Abstract
Objective
Jatropha podagrica is extensively used in traditional medicine to cure several illnesses, including infectious and non-communicable diseases. This current study investigated the effects of extracts from different parts of this plant on the bacteria causing intestinal diseases, including Bacillus cereus, Escherichia coli, and Salmonella spp., thereby unveiling a potential therapeutic approach to these bacteria.
Methods
Jatropha podagrica leaves, stems, and roots were extracted using 70% ethanol. The extract was then fractionated with n-hexane and ethyl acetate. Antibacterial assays, including agar well diffusion and MIC - MBC determination, were carried out. Bioactive compounds were docked with bacterial target proteins using AutoDock Vina. Binding interactions were analyzed. SwissADME, ProTox-3.0, and STopTox were used to assess the drug-likeness and toxicity of selected compounds. Data were expressed as mean ± SD using Microsoft Excel 2010.
Results
Extracts of J. podagrica roots and stems exhibited antibacterial activities against the selected pathogens, with the ethyl acetate fraction showing the highest potential. The extracts had higher effectiveness towards Gram-positive bacteria than Gram-negative bacteria. Further experiments demonstrated that the ethyl acetate fraction extract of J. podagrica roots identified bactericidal effects against selected pathogens, while the n-hexane fraction extract exhibited bacteriostatic effects against these pathogens based on MIC-MBC ratio. Molecular docking analysis indicated that the compound jatrophone had adequate binding affinity to ADP and higher binding affinity to PBP1a and PBP5 compared to the standard drug.
Conclusion
These findings suggest that J. podagrica could be a potential source of antibacterial agents, warranting further investigation for therapeutic applications.
Introduction
Gastrointestinal (GI) illnesses are among the leading causes of morbidity and mortality. 1 In low- and middle-income countries, they have been linked to at least 8 million deaths and at least 6 trillion dollars in economic losses. In 2015, it was the ninth leading cause of death for all ages; in 2017, among children under five, it was the third leading cause of death.2,3 Diarrheagenic Escherichia coli (DEC), Salmonella spp., and Bacillus cereus are some of the commonly found pathogens linked to acute infectious diarrhea.4,5
The main component of the bacterial cell wall is peptidoglycan (PG). This crucial macromolecule gives bacteria resistance to osmotic pressure, shapes them, and supports surface molecules. 6 Penicillin-binding proteins (PBPs) make up the majority of the proteins engaged in the enzymatic activities that synthesize and modify PG. 7 They are categorized as low MW PBPs and high MW PBPs based on their molecular weights (MW), and both are further subdivided into A, B, and C groups based on sequence similarity. 8 Bacillus cereus’ alkaline D-peptidase (ADP) is a class C PBP of the AmpH type. The role of ADP as a catalyst in the synthesis of D-Phe oligopeptides, or, with the aid of substrate mimetics, L-amino acid peptides and proteins, has attracted increasing attention. 9 PBP5 is a class B transpeptidase with a low affinity that gives E. coli intrinsic beta-lactam resistance and preserves cell shape.10,11 PBP1a, a vital PBP of S. typhimurium, is a class A high molecular weight (HMW) bifunctional PBP that is expected to have both transpeptidase activity, which is necessary for the cross-linking of peptide side chains on PG strands, and glycosyltransferase activity, which is essential for PG strand polymerization.12,13 Because they play a crucial role in bacterial cell wall formation, PBPs are an attractive target for antibiotics, especially beta-lactam antibiotics. However, there is a constant quest for novel active antibiotics since pathogens eventually develop antibiotic resistance. 14
The emergence of multidrug-resistant bacterial strains, especially intestinal pathogenic bacteria, is a major concern worldwide. 15 Over the past few decades, a rising number of resistance genes have been found in E. coli isolates, and a large percentage of these resistance genes were acquired by horizontal gene transfer. E. coli has been observed to be resistant to a wide range of antibiotics, namely broad-spectrum β-lactam and fluoroquinolones. 16 More than 1.2 million illnesses are linked to Salmonella each year, with at least 100 000 caused by antibiotic-resistant strains, including those resistant to clinically significant drugs such as ciprofloxacin (33 000 cases per year) and ceftriaxone (36 000 cases per year). 17 B. cereus, an opportunistic pathogen primarily associated with foodborne illnesses, is generally resistant to β-lactam antibiotics and can develop resistance to other common antibiotics, including ciprofloxacin, cloxacillin, erythromycin, tetracycline, and streptomycin. 18 Consequently, these call for the development of novel therapeutic approaches.
