Abstract
Introduction
Coronavirus disease 2019 (COVID-19), which is caused by infection with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has rapidly spread worldwide since the end of 2019. The cumulative global number of confirmed cases is hundreds of millions, with the disease affecting most of the countries and regions. As COVID-19 poses a serious threat to human health, there is an urgent need for developing a drug against it.
Recent reports have highlighted that chemicals derived from medicinal plants, including traditional Chinese medicine, can play an important role in the fight against COVID-19. 1 The medicinal plant, Polygonum cuspidatum, has been helpful in preventing and controlling epidemics. The functions of P cuspidatum, as described in traditional Chinese medicine records, include clearing away of heat, detoxification, elimination of dampness, alleviation of jaundice, dispersing of stasis, resolution of pain, relieving cough, and resolving phlegm . 2 Modern pharmacological research has shown that this plant can also be used to treat viral infections and that it has antitussive and anti-asthmatic effects. 2 A number of medicines that may be effective in treating patients with COVID-19 have been discovered at Shanghai Institute of Materia Medica of Chinese Academy of Sciences and ShanghaiTech University using virtual screening and enzyme testing. Polydatin, a compound in P cuspidatum, was found to combine with the SARS-CoV-2 3CL hydrolase, leading to virus repression. 3 Polydatin is also isolated from many plants, such as peanuts, grapes, red wines, and several daily diets , 4 which plays a role in prevalent diseases through their antioxidant and anti-inflammatory effects, regulation of autophagy and apoptosis, maintenance of mitochondrial function, and lipid regulation . 5 Accordingly, the mechanism of polydatin against COVID-19 needs to be further explored.
With the accumulation of data and the progress in computational technology, several bioinformatics methods have come to prominence. Recently, they have been widely used in exploring the mechanism of phytochemicals against COVID-19. Phytochemicals derived from ayurvedic and African medicinal plants, including Picrorhiza kurroa, Cyperus rotundus, Aegle marmelos, Vetiveria zizanoides, Moringa olifera, Zingiber officinale, and Anacardium occidentale, have antiviral potential, which was validated through in silico methods, including molecular docking and molecular dynamic simulation.6–10 As network pharmacology is an analytical method based on the interactive network of diseases, targets, and drugs, it was employed in the present study. This method integrates bioinformatics, systematic biology, molecular biology, and other disciplines. 11 We were able to explore the mechanism of action of polydatin in COVID-19 treatment using computational analysis and provide a reference for further research.
Methods
Collection of Drug Targets and Analysis of Their spectrum of Disease
The two-dimensional (2D) and three-dimensional (3D) structural models of polydatin were obtained from PubChem (CID:5281718), 12 wherein the polydatin targets were identified from TCMSP, 13 TCMIP v2.0, 14 and CTD 15 databases; the targets were named in accordance with those in UniProt. 16 After target collection, Metascape 17 was used to analyze the disease spectrum associated with the polydatin targets.
Determining Targets of Polydatin Against COVID-19
Using DisGeNET (N.PMIDs≥20), 18 Genecards (relevance score>5), 19 and CTD 15 databases (top 100), the targets of novel coronavirus pneumonia or COVID-19 were identified and named according to those in UniProt. 16 Moreover, in concordance with existing reports, 20 genes (P value<0.05) that correlated with the angiotensin-converting enzyme 2 (ACE2) and are expressed in colonic epithelial cells were also identified and named according to those in UniProt.
Venny v2.1 software 21 was used to generate a Venn diagram of the shared targets between polydatin and those of COVID-19. STRING v11 22 was used to analyze the protein–protein interactions (parameter settings: minimum required interaction score, 0.40), Cytoscape 3.7.2 23 was used to draw the target diagrams, and to obtain the hub targets of polydatin against COVID-19. Venny v2.1 was also used to produce a Venn diagram depicting the common targets between polydatin and those correlated with ACE2 expression in colonic epithelial cells; STRING v11 was used to draw diagrams of the common targets (parameter settings: minimum required interaction score, 0.40).
