Abstract
Introduction
Staphylococci (Staphylococcus) bacteria are a common cause of human illness. Frequently forming clusters that resemble grape bunches, most Staphylococcus are found on the skin and mucous membranes.1–3 One especially prevalent species, S. aureus, causes osteomyelitis, endocarditis, sepsis, meningitis, scalded skin syndrome, toxic shock syndrome, and food poisoning, among other serious and deadly disorders.2,4,5 This species usually parasitizes the skin and nasopharynx, causing disease in immunodeficient people. 6 Approximately 50% of current S. aureus infections are resistant to multiple antibiotics, including penicillin, methicillin, tetracycline, ampicillin, and erythromycin.1,3 The use of multiple antibiotics has promoted the development of various resistant S. aureus strains, including to linezolid, quinupristin, and dalfopristin.1,7 S. aureus sortase A is regarded as a promising antibacterial drug target.8,9 This protein is critical for bacterial adhesion, biofilm formation, host cell entry, nutrient acquisition from the host, and immune evasion and suppression, all of which are crucial steps in the infection process. 10 Sortase A is a prospective target because it is situated on the extracellular side of the cell membrane, making it accessible without the need to penetrate the bacterial cell wall, which increases the inhibitor's bioavailability. Additionally, sortase A does not have human homologs; selective inhibitors of this enzyme should therefore be less harmful. Sortase A is not essential for bacterial growth and viability, and its inhibition is therefore anticipated to result in less selection pressure for the emergence of resistance. 11
The need to develop new and more potent antimicrobials as adjuncts or alternatives to antibiotic treatments has never been greater, as antibiotic resistance is increasing dramatically. For thousands of years, nature has been a source of medicinal compounds, which evolved various antimicrobial resistance mechanisms against bacterial and fungal pathogens. Mushrooms contain a wide range of bioactive compounds with many chemical configurations, and chemicals isolated from mushrooms could be valuable in the search for novel antibacterial drugs. Ganodermataceae species (Polyporales) are medicinal mushrooms that have been used in traditional medicine in China and other Asian countries for approximately 2000 years.12–14
Drug discovery is a time-consuming and costly process. Recently, computational methods have provided a better understanding of the interactions between herbal medicine drug targets to help clinical practitioners design herbal formulae rationally. 15 In this study, we investigate anti-S. aureus activity of Ganoderma sp. using a virtual screening method. Molecular docking and dynamic simulation were also used to assess the stability of the ligand and protein complex. In addition, 2 natural Ganoderma strains, G. multiplicatum VNKKK1901 and G. sinense VNKKK1902 were selected for in vitro tests to further reinforce the computational results.
Results
Virtual Screening and Molecular Docking Analysis
A list of natural compounds isolated from Ganoderma sp. was collected for the molecular docking process (see Supporting Information section). The co-crystallized ligand presenting in the respective binding site of the protein (PDB ID: 2MLM) was re-docked to validate the precision docking protocol. The crystal ligand docked in the same binding pose as the native along with a docking score of −6.7 kcal/mol. The root-mean-square deviation (RMSD) between the redocked and native ligand conformation was calculated using Pymol (0.296 Å) (Supporting Information section). The co-crystallized ligand was also selected as a reference compound targeting protein sortase A. Notably, most of the selected compounds had docking scores higher than that of the reference compound, −6.7 kcal/mol. These results are expected to be due to the synergistic effect of the components in the crude extract in inhibiting the growth of bacteria. In this study, the components were investigated for mechanisms that augur well for their potential for isolation and development into drugs capable of treating S. aureus infections. The top compounds with good docking scores are shown in Table 1.
Docking Results of Top-Ranked Ganoderma sp. Compounds with Sortase A.

