Abstract
Introduction
Coronavirus disease 2019 (COVID-19), formally known as novel coronavirus pneumonia, is an infectious disease caused by the SARS-CoV-2 virus, and its source is individuals infected with novel coronaviruses, which are primarily transmitted through the respiratory tract, close contact, aerosol transmission in confined spaces, and contact with contaminated objects.1,2 SARS-CoV-2 invades cells via angiotensin-converting enzyme 2, thereby directly damaging lung tissues, triggering cytokine storms, worsening inflammatory responses, and resulting in conditions such as acute respiratory distress syndrome (ARDS). In severe cases, it results in respiratory–circulatory failure and even systemic inflammatory response syndrome, which is characterized by shock, vascular leakage, disseminated intravascular coagulation, and multiple organ failure (MOF). 3
Traditional Chinese medicine (TCM) has been used for thousands of years to treat epidemic diseases, and its preventive and therapeutic effects are well-documented. TCM-based approaches have shown initial success in preventing, treating, and rehabilitating COVID-19. However, most of the TCM-related studies rely on clinical experience or involve small sample sizes, with insufficient research conducted on mechanisms underlying TCM treatment for COVID-19. 4 This study introduces a novel approach by combining network pharmacology and experimental validation to comprehensively explore the therapeutic mechanisms of andrographolide, a key bioactive compound from Andrographis paniculata, against COVID-19. By integrating bioinformatics predictions with laboratory experiments, this research provides a unique insight into both the anti-inflammatory and antiviral potential of andrographolide, aiming to bridge the gap between traditional knowledge and modern scientific evidence.
Andrographis paniculata (Burm. f.) Nees., commonly referred to as “King of Bitters” or “Indian Echinacea,” belongs to the Acanthaceae family. Andrographolide, a primary active compound derived from the entire herb or its leaves, possesses detoxification, antimicrobial, anti-inflammatory, antitussive, and antidiarrheal potential. 5 Andrographis paniculata is widely used in Asian countries including China to treat diseases associated with inflammation, including upper respiratory tract infections, viral pneumonia, and bacillary dysentery. 6 A study has shown that andrographolide can be used in the clinical treatment of COVID-19. 7 Thus, herein, we elucidated the mechanism of action through which andrographolide helps in COVID-19 treatment using network pharmacology and subsequently validated its effects by performing relevant experiments.
Materials and Methods
Target Acquisition and Screening
The PubChem database (https://pubchem.ncbi.nlm.nih.gov) was used to obtain the molecular structure and SMILES notation of andrographolide. 8 Further, its potential protein targets were acquired from four databases, TCMSP (http://lsp.nwu.edu.cn/tcmsp.php), 9 DrugBank (https://www.drugbank.ca), 10 STITCH (http://stitch.embl.de/), 11 and SwissTargetPrediction (http://swisstargetprediction.ch/). 12 The identities of these potential targets were validated by cross-referencing using UniProt, and the final set of targets was established after eliminating duplicates. COVID-19-related targets were acquired by searching the following databases: PubChem (https://pubchem.ncbi.nlm.nih.gov), 8 Therapeutic Target Database (TTD, http://db.idrblab.net/ttd/), 13 GeneCards (https://www.genecards.org/), 14 DisGeNET (https://www.disgenet.org/), 15 DrugBank (https://go.drugbank.com/), 10 and PharmGKB (https://www.pharmgkb.org/) 16 using the keywords “COVID-19” and “SARS-CoV-2”. A Venn diagram illustrating the overlap between the andrographolide and COVID-19 targets was created.
Protein–Protein Interaction (PPI) Network Construction
The catalog of shared targets between andrographolide and COVID-19 was submitted to the STRING database (version 11.5, https://string-db.org). 17 “Homo sapiens” was selected as a species, a confidence threshold of ≥0.4 was applied, and the default settings were used for the remaining parameters. The PPI was extracted from STRING, exported in the tab-separated value format, and subsequently visualized using Cytoscape (version 3.7.0) to construct a PPI network diagram. 18 “Network Analyzer” in Cytoscape was used to analyze network topology parameters. A node filtration process was followed to find out the top 10 targets exhibiting the highest degree of centrality, thereby identifying prospective key targets occupying central positions within the network.