Enrichment Analysis of Drug–Disease Targets
To elucidate the mechanism of action of polydatin against COVID-19, enrichment analyses were performed using the Metascape database, with the following ontology sources: KEGG Pathway, GO Biological Processes, Reactome Gene Sets, Canonical Pathways, and WikiPathways. 17 To analyze the common targets between polydatin and those correlated with ACE2 expression in colonic epithelial cells, KEGG pathway analysis was conducted using STRING v11 (parameter settings: minimum required interaction score, 0.40). 22
Molecular Docking
To validate further the polydatin targets in COVID-19, we analyzed the docking scores of polydatin, with the following potential target proteins: SARS-CoV-2 Mpro (PDB: 6LU7), SARS-CoV-2 RdRp (PDB: 6M71), and human ACE2 (PDB: 6M17). The crystalline structures of the candidate proteins were obtained from the Protein Data Bank, 24 and were then modified by removing water, adding hydrogen, and optimizing the structure. The protein structures were subjected to a series of restrained minimizations to relieve any static clashes using an OPLS2005 force field up to 0.30 Å RMSD. The molecular docking was performed using Schrödinger Maestro software v11.4. 25
Cell and Pseudovirus Preparation
HEK293T-ACE2, a stable cell line overexpressing human ACE2, was obtained from Obio Technology (Shanghai) Corp., Ltd. The cell line was constructed via lentiviral infection combined with puromycin (Amresco) screening. Cells were cultured using DMEM (Gibco) containing 10% fetal bovine serum (FBS) (Gibco) and 1% penicillin/streptomycin (Gibco). The cells were maintained at 37 °C in a 5% CO2 atmosphere. SARS-CoV-2 spike pseudovirus was obtained from Obio Technology (Shanghai) Corp., Ltd; it can be used to infect cells expressing ACE2, takes lentivirus as the basic vector, and replaces the VSV-G envelope protein of lentivirus with the spike protein of SARS-CoV-2. The pseudovirus also carries enhanced green fluorescent protein (EGFP) gene and luciferase reporter gene, which facilitate the detection in the subsequent experiment.
CCK8 Assay
To assess cell viability, HEK293T-ACE2 cells were seeded in 96-well plates. The cells were incubated with different concentrations of polydatin (MedChemExpress) for 48 h. The CCK8 reagent (Dojindo) was then added to each well and cells were incubated for 1 to 4 h. During incubation, the absorbance of cells in each well at 450 nm was measured using a microplate reader (Tecan).
Flow Cytometry
HEK293T-ACE2 cells were treated with SARS-CoV-2 spike pseudovirus and polydatin. HEK293T-ACE2 cells in the pseudovirus group only were infected with SARS-CoV-2 spike pseudovirus for 8 h. After 8 h of pseudovirus infection, the medium was replaced with a culture medium without pseudovirus and polydatin. HEK293T-ACE2 cells in the polydatin + pseudovirus group were pretreated with polydatin at different concentrations for 4 h, and then infected with SARS-CoV-2 spike pseudovirus for 8 h. After 8 h of pseudovirus infection, the medium was replaced with a culture medium containing polydatin. HEK293T-ACE2 cells in the pseudovirus + polydatin group were first infected with SARS-CoV-2 spike pseudovirus for 8 h, and then polydatin at different concentrations was added after 4 h of pseudovirus infection. After 8 h of pseudovirus infection, the medium was replaced with a culture medium containing polydatin. After incubation for 48 h in a 5% CO2 cell incubator at 37 °C, the cells were gently harvested and prepared as a single cell suspension for counting. To validate whether or not polydatin decreases SARS-CoV-2 spike pseudovirus infection in HEK293T-ACE2 cells, the level of green fluorescence in the cells was determined using flow cytometry. The cells were transferred to the tubes for flow cytometry and evaluated using FACSCalibur (BD).
Luciferase Assay
Cell preparation and treatments for the luciferase assays were the same as those described for the flow cytometry analysis. Luciferase activity was detected using a dual-luciferase reporter assay system (Promega) according to the manufacturer's specifications. After the appropriate pseudovirus and/or polydatin treatment, the cells were gently harvested at a density of 1 × 106 cells of each sample and resuspended in 100 μL of 1 × passive lysis buffer (PLB), which was provided with the assay system. Thereafter, 10 μL of lysis buffer was transferred into 96-well plates and 100 μL of luciferase assay reagent II (LAR II) was added. Fluorescence was detected using a microplate reader (Tecan). The experiment was performed in the dark.
