Abstract
The network pharmacological regulation mechanism has been studied in which Qing-Fei-Pai-Du Decoction (QFPDD) can be used for treating diabetic patients suffering from coronavirus disease 2019 (COVID-19). In this study, we retrieved the target genes of QFPDD along with its effective components from the Traditional Chinese medicine systems pharmacology database. The target genes of diabetes and COVID-19-associated diseases were searched based on the GeneCards database, and the co-target genes of QFPDD were obtained. The target genes were analyzed using the Gene Ontology and the Kyoto Encyclopedia of Genes and Genomes. Cytoscape software was used for constructing the “herb-ingredient-target” network. Then, the co-target genes were imported into the Metascape database to analyze protein–protein interaction. Based on the results of the analysis of the hub genes from each database, the key target genes and the active components of QFPDD effective for treating diabetic COVID-19 patients were obtained. The crystal structures of the key target genes were retrieved from the protein data bank database, and molecular docking (MD) was performed using the AutoDock Tool, PyMoL, Chen3D, and other software. In total, 305 active ingredients and 274 target genes were identified among 19 traditional Chinese medicines present in QFPDD. We found 4585 COVID-19 target genes, 16 660 diabetes target genes, and 60 drug-disease target genes. The results of the Gene ontology and Kyoto Encyclopedia of Genes and Genomes analyses showed that the main function of these co-target genes was immune regulation. Additionally, protein interaction analysis and cluster analysis were performed to obtain a protein interaction network, and 3 core proteins and 4 core active components were found. According to the results of MD, the main effective components (quercetin, kaempferol, and wogonin) present in QFPDD were found to bind strongly to the receptor. The main active components of QFPDD were effective in treating diabetes complicated with COVID-19 through their actions on multiple biological pathways.
Introduction
Coronavirus disease 2019 (COVID-19) is an acute infectious disease caused by severe acute respiratory syndrome-coronavirus 2 (SARS-COV-2). 1 Since December 2019, COVID-19 has led to significant human losses globally. SARS-COV-2 infection has affected over 500 million individuals worldwide, resulting in over 6 million deaths. 2 Individuals infected with SARS-CoV-2 often experience fever, dry cough, and a lack of energy. However, the severity of the disease can rapidly intensify and lead to the development of sepsis shock, acute respiratory distress syndrome, multiple organ dysfunction syndrome, blood coagulation disturbance, and difficulty in correcting metabolic acidosis. 3 Additionally, the risk of severe illness and death is considerably higher among elderly patients with diabetes. 4
Traditional Chinese medicine (TCM) has a strong theoretical foundation for preventing and treating diseases, and it has a significant impact on COVID-19 treatment. 5 Qing-Fei-Pai-Du Decoction (QFPDD) is a TCM formulation consisting of 21 Chinese herbs. It helps to relieve lung congestion and cough, detoxify dampness, invigorate the spleen, and improve health in general. 6 During the outbreak of COVID-19 in Wuhan (China) in 2020, QFPDD was administered to effectively control disease progression in COVID-19 patients. Among 701 patients treated with QFPDD, 130 were discharged with complete disease remission, 51 patients died, 268 patients showed improvements in their symptoms, and 212 patients had stable symptoms. These results indicated good clinical efficacy of QFPDD. 7 In another study conducted with 214 patients, the overall response rate for treating COVID-19 with QFPDD was higher than 90%. 8 QFPDD can significantly reduce the probability of severe diseases in mild patients. Therefore, when the “COVID-19 Diagnosis and Treatment Protocol (Trial Edition 7) in China” was released, QFPDD was listed as the first choice for the clinical treatment of TCM for COVID-19, suitable for mild, ordinary, and severe patients, and was administered for treating critical cases. 9
TCM is a complex system, and the therapeutic effects of QFPDD on COVID-19 patients have not been fully elucidated. It can effectively control the inflammatory response in diabetic patients with COVID-19 and has a significant positive effect on the treatment of diabetic patients with COVID-19; however, its regulatory mechanism is not clear. In this study, we investigated the known active ingredients and the targets of QFPDD from a systematic and holistic perspective by applying network pharmacology (NP). We also analyzed the mechanism by which QFPDD can be used to treat diabetic patients suffering from COVID-19, based on network topology analysis and molecular docking (MD), gene ontology (GO), and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses. Our findings provided the theoretical foundation for treating diabetes combined with COVID-19 by administering QFPDD.