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Enrichment Analyses
Intersecting targets were uploaded to Metascape (http://metascape.org) for GO and KEGG pathway enrichment analyses, 19 with “Homo sapiens” specified as a species. GO analysis focused on biological processes, cellular components, and molecular function categories. The top 10 terms obtained by calculating −log10 (P-value) values for the GO and KEGG pathway enrichment results were plotted as bar graphs using R software (version 4.1.1). To explore relationships between the targets, we constructed a function–target–pathway network using Cytoscape (version 3.7.0) and analyzed it for properties including degree, betweenness centrality, and closeness centrality. The andrographolide and COVID-19 target intersection results were integrated during this network visualization, providing insights into mechanisms underlying the effects of andrographolide on COVID-19.
Molecular Docking
The three-dimensional (3D) protein structures of key targets were obtained from the Protein Data Bank (PDB; http://www.rcsb.org) in the PDB format. 20 Furthermore, homology modeling was performed for proteins with incomplete structural data using SwissDock (https://swissmodel.expasy.org), 21 an automated protein structure homology-modeling server, to complete missing loops and domains. Additionally, the 3D structures of the top 10 active compounds by degree centrality were downloaded from PubChem in the PDB format. The molecular docking of these active compounds to the protein targets was performed using Autodock Tools 1.5.6 and Autodock Vina 1.1.2, 22 with the docking box center defined as the coordinates of the original ligand in the receptor structure. The molecular docking results were visualized and schematically represented using PyMol 2.4.0. 23
Drug Property Prediction
The absorption, distribution, metabolism, and excretion parameters, pharmacokinetic properties, drug-likeness, and medicinal chemistry friendliness of the compound were analyzed using SwissADME. 24
Cell Culture and Cytotoxicity Assays
The human bronchial epithelial cell line BEAS-2B was procured from the Chinese Typical Culture Collection Center (Wuhan, China). BEAS-2B cells were maintained in the Roswell Park Memorial Institute 1640 medium supplemented with 10% fetal bovine serum at 37 °C in a humidified 5% CO2 cell culture incubator. For cytotoxicity assays, 100 μL of a cell suspension containing 5 × 10 4 cells/mL was seeded into each well of 96-well plates. After 24 h of incubation, andrographolide (Sigma-Aldrich, St. Louis, MO, USA) dissolved in the culture medium was added to each well at final concentrations of 10, 20, 40, 60, 80, 100, and 160 μmol/L. Cells cultured without any added drug were used as the negative control. After additional incubation for 24 h, 10 μL of cell counting kit-8 (CCK-8) solution (Beyotime Biotechnology, Shanghai, China) was added to each well and incubated for 2 h. Optical density at 450 nm was measured for each well. The cell survival rate (%) of each experimental group was calculated as follows: cell survival rate (%) = OD (test − blank wells)/OD (control − blank wells) × 100%. The experiment was repeated thrice with five parallel wells each time.
Quantitative Polymerase Chain Reaction (qPCR) of IL-6, IL-1β, TNF-α, and STAT3 Genes
The cells were collected and total RNA was extracted using the ES Science RNA Rapid Extraction Kit per the manufacturer's protocol. RNA purity and concentration were assessed by spectrophotometry. Complementary DNA (cDNA) was synthesized from the total RNA using the Thermo Scientific reverse transcription kit. qPCR was performed on the cDNA using PowerUp SYBR Green Master Mix (Thermo Scientific, American) to evaluate the mRNA expression of IL-6, IL-1β, TNF-α, and STAT3. The 2−ΔΔCt method was used to calculate relative mRNA expression normalized to that of an internal control gene. The details of primers and the reference gene are provided in Table 1.
Details of Primers of IL-6, IL-1β, TNF-α, and STAT3 and the Reference Gene (β-Actin) Used in the Study.

Pseudovirus Infection
Pseudovirus-SARS-CoV-2 (B.1.617.2 variant) harboring RNA sequences encoding green fluorescent protein (GFP) and luciferase were procured from General Biology (Anhui, China). 25 Infection efficiency was analyzed by visualizing GFP expression via fluorescence microscopy and luciferase activity was quantified via a reporter assay. BEAS-2B cells were seeded in 96-well plates at a density to achieve 40% confluence. The cultures were pre-treated with varying concentrations of andrographolide (0, 10, 20, and 40 μmol/L) for 6 h, followed by pseudovirions (10 μL/well) supplementation and co-incubation with the cells for 8 h. After incubating for 48 h, GFP fluorescence was imaged using an epifluorescence microscope. The luciferase assay was performed using a commercial system as per the manufacturer's instructions. A total of 20 μL of the luciferin reagent was added to each well to determine firefly luciferase activity. The results were expressed as the ratio of experimental reporter luminescence to control reporter luminescence, which was normalized to that observed in the vehicle control wells. Each condition was examined in triplicate. these experiments helped understand the effects of andrographolide on SARS-CoV-2 pseudovirus infection in the BEAS-2B cells.