Ganosinensin B generated the highest binding energy with sortase A, with a docking score of −8.4 kcal/mol. Ganosinensin B was found to interact with sortase A with 3 hydrogen bonds at Cys126, Val108, and Leu11, and via hydrophobic interactions at Ile141 (Figure 1A). Ganosinoside A showed interactions with residues Glu113, Val108, Ile124, and Trp136 (Figure 1B). Notably, the residue Leu111 contributed a hydrogen bond with 3 candidates in the top 5 ranked natural compounds. This finding suggests a role for this amino acid in the sortase A protein's ability to bind to inhibitors. In future studies, this residue site could be mutated to compare the interaction of inhibitors and their binding capacity to the protein. In addition, the active site of sortase A is a long, narrow groove (Figure 1C); this matches the poses of molecules with large lengthwise structures, such as ganosinensin B and ganosinoside A. These compounds contain a similar structural framework, consisting of 4 linked aromatic rings. Overall, the molecular docking results showed that Ganodermataceae sp. are promising for the inhibition of S. aureus strains.

Interaction diagram of (A) ganosinensin B, (B) ganosinoside A, and (C) active site groove of sortase A.
Molecular Dynamic Simulation
For further investigation, the top 2 sortase-A-binding compounds, ganosinensin B and ganosinoside A, were studied using molecular dynamic simulation to evaluate the stability of the protein-ligand complexes at different temperatures and pressure conditions. Small molecules interacting with protein surfaces can cause large tertiary structural changes, which can lead to drug design improvements. A major benefit of molecular dynamics (MD) simulation is its capacity to detect the flexibility of the protein-ligand complex. This allows MD studies to precisely assess the thermodynamics and kinetics that occur during drug-enzyme binding. The use of RMSD is a well-known measure for analyzing protein-residue-based fluctuations. To compare these dynamics, we estimated the RMSD of apo protein and ligand-bound protein. Solvent accessible surface area (SASA) indicates the surface area of a protein which interacts with its solvent molecules (Figures 2F and 3F).

Dynamics of ganosinensin B binding to sortase A during a simulation time of 100 ns. (A) RMSD, (B) RMSF, (C) the average center of mass distance between ligand and protein, (D) hydrogen bonds (protein-ligand), (E) radius of gyration of protein-ligand complexes, and (F) solvent accessible surface area. Abbreviations: RMSD, root-mean-square deviation; RMSF, root-mean-square fluctuation.