Results and Discussion
Diabetes is a chronic metabolic disorder. Several studies have shown that diabetes is associated with the adverse prognosis of COVID-19, its severity, and the risk of higher mortality.10,11 In diabetic patients, SARS-COV-2 infection can induce a fatality rate of 20.3%, which is much higher than that of the general population (10.5%).12,13 The death rate due to chronic noncommunicable diseases increased by 21% in COVID-19 patients, including an 83% increase in the death rate due to diabetes. 14
Several studies have reported a complex and bidirectional relationship between COVID-19 and diabetes. 15 First, diabetes-related complications further increase the incidence of severe COVID-19. Second, severe COVID-19, along with steroid therapy, promotes diabetes and exacerbates hyperglycemia by enhancing insulin resistance while decreasing β-cell secretion. SARS-COV-2 infection increased the incidence of diabetes by 40% in 1 year, compared to the incidence of diabetes in uninfected individuals. 16
We obtained information on the main Chinese medicine, active ingredients, and the corresponding target genes of QFPDD from the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) database (https://old.tcmsp-e.com/tcmsp.php). 17 In total, 274 potential target genes were identified. We also identified the potential target genes of diabetes and COVID-19 from the GeneCards database (https://www.genecards.org/) 18 and identified 16 660 and 4585 potential target genes for diabetes and COVID-19, respectively. These potential target genes were analyzed in the Venn diagram database, and 60 co-target genes were identified. Next, the functions of the co-target genes were determined by performing the GO and KEGG signaling pathway analyses. The protein–protein interaction (PPI) analysis was performed for these co-target genes to identify the key regulatory target genes. These key regulatory target genes and the active substances in the drugs were analyzed by MD, and the regulatory target genes and effects were investigated (Figure 1).

Workflow chart of the present article.
First, we used the GeneCards database to identify the genes related to COVID-19 and diabetes and found that there were 3028 co-related genes, accounting for 66.04% of the genes related to COVID-19 and 18.18% of the genes related to diabetes. The results of the GO and KEGG analyses showed that these genes were significantly associated with cellular immune regulation and virus infection (Supplementary Table 1). QFPDD can effectively treat diabetic patients suffering from COVID-19 by significantly regulating the expression of inflammatory cytokines and reducing the damage caused by inflammation. 19 QFPDD also shortens the time of the virus in vivo and has a good prognosis.19,20 Using the TCMSP database, we found 305 effective components based on oral bioavailability (OB) ≥30% and drug-likeness (DL) ≥0.18 (Supplementary Table 2) from 19 Chinese medicines of QFPDD. These 305 active ingredients were screened for target genes, and 274 targets were identified. Additionally, 4585 targets were retrieved using the keyword “COVID-19,” and 16 660 target genes were retrieved using the keyword “diabetes,” which were found in combination with the drug targets. The co-target genes of QFPDD, COVID-19, and diabetes were analyzed using Venn diagrams (Figure 2). Through NP analysis, 60 co-targeted genes were found to be associated with cytokine activity, the phosphoinositide 3-kinase/Akt (PI3K/Akt) pathway, the tumor necrosis factor (TNF) pathway, the interleukin 17 (IL-17) pathway, and the mitogen-activated protein kinase (MAPK) pathway, as indicated by the results of the GO and KEGG analyses (Figure 3 and Supplementary Table 3).

Venn diagram analysis of the co-target proteins of COVID-19, diabetes, and QFPDD. (A) Co-target genes of Qing-Fei-Pai-Du Decoction (QFPDD) and COVID-19; (B) Co-target genes of COVID-19 and diabetes; and (C) Co-target genes of COVID-19, diabetes, and QFPDD.

GO and KEGG analyses of the co-target genes of COVID-19, diabetes, and Qing-Fei-Pai-Du Decoction (QFPDD). (A) gene ontology-molecular function (GO-MF) terms of the co-targets; (B) gene ontology-biological process (GO-BP) terms of the co-targets; (C) gene ontology-cellular component (GO-CC) terms of the co-targets; and (D) KEGG bubble diagram showing the key pathways of the co-targets.
Next, we built the network of herbs–ingredients–targets (Figure 4A and Supplementary Table 4) using Cytoscape software. We also used the CytoHubba plugin to examine the network hub components (Figures 4B-D and Table 1). The key target genes of the herb–ingredient–target network were tyrosine-protein phosphatase non-receptor type 1 (PTPN1), tumor necrosis factor (TNF), cAMP-dependent protein kinase catalytic subunit alpha (PRKACA), apoptosis regulator BAX (BAX), glycogen synthase kinase-3 beta (GSK3B), and androgen receptor (AR) (Figure 4F). The interactions and functions of the 60 co-targeted genes were investigated using the Metascape database (Figure 5). The functions were associated with signaling by ILs (IL-4 and IL-13) and cytokines, the inflammatory response, the Th17 cell differentiation pathway, IL-10 signaling, and the apoptotic signaling pathway (Figure 5A). We also identified 60 co-target genes that interacted with one another, as well as the genes and categories of related functional modules (Figure 5B). IL signaling, chemical carcinogens-reactive oxygen species, cytokine signaling associated with the immune system, the hypoxia-inducible factor 1 (HIF-1) pathway, the insulin pathway, protein binding regulation, control of immune tolerance by vasoactive motility peptides, and IL-4/IL-13 signaling were identified by the molecular complex detection (MCODE) algorithm analysis of these genes (Table 2).