Statistical Analysis
Data were analyzed by one-way analysis of variance using GraphPad Prism 8. Differences at P < 0.05 were considered statistically significant.
Results
Prediction of the Target of Action of Andrographolide
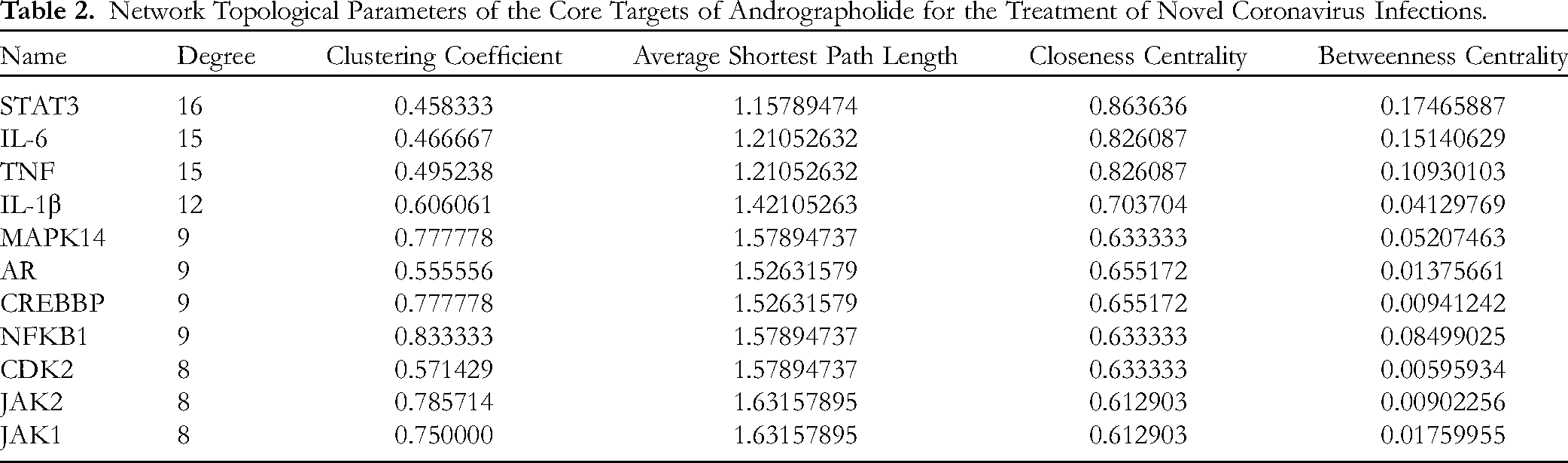
In silico target identification was performed by mining BindingDB, DrugBank, and SwissTargetPrediction, and 50 putative targets for andrographolide were identified. The keywords “COVID-19” and “SARS-CoV-2” were used to query GeneCards, TTD, DisGeNET, DrugBank, PharmGKB, and PubChem to obtain putative targets implicated in coronavirus disease, yielding 1,346, 65, 765, 21, 4, and 21 hits, respectively. Retrieved proteins were verified against those deposited in UniProt, obtaining 2004 unique COVID-19-related human proteins. The predicted andrographolide targets intersected with the SARS-CoV-2 interactome, identifying 23 proteins at the interface between the natural product and the infectious agent (Figure 1). Network analysis was performed by mapping the 23 intersected targets onto the STRING PPI network and visualizing the composite network using Cytoscape. The topological analysis of the resulting andrographolide–COVID-19 interactome network showed 10 highly connected nodes (hub proteins), namely signal transducer and activator of transcription 3 (STAT3), interleukin-6 (IL-6), tumor necrosis factor (TNF), interleukin-1 beta (IL-1Β), mitogen-activated protein kinase 14 (MAPK14), androgen receptor (AR), CREB binding protein (CREBBP), nuclear factor kappa B subunit 1 (NFKB1), cyclin-dependent kinase 2 (CDK2), and Janus kinases 2 and 1 (JAK2 and JAK1), which were predicted as key targets (Table 2 and Figure 1).

Disease-drug target Venn diagram and the core target PPI network.
Network Topological Parameters of the Core Targets of Andrographolide for the Treatment of Novel Coronavirus Infections.