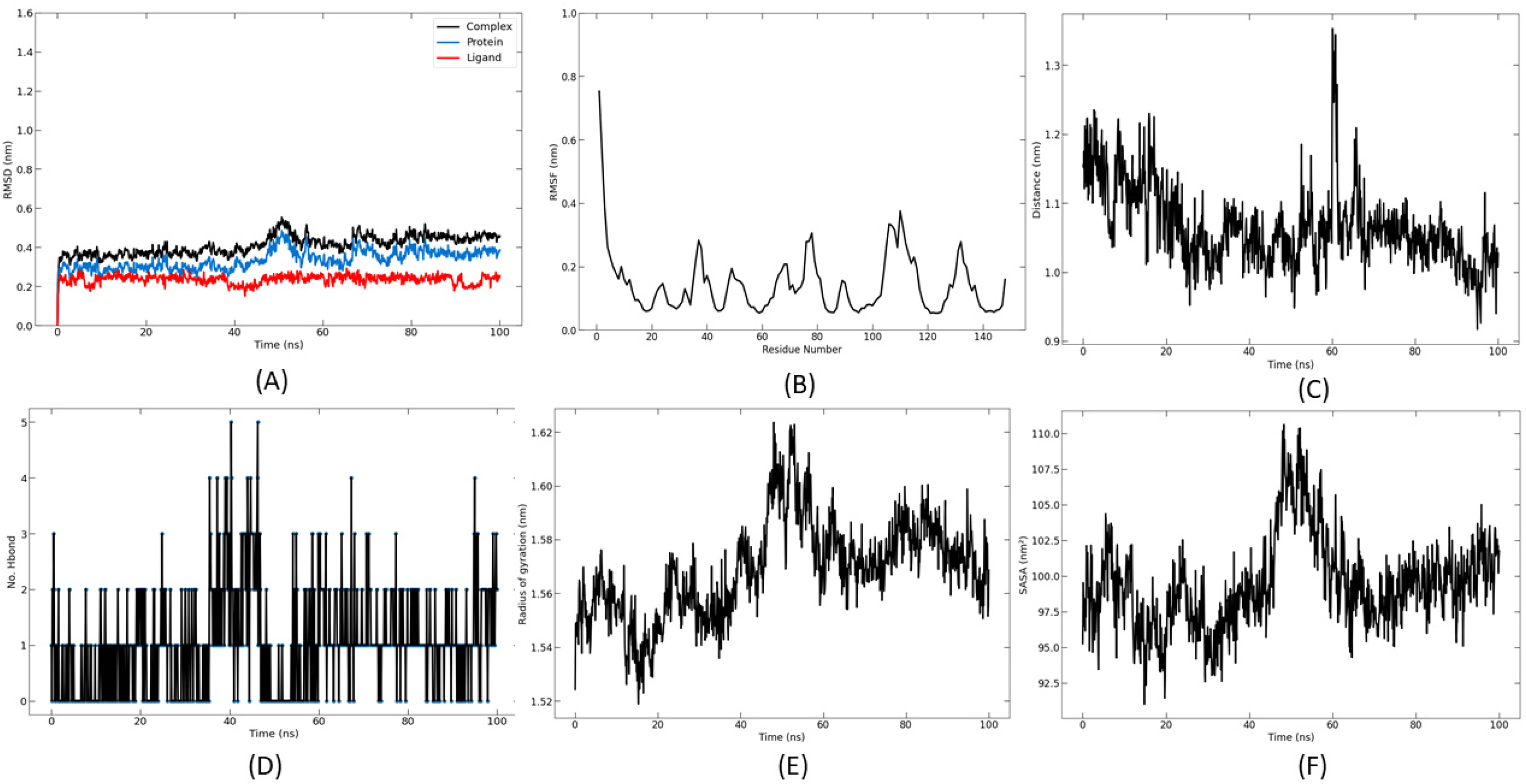
Dynamics of ganosinoside A binding to sortase A during a simulation time of 100 ns. (A) RMSD, (B) RMSF, (C) the average center of mass distance between ligand and protein, (D) hydrogen bonds (protein-ligand), (E) radius of gyration of protein-ligand complexes, and (F) solvent accessible surface area. Abbreviations: RMSD, root-mean-square deviation; RMSF, root-mean-square fluctuation.
For the complex between sortase A and ganosinensin B, the results showed that the structure remained stable throughout the simulation time, with fluctuation within the range of 0.4 Å (Figure 2A). The root-mean-square fluctuation (RMSF) results also demonstrated that the ligand stabilized several residues of interaction at the active site (Figure 2B). Overall, the fluctuation intensity remained below 2.5 Å. The average center-of-mass distance between the ligand and sortase A protein during the simulation time is shown in Figure 2C. The results indicated that the ligands showed a stable distance with <0.5 Å of fluctuation. In addition, the total number of hydrogen bonds formed between ligand and protein during 100 ns was also calculated. During the molecular dynamics trajectory, ligands almost always formed at least 1 to 2 hydrogen bonds with proteins (Figure 2D). The radius of gyration (Rg) is defined as the distribution of atoms of a protein around its axis. The fluctuations of Rg indicate that the ligand is changing conformation within the interaction pocket. There was a change in the conformation of ganosinensin B from 80 ns. This also indicates small fluctuations in RMSD in the same period.
The results of the MD simulation of sortase A with ganosinoside A show that the complex remained stable throughout the simulation (Figure 3A). The RMSF results also demonstrated that the ligand stabilized several residues of interaction at the active site (Figure 3B). Overall, the fluctuation intensity remained below 2.5 Å. The average center-of-mass distance between ligand and protein during the simulation time is shown in Figure 3C; the results indicated that the ligands showed a stable distance. In addition, the total number of hydrogen bonds formed between ligand and protein during 100 ns was also calculated. During the molecular dynamics trajectory, ligands almost always formed at least 1 hydrogen bond with the protein (Figure 3D). There was a change in Rg when the ligand bound to the protein, which changed the conformation of ganosinoside A from 40 to 60 ns. This also indicates a notable variation in the accompanying results.