Natural antimicrobial products have garnered significant attention in science because of their vast chemical diversity and inherent qualities, which have encouraged research into them as potential medicinal agents in recent years. 19 Given that they are made from natural resources, plant-based medications, and herbal therapies are regarded as pure, safe, and healthful. 20 The Jatropha species, a member of the tribe Joanneasiae of Crotonoideae in the family Euphorbiaceae, are employed in traditional medicine to treat a variety of illnesses in Asia and Africa.21,22 Jatropha podagrica, a member of the genus Jatropha, has been widely used to successfully treat a variety of conditions, including gout, paralysis, gonorrhea, skin infections, jaundice, and fever. 23 The extracts of different parts of J. podagrica have been proven to possess significant medicinal potential. The stem and stem bark extracts in three distinct solvents (chloroform, methanol, and hexane) have been shown to effectively inhibit clinical isolates of S. aureus, E. coli, Candida albicans, and several Gram-positive bacteria, indicating the plant's capability as an antimicrobial and antifungal agent.24–26 At a concentration of 100 µg/mL, J. podagrica extract exhibits anti-proliferative effects against PC12 and A549 cells, reducing cell viability by more than 80%. It also exhibited effective protective qualities against DNA damage resulting from hydroxyl radicals and in isolated brain tissue, the extract reduces protein carbonylation and lipid peroxidation. 27
Although J. podagrica has been extensively studied, its effects on intestinal infection treatment and its molecular mechanisms remain unreported. Moreover, the requirement of finding broad-spectrum antibiotics to treat antibiotic-resistant bacteria is a priority solution. Therefore, in this research, we aim to elucidate the antimicrobial properties of the extracts of stem, bark, and leaves of this herb against three common intestinal pathogens E. coli, Salmonella spp., and B. cereus by using in vitro investigations. Molecular docking was also applied to thoroughly evaluate the interaction between the main bioactive compounds in J. podagrica and selected penicillin-binding proteins (PBPs) and alkaline D-peptidase (ADP).
Materials and Methods
Materials
Collection of Plant Samples
The Jatropha podagrica was collected in October 2022 from Phu Lap commune, Tan Phu district, Dong Nai province, Vietnam. The plant voucher specimen was identified by Dr Loi Huynh, and the sample is currently stored in the Laboratory of the Department of Pharmacognosy and Drug Control at the School of Medicine and Pharmacy, The University of Danang, Vietnam.
Microorganisms
Escherichia coli ATCC 17802, Bacillus cereus ATCC 10876, and Salmonella enterica subsp. enterica serovar Enteritidis ATCC 13 076 were cultured and preserved at the Department of Biotechnology, Faculty of Chemical Engineering, University of Science and Technology, The University of Danang, Viet Nam.
Chemicals and Reagents
The chemicals and reagents used in the research included: distilled water, ethanol 96%, DMSO 100% (Sigma), Mueller-Hinton broth (MHB) and Mueller-Hinton agar (MHA) (TM media), Ampicillin 50 μg/mL (Merck), Resazurin 0.015% (Santa Cruz).
Methods
Preparation of Jatropha podagrica Extracts
Different Jatropha podagrica fractions were prepared following a previous protocol. 28 Briefly, fresh J. podagrica leaves, stems, and roots were crushed into a powder after being shade-dried. 50 g powder samples of leaves, 50 g of stems, and 50 g of roots were extracted using 70% ethanol (250 mL × 3 times). After extraction, the mixture was collected and passed through filter paper for filtration. A rotary vacuum evaporator was then used to evaporate the solvent from the filtered solution under low pressure, producing a 70% ethanol extraction residue from the leaves, stems, and roots. The n-hexane and ethyl acetate fractions were obtained by dissolving this extraction residue in 200 mL of warm water and fractionating using these two solvents. The fractionated samples are stored at 4 °C.
Antibacterial Assays
Agar Well Diffusion Method
The modified methodology was applied in this study.
26
Bacterial cultures were grown overnight in Mueller-Hinton broth (MHB) at 37 °C with shaking at 200 rpm. The overnight culture of bacteria was then diluted with MHB to adjust its turbidity to 0.5 McFarland standard. 100 μL of the diluted bacterial suspension was surface-inoculated on Mueller-Hinton agar (MHA) using the streak plate method. On each plate, 40 μL of ethanolic extract, n-hexane, and ethyl acetate fractions were loaded into separated 6-mm wells. Ampicillin (50 μg/mL), and DMSO 10% were used as a positive and negative control, respectively. The antibacterial activity of these extracts was evaluated based on the inhibition zone using the following formula: D: Diameter of the inhibition zone (mm), d: Diameter of the agar well (d = 6 mm)
The experiments were triplicated.
Determination of Minimum Inhibitory Concentration
The broth microdilution technique in 96-well flat bottom microtiter plates was used to determine the minimum inhibitory concentration (MIC) of the samples. 29 The extracts were twofold serially diluted from 100 mg/mL to 0.1 mg/mL using MHB. 50 μL of the extract at different concentrations and 50 μL of the bacterial suspensions (106 CFU/mL) were loaded into each well. The negative control wells individually contained 50 μL of the extracts and 50 μL of MHB. The plates were incubated at 37 °C overnight, followed by the addition of 30 μL of Resazurin 0.015% (Santa Cruz, CAS 62758-13-8). The plates with resazurin were then incubated at 37 °C for 20 min. Changing of resazurin from blue to pink was then observed and the lowest concentration of the extract prior to color change was considered as MIC value. The experiments were triplicated.