Herb–ingredient–target network of a key component of Qing-Fei-Pai-Du Decoction (QFPDD). (A) Herb–ingredient–co-target regulatory network of QFPDD. (B–E) CytoHubba analysis of the TOP 30 components in the network; (B) Matthews correlation coefficient (MCC)-TOP30, (C) Degree-TOP30, (D) Closeness-TOP30, and (E) Betweenness-TOP30. (F) Co-target genes associated with the TOP 30 components in the network of QFPDD.

Protein–protein interaction (PPI) and enriched terms analyses of 60 target genes were performed using Metascape. (A) The enriched terms of the signaling pathway analysis and the typical terms in the full cluster were converted into the network layout. Cytoscape was used to visualize a network that had a “force-directed” layout and edge bundled for clarity. The nodes with different colors are based on P values in the network. A darker color indicates a higher node significance. (B) The PPIs across input genes were obtained from the PPI database and the PPI network was constructed. The MCODE algorithm was used for identifying neighborhoods with densely connected proteins.
Top 30 Effective Ingredients in the Herb–Ingredient–Target Network of QFPDD.
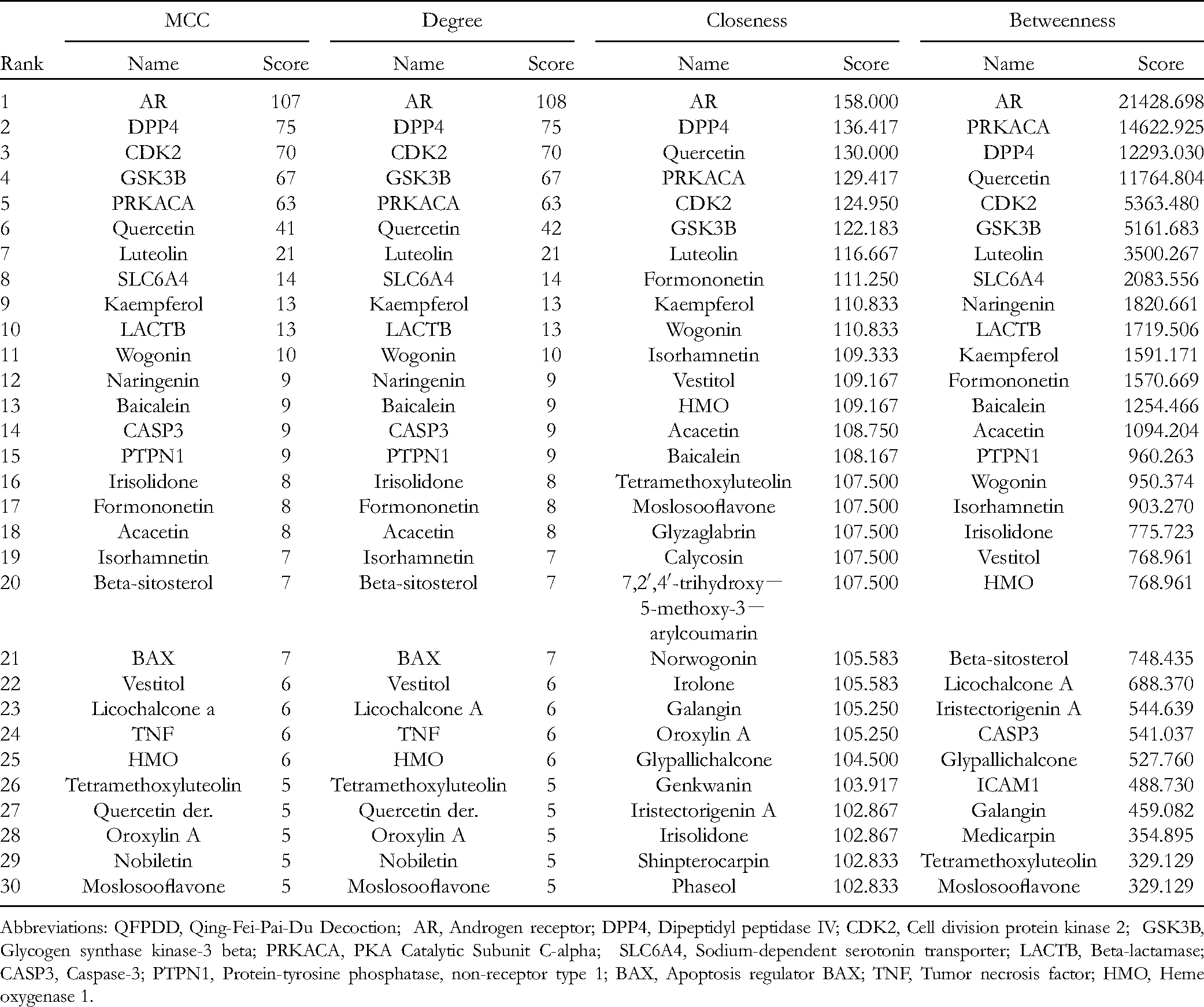
Abbreviations: QFPDD, Qing-Fei-Pai-Du Decoction; AR, Androgen receptor; DPP4, Dipeptidyl peptidase IV; CDK2, Cell division protein kinase 2; GSK3B, Glycogen synthase kinase-3 beta; PRKACA, PKA Catalytic Subunit C-alpha; SLC6A4, Sodium-dependent serotonin transporter; LACTB, Beta-lactamase; CASP3, Caspase-3; PTPN1, Protein-tyrosine phosphatase, non-receptor type 1; BAX, Apoptosis regulator BAX; TNF, Tumor necrosis factor; HMO, Heme oxygenase 1.