To further evaluate the interaction between the ligands and the sortase A protein, a contact frequency analysis was performed using molecular dynamics (VMD) with a cutoff of 4 Å. The results are shown in Figure 4. This shows that the residues were Ala34, Ala46, Leu111, Cys126, ILE124, Trp136, and Arg139. The residues played an important role in forming interactions with ligands. These residues were similar to the key amino acids discussed in the molecular docking analysis. In future studies, these amino acids could be mutated to evaluate their specific role in sortase A activity. In general, both ganosinensin B and ganosinoside A demonstrated strong and stable binding capacity to proteins. This showed the potential to develop these hit compounds into drugs capable of inhibiting proteins for use in the treatment of S. aureus infections. In addition, the results of this study may provide good information for the development of new antibacterial treatments using fungal extracts.

CF analysis in ligand-protein complexes. Abbreviation: CF, contact frequency.
Antibacterial Activity of Extracts
Subsequently, to assess the molecular docking analysis results, G. multiplicatum VNKKK1901 and G. sinense VNKKK1902 were selected for in vitro tests to reinforce the computational results. Their extracts were tested in vitro to validate inhibitory activity on S. aureus ATCC 29213 and some other bacterial strains, such as Bacillus cereus ATCC 11778, Pseudomonas aeruginosa ATCC 27853, Escherichia coli ATCC 35218, and Klebsiella pneumoniae ATCC 750603. The antibacterial activities of these extracts are shown in Tables 2 and 3.
Diameter (mm) of ZOI of Different Extracts on the Growth of Staphylococcus aureus ATCC 29213.

*Values are expressed as mean ± SD, analysis was performed with one-way ANOVA followed by Tukey post hoc test.
(+) Antibiotic positive control, ciprofloxacin (5 μg/filter disk).
(−) Means DMSO 5%.
Abbreviations: ANOVA, analysis of variance; DMSO, dimethyl sulfoxide; SD, standard deviation; ZOI, zones of inhibition
MIC and MBC of Extracts on Staphylococcus aureus ATCC 29213.

Values are expressed as average of triplicated measures; analysis was performed with one-way ANOVA followed by Tukey post hoc test.
Positive control.
**P < .01; *P < .05.
Abbreviations: ANOVA, analysis of variance; CIP, ciprofloxacin (5 μg/filter disk); MBC, minimum bactericidal concentration; MIC, minimum inhibitory concentration.
All of the extracts (from 1–4 mg/disk) exhibited zones of inhibition (ZOI) against bacteria in a dose-dependent manner. Notably, G. multiplicatum VNKKK1901 demonstrated the greatest antibacterial activity against S. aureus, beginning at 1 mg/filter disk, followed by G. sinense VNKKK1902, which demonstrated antimicrobial ability against S. aureus from 2 mg/filter disk. The positive-control antibiotic, ciprofloxacin, inhibited the growth of S. aureus at 5 μg/filter disk, with ZOI above 30 mm (Table 2).
The minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) values also revealed varied susceptibility of the tested bacterial strains to the 2 extract concentrations. As shown in Table 3, G. sinense VNKKK1902 was more potent than G. multiplicatum VNKKK1901 against S. aureus, with the lowest MIC of 1.875 mg/mL. The lowest concentration of extract at which 99.9% of bacteria (5 × 105 colony forming unit [CFU]/mL) were killed (MBC) was that of G. sinense VNKKK1902, at 3.75 mg/mL for S. aureus, followed by G. multiplicatum VNKKK1901 at 7.5 mg/mL. The MIC for ciprofloxacin was 0.0125 mg/mL and the MBC was 0.05 mg/mL. Overall, the G. sinense VNKKK1902 extract was more promising against S. aureus than that of G. multiplicatum VNKKK1901. The extracts of both fungi were bactericidal, as their ratios of MBC to MIC were 2. 16
Discussion
Botanical antibiotics have received substantial attention recently for their potential ability to inhibit the survival of S. aureus, a major nosocomial pathogen responsible for many serious infectious diseases in humans. In recent years, many studies have reported the antimicrobial activities of Ganodermataceae species. Among these, 20 species have shown significant antimicrobial activity, of which G. lucidum has received particular attention.16,17 Previously, the methanol extract of G. lucidum collected in India showed activity against resistant E. coli MTCC 443, Salmonella typhimurium MTCC 98, and Bacillus subtilis MTCC 411 at 10 mg per well. 18 The mycelium of G. lucidum RCKB-2010 was cultured using a submerged fermentation method to obtain ganoderic acid, which was effective against E. coli, P. aeruginosa, S. aureus, and S. epidermidis at a concentration of 0.25 mg/mL. G. lucidum extract also inhibited the growth of Helicobacter pylori, which causes stomach ulcers and stomach cancer.19,20 The chloroform extract of G. lucidum, collected in Iran, showed an ability to resist B. subtilis ATCC 456, with an MIC of 8 mg/mL and an MBC of 16 mg/mL, and S. aureus ATCC 25923, with MIC and MBC values of 8 mg/mL. 21 The hot aqueous extract of G. lucidum from Korea was tested in vitro against 15 bacterial strains. Furthermore, G. lucidum extract in combination with cefazolin showed antibacterial effects against B. subtilis and Klebsiella oxytoca. 22