Determination of Minimum Bactericidal Concentration
To determine the minimum bactericidal concentration (MBC), 50 μL of the extracts at the concentrations of 1xMIC, 2xMIC, 4xMIC, and 8xMIC were added to each tube containing 50 μL of the bacteria suspension. The samples were then incubated overnight at 37 °C. Following that, the mixtures were distributed on MHA plates and incubated overnight at 37 °C. The MBC value is the lowest concentration of the extracts that can kill 99.9% of the examined bacteria. 30
Molecular Docking Analysis
Ligand and Protein Preparation
A list of bioactive compounds isolated from J. podagrica roots was obtained from our previous study (Supplementary Table 1). 23 The structures of these compounds (ligands) were retrieved in SMILES strings from the PubChem chemical database. Their structures were drawn using ChemDraw for bioactive compounds that were not yet available on PubChem. The 3D structures were then built from obtained SMILES strings using Chimera. The tool Dock Prep was used to prepare structures for molecular docking: after merging non-polar hydrogens, polar hydrogen was added along with Gasteiger charges. The structures were minimized to obtain stable conformations and then were saved in PDB format.
The structures of PBP5 of E. coli (PDB ID: 3MZD), PBP1a of Salmonella sp. (PDB ID: 2OQO), and Alkaline D-peptidase of B. cereus (ADP) (PDB ID: 1IKG) were retrieved from RCSB Protein Data Bank. The ligand, water molecule, heteroatoms, and co-crystallized solvents were removed to prepare the target proteins. Polar-charged hydrogen and Gasteiger charges were added after non-polar hydrogens were combined. The binding sites of PBP5 and PBP1a were selected based on active sites of proteins in previous publications.31,32 For ADP, the binding sites were calculated using PrankWeb. The protein structures were then saved in PDB format.
Molecular Docking
An analysis of the binding kinetics between the target protein and compounds from J. podagrica through molecular docking was conducted using the AutoDock Vina software program (version 1.2.5). The predicted binding sites, centers of rotation, and grid box sizes of the docked combinations were recorded (Table 1).
The Center of Rotation and Grid Box Size of the Docked Combinations.

Evaluation of Physicochemical Properties and Toxicity
The physicochemical and toxicological parameters of the bioactive compounds isolated from J. podagrica with the highest potential were analyzed using SwissADME and DL-AOT Prediction Server. ProTox-3.0 and STopTox were utilized to get more detailed information about the compound's lethal dose and its toxicity towards organs.
Statistical Analysis
Results were calculated as the average of three replicates per experiment. The results were displayed as mean ± standard deviation (SD) using Microsoft Excel 2010.
Results
Antibacterial Activity of the Extracts Isolated from Different Parts of J. podagrica
The antibacterial activities of the extracts by different solvents were evaluated based on the diameters of the inhibition zones. It can be observed that the extracts of J. podagrica root exhibited antibacterial activities against the selected pathogens, with the ethyl acetate fraction extract showing the highest antimicrobial potential. The extracts of J. podagrica stem also demonstrated antimicrobial properties against the pathogens, with the ethyl acetate fraction being the most consistent, exhibiting moderate activity against E. coli and B. cereus (Figure 1 and Table 2).

Zones of Inhibition of Different Extracts from Jatropha podagrica Against Different Intestinal Bacteria. Upper-Root Extracts and Lower-Stem Extract,
Zones of Inhibition of 100 mg/mL Extracts from Jatropha podagrica Stem and Root Against Bacillus Cereus, Escherichia Coli, and Salmonella spp

As described in Table 2, at a concentration of 100 mg/mL, ethyl acetate extract from the stem exhibited a maximum ZOI value of 8.7 ± 1.2 mm in inhibitory activity against B. cereus, followed by E. coli (8.3 ± 0.6 mm). In the case of root extracts, the n-hexane fraction had the highest ZOI value of 9.0 ± 1.0 mm against B. cereus, followed by the ethyl acetate fraction against B. cereus (6.7 ± 0.6 mm). However, all extracts isolated from J. podagrica leaves did not show any visible antibacterial potential against these testing pathogens.
The minimum Inhibitory Concentrations and the minimum Bactericidal Concentrations of J. podagrica Root Extracts
The minimum inhibitory concentration (MIC) of an antibacterial agent is the concentration at which the test strain of an organism cannot grow at all under carefully regulated in vitro conditions. 33 Based on the color change of the resazurin reagent (blue) into resorufin (pink) due to the reduction caused by bacterial activity, the MIC value of the extract can be determined. In the ethyl acetate fraction of J. podagrica root, for B. cereus, the wells supplemented with extract concentrations from 50 mg/mL to 6.25 mg/mL still retained the blue color, proving that the bacteria were inhibited. Meanwhile, the color change of the indicator began to appear in the wells with extract concentrations less than or equal to 3.13 mg/mL (Figure 2A). For E. coli and Salmonella, the MIC values were both measured at the concentration of 12.5 mg/mL (Figure 2B and C).