GO and KEGG Enrichment Analyses
The functional analysis of the 23 intersected targets was performed using Metascape, a bioinformatics platform, to identify enriched GO and KEGG pathways. The results of the GO biological process enrichment analysis revealed that the andrographolide-COVID-19 target network was highly enriched in processes related to responses to lipopolysaccharides, bacterial molecules, hormones, and other endogenous and exogenous stimuli. Pathways related to hepatitis B infection, advanced glycation end-product (AGE)–receptor for AGE signaling in diabetic complications, lipid and atherosclerosis, COVID-19, and influenza A infection were the top 5 statistically enriched KEGG pathways. These results indicated the identified molecular targets of andrographolide in modulating critical pathways associated with host responses to SARS-CoV-2 infection, suggesting potential mechanisms by which this natural product exerted therapeutic effects against COVID-19 (Figure 2). In summary, this in silico study showed multiple andrographolide targets at the interface of host–virus interaction networks and biologically relevant signaling cascades.

Results of GO and KEGG analysis. (A) Bar chart of the molecular function categories. (B) Bar chart of biological process categories. (C) Bar chart depicting the cellular component categories. (D) Bar chart depicting the KEGG pathways.
Network Analysis
The topological analysis of biological networks can provide critical insights into mechanisms underlying drug–target interactions. An integrative network was constructed using Cytoscape 3.7.0 that showed intersections between andrographolide targets, their biological functions, and pathways associated with COVID-19 pathogenesis (Figure 3). This system-level visualization comprised 67 highly interconnected nodes mediated through 308 edges. Within the network, the non-uniform connectivity distribution indicated the presence of highly concentrated hub regions, suggesting the tendency of andrographolide to modulate specific proteins and pathways over others.

The compound-target-pathway network.
Molecular Docking Analysis
As shown in Table 3, the binding energies of andrographolide with NFKB1, JAK2, IL-6, AR, TNF, IL-1β, CREBBP, CDK2 and MAPK14 were less than −5.0 kcal·mol−1, indicating a good binding force. The binding energies of andrographolide with CDK2, JAK2, IL-1β, and MAPK14 are shown in Figure 4.

Molecular docking diagram of andrographolide and its target.
Docking Results of Andrographolide and Target Proteins.

Andrographolide Inhibits the mRNA Expression of Inflammatory Factors
In the CCK-8 cytotoxicity assay, treatment with andrographolide resulted in a dose-dependent decrease in the viability of BEAS-2B bronchial epithelial cells. The cell viability was >90% at andrographolide concentrations up to 60 μmol/L, suggesting the selective cytotoxicity of the compound (Figure 5A). qPCR revealed that andrographolide dose-dependently inhibited the mRNA expression of the proinflammatory mediators IL-6, TNF-α, IL-1β, and STAT3 compared with untreated controls (Figure 5B). These results suggest the concentration-dependent anti-inflammatory effects of andrographolide at the transcriptional level. Collectively, our data indicate that andrographolide suppresses inflammatory signaling in lung epithelial cells, with minimal toxicity at pharmacologically relevant doses.

Cellular assay to analyze the action of andrographolide on inflammatory factors and pseudoviral infections. (A) The viability of BEAS-2B cells was measured after 24 h treatment with different concentrations of andrographolide. (B) The mRNA expression of target proteins after the effect of different concentrations of andrographolide. (C) Andrographolide drug property prediction using the SwissADME web tool. The BioavailabilityRadar displays the drug-likeness of a molecule. The pink area represents the optimal range for each property. Six physicochemical properties are considered: lipophilicity, size, polarity, solubility, flexibility, and saturation. (D) The luciferase activities of the reporter genes in each group were measured using the luciferase assay kit after infection with SARS-CoV-2 pseudovirus and treatment with different concentrations of andrographolide. When compared with the blank group: *P < 0 .05, **P < 0.01. (E) Fluorescence photos of SARS-CoV-2 pseudovirus cells infected with different concentrations of andrographolide (100×).
Inhibition of SARS-CoV-2 Pseudovirus Infection by Andrographolide
Compared with uninfected controls, SARS-CoV-2 pseudovirus-infected BEAS-2B cells exhibited robust enhancements in bioluminescence and fluorescence emissions; this indicates efficient viral binding and replication (Figure 5D and 5E). Treatment with increasing concentrations of andrographolide markedly suppressed the luminescence emissions in a dose-responsive manner, suggesting the ability of andrographolide to interfere with pseudovirus entry and intracellular replication cycles. As such, our results reveal the antiviral properties of andrographolide against the novel coronavirus. Furthermore, in silico pharmacological profiling predicted advantageous drug-like attributes for andrographolide, further highlighting its potential as a therapeutic candidate against COVID-19 (Figure 5C).