The membrane and cell wall of gram-negative bacteria have a more complex structure than those of gram-positive bacteria. The peptidoglycan of gram-negative bacteria is surrounded by an outer membrane, which restricts diffusion through its lipopolysaccharide, thus preventing the inroad of compounds that are harmful to the membrane. On the other hand, gram-positive bacteria have a simpler wall structure, which is thick and hydrophilic, and the main structure is peptidoglycan, so compounds are more easily absorbed. Therefore, gram-negative bacteria should be more sensitive to contact with compounds isolated from mushrooms. 23 The antibacterial mechanism of Ganoderma has its primary effect on the membrane and cell wall. Because the principal function of the membrane and cell wall is to provide shape and integrity to the cell, their role as an osmotic barrier protects the cell. The transport process, regulation, respiration, biosynthesis and cross-linking of peptidoglycan, and lipid synthesis are essential functions that are regulated by the bacterial membrane. Therefore, the interruption or destruction of the membrane can directly or indirectly cause cell inactivation or cell death. Consequently, the mode of operation of antibacterial agents often relates to involvement in cell membrane synthesis, alteration of membrane permeability, and interference in the replication of genetic material or protein synthesis. 18 Ganoderma sp. have many antibacterial compounds, of which triterpenes and polysaccharides are the 2 main biologically active classes.18,24
In our study, we first chose triterpenoids isolated from Ganoderma sp. from the structural bank. These constructs were included in the prediction of interactions with the potential target of S. aureus sortase A. This protein target is a membrane-anchored enzyme that is required to sort surface proteins to the cell wall envelope. 25 S. aureus can display up to 21 cell surface proteins containing the conserved peptide motif (LPXTG) proteins on its surface, anchored to the cell wall. Sortase A recognizes the LPXTG motif present at the C-terminus of cell surface proteins and catalyzes a ligation reaction between a 5-amino-acid substrate motif and oligoglycine nucleophiles to recruit them as building blocks for the peptidoglycan cell wall. 26 Therefore, the study of sortase A inhibitor molecules is a promising strategy to attenuate S. aureus virulence and biofilm formation. 27 Terpenoids are natural compounds, and most terpenoids comprise a cyclized hydrocarbon backbone. Terpenoids have demonstrated bactericidal and bacteriostatic activities by the following mechanisms: (1) Inhibition of oxygen uptake and oxidative phosphorylation in bacteria 23 ; (2) alteration of membrane fluidity by interacting with the hydrophobic tails and hydrophilic heads of phospholipids, thereby influencing the nonpolar and polar transmembrane pathways 28 ; (3) modulation of the bacterial efflux pump; (4) inhibition of virulence factors; and (5) inhibition of biofilm formation.29,30
Small molecules interacting with protein surfaces can cause large tertiary structural changes, which can lead to drug design improvements. Accordingly, based on our docking simulation results, we selected the active compounds that had the strongest interaction with sortase A and then analyzed the physical movements of their atoms and molecules using MD. A major benefit of MD simulation is its capacity to detect the flexibility of the protein-ligand complex. This benefit allows MD studies to precisely assess the thermodynamics and kinetics that occur during drug-enzyme binding. The use of RMSD is a well-known measure for analyzing protein-residue-based fluctuations. To compare these dynamics, we estimated the RMSD of apo protein and ligand-bound protein. SASA was shown to indicate the surface area of a protein which interacts with its solvent molecules. The selected compounds from Ganodermataceae contain a similar structural framework, consisting of 4 linked aromatic rings, and are found in many species of Ganoderma. Therefore, to gain more confidence in the binding stability, we undertook MD simulation of the top 2 sortase-A-binding compounds, ganosinensin B and ganosinoside A. Ganosinensin B and ganosinoside A were identified as the most highly active ingredients of Ganoderma sp. against sortase A. Molecular docking results showed that they bound tightly to sortase A, which is the most central and common therapeutic target of S. aureus. Interestingly, both ganosinensin B and ganosinoside A are the main secondary metabolites from the fruiting bodies of G. sinense. These compounds are unique, with a lanostane-type triterpenoid having farnesyl hydroquinone moieties.31,32