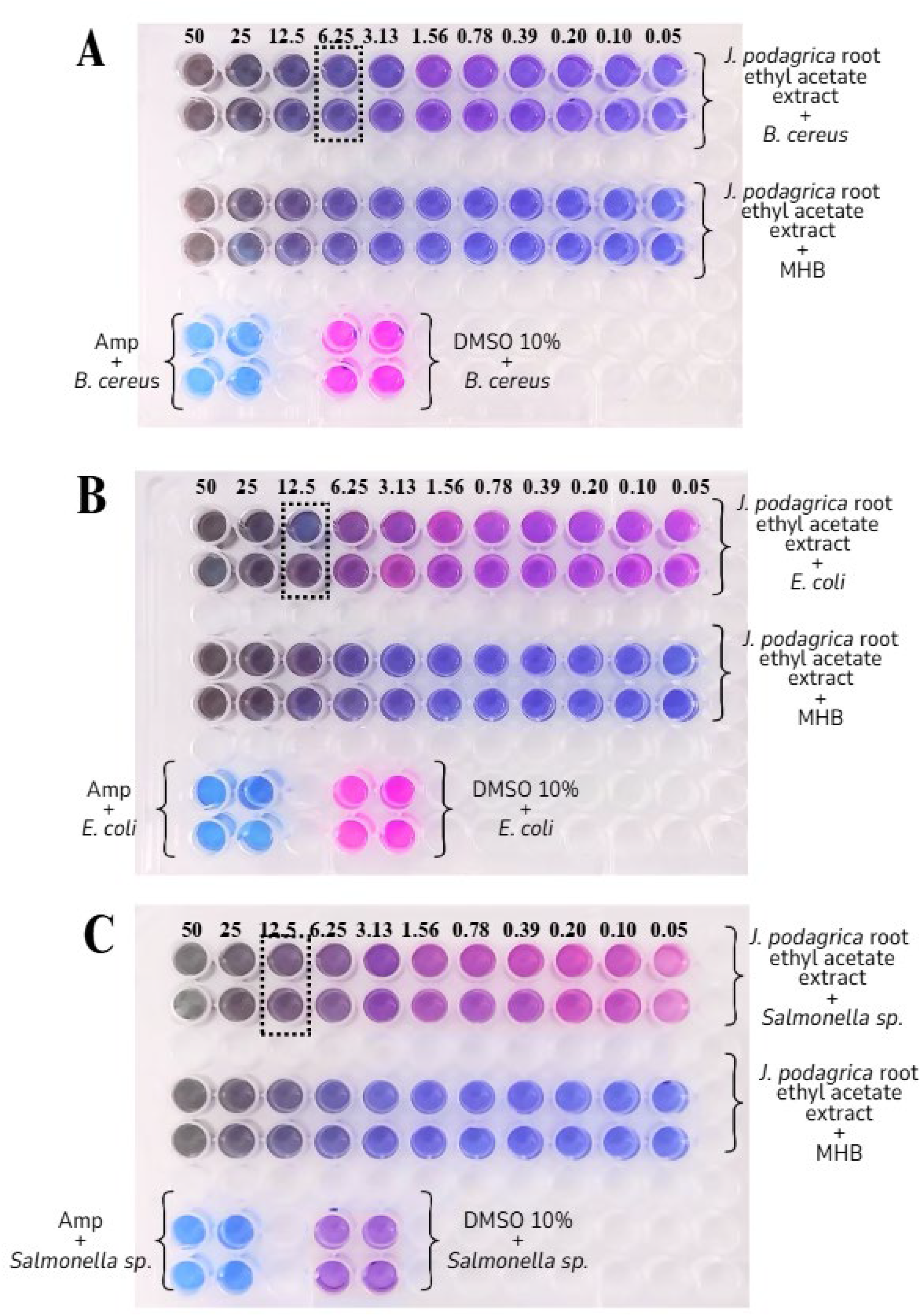
Determination of MIC Values of Ethyl Acetate Extract from Jatropha podagrica Root Against Bacillus cereus (A); Escherichia coli (B), and Salmonella spp (C). The Fractions were Two-fold Serial Diluted from the Concentration of 100 mg/mL to 0.1 mg/mL Before Being Added to Each Well. The Numbers Indicated the Final Concentration of Extracts in Each Well. After 24hr Incubation at 37 °C, Resazurin Dye was Supplied, and Color Change was Observed 20 min Later. Ampicillin (50 μg/mL) was a Positive Control, and 10% DMSO was a Negative Control.
Consistent with the results of the agar diffusion assay (Figure 1), n-hexane fraction of J. podagrica root exhibited weaker activity compared to the ethyl acetate fraction, with MIC value of 12.5 mg/mL (Supplementary Figure 1). It can be seen that the MIC values of the ethyl acetate and n-hexane extracts from J. podagrica roots were lower when against Gram-positive bacteria compared to Gram-negative strains, indicating that they had higher effectiveness towards Gram-positives than Gram-negatives (Table 3).
MBC/MIC Ratio of J. podagrica Ethyl Acetate Root Extract Against Selected Pathogens.

The minimum bactericidal concentration (MBC) of J. podagrica ethyl acetate and n-hexane fractions against three selected pathogens were all recorded at 25 mg/mL and higher than 50 mg/mL, respectively (Supplementary Figure 2 and Figure 3). The ethyl acetate fraction showed bactericidal effects against B. cereus, E. coli, and Salmonella as the ratio of the MBC to the MIC is equal to or lower than 4.0. Meanwhile, the n-hexane fraction of J. podagrica root only showed bacteriostatic effects against the tested pathogens as the MBC/MIC ratios were all over 4 (Table 3). From the MBC results, it was clear that the ethyl acetate fraction of J. podagrica was more effective as a bactericidal agent than the n-hexane fraction.