Statistical Analysis
Experimental data are presented as the mean ± SEM. The data were analyzed by Student's two-tailed t-test using GraphPad Prism 8.0 (www.graphpad.com). Error bars in the figures represent SEM. Statistical significance was accepted at *P < 0.05, **P < 0.01,***P < 0.001, and **** P < 0.0001.
Results
Molecular Structure, Targets, and Disease Spectrum of Polydatin
The 2D and 3D structures of polydatin are shown in Figure 1, and the top 20 diseases targeted by polydatin ranked according to log10(P) or log10(q) are summarized in Figure 2 and Supplementary Table 1. Log10(P) is the P value in log base 10, whereas log10(q) is the multi-test adjusted P value in log base 10. The spectrum of diseases that can be treated using polydatin includes diabetes mellitus, fatty liver disease, lung diseases, respiratory distress syndrome, endothelial dysfunction, cerebral infarction, pancreatitis, hepatitis C, Helicobacter pylori infection, and malignant neoplasm of the skin, mouth and kidney. Overall, these diseases involve aspects of metabolism, lung, blood vessels, inflammation, infection, and cancer.
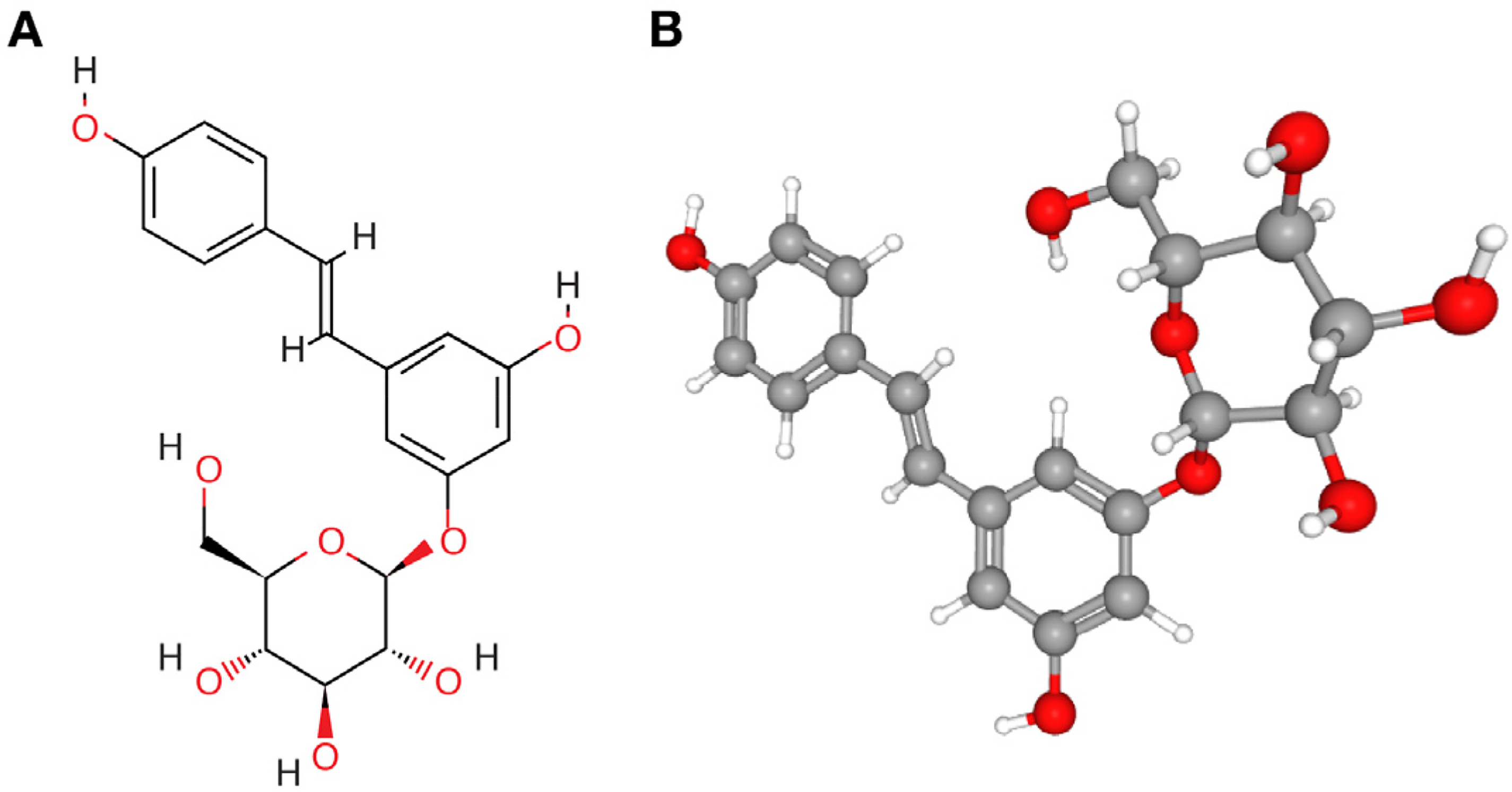
(A) Two- and (B) three-dimensional (2D and 3D) structural models of polydatin obtained from PubChem.