Several MD factors were analyzed, such as the RMSD, RMSF, average center-of-mass distance between ligand and protein, hydrogen bonds (protein-ligand), radius of gyration of protein-ligand complexes, and SASA. In both cases, the RMSD values moving from 0.2 to 0.6 nm denoted the average variance between the atoms of the trajectory frames, which indicated that the protein-ligand complexes remained constant throughout the simulation time. Meanwhile, the RMSF index was within the range of 0.1 to 0.8 nm, indicating the stability of the complexes and confirming the dynamic behavior of the amino acid residues and the flexibility of the protein structure during the interaction phase of the simulation. Additionally, SASA was used in determining the stability of the protein's hydrophobic core and the biomolecules.
Some mushroom extracts did not show an antibacterial zone in the agar plate diffusion method but showed activity in the broth microdilution method. This can be explained by the high molecular weight of the polysaccharide in the extract, which slowed its ability to diffuse into the agar medium. In contrast, the broth microdilution method suspended the compounds and bacteria in the liquid medium, enabling the compounds to quickly affect the membrane and wall structure of the bacteria, inhibiting, and killing them more effectively. Therefore, more clinical evidence needs to be gathered to further investigate the relationship between the compounds from Ganoderma sp. and S. aureus. Our study relied on data mining and analysis, but further validation using experimental evidence is required. More databases containing composition and target information on Ganoderma sp. should be included, which could increase the reliability of our results.
Materials and Methods
Data Preparation
In this study, we used a data mining method from Pubmed database (https://pubmed.ncbi.nlm.nih.gov/) to build a dataset of compounds extracted from Ganoderma strains. A list of compounds from Ganoderma sp. was collected for the molecular docking process. The 3-dimensional (3D) structures of the compounds were obtained from PubChem (http://pubchem.ncbi.nlm.nhi.gov), and prepared by MarvinSketch (ChemAxon, Cambridge, MA, USA).
Protein Preparation
The crystal structure of sortase A (PDB ID: 2MLM) was obtained from the Research Collaboratory for Structural Bioinformatics Protein Data Bank. Existing ligands and water molecules were removed using Discovery Studio. Polar hydrogen and Kollman charges were added to the protein using Autodock tools (v. 1.5.6). Finally, the protein was converted to pdbqt format for molecular docking analysis.
Molecular Docking Study
The selected compounds were screened virtually on sortase-A proteins using AutoDock Vina 1.1.2. The grid box covering the active site of the protein was selected from the interactive site of the crystal ligand. Docking scores, reported in kcal/mol, were used to rank the compounds. Finally, the molecular interactions between proteins and selected ligands were visualized by Discovery Studio Visualizer 2020, and their 3D and 2D interaction plots were derived. 15
MD Simulations
The system was prepared using CHARMM-GUI interface with the CHARMM36 force field.33,34 All the simulation was made using the NAMD 2.13 package. The TIP3P explicit solvation model was used. 35 The simulation box dimensions were 76 Å, 76 Å, and 76 Å in x, y, and z. The parameters for the top docking results were generated using CHARMM general force field. Afterwards, the system was neutralized using Na+ ions. The protocols involved minimization, equilibration, and production. A 2-fs time step of integration was chosen for all MD simulations, the equilibration was carried in the canonical (NVT) ensemble, and the isothermal-isobaric (NPT) ensemble was used for the production. Through the 100 ns of MD production, the pressure was set at 1 atm and the temperature at 298.15 K. Finally, the protocol was applied to the 2 selected candidates.
