecular structure of acacetin was retrieved from the PubChem database (https://pubchem.ncbi.nlm.nih.gov) and exported in MOL2 format. The 3D structures of the core proteins were obtained from the Protein Data Bank (https://www.rcsb.org/). Ligands and protein structures were imported into AutoDock Tools software. 20 The crystal structures of the target proteins were prepared by removing water molecules and adding hydrogen atoms to ensure compatibility with the ligand's low-energy conformation. Molecular docking between acacetin and the core proteins was then conducted, and the results were visualized using PyMOL. 21 Binding activity was evaluated based on binding energy: a binding energy ≤−5.0 kcal/mol (1 cal = 4.2 J) indicated good binding affinity, while a binding energy ≤ −7.0 kcal/mol suggested strong binding activity. 22
Molecular Dynamics Simulation
To explore the stability of the protein-ligand interactions in greater depth, molecular dynamics simulation was performed on acacetin-MMP9 complexe, using GROMACS 2023.2 software. 23 The CHARMM 36 force feld was employed for the topology generation of the ligand and protein. 24 The complex was entered into the Solution Biulder module of CHARMM-GUI to establish a solvent box for the system at least 2 Å away from the complex boundary.25,26 TIP3, sodium ion and chloride ion were added to neutralize the system. Energy minimization was performed for 50 000 steps, followed by equilibration through a 125 ps simulation in the NPT ensemble at fixed volume and constant heating rate with a 2.0 fs time step. Finally, MD simulation of the docking complex was carried out for 200 ns in 303.15 K temperature and 1 bar pressure with a time step of 2 femtoseconds. Based on the results of the MD simulation, we calculated the root mean square deviation (RMSD), root mean square fluctuation (RMSF), radius of gyration (Rg) values, and hydrogen bonds (H-bonds) of the complex. Gibbs free energy landscape was drawn.
In Vitro Validation Experiment
Materials
A variety of reagents and materials were used in this study. Acacetin (Lot No.: 8012257C294), dimethyl sulfoxide (DMSO; Lot No.: D0798), and the Cell Counting Kit-8 (CCK-8; Lot No.: MA02181108H) were procured from Report Company. Hoechst 33258 stain (Lot No.: P9002-B0029) was supplied by RedPartyTech Company. The SparkZol Reagent (Lot No.: AC0101) was provided by SparkJade. The Reverse Transcription Kit (Lot No.: BL696A) and Universal SYBR qPCR Master Mix (Lot No.: BL697A) were obtained from BioSharp.
Cell Culture
Hep-G2 human liver cancer cells (ATCC: HB-8065; lot number: 20240313) were purchased from Shanghai Yizefeng Biotechnology Co., Ltd Cell culture materials, including MEM medium (lot number: 2403033) and fetal bovine serum (lot number: 2342332), were obtained from Vivacell.
Culture Conditions of Hep-G2 Human HCC Cells:Hep-G2 cells were cultured in MEM complete medium containing 10% fetal bovine serum (FBS). Cells were incubated at 37 °C in a sterile CO2 incubator with 5% CO2. Passaging was performed every 48 h by treating the cells with trypsin and splitting at a 1:2 ratio.
Drug Concentration Setting
The acacetin treatment concentrations were selected based on previous literature 27 and further optimized through preliminary experiments. The control group received cell medium containing 0.3% DMSO, while the treatment groups were exposed to varying concentrations of acacetin (5, 10, 15, 20, and 25 μg/mL). Following the CCK-8 assay, the most appropriate concentration was chosen for subsequent experiments.
Cell Viability Assays
The CCK-8 assay kit was employed to evaluate the effect of acacetin on the proliferation of Hep-G2 human HCC cells. Cells were seeded into 96-well plates and allowed to adhere for 24 h. Subsequently, acacetin was administered at concentrations of 5, 10, 15, 20, and 25 μg/mL, with a control group treated with 0.3% DMSO-containing medium. Each group included six replicate wells. After a 24-h incubation, cell viability was assessed by measuring the absorbance at 450 nm. The following formula was used to calculate cell viability: Cell viability (%) = (ODacacetin - ODblank) / (ODcontrol - ODblank) × 100%. 28 All experiments were conducted in triplicate to ensure reproducibility.
Hoechst 33258 Staining
A suspension of Hep-G2 cells at a density of 1 × 105 cells/mL was seeded into 12-well plates. Cells were treated with acacetin at concentrations of 10, 15, and 20 μg/mL, with three replicate wells per concentration, followed by a 24-h incubation. After incubation, the culture medium was carefully removed, and the cells were washed twice with pre-cooled PBS. The cells were then fixed with 1 mL of 4% paraformaldehyde for 15 min. Following fixation, the cells were washed once with pre-cooled PBS. Subsequently, the cells were stained with 500 μL of Hoechst 33258 (10 μg/mL) for 10 min at 25 °C. After staining, the dye was removed, and the cells were washed twice with PBS to eliminate residual stain. Morphological changes were observed using an inverted fluorescence microscope. Under fluorescence microscopy, viable cells exhibited uniformly stained blue nuclei, while apoptotic cells showed intensely fluorescent blue nuclei with condensed chromatin. 29 These observations were used to evaluate the apoptotic effects induced by acacetin. All conditions were assessed in triplicate independent experiments.
RNA Extraction and qRT-PCR
Total RNA was extracted with SparkZol reagent, according to the manufacturer's instructions. RNA concentration was quantified using a microplate reader. Subsequently, cDNA synthesis and quantitative PCR (qPCR) were performed by the Reverse Transcription Kit and Universal SYBR qPCR Master Mix, respectively, each according to the manufacturer's instructions. Relative expression levels of MMP9 was quanti-fied according to the 2−ΔΔCt. The mRNA levels of tested genes were normalized glyceraldehyde-3-phosphate dehydrogenase (Gapdh) as an internal reference. Quantitative real-time PCR (qRT-PCR) assays were conducted with three independent biological replicates for each treatment group to ensure data reliability.