Top 20 potential diseases targeted by polydatin according to the enrichment analysis based on Metascape.
Analysis of the Polydatin Mechanism of Action Against COVID-19
Targets of polydatin and COVID-19 were identified. The Venn diagram of the shared targets of polydatin and COVID-19 is shown in Figure 3. The network diagram of the common targets is shown in Figure 4 with the targets listed in Supplementary Table 2. In addition, hub 10 targets and hub 6 targets were obtained based on the degree values of protein–protein interactions. These targets, including MAPK3, TP53, TNF, IL6, IL1B, PTGS2, CCL2, RELA, TGFB1, and CXCL8, are important potential targets of polydatin against COVID-19.

Venn diagram of the shared targets of polydatin and COVID-19 generated using Venny.

(A) Protein–protein interaction network of potential targets of polydatin against COVID-19, identified from STRING and visualized using Cytoscape. Protein–protein interaction network of (B) hub 10 targets and (C) hub 6 targets of polydatin against COVID-19 generated using Cytoscape. Rectangular nodes indicate targets, and the links represent different interactions.
The results of enrichment analysis regarding the mechanism of action of polydatin against COVID-19 are shown in Figure 5 and Supplementary Table 3. The main potential mechanisms involve the IL-17 signaling pathway, regulation of protein phosphorylation, folate metabolism, lipopolysaccharide-mediated signaling pathway, FOXO signaling pathway, T-cell activation, cytokines and inflammatory response, SARS-CoV-2 innate immunity evasion, and cell-specific immune response.

Top 20 terms identified from the pathway and process enrichment analyses of polydatin against COVID-19 based on Metascape.
Pathway Analysis Revealed the Common Targets Between Polydatin and Those Correlated With ACE2 Expression in Colonic Epithelial Cells
The identified common targets between polydatin and those correlated with ACE2 expression in colonic epithelial cells are shown in Figure 6A, whereas the network diagram of the shared targets including MAPK3, EGFR, IL4, and IFNB1 is shown in Figure 6B. The top 20 terms identified from the KEGG pathway analysis of the common targets are summarized in Table 1. The main enrichment terms involve cAMP signaling pathway, cytomegalovirus infection, tuberculosis, Rap1 signaling pathway, pertussis, shigellosis, Ras signaling pathway, bile secretion, pancreatic secretion, C-type lectin receptor signaling pathway, and T cell receptor signaling pathway.