Sample Collection and Extract Preparation
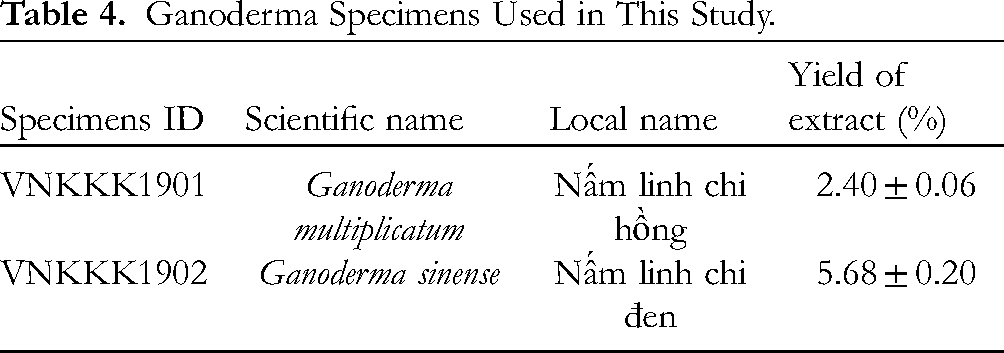
G. multiplicatum VNKKK1901, and G. sinense VNKKK1902 were collected in Vietnam (Table 4). All specimens were identified by morphological and phylogenetic (5.8S ITS rDNA gene) methods. Fruiting bodies were dried at 50 °C to a constant weight in a heating/drying oven (Memmert UFE 600, Buechenbach, Germany). Pulverized basidiocarps (100 g) were extracted with 1500 mL 70% ethanol by magnetic stirring (120 rpm) for 72 h. The solid and liquid fragments were separated by a fine-mesh strainer (0.2 cm), and the solids were extracted again with 1500 mL 70% ethanol. The liquid extract was filtered through Whatman No. 4 filter paper, concentrated under reduced pressure in a rotary evaporator (RV 10 digital V-C, IKA, Königswinter, Germany) at 40 °C to dryness, and dissolved in 5% (v/v) dimethyl sulfoxide (DMSO) solution (Merck, Darmstadt, Germany) to an initial concentration of 100 mg/mL. The ratio of dry fungal biomass before extraction to dry extract weight after evaporation was used to calculate the extraction yield. Among the 2 strains of Ganoderma used in this study, the yield of extracts of G. multiplicatum VNKKK1901, and G. sinense VNKKK1902 were 2.40 ± 0.06%, and 5.68 ± 0.20%, respectively (Table 4).
Ganoderma Specimens Used in This Study.

Microorganisms and Culture Conditions
S. aureus ATCC 29213 and the other bacterial strains were supplied by the Institute of Public Health, Ho Chi Minh City, Vietnam [Draft Genome Sequence of Methicillin-Sensitive Staphylococcus aureus ATCC 29213]. Bacteria were grown on nutrient agar and cultivated on Mueller–Hinton agar (MHA, Himedia, Mumbai, India) at 37 °C for 24 h. Turbidity was determined spectrophotometrically using the 0.5 McFarland turbidity standard (at 625 nm). The bacterial suspensions were adjusted with sterile saline solution (0.85%) to a final concentration of approximately 106 CFU/mL.
Disk Diffusion Method
The modified disk diffusion method was used in this study with slight modification.36,37 And 1 mL of a cell suspension of 106 CFU/mL S. aureus ATCC 29213 was spread evenly over the surfaces of MHA plates using sterile swabs. Sterile filter paper discs (6.0 mm in diameter) were impregnated with different given volumes of the test extract to obtain different final amounts (1, 2, 3, and 4 mg). These discs were placed on top of the prepared agar plates. Ciprofloxacin (5 μg/filter disk) was used as the positive control and 5% DMSO as the negative control. These plates were incubated at 4 °C for 4 h for metabolite diffusion and incubated at 35 ± 2 °C for 24 h. The diameter of the zone of bacterial growth inhibition (ZOI) was measured using a ruler and considered as an indication of the antibacterial activity of the extracts. The experiments were replicated 3 times.