Statistical Analysis
In the network pharmacology analysis, all statistical evaluations were performed using the DAVID database. For
Results
Common Potential Targets of Acacetin and HCC
A total of 125 potential targets of acacetin were identified using the PharmMapper, Swiss Target Prediction, and TCMSP databases. Additionally, 2228 HCC-related targets were obtained from the OMIM, TTD, and GeneCards databases. As was observed in Figure 1, 77 overlapping targets were identified through Venny analysis.

Venn Diagram of Common Targets of Acacetin and HCC. HCC, Hepatocellular carcinoma.
PPI Network Construction and Acquisition of Core Genes
77 potential targets of acacetin in the treatment of HCC were imported into the STRING database to construct a PPI network. According to Figure 2, there were 77 nodes and 662 edges in PPI interaction (
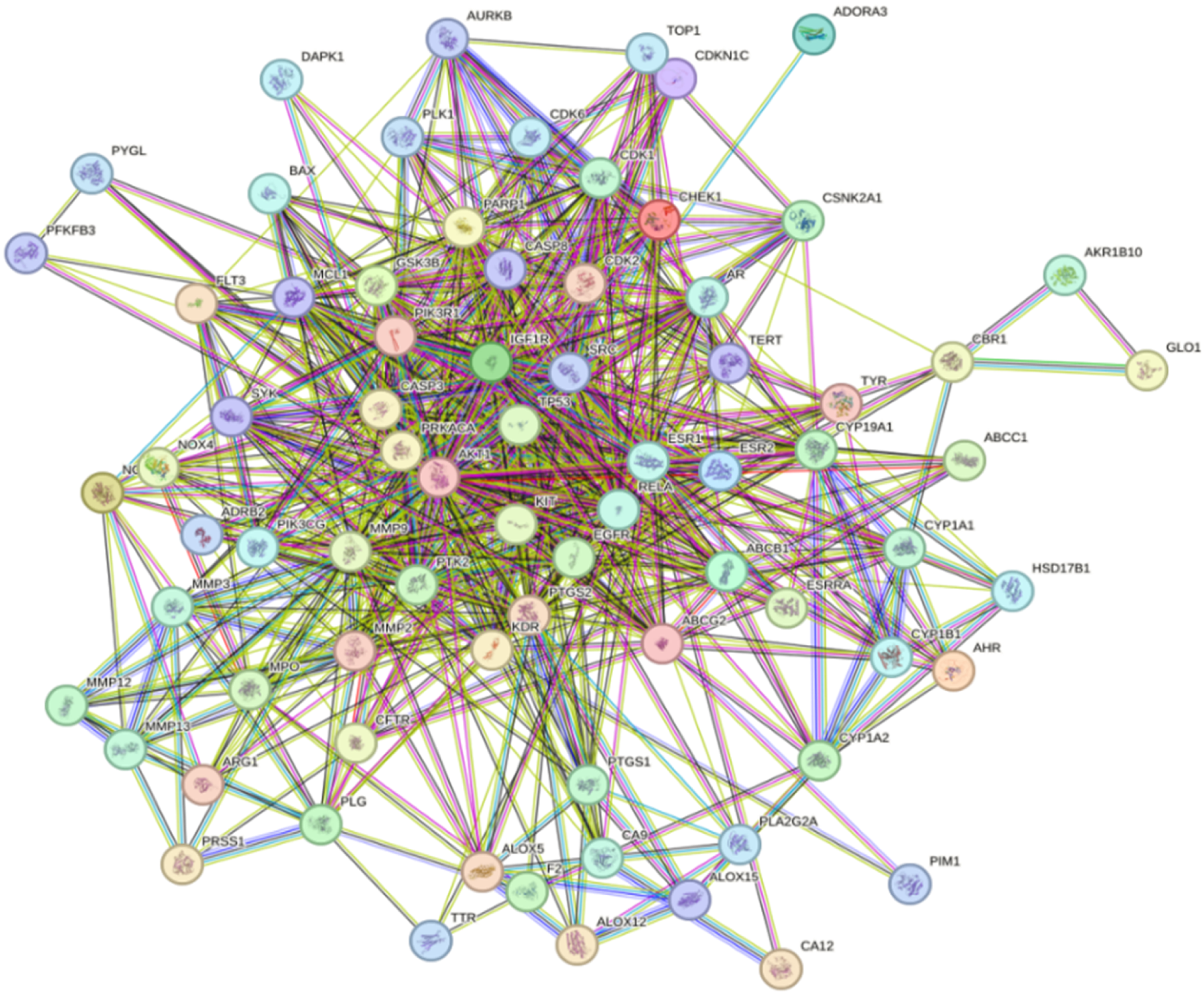
PPI Network of Common Targets of Acacetin and HCC. HCC, Hepatocellular carcinoma; PPI, protein-protein interaction.

PPI Network Diagram of Core Targets. PPI, protein-protein interaction.