Discussion
In TCM, Andrographis paniculata is ascribed heat-clearing, detoxifying, blood-cooling, and swelling-reducing properties. Modern pharmacological studies have continuously validated the broad anti-inflammatory, antibacterial, and antiviral bioactivities of this plant.26–28 Andrographolide preparations exhibit considerable antipyretic and anti-inflammatory effects, with substantial clinical evidence that they ameliorate upper respiratory tract infections, viral pneumonia, chronic obstructive pulmonary disease, bronchitis, and other inflammatory disorders. 29 The drug may attenuate pathological inflammation and improve symptoms of cough, fever, and pulmonary rales while delaying disease progression in the lungs. 30 Unlike corticosteroidal immunomodulators, andrographolide mediates anti-inflammatory effects via unique mechanisms that spare immunosuppression and minimize adverse reactions. Previous studies have demonstrated the inhibitory effects of andrographolide against SARS-CoV viral replication. 31 In our previous study, we demonstrated that andrographolide effectively inhibited SARS-CoV-2 infection by downregulating the expression of ACE2. 32 Along with the characterized antiviral mechanisms of andrographolide, these findings suggest it exhibits potential anti-SARS-CoV-2 activity.
In the present study, network pharmacology was used to determine the putative mechanisms of action underlying the therapeutic effects of andrographolide against COVID-19.33,34 The construction of the PPI networks led to the identification of the key targets of andrographolide involved in COVID-19 treatment: STAT3, IL-6, TNF, IL-1β, MAPK14, AR, CREBBP, NFKB1, CDK2, JAK2, and JAK1. Upon infection with SARS-CoV-2, although the immune system acts as the first line of defense, excessive inflammation, and impaired adaptive responses can precipitate systemic damage. 35 In particular, a cytokine or chemokine storm reflects the extensive dysregulation of host immunity, potentially resulting in ARDS, MOF, and death. 36 The major cytokines involved in COVID-19 hyperinflammation are interleukins, interferons, colony-stimulating factors, and monocyte chemoattractant protein-1.37,38 Notably, the progressive increase in IL-6 levels serves as an important marker for disease progression. 36 Because IL-6 is secreted by immune cells, it activates the JAK–STAT signaling pathway; the overactivation of this pathway can lead to aberrant inflammation via widespread cytokine release. 39 The JAK–STAT pathway represents a vital signaling conduit for myriad cytokines involved in physiological and pathological processes, including the cytokine storm. 40 In this pathway, the kinase JAK2 and its effector STAT3 represent key regulatory nodes because JAK2-mediated STAT3 phosphorylation facilitates its nuclear translocation and the transcriptional regulation of proinflammatory genes. 41 Both the JAK2–STAT3 and MAPK signaling pathways play key roles in controlling immune responses and inflammation. 42 Moreover, molecular docking revealed potent binding between andrographolide and the identified protein targets, suggesting that the compound exerts anti-inflammatory effects in COVID-19 by modulating these central networks. In summary, these vital immune pathways may serve as high-value therapeutic targets for ameliorating hyperinflammation and improving COVID-19 outcomes.
GO enrichment analysis of the intersecting targets revealed the involvement of the pathways controlling cellular responses to lipopolysaccharide and bacterial molecules; this suggests that anti-inflammatory and antibacterial mechanisms support the bioactivities of andrographolide. Furthermore, KEGG analysis revealed the enrichment of the pathways intimately associated with viruses, influenza, and inflammation. Notably, the top enriched pathways included influenza A infection, Mycobacterium tuberculosis infection, SARS-CoV-2 infection, and T helper 17 (Th17) cell differentiation, implicating the key roles of this natural product in modulating inflammation and antiviral immunity. SARS-CoV-2 induces the substantial release of inflammatory mediators; this “cytokine storm” drives tissue damage in the liver, lungs, and other organs.43,44 Andrographolide may mitigate COVID-19 severity and multiorgan injury by attenuating aberrant inflammation and immune hyperactivation. Th17 cells, a recently discovered effector T cell subset, can effectively induce inflammation by secreting different inflammatory cytokines and chemokines. 45 As such, Th17 cells play vital roles in maintaining pathological inflammation and autoimmunity. 46 The putative effects of andrographolide on Th17-mediated inflammation may support its therapeutic efficacy in inhibiting the cytokine storm and hyperinflammation associated with poor COVID-19 prognosis. In summary, andrographolide may improve COVID-19 outcomes via the coordinated inhibition of inflammatory cascades and restoration of balanced antiviral immunity.