Minimum Inhibitory Concentration and Minimum Bactericidal Concentration Determination
The MIC was used to identify the lowest concentration of the extract that inhibits the microbial growth after 24 h of incubation using broth microdilution in 96-well microtiter plates; the total volume in each well was 100 μL.38,39 The bacterial suspension was adjusted with saline to 1 × 106 CFU/mL. Mushroom extracts were dissolved in 5% DMSO solution and added to Mueller–Hinton broth (MHB, Himedia, Mumbai, India) with bacterial inoculum (5 × 105 CFU per well) to achieve the desired final concentrations of extracts (0.05-100 mg/mL). Microtiter plates with bacteria were incubated at 35 ± 2 °C for 24 h. The MIC value was determined by a colorimetric bacterial viability assay based on reducing either 7-sodiooxy-3H-phenoxazin-3-one 10-oxide or resazurin (VWR Chemicals, Radnor, PA, USA) using a microplate reader (BMG Labtech CLARIOstar®, Ortenberg, Germany). MBC was defined as the lowest concentration of extract that did not exhibit any bacterial growth after reinoculation of 2.0 μL of the mixture on MHB, indicating 99.9% killing of the original inoculum. Ciprofloxacin (Sigma, Ronkonkoma, NY, USA) was used as the positive control and 5% DMSO as the negative control. The experiments were repeated 3 times.
Statistical Analysis
Quantitative data were obtained by triplicate experiments and analysis was performed with one-way analysis of variance (ANOVA) followed by Tukey Post Hoc test in Excel 2020.
Conclusions
In this study, a list of natural compounds from Ganodermataceae fungal species was collected and virtually screened in sortase A complexes using docking simulations. Ganosinensin B and ganosinoside A showed the highest binding energies. Molecular dynamics simulation also showed the stability of ligand and sortase A complexes in 100 ns. The extracts of G. multiplicatum VNKKK1901 and G. sinense VNKKK1902 collected in Vietnam were tested against S. aureus to strengthen the computational results. Our findings demonstrate the potential antibacterial properties of these extracts against S. aureus. This study highlights the potential value of fungal extracts and their secondary metabolites as sources of antimicrobials. However, further mechanism analysis of the antibacterial activity of these fungal extracts and secondary metabolites is needed.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X231167289 - Supplemental material for Integrating in Silico and In Vitro Studies to Screen Anti-Staphylococcus aureus Activity From Vietnamese Ganoderma multiplicatum and Ganoderma sinense
Supplemental material, sj-docx-1-npx-10.1177_1934578X231167289 for Integrating in Silico and In Vitro Studies to Screen Anti-Staphylococcus aureus Activity From Vietnamese Ganoderma multiplicatum and Ganoderma sinense by Trang Thi Thu Nguyen, Trinh Thi Tuyet Nguyen, Hoang Duc Nguyen, Tan Khanh Nguyen, Phu Tran Vinh Pham, Linh Thuoc Tran, Linh Thuy Thi Tran and Manh Hung Tran in Natural Product Communications
Footnotes
Acknowledgements
Trang Thi Thu Nguyen and the research group are grateful for the funding support of the Vingroup Joint Stock Company PhD Scholarship Programme. This work was funded by the PhD Scholarship Programme of the Vingroup Innovation Foundation (VINIF), Vingroup Big Data Institute (VINBIGDATA), code VINIF.2020.TS.68 (TTTN). This research is also funded by Vietnam National University, Ho Chi Minh City (VNU-HCM) under grant number C2021-18-12.
Author Contributions
TTTN, LTT, and MHT designed the study. TTTN, TTTN, DHN, TKN, and PTVP carried out the experiments. TTTN and MHT drafted the manuscript and conducted experiments. TKN analyzed the data. PTVP, LTTT, TKN, and MHT revised the manuscript. All authors read and approved the final manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Statement of Informed Consent
There are no human subjects in this article, and informed consent is not applicable.
ORCID iDs
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