GO Functional Analysis and KEGG Pathway Analysis
Gene Ontology (GO) enrichment analysis showed that the 77 intersecting genes were significantly enriched in 453 GO terms (
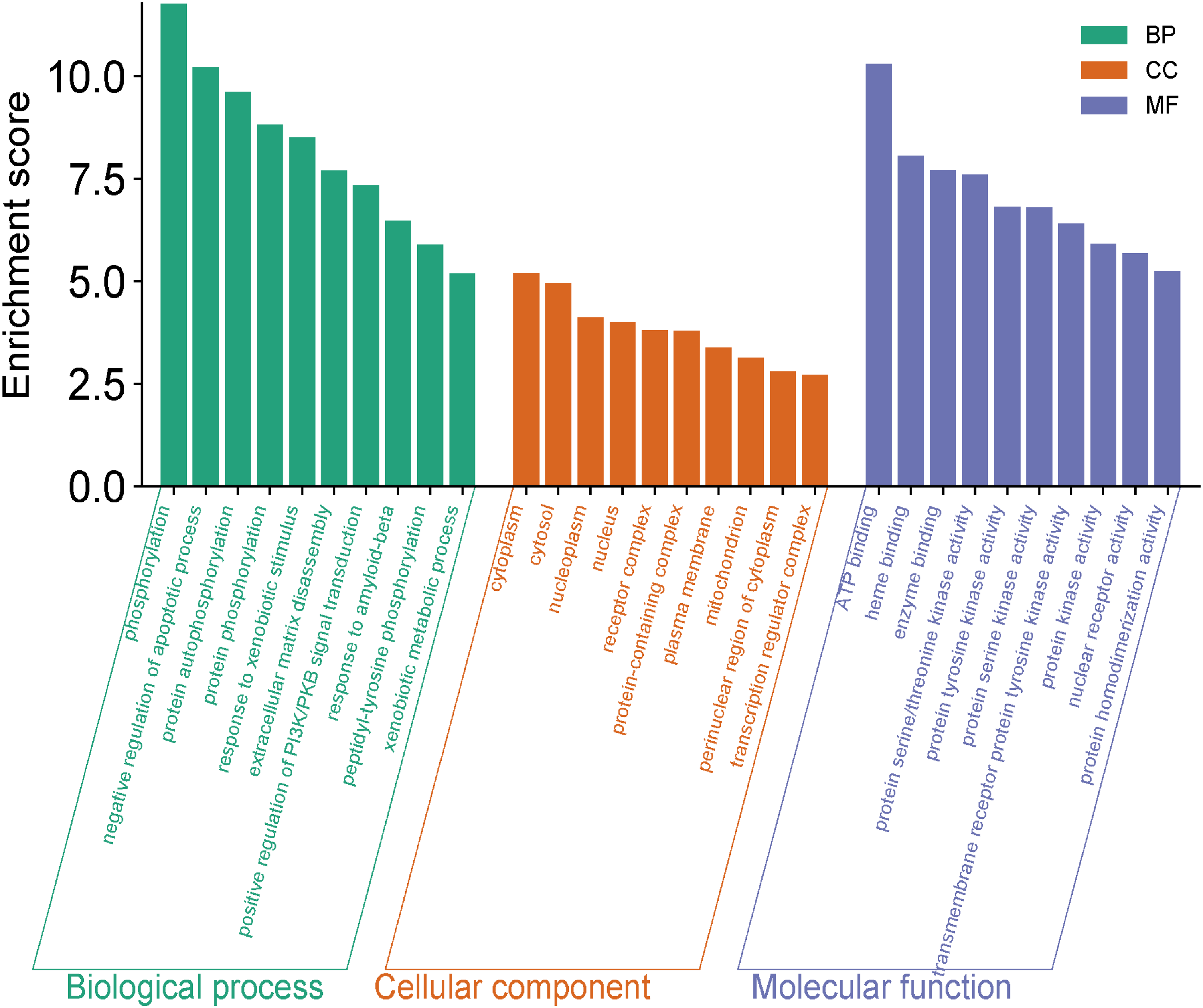
Histogram of GO Functional Enrichment Analysis.
To investigate the representative signaling pathways associated with the key targets, KEGG enrichment analysis was performed. A total of 116 significantly enriched signaling pathways were identified. Based on statistical data, pathways with smaller
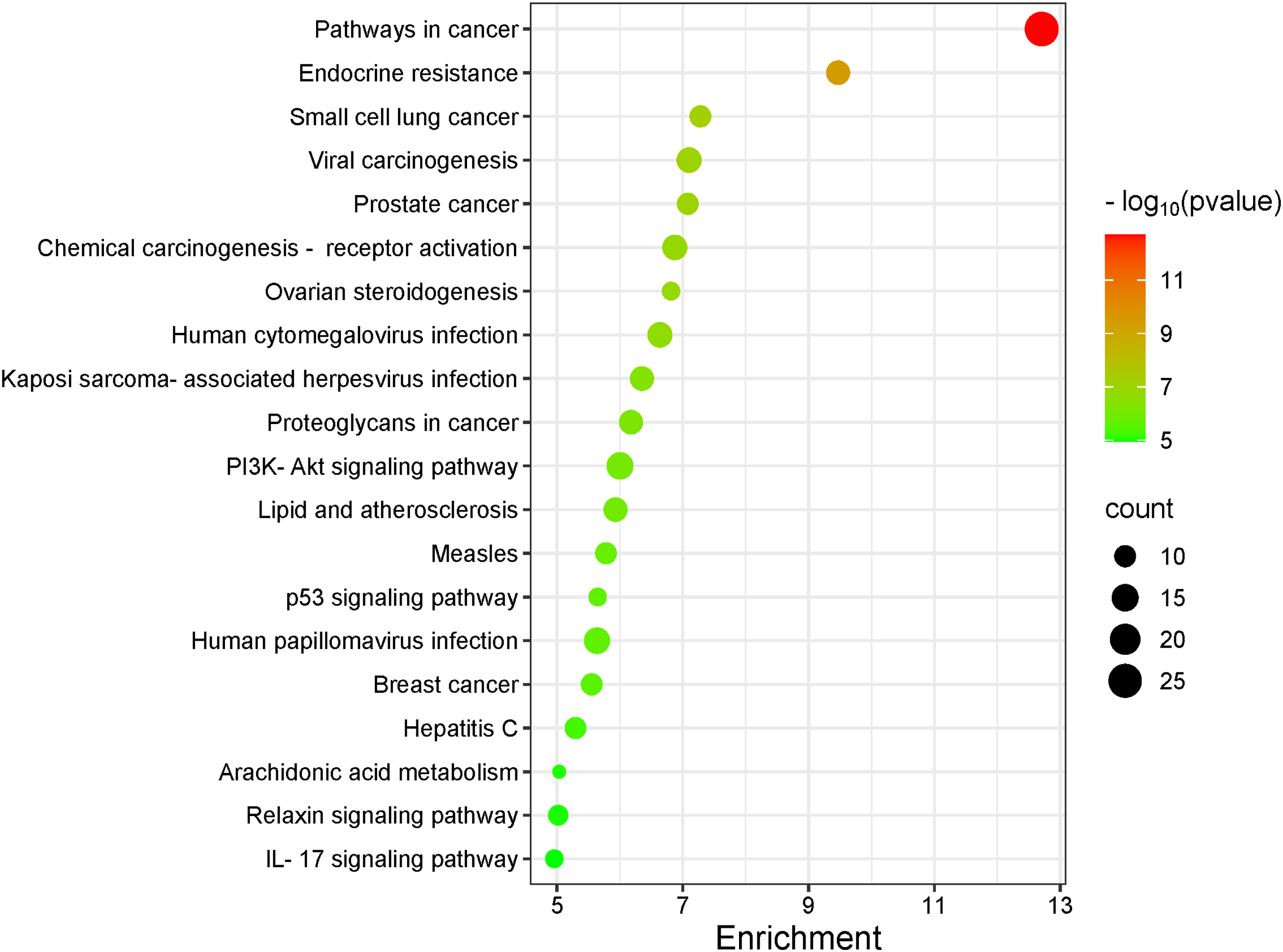
Bubble Diagram of KEGG Channel Enrichment Analysis. KEGG, Kyoto Encyclopedia of Genes and Genomes.