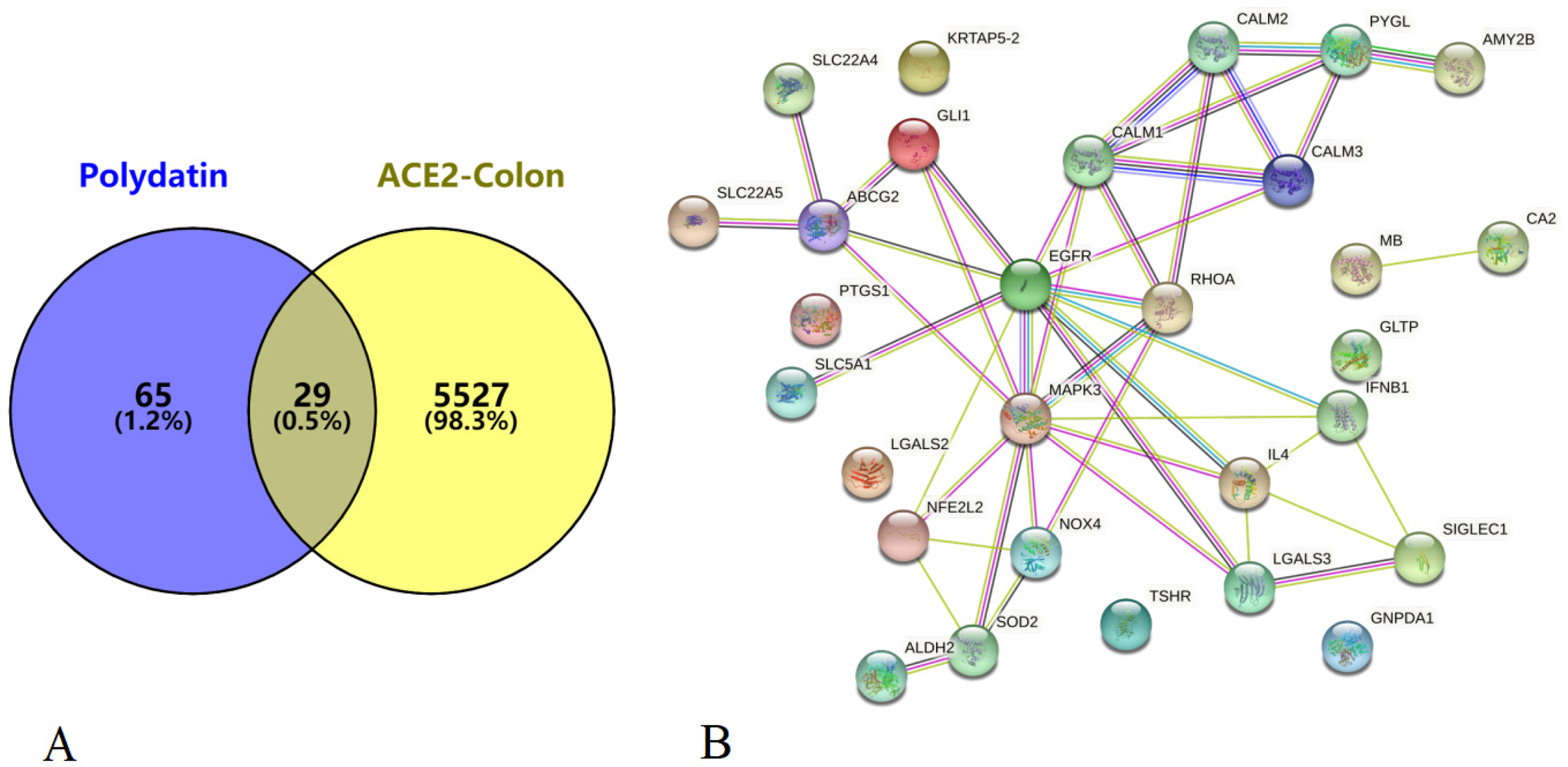
(A) Venn diagram depicting the targets of polydatin and the targets correlated with ACE2 expression in colonic epithelial cells generated using Venny. (B) The network diagram of the shared targets between polydatin and those correlated with ACE2 expression in colonic epithelial cells generated using STRING.
Top 20 Terms Identified From KEGG Pathway Analysis of the Common Targets Between Polydatin and Those Correlated With ACE2 Expression in Colonic Epithelial Cells, Which are Ranked Using the False Discovery Rate (FDR).

Molecular Docking Analysis of Polydatin and its Targets
The molecular docking results of 2D and 3D structural models of polydatin and potential target proteins, SARS-CoV-2 Mpro, RdRp, and human ACE2 are shown in Figure 7. Table 2 presents the docking score, which refers to the binding affinity between the compound and the target protein. The higher the absolute docking score, the greater the binding affinity. As can be seen from the analysis results, polydatin exhibited the potential to bind to all three of these target proteins related to COVID-19.