Virtual Screening Results of Bioactive Compounds Isolated from J. podagrica Roots with Protein Targets of Investigated Pathogens
To validate the docking protocol, we performed a re-docking experiment using the crystal structure of PBP5 (PDB ID: 3MZD). The co-crystallized ligand was extracted and re-docked into its original binding site. The resulting docked pose showed a root-mean-square deviation (RMSD) of approximately 1.36 Å compared to the crystallographic pose, indicating good agreement. This result confirms the reliability and accuracy of our docking protocol. The binding affinity values of bioactive compounds isolated from J. podagrica roots were adequate, with 2-hydroxyisojatrogrossidion (−8.36 Kcal/mol) and jatrophone (−8.41 Kcal/mol) being the closest to that of ceftobiprole (−8.98 Kcal/mol). In this experiment, Ceftobiprole's inhibitory effect was achieved by binding to abnormal PBPs, 34 therefore, ceftobiprole was chosen as the positive control in the following molecular docking studies (Figure 3 and Supplementary Table 2).
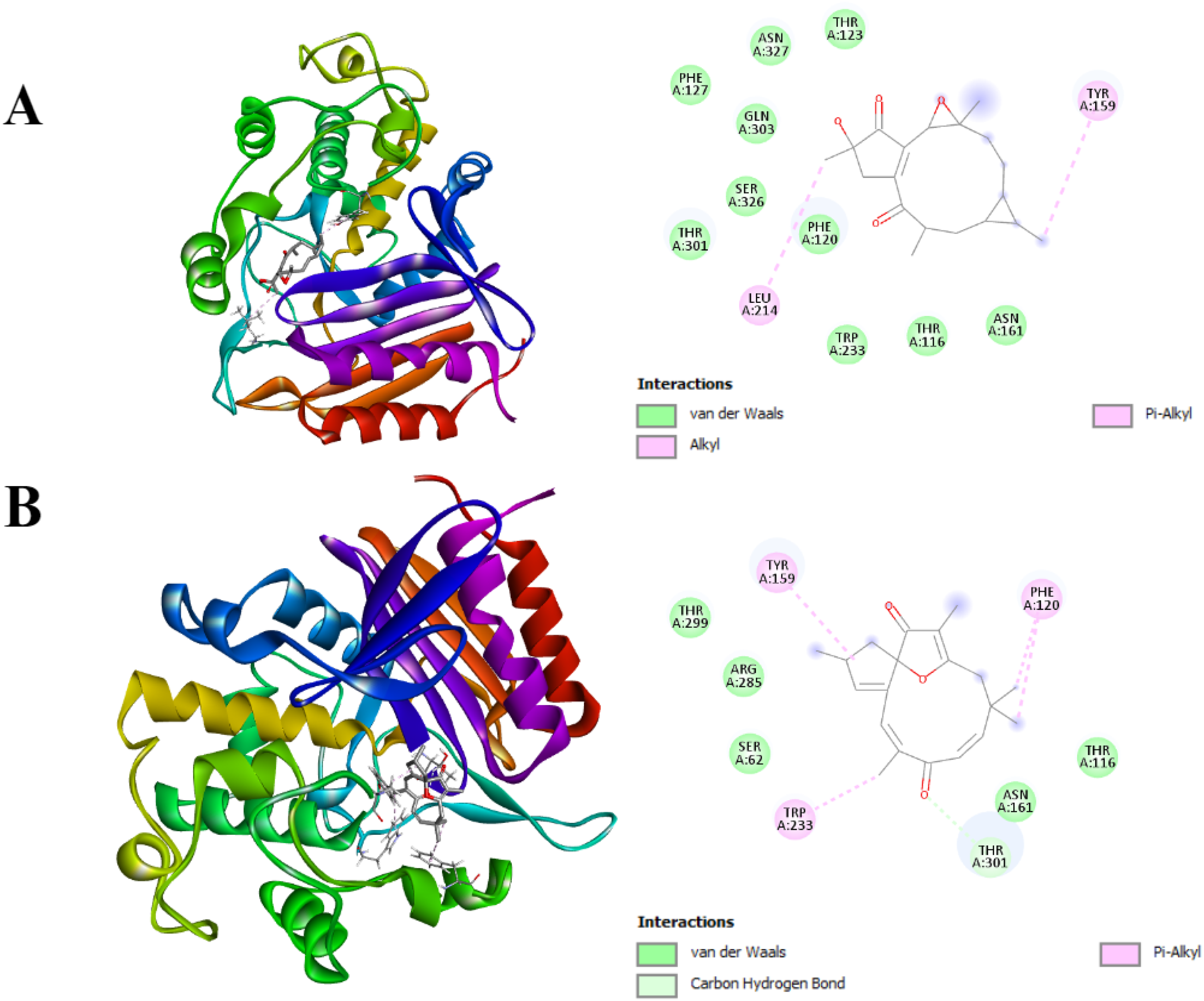
Molecular Docking Illustrations of Binding Activity Between Bioactive Compounds and ADP in B. cereus. A. 2-hydroxyisojatrogrossidion
The binding affinities of two bioactive compounds, including jatrophone (−6.97 Kcal/mol) and 2-hydroxyisojatrogrossidion (−6.58 Kcal/mol) with PBP5 were greater than ceftobiprole (−6.52 Kcal/mol) (Figure 4 and Supplementary Table 3).

Molecular Docking Illustrations of Binding Activity Between Bioactive Compounds with PBP5 in E. coli.