The “Drug-Target-Pathway-Disease” Network
Cytoscape 3.10.2 software was used to construct a network diagram illustrating the relationships among acacetin, 77 common targets, the top 20 signaling pathways, and HCC, as illustrated in Figure 6. The resulting “drug-target-pathway-disease” network consisted of 99 nodes and 132 edges. In the diagram, red represented the disease, blue indicated acacetin, green denoted the top 20 signaling pathways, and orange corresponded to the 77 core targets.

The “Drug-Target-Pathway-Disease” Network.
Molecular Docking
Molecular docking studies were performed using acacetin as the ligand and six key protein targets—TP53, AKT1, EGFR, CASP3, ESR1, and MMP9—as receptors. A greater absolute value of binding energy indicated a more stable interaction between the ligand and the receptor. The results demonstrated that acacetin exhibited favorable binding affinities with all six key targets (Figure 7). Specifically, the binding energy between acacetin and TP53 was −8.68 kcal/mol, with hydrogen bonds formed at PRO-152 and LEU-145. For AKT1, acacetin showed a binding energy of −6.27 kcal/mol, interacting via hydrogen bonds at SER-56, ALA-58, LEU-78, GLN-59, ARG-76, and GLN-61. The binding energy with EGFR was −7.24 kcal/mol, involving hydrogen bonds at LEU-445, ALA-448, and ASP-92. In the case of CASP3, the binding energy was −6.28 kcal/mol, with hydrogen bonds at PRO-42, ASN-80, MET-39, and HIS-277. Acacetin bound to ESR1 with a binding energy of −8.03 kcal/mol, forming hydrogen bonds at THR-347, LYS-531, and ASN-532. The strongest binding was observed with MMP9, where acacetin exhibited a binding energy of −10.18 kcal/mol and formed hydrogen bonds at ALA-242, MET-247, and ARG-249. Among all, MMP9 showed the most robust binding affinities, suggesting their critical involvement in the therapeutic effects of acacetin against HCC.
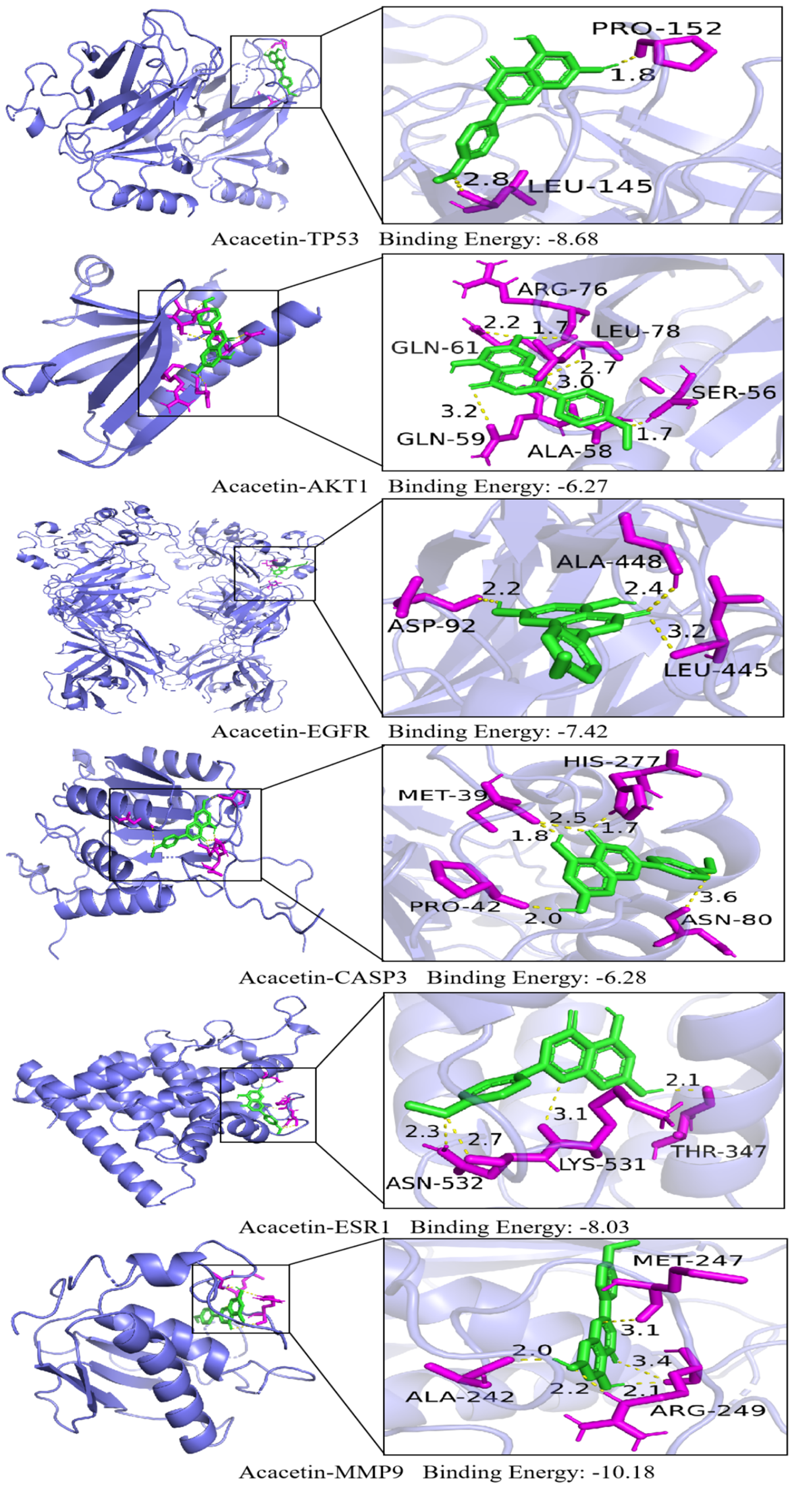
Molecular Docking Diagram of Acacetin and Core Target.