(A) Two- and (B) three-dimensional (2D and 3D) molecular docking models depict polydatin binding to the target proteins generated using the Schrödinger Maestro software.
Molecular Docking Results of Polydatin and its Candidate Target Proteins, Determined Using the Schrödinger Maestro Software.

Effects of Polydatin on the Viability of HEK293T-ACE2 Cells
In the CCK8 assay, we detected the viability of HEK293T-ACE2 cells with an increase in polydatin dose. The results showed that the cell viabilities of the Group Polydatin (PLD) 0.781 μM, PLD 3.125 μM, PLD 12.5 μM, PLD 50 μM, PLD 200 μM, and PLD 400 μM were 102.95% (P > 0.05), 96.69% (P > 0.05), 95.93% (P < 0.05), 83.72% (P < 0.05), 73.66% (P < 0.05), and 58.69% (P < 0.05), respectively relative to the Control (Figure 8). This suggested that 12.5 μM polydatin exerted a slight inhibitory effect on the viability of HEK293T-ACE2 cells; the higher the concentration of polydatin, the more significant is the inhibitory effect.

Effects of polydatin on the cell viability of HEK293T-ACE2 cells. Cells were cultured with different concentrations of polydatin for 48 h.
Effects of Polydatin on HEK293T-ACE2 Cells Infected With SARS-CoV-2 Spike Pseudovirus
The green fluorescence of HEK293T-ACE2 cells was detected using flow cytometry. The results showed that polydatin could decrease the infection of SARS-CoV-2 spike pseudovirus in HEK293T-ACE2 cells (Figure 9A and B). The infection rate in the groups that were pretreated with 3.125 and 12.5 μM polydatin decreased by 20.80% and 24.32%, respectively, compared with that in the positive control group, which was treated with the pseudovirus only. Moreover, the infection rate in the groups that were treated with 3.125 and 12.5 μM polydatin after 4 h of pseudovirus infection decreased by 7.45% and 11.76%, respectively, compared with that in the positive control group.

Effects of polydatin on HEK293T-ACE2 cells infected with SARS-CoV-2 spike pseudovirus detected using (A, B) flow cytometry and (C) luciferase assay.
The evaluation of luciferase fluorescence led to a similar conclusion (Figure 9C). The infection rate in the groups pretreated with 3.125 and 12.5 μM polydatin decreased by 26.24% and 45.34%, respectively, compared with that in the positive control group. Moreover, the infection rate in the groups treated with 3.125 and 12.5 μM polydatin after 4 h of pseudovirus infection decreased by 28.07% and 41.32%, respectively, compared with that in the positive control group.
Discussion
SARS-CoV-2 is a newly identified coronavirus strain that was previously not observed in humans. The genome of SARS-CoV-2 shares approximately 80% identity with that of SARS-CoV. 26 P cuspidatum has a long history of use in traditional Chinese medicine, with its active components reported to inhibit the replication of the influenza virus. 27 In the current study, the enrichment analysis of potential targets showed that polydatin may be effective against metabolic diseases, lung diseases, inflammation, infectious diseases, and tumors, which are also closely related to COVID-19. Acute respiratory distress syndrome (ARDS) is a common complication as a result of severe COVID-19 infection caused by SARS-CoV-2, which can induce hyperinflammation and cytokine storm, thereby, causing lung damage. 28 The severity and death due to COVID-19 are also related to cardiovascular disease, obesity, and metabolic disorders.29,30 In addition, the expression of ACE2 is higher in cervical squamous cell carcinoma, endocervical adenocarcinoma, pancreatic adenocarcinoma, rectum adenocarcinoma, kidney renal papillary cell carcinoma, kidney renal clear cell carcinoma, and other tumor tissues relative to that in the surrounding tissues. This has led to the hypothesis that patients with any one of these cancers may be susceptible to SARS-CoV-2 infection. 31 This hypothesis was partially supported by our analysis of the spectrum of diseases targeted by polydatin.