To further determine the immunomodulatory effects of andrographolide, qPCR was performed to evaluate its effect on the mRNA expression of IL-6, TNF-α, IL-1β, and STAT3 in BEAS-2B bronchial epithelial cells. Treatment with andrographolide significantly inhibited the expression of these proinflammatory mediators, confirming the anti-inflammatory effects of andrographolide. In addition, we examined the direct antiviral activity of andrographolide against SARS-CoV-2 using a pseudovirus system incorporating the B.1.617.2 mutant spike glycoprotein. This pseudovirus displayed the variant spike protein and encapsidated reporter genes encoding GFP and luciferase. Andrographolide effectively suppressed pseudovirus entry and infection in HEK293 T cells, engineered to express angiotensin-converting enzyme 2, a SARS-CoV-2 receptor. Taken together, these findings suggest that andrographolide exhibits both anti-inflammatory and antiviral effects against SARS-CoV-2 in vitro, thereby inhibiting virus entry and replication and proinflammatory signaling cascades.
However, several limitations to this research remain. The utilization of network pharmacology, albeit an innovative strategy for generating hypotheses, tends to oversimplify the vast and complex biological systems and interactions. Consequently, this might lead to an exaggerated perception of andrographolide's efficacy, neglecting the dynamic and intricate nature of biological networks in the human body. Additionally, the study's experimental framework, which predominantly leans on in vitro assays using a solitary cell line, may not offer a true reflection of the in vivo situation. Such this cell culture model falls short in mirroring the intricate interplay among various cell types and the immune response observed in a living organism. While the pseudovirus system offers insight into viral entry mechanisms, it does not encompass the full complexity of SARS-CoV-2's pathogenesis. Thus, while the study provides a preliminary groundwork for further exploration, its limitations underscore the critical need for comprehensive in vivo investigations and clinical trials to confirm andrographolide's potential as a therapeutic agent against COVID-19.
Conclusion
This study aimed to explore the therapeutic potential of andrographolide against COVID-19 using a combination of network pharmacology and experimental validation. We identified key molecular targets of andrographolide that are implicated in the pathogenesis of COVID-19, including STAT3, IL-6, TNF, and IL-1β, among others. Our in vitro experiments demonstrated that andrographolide effectively inhibits the mRNA expression of these proinflammatory mediators and suppresses pseudovirus entry and replication, suggesting its dual role in modulating inflammation and antiviral immunity. These findings suggest that andrographolide could serve as a promising therapeutic agent for COVID-19, with its ability to target critical inflammatory and immune pathways. However, while our results provide a strong foundation for further exploration, the study's reliance on in vitro assays highlights the need for in vivo validation and clinical trials to fully elucidate the therapeutic potential of andrographolide in a clinical context. In summary, this study not only meets the objectives set forth to understand the mechanisms underlying andrographolide's effects against COVID-19 but also offers insights into its possible application as a treatment option. Future research should focus on validating these findings in vivo and exploring the clinical feasibility of andrographolide in treating COVID-19 patients.
Footnotes
Acknowledgments
The authors are grateful to all those who helped in this study.
Authors Contributions
Jingyi Zhang and Hongmei Lu participated in the experiments, collated data, visualized data, and performed statistical analysis. Hongmei Lu and Yongdui Ruan reviewed the manuscript and provided funding. Wentao Guo was responsible for research oversight. Shiyi Huang and Siying Deng provided suggestions and ideas for manuscript writing. Zuguo Zhao and Long Feng contributed to research execution, management, and coordination. Jingyi Zhang, Hongmei Lu, and Yongdui Ruan contributed equally to this work. All authors contributed to the article and approved the final manuscript version.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical approval is not applicable for this article.
Funding
This study received funding support from several sources, including the Guangdong Provincial Bureau of Traditional Chinese Medicine Scientific Research Project (20232097), the Guangdong Provincial Bureau of Traditional Chinese Medicine Scientific Research Project (20211220), the Science and Technology Project of Dongguan (20211800905542), the Guangdong Medical University Scientific Research Project (GDMUQ2021005), and the Discipline Construction Project of Guangdong Medical University (4SG21229GDGFY01), Henan Natural Science Foundation (232300421184), National Natural Science Foundation of China (81503677), Postgraduate Education Reform and Quality Improvement Project of Henan Province (YJS2023ZX15).
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