Molecular Dynamics Simulation
To further evaluate the effects of acacetin on MMP9 protein, molecular dynamics (MD) simulation analysis was conducted to assess the stability of the protein-ligand complexe over a 200-nanosecond (ns) period. The simulation analysis included the evaluation of RMSD, RMSF, radius of gyration (Rg), hydrogen bonds, and three-dimensional (3D) free energy topography. These analyses were performed to determine whether the interactions between the ligand and receptor were maintained and remained stable throughout the simulation period. The stability and persistence of these interactions were critically examined to gain insights into the binding mechanisms and conformational dynamics of the complexe.
Root Mean Square Deviation (RMSD)
RMSD values were employed to assess whether the simulation systems reached equilibrium, with RMSD values below 1 nm considered indicative of relatively stable protein-ligand interactions under physiological conditions. 31 As depicted in Figure 8A, the RMSD trajectories of both the MMP9-acacetin complex and the MMP9 protein exhibited a slight increase during the initial phase of the simulation and stabilized after 80 ns. The average RMSD value of the Acacetin-MMP9 complex was 0.36 ± 0.03 nm, with a total fluctuation range of less than 0.1 nm. These results suggest that binding of the small-molecule ligand did not induce significant conformational changes in the protein, indicating a relatively stable interaction between the two molecules.

The MD Simulation Results of Acacetin and MMP9. (A) the RMSD of Acacetin with MMP9; (B) the RMSF of Acacetin with MMP9; (C) the Rg of Acacetin with MMP9; (D) the Hydrogen Bond Number of Acacetin with MMP9; (E) 3D Gibbs Free Energy Landscape. RMSD, root mean square deviation; RMSF, root mean square fluctuation.
Root Mean Square Fuctuation (RMSF)
RMSF was employed as a statistical measure in molecular dynamics simulations to quantify the positional deviations of atoms over time. It represented the extent to which an atom fluctuated around its average position. Higher RMSF values indicated greater atomic mobility in specific regions, suggesting increased flexibility or involvement in dynamic processes. The RMSF plot in Figure 8B demonstrated that the fluctuation in amino acid residues with the MMP9 protein ranged from 0.1 to 0.8 nm. Notably, residues within the 170-220 region exhibited more pronounced fluctuations, suggesting that these regions may have exhibited unstable conformations. This local instability was likely due to frequent non-covalent interactions between acacetin and surrounding residues, which led to increased flexibility in these regions. Based on these observations, it was hypothesized that these regions could represent potential binding sites for the protein.
The Radius of Gyration (Rg)
The radius of gyration (Rg) was employed to evaluate the folding behavior of protein-ligand complexes during molecular dynamics (MD) simulations. It provided insights into the overall compactness of the complexes. For molecules of a given mass, more compact structures exhibited lower Rg values, whereas extended or loosely packed structures showed higher Rg values, reflecting greater flexibility and internal motion. As illustrated in Figure 8C, the complexe exhibited stable fluctuations throughout the 0-200 ns simulation period. Specifically, the Rg values of the MMP9-acacetin complex remained steady between 1.5 and 1.6 nm. This finding was consistent with the RMSD results, indicating that the protein maintained a stable and compact conformation. Furthermore, the binding of acacetin did not significantly affect the structural stability of the protein.
Hydrogen Bonds
The number of hydrogen bonds was recognized as an important indicator for maintaining the relative orientation between molecules, as hydrogen bonds were among the most common and readily formed non-covalent interactions. In biomacromolecular systems—such as enzyme-substrate complexes, receptor-ligand interactions, and drug-target binding-hydrogen bonds were considered critical contributors to specific molecular recognition, helping to define the binding mode and affinity between interacting partners. The formation of hydrogen bonds between acacetin and MMP9 was analyzed throughout the simulation, as depicted in Figure 8D. It was observed that, in most cases, 2-3 hydrogen bonds were formed. These interactions contributed to stabilizing the ligand-receptor complex and maintaining the structural integrity of the system.
3D Free Energy Topography Analysis
The Gibbs free energy landscape was used to evaluate the conformational stability of the complex. 32 RMSD and Rg were selected to construct the energy landscape and explore the steady-state structures of the system. As shown in Figure 8E, the Gibbs free energy landscape of the MMP9-acacetin complex revealed that the blue and purple regions corresponded to lower energy states, indicating that the complex adopted stable conformations in these free energy minima. 33 The 3D free energy surface demonstrated a significant energy well in the molecular dynamics (MD) simulations, suggesting that the system reached its most stable structural states. Specifically, when the Rg value ranged from 1.48 to 1.56 nm and the RMSD value from 0.25 to 0.40 nm, the free energy of the MMP9-acacetin complex reached its minimum.