Compared to ceftobiprole (−6.54 Kcal/mol), two bioactive compounds jatrophone and 2-hydroxyisojatrogrossidion (−6.79 Kcal/mol) exhibited higher binding affinities for PBP1a (Figure 5 and Supplementary Table 4).
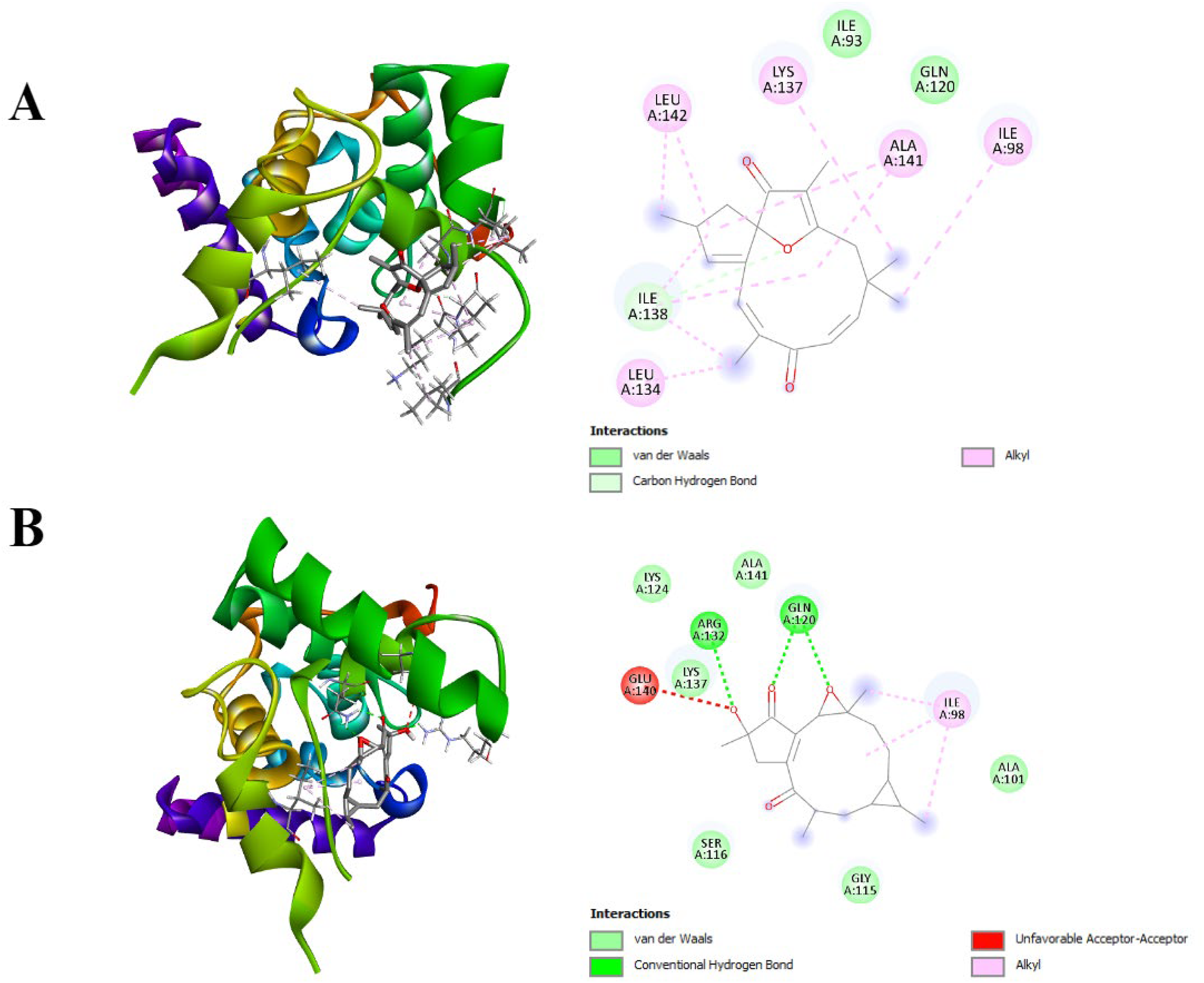
Molecular Docking Illustrations of Binding Activity Between Bioactive Compounds with PBP1a in Salmonella.
In detail, jatrophone bound to ADP with one carbon-hydrogen bond at THR301 and three hydrophobic interactions at PHE120, TYR159, and TRP233. Additionally, the compound showed interaction with PBP5 with one carbon-hydrogen bond at PHE262, one conventional hydrogen bond at ARG302, and four hydrophobic interactions at PRO10, MET8, PRO301, LYS6. It is also noted that jatrophone bound to PBP1a with one carbon-hydrogen bond at ILE138 and five hydrophobic interactions at LEU142, LYS137, ALA141, ILE98, and LEU134.