The overactivation of T cells may occur in patients with COVID-19, leading to severe immune-related damage. 32 The cytokine storm induced by SARS-CoV-2 infection also attacks healthy cells in the lungs, leading to respiratory failure. In the current study, we predicted therapeutic targets of polydatin against COVID-19, including MAPK3, TP53, TNF, IL6, IL1B, PTGS2, CCL2, RELA, TGFB1, and CXCL8. The tissue- and gender-specific regulation of ACE2 by p53 was reported in pigs. The absence of the TP53 binding site increases the activity of ACE2 promoter in porcine kidney cells. 33 An earlier study reported that p53 is one of the important targets of traditional Chinese herbal compound prescription, Xuanfei Baidu Tang, in the treatment of COVID-19. 34 TNF-α and IFN-γ can cause a lethal cytokine shock and inflammatory cell death in mice with COVID-19. 35 The enrichment analyses also revealed that the potential therapeutic mechanisms of polydatin against COVID-19 involve various pathways and processes, including the IL-17 signaling pathway, T-cell activation, cytokines and inflammatory response, lipopolysaccharide-mediated signaling pathway, folate metabolism, and FOXO signaling pathway, as well as SARS-CoV-2 innate immunity evasion and cell-specific immune response. Studies have also confirmed that polydatin has the function of regulating inflammatory mediators and immune response.36,37 These results suggest that polydatin may act against COVID-19 through these targets and mechanisms, indicating that the functions related to immune and inflammatory responses are largely associated with the activities of polydatin against COVID-19. The activation of G6PD is related to increased oxidative stress and inflammation in acute pulmonary injury. Polydatin can be used for patients in severe stages of COVID-19 as a potential G6PD inhibitor. 4 Polydatin can moderate acute clinical, chronic subclinical inflammation, and pneumonia or lung inflammation processes through the TLR4-MyD88-NF-κB pathway. 38 It has also been reported that glucose and folate are depleted in cells infected with SARS-CoV-2, and viral replication is exquisitely sensitive to inhibitors of folate and one-carbon metabolism. 39 Decreased levels of serum folate are common among inpatients with COVID-19. 40 Multiple strategies enable SARS-CoV-2 to evade the innate immunity; these include low cytosine–phosphate–guanosine levels in the genome and suppression of IFN-α/β production at the earliest stage of infection.41,42
According to a classic theory of traditional Chinese medicine, the lung and large intestine are “interior–exteriorly” related. It has been demonstrated that the frequency of gastrointestinal syndromes is high in patients with COVID-19. 43 Clinical data revealed diarrhea in a small number of patients with COVID-19, and SARS-CoV-2 was detectable in their stool samples.44,45 Furthermore, gastrointestinal infection can lead to the progression of COVID-19 and the gut microbiota may influence the expression level of colonic ACE2. 43 As ACE2 is typically highly expressed in the colonocytes and is involved in intestinal inflammation, the candidate targets identified in the present study were reported to correlate with ACE2 expression in colonic endothelial cells. 20 Our enrichment analyses indicate that the common targets between polydatin and those correlated with ACE2 expression in colonic epithelial cells potentially function in cAMP, Ras, and Rap1 signaling pathways, as well as in immune mechanisms, such as C-type lectin receptor and T cell receptor signaling pathway. Potential mechanisms include digestive mechanisms, such as adherens junction, bile secretion, pancreatic secretion, and colorectal cancer, as well as infectious diseases, such as tuberculosis, pertussis, shigellosis, and cytomegalovirus infection. Intestinal mucosal epithelial cells maintain their barrier function by forming complex protein networks, including adherens junctions, tight junctions, and gap junctions. Polydatin may alleviate mucosal oxidative stress and immunological damage and may improve the intestinal epithelium barrier.46,47
SARS-CoV-2 Mpro (also known as 3C-like protease) is an important enzyme for the replication of the coronavirus and is responsible for processing the polypeptide into functional proteins. 48 Furthermore, SARS-CoV-2 RdRp can also play a primary role in coronavirus replication and transcriptional machinery. 49 Human ACE2 is a cellular receptor for both SARS-CoV and SARS-CoV-2. 26 Accordingly, these three proteins have been reported to be potential targets for the development of antiviral drugs. Our molecular docking simulations suggest the potential of polydatin binding to the target proteins directly related to COVID-19, including SARS-CoV-2 Mpro, RdRp, and human ACE2, thereby, playing either a regulatory or therapeutic role. The potential binding affinity was the highest between polydatin and SARS-CoV-2 Mpro. A previous report based on molecular docking analyses suggests that the main bioactive compounds of Shufeng Jiedu capsules, including quercetin, wogonin, and polydatin, can bind directly to SARS-CoV-2 Mpro. However, the computational results in this report have not been verified experimentally. 50