Effects of Acacetin on Proliferative Activity of Hep-G2 Cells
The effect of acacetin on the proliferative activity of Hep-G2 cells was assessed using the CCK-8 assay after 24 h of treatment with various concentrations of the compound. Compared with the control group, acacetin significantly inhibited the viability of Hep-G2 cells in a concentration-dependent manner (
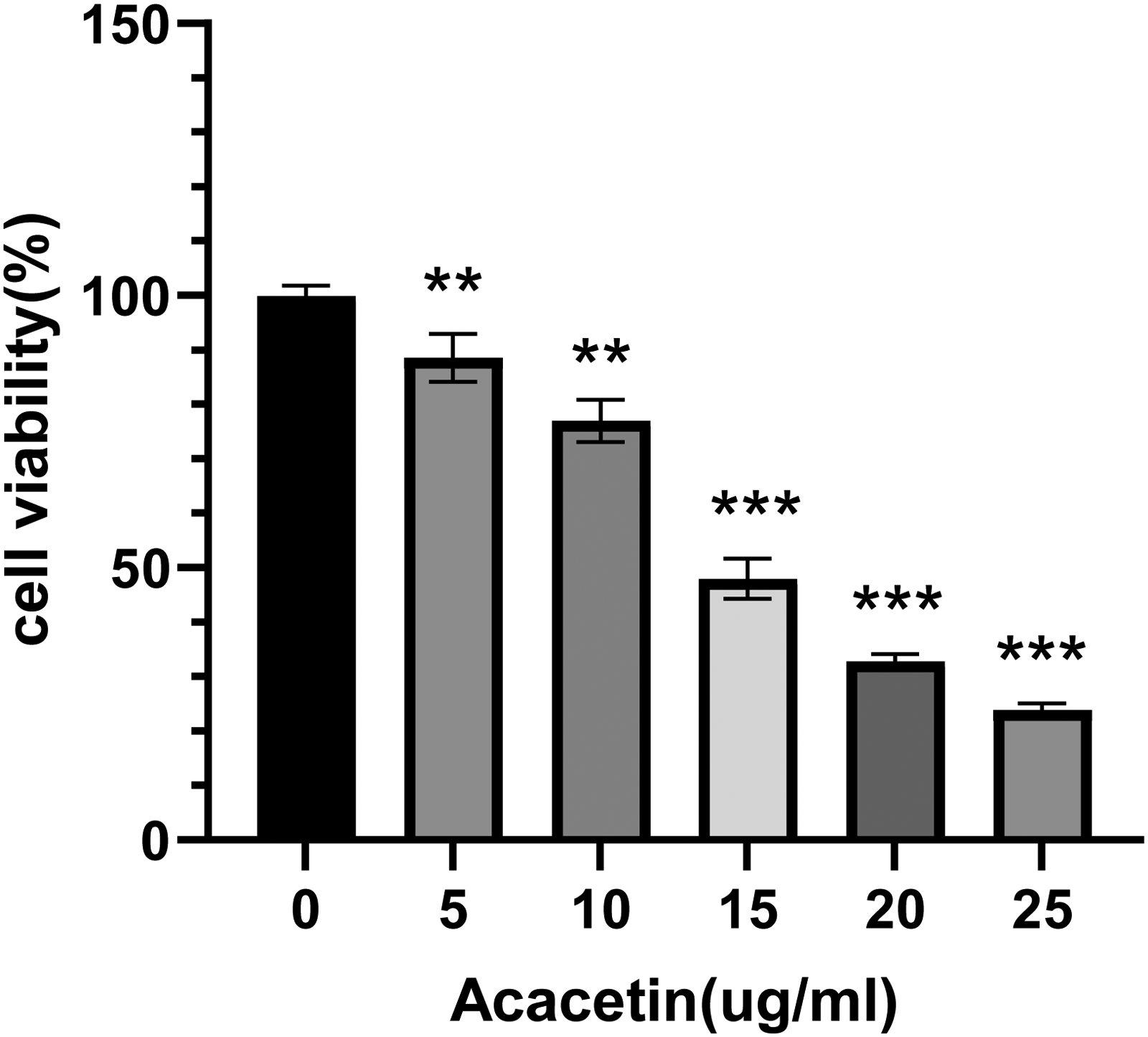
Inhibitory Effect of Acacetin on the Proliferation of Hep-G2 Cells. Error Bars Denote Standard Deviation (SD) from Biological Replicates (n = 3). **
Effects of Acacetin on Apoptosis of Hep-G2 Cells
Fluorescence microscopy revealed that acacetin significantly induced apoptosis in Hep-G2 cells. Compared with the untreated control group, cells treated with increasing concentrations of acacetin (Figure 10) exhibited progressively more pronounced chromatin fragmentation (intense blue fluorescence) and nuclear condensation—hallmarks of apoptotic morphology. These results suggest that acacetin induces apoptosis in Hep-G2 cells in a concentration-dependent manner.