Physicochemical Properties of Jatrophone - a Potential Bioactive Compound Found in Abundance in J. podagrica
In addition, this study assessed the physicochemical parameters of aqueous solubility (log S), total polar surface area (TPSA), and number of rotatable bonds. To guarantee optimal intestinal absorption and oral bioavailability, the TPSA value should not rise over 140 Ų and the number of rotatable bonds should not surpass 10. Jatrophone has no rotatable bond and has a positive TPSA value of 43.37 Ų. It was anticipated that this substance would have a high rate of gastrointestinal absorption and blood-brain barrier penetration. Drug biotransformation is mostly dependent on a number of cytochrome P enzymes, including CYP1A2, CYP2C19, CYP2C9, CYP2D6, and CYP3A4. It was predicted that jatrophone would inhibit CYP2C19, CYP2C9, and CYP3A4. Furthermore, it is not anticipated that they will be P-glycoprotein (P-gp) substrates. Using the DLAOT Prediction Server, the LD50 value of jatrophone was determined to be 130 mg/mL and its toxicity was categorized as “Caution”. Comprehensive data can be observed in Table 4.
Physicochemical Properties and ADME Predictions Computed by SwissADME.

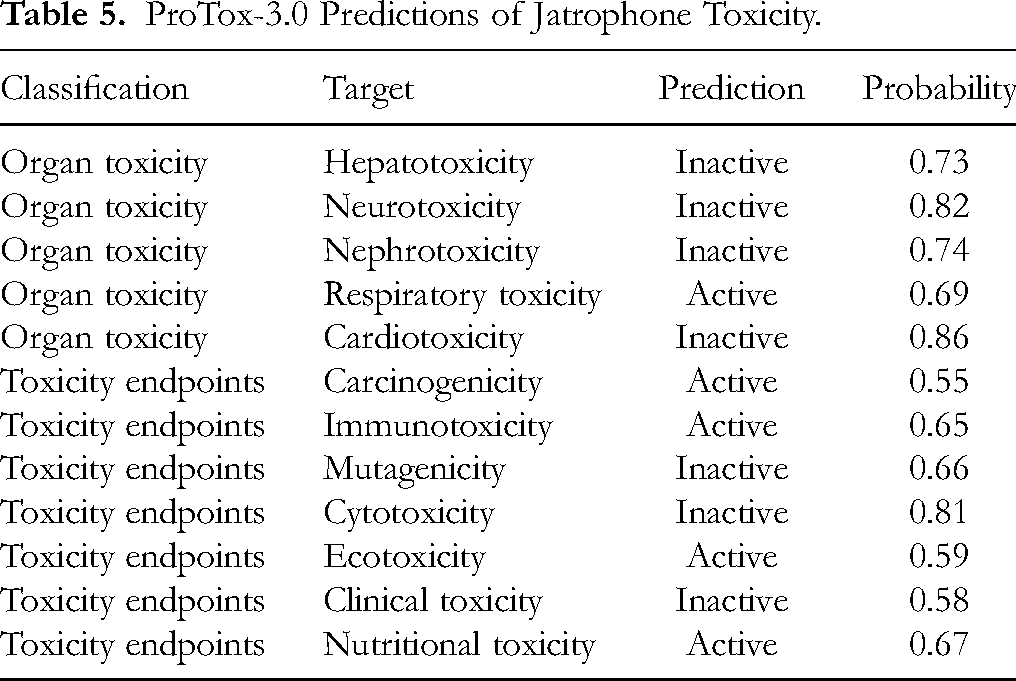
ProTox-3.0 and STopTox were also utilized to predict the toxicological properties of jatrophone. The results varied among different targets. Jatrophone was predicted to be inactive in terms of hepatotoxicity, neurotoxicity, nephrotoxicity, cardiotoxicity, mutagenicity, cytotoxicity, and clinical toxicity (Table 5).
ProTox-3.0 Predictions of Jatrophone Toxicity.
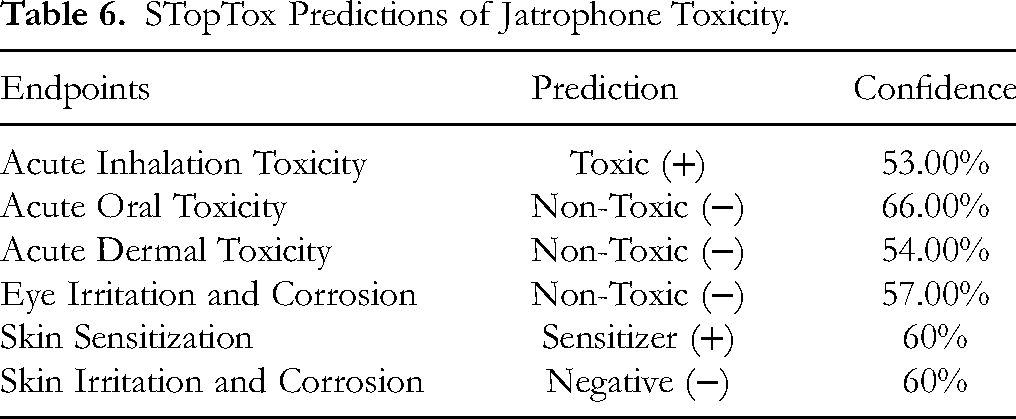
In addition, further predictions showed that jatrophone is non-toxic with regard to acute oral toxicity, acute dermal toxicity, eye irritation and corrosion, and skin irritation and corrosion (Table 6).
STopTox Predictions of Jatrophone Toxicity.