The expression of the ACE2 receptor has been observed in multiple tissues including the lung, heart, kidney, endothelium, and intestine.51,52 ACE2 is one of the crucial enzymes in the renin–angiotensin system (RAS), which can regulate blood pressure, isohydria, inflammation, and vascular resistance. The SARS-CoV-2 spike protein is able to directly bind with the ACE2 receptor of the host cell surface, leading to virus entry and ultimately multiple organ damage. 53 Considering the importance of ACE2 and the analysis results of our network pharmacology and molecular docking analyses, we carried out experimental verification of our findings. The experimental results demonstrated that polydatin reduced the infection of the SARS-CoV-2 spike pseudovirus in HEK293T-ACE2 cells, which overexpress ACE2. We suggest that polydatin may retard the entry of SARS-CoV-2 into cells by competitively binding to human ACE2 (Figure 10). Molecular docking simulations in another report suggest that polydatin can bind to the SARS-CoV-2 Spike and ACE2 proteins. These simulations were confirmed by biochemical assays using the purified ACE2 protein and biotinylated SARS-CoV-2 Spike S1 protein. 54 This report corroborates our findings. We performed different experimental assays, using the SARS-CoV-2 spike pseudovirus and HEK293T-ACE2 cells, to better simulate the process of virus infection of human cells. However, owing to the limitations of the current research conditions employed, the mechanisms underlying the action of polydatin in the treatment of COVID-19 are worthy of further research.

Potential mechanism of action of polydatin against COVID-19. Polydatin has the potential to reduce the entry of SARS-CoV-2 into cells by competitively binding to human ACE2.
Conclusions
In this study, we explored the potential therapeutic mechanisms of polydatin action against COVID-19 using a combined approach of network pharmacology, molecular docking, and experimental verification. Polydatin may exhibit a functional role in the lungs and intestine, which are mainly involved in regulating immune functions and inflammatory responses. Polydatin may also exhibit potential regulatory or therapeutic effects by binding to the target proteins that are directly related to COVID-19, such as SARS-CoV-2 Mpro, RdRp, and human ACE2. Our experimental results show that polydatin can decrease the infection of the SARS-CoV-2 spike pseudovirus in HEK293T-ACE2 cells. Hence, polydatin may retard the entry of the virus into cells by competitively binding to human ACE2. These results suggest the potential of developing polydatin as a treatment for patients with COVID-19.
Supplemental Material
sj-doc-1-npx-10.1177_1934578X221095352 - Supplemental material for Combined Network Pharmacology, Molecular Docking, and Experimental Verification Approach to Investigate the Potential Mechanisms of Polydatin Against COVID-19
Supplemental material, sj-doc-1-npx-10.1177_1934578X221095352 for Combined Network Pharmacology, Molecular Docking, and Experimental Verification Approach to Investigate the Potential Mechanisms of Polydatin Against COVID-19 by Meng Wang, Kaijian Qin and Xiaofeng Zhai in Natural Product Communications
Footnotes
Acknowledgments
The authors would particularly like to acknowledge the team members for their wonderful collaboration and patient support.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by Shanghai major clinical research project on traditional Chinese medicine [grant number ZY(2018-2020)-CCCX-4003] and Three-year Action Plan for the Development of TCM in Hongkou District, Shanghai [grant number HGY-KY-2018-01].
Authors Contributions
Meng Wang: Conception of the study, analysis and interpretation of the data, and drafting the article. Kaijian Qin: Acquisition of data and revising the article critically. Xiaofeng Zhai: Design of the study and final approval of the version to be submitted.
Data Availability
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
Ethical Approval
Not applicable, because this article does not contain any studies with human or animal subjects.
Informed Consent
Not applicable, because this article does not contain any studies with human or animal subjects.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