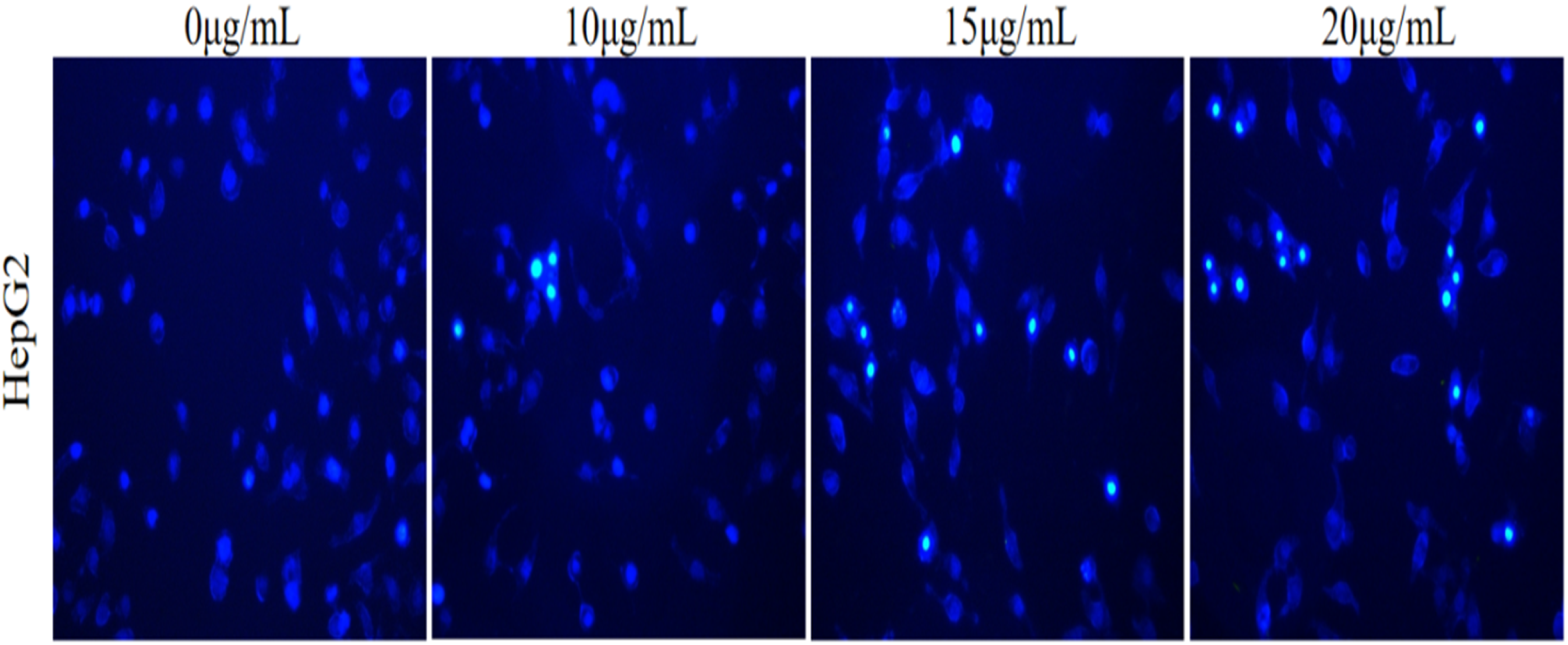
Hep-G2 Cell Apoptosis Promoted by Acacetin.
Effects of Acacetin on Target Gene MMP9
To investigate the effect of acacetin on MMP9 mRNA expression in Hep-G2 cells, we performed quantitative reverse transcription PCR (qRT-PCR). Hep-G2 cells were treated with varying concentrations of acacetin (0, 10, and 20 μg/mL) for 24 h. qRT-PCR analysis revealed that acacetin significantly suppressed MMP9 mRNA expression in a dose-dependent manner compared to the untreated control (Figure 11).

Acacetin Decreased MMP9 Expression in Hep-G2 Cell (***
Discussion
HCC was the sixth most common malignant tumor and the third leading cause of cancer-related death worldwide. Approximately 650 000 patients died from HCC each year, with 75% to 80% of these cases occurring in Asian countries, and 55% of all HCC cases reported in China. 34 Currently, surgical resection was still the primary treatment modality for HCC. However, due to the insidious onset of the disease, most patients were diagnosed at intermediate or advanced stages. Fewer than 20% of patients were eligible for surgical resection, and the postoperative recurrence rate remained high, severely affecting prognosis and overall survival. 35 Therefore, drug therapy was still the mainstay for patients with intermediate and advanced HCC. Compared to conventional chemotherapeutic agents, TCM had been widely employed in the treatment and prevention of liver diseases—particularly liver cancer—due to its multi-component, multi-target mechanisms, low toxicity, and reduced incidence of side effects. The use of TCM could prolong the median survival of liver cancer patients and improve their overall survival. TCM had demonstrated the ability to prolong median survival and improve overall outcomes in liver cancer patients. Moreover, combining TCM with Western medicine could mitigate drug resistance in liver cancer treatment. 36 With the progress of pharmacological research, TCM had gained increasing recognition as a valuable source of alternative and complementary antineoplastic agents. 5
As a natural flavonoid compound, acacetin was recognized for its diverse biological activities, including antioxidant, anti-inflammatory, antibacterial, antiviral and antitumor effects.
7
Acacetin had been demonstrated to possess potential pharmacological effects against HCC. Previous studies revealed that its anti-HCC mechanisms involved the suppression of HepG2 cell proliferation by blocking cell cycle progression at the G1 phase.
37
Additionally, acacetin was found to induce apoptosis in HCC cells by antagonizing the non-genomic effects of retinoic acid receptor gamma (RARγ) on AKT and p53 signaling.
38
Furthermore, it was reported that acacetin inhibited the proliferation and migration of Hep-G2 cells and promoted apoptosis by activating mitochondrial autophagy via the PINK1/Parkin pathway.
39
Based on these findings, we speculated that acacetin may exert preventive and therapeutic effects against HCC by regulating multiple targets, signaling pathways, and biological processes. However, existing research has focused on only a limited number of targets and pathways, and the underlying mechanisms remain incompletely understood. Therefore, a systematic investigation into the pharmacological mechanisms of acacetin was warranted. In this study, we employed a network pharmacology approach and conducted a comprehensive computational analysis. Several potential targets and signaling pathways were predicted and subsequently validated through molecular docking and molecular dynamics simulations, confirming the spatial and energetic compatibility between acacetin and its putative targets. Furthermore,
In this study, the potential targets and associated signaling pathways of acacetin in the treatment of HCC were explored using network pharmacology. A total of 125 targets of acacetin, 2228 HCC-related targets, and 77 overlapping targets were identified. A PPI network was constructed based on the intersecting genes shared by acacetin and HCC. Subsequent analysis revealed that TP53, AKT1, EGFR, CASP3, ESR1, and MMP9 were significantly prioritized as key nodes. These findings provide critical insights for further investigation into the potential mechanisms underlying acacetin's regulatory effects on HCC.