Discussion
Overall, the extracts suppressed more effectively against Gram-positive bacteria (B. cereus) than Gram-negative bacteria (E. coli and Salmonella sp.) (Figure 1 and Table 2). This is due to the fact that the outer membrane of Gram-negative bacteria provides extra protection against antibiotics, making them more resistant to them than Gram-positive bacteria. 35 The n-hexane and ethyl acetate fractions of J. podagrica root were selected to conduct further studies due to their great potential in inhibiting the tested pathogens and the abundance of experiments on J. podagrica stem bark.36–38 So far, there has been only one study focusing on the antibacterial activity of J. podagrica root extracts, in which the hexane, chloroform, and methanol extracts were proven to have broad-spectrum antimicrobial activity at a dosage of 20 mg/mL. The yellow root bark hexane extract exhibited the highest level of activity. Its level of activity was similar to that of gentamicin, but it performed better in terms of controlling B. cereus. 26 In another study, the hexane extract of J. podagrica stem bark demonstrated good inhibitory activity against MRSA and E. coli, with the inhibition zone of 15−18 mm. 24 Yet, no research has been found to conduct experiments on the ethyl acetate fraction of J. podagrica roots or on J. podagrica extract against Salmonella spp. The in vitro analysis of J. podagrica root ethyl acetate fraction in this research proved that the extract had potent antibacterial activities against both Gram-positive and Gram-negative pathogens, as either a bacteriostatic or bactericidal agent (Figure 2 and Table 3).
Alkaline D-peptidase (ADP) from B. cereus is an AmpH-type class C PBP. 9 While endopeptidases themselves are vital for bacterial growth, exposure to antibiotics that inhibit peptidoglycan synthases can lead to deadly cell wall disintegration. Therefore, emerging antibiotics and their adjuvants find endopeptidases to be interesting targets. 39 The dd-carboxypeptidase E. coli penicillin-binding protein 5 (PBP5), which is encoded by the dacA gene, is essential for maintaining cell shape. In addition to preserving cell shape, PBP5's role in preserving intrinsic beta-lactam resistance is a significant physiological finding. As a result, eliminating PBP5 could be a useful method for introducing beta-lactam susceptibility in the lab. 40 PBP1a is a bifunctional PBP implicated in the process of drug resistance and is essential for the septum's development during the cycle of bacterial cell division. PBP1a and PBP2a work together to maintain cell viability. 41 Based on the aforementioned in silico results (Figure 3, 4, and 5), jatrophone can be regarded as the most capable bioactive compound isolated from J. podagrica roots since it showed consistently satisfactory binding ability to different bacterial protein targets compared to the standard drug and the compound 2-hydroxyisojatrogrossidion, which also exhibited adequate binding affinity.
Jatrophone is a macrocyclic diterpene that has been isolated from many Jatropha species. Its derivatives have demonstrated a variety of biological characteristics, such as the ability to suppress insulin, lymphocyte activation, tumor cell growth, molluscicidal activity, and gastroprotective advantages. 42 The prominent “Lipinski's Rule of Five” has long been used as a standard to determine how drug-like a substance is. Jatrophone molecule weights are less than 500 Daltons. It also shows log P values less than 5, less than 10 hydrogen bond acceptors, and less than 5 hydrogen bond donors. The combination of these mentioned parameters and our results demonstrated that jatrophone has promising drug-like qualities due to the chemical satisfying Lipinski's criterion.
Conclusion
In this study, the acquired data demonstrated that J. podagrica root and stem extracts had good antibacterial activity against the chosen pathogens, with the root extract from the ethyl acetate fraction demonstrating the highest potential. Salmonella spp., E. coli, and B. cereus were all susceptible to the bactericidal properties of the J. podagrica root ethyl acetate fraction extract. Meanwhile, the n-hexane fraction extract of J. podagrica roots exhibited bacteriostatic effects against all test pathogens. Molecular docking analysis of the bioactive compounds extracted from J. podagrica roots showed that jatrophone had the greatest potential as a novel drug for the treatment of the studied bacterial infections. It showed consistently satisfactory binding affinity values compared with the standard drugs and other experimentally bioactive compounds. ADME analysis and toxicity predictions also demonstrated the potential of jatrophone to become a novel drug for the treatment of targeted infections. The results of this study will offer helpful information to researchers on the use of herbal-based supplements as a new approach to treating intestinal infections caused by antibiotic-resistant bacteria.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X251342461 - Supplemental material for Broad-Spectrum Antimicrobial Properties of Jatropha podagrica Extracts Against Intestinal Bacteria: A Combined Laboratory and Computational Study
Supplemental material, sj-docx-1-npx-10.1177_1934578X251342461 for Broad-Spectrum Antimicrobial Properties of Jatropha podagrica Extracts Against Intestinal Bacteria: A Combined Laboratory and Computational Study by Van Ngo Thai Bich, Y Ngo Huynh Thien, Hieu Phu Chi Truong, Tan Khanh Nguyen and Phu Tran Vinh Pham, Manh Hung Tran in Natural Product Communications
Footnotes
Acknowledgments
The authors appreciate the support of University of Science and Technology, School of Medicine and Pharmacy, and VN-UK Institute for Research and Executive Education, The University of Danang, Viet Nam for providing finance, infrastructure, and facilities to perform this study.
Author Contributions
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Science and Technology Development Fund of the University of Danang, Viet Nam under grant code B2022-DN01-02.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