In the present study, molecular docking was employed to evaluate the interaction and binding force between acacetin and key protein targets. The protein-ligand complexe with the highest binding affinities was subsequently selected for molecular dynamics simulation (MDS) to assess the stability of these interactions and to preliminarily validate the predicted targets. Docking results revealed that the binding energies between acacetin and the key targets (TP53, EGFR, ESR1, MMP9) were all ≤−7.0 kcal/mol, indicating stable interactions. Notably, MMP9 exhibited the strongest binding affinities. Furthermore, MDS confirmed the structural stability of the acacetin-MMP9 complexe. These findings suggested that the complexe remained stable once formed, further supporting the effective interaction between acacetin and MMP9. Collectively, molecular docking and MDS provided preliminary validation of the predicted targets, indicating that acacetin may exert its anticancer effects against HCC by modulating these key proteins.
The TP53 protein is a major component that regulates various pathways of cellular stress response, and was a common tumor suppressor gene that can regulate apoptosis and aging of tumor cells. 40 TP53 mutations are closely related to various tumors and play a negative regulatory role in anti-tumor immunity, making it one of the most common genetic changes in HCC. 41 Previous studies confirmed that acacetin could specifically bind to TP53 in gastric cancer (GC) cells, inhibiting GC cell viability by promoting TP53 phosphorylation. 42 Matrix metalloproteinases (MMPs) family is a zinc-dependent enzyme which is involved in the degradation of the extracellular matrix both in physiological and pathological processes, such as embryonic development, tissue remodeling, arthritis and tumor progression and metastasis. 43 MMP9 is one of key member of MMPs family. Elevating MMP9 expression plays a key role in the progression of many tumors, such as bladder cancer, esophageal squamous cell carcinoma, and intrahepatic cholangiocarcinoma. It contributes to tumor angiogenesis by regulating growth plate angiogenesis and intervening in endothelial stem cell recruitment. 44 It had also been reported that acacetin reduces the expression of MMP2, MMP9, and u-PA by inhibiting the p38 MAPK signaling pathway and suppressing the activity of NF-κB and AP-1, thereby attenuating tumor cell invasion and migration. 45
The results of GO enrichment analysis primarily involved the negative regulation of the apoptosis signaling pathway. CCK-8 assay and Hoechst staining further confirmed that acacetin inhibited the proliferation and promoted the apoptosis of Hep-G2 cells in a concentration-dependent manner. KEGG pathway enrichment analysis revealed that acacetin may regulate cancer-related pathways, including hepatitis C, proteoglycans in cancer, the PI3K/Akt signaling pathway, p53 signaling pathway, and IL-17 signaling pathway, all of which are closely associated with HCC and inflammation. These findings suggested that acacetin exerted its anti-liver cancer effects by modulating multiple signaling pathways. The PI3K/Akt signaling pathway was known to participate in the development of various cancers by influencing apoptosis, autophagy, and cell survival. 46 Increasing evidence had demonstrated that this pathway played a pivotal role in the anticancer activity of acacetin. For instance, acacetin was reported to inhibit gastric cancer invasion, metastasis, and TGF-β1-induced epithelial-mesenchymal transition through the suppression of the PI3K/Akt/Snail pathway. 47 Moreover, acacetin was found to counteract mesenchymal cell-induced malignancy in ovarian cancer by blocking the RAGE-PI3K/Akt signaling pathway activated by lysophosphatidic acid. 48 Additionally, acacetin, as a major active component of Coix seed, was shown to suppress the proliferation and migration of HT-29 cells while inducing apoptosis via the PI3K/Akt/p53 signaling pathway. 49
The strength of this study lay in its inaugural application of a bioinformatics approach integrating network pharmacology, molecular docking, and molecular dynamics, which enabled a comprehensive and systematic investigation into the mechanism of action of acacetin against HCC. This study establishes a solid theoretical foundation for investigating the anti-HCC mechanisms of acacetin, providing valuable references and evidence for subsequent experimental research.
However, the study had certain limitations. First, due to the inherent constraints of bioinformatics methods, this work served only as a preliminary exploration of the potential mechanisms of acacetin in HCC. Notably, the network pharmacology predictions require further validation through in vitro or in vivo experiments to enhance the robustness of the findings. Second, although MMP9 was identified as a key target, the validation of the key target only examined the changes in mRNA transcriptional levels. Follow-up animal studies will be necessary to assess protein-level changes and signaling pathway activities to clarify their mechanistic involvement. Lastly, despite its therapeutic potential, acacetin suffers from poor solubility, low chemical stability, and limited bioavailability, which substantially impede its clinical application. Therefore, extensive further research is needed before acacetin can be advanced to clinical trials.
Conclusions
This study provided an initial theoretical investigation into the mechanism of action of acacetin in the treatment of HCC. The relationship between acacetin and HCC targets was elucidated through network pharmacology and visualized via network diagrams, predicting that the mechanism might involve regulation of signaling pathways such as PI3K/Akt and p53. Molecular docking and molecular dynamics simulation were performed to verify the binding interactions between acacetin and key targets, suggesting that MMP9 may play pivotal role in the therapeutic effects of acacetin against HCC. Subsequently, preliminary validation was conducted via
Footnotes
Acknowledgments
The authors express their sincere gratitude to Jing Bai and team members for their experimental support. Special thanks are extended to Rui Feng and Jintuo Yin for their expert guidance and constructive feedback throughout this research.
Ethical Considerations
This study did not involve human participants or animal subjects, and thus did not require ethical approval.
Author contributions
LH and BJ conceived and designed the experiments, and drafted the manuscript; FR and YJT monitored the study and edited the ffnal version; WZY and SSW participated in the experiment and bioinformatics analysis; SXL and HLM assisted in preparing the manuscript; XYM revised it critically for important intellectual content. All authors agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by Natural Science Foundation of Hebei Province, China [H2022206533].
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
For reasonable requirements, the data related to this study can be requested from the corresponding author.