Description of MCODE for Target Genes of QFPDD, as Determined by the Metascape Analysis.

Abbreviations: QFPDD, Qing-Fei-Pai-Du Decoction; GO, Gene ontology; CXCL10, C-X-C motif chemokine 10; CCL2, C-C motif chemokine 2; MAPK3, Mitogen-activated protein kinase 3; MAPK1, Mitogen-activated protein kinase 1; STAT1, Signal transducer and activator of transcription 1; STAT3, Signal transducer and activator of transcription 3; TNF, Tumor necrosis factor ; CXCL2, C-X-C motif chemokine 2; IL1A, Interleukin-1 alpha; IFNG, Interferon gamma; IL1B, Interleukin-1 beta; IL2, Interleukin-2; IL6, Interleukin-6; PARP1, Poly [ADP-ribose] polymerase 1; HIF1A, Hypoxia-inducible factor 1-alpha; HMOX1, Heme oxygenase 1; VEGFA, Vascular endothelial growth factor A; PTPN1, Protein-tyrosine phosphatase, non-receptor type 1; GJA1, Gap junction alpha-1 protein; EGFR, Epidermal growth factor receptor; GSK3B, Glycogen synthase kinase-3 beta; PRKACA, PKA Catalytic Subunit C-alpha; BAX, Apoptosis regulator BAX; SREBF1, Sterol regulatory element-binding protein 1; FASN, Fatty acid synthase; CAV1, Caveolin-1; ICAM1, Intercellular adhesion molecule 1; IL10, Interleukin-10; IL1, Interleukin-1; TGFB1, Transforming growth factor beta-1.
We constructed an ingredient–target interaction network diagram of the 6 hub genes (Figure 6A and Supplementary Table 5). The key components were investigated using the CytoHubba algorithm (Figures 6B–E). The flavonoid, quercetin, is found in Chinese herbal medicines. 21 It shows anti-inflammatory, anticancer, antioxidant, and antiapoptotic effects, mainly by inhibiting inflammatory factors and their expression. Quercetin can block the activity of Middle East respiratory syndrome coronavirus 3C-like (MERS-CoV 3C-like) protease. 22 It can also suppress the influenza virus, the respiratory syncytial virus, and other disease-causing agents. Kaempferol is an effective antiviral component23,24 that can strongly inhibit the synthesis of the messenger RNA (mRNA) of the M2 virus, thus, acting as an anti-influenza virus agent. It also has anti-inflammatory and antiviral effects, improves immunity, and exhibits other pharmacological effects. Quercetin combined with vitamins C and D can effectively treat COVID-19 and improve inflammation in patients. Quercetin might act by effectively binding to key proteins, including angiotensin-converting enzyme 2 (ACE2), and prevent the virus from entering the body. 25 The anti-inflammatory effect of quercetin and kaempferol might be due to their ability to block nuclear factor kappa B (NF-κB) activation, which leads to the suppression of mRNA and protein expression of Cyclooxygenage-2 (COX-2) and C-reactive protein (CRP). 24 Wogonin is a major effective component found in Scutellaria baicalensis and has antibacterial and antiviral effects, eliminating oxygen free radicals, promoting antioxidation, antipyresis, analgesia, and anti-inflammation, and regulating immune functions. 26 Isorhamnetin acts as a cough reliever and expectorant, and also has anti-inflammatory, antioxidative, and antiapoptotic effects. 27 In the herb–ingredient–target experiment, quercetin was found to be associated with AR and PRKACA, kaempferol with AR and PRKACA, wogonin with AR, PRKACA, and GSK3B, and isorhamnetin was significantly associated with AR and GSK3B (Figure 6F). We conducted molecular pairing experiments on these related molecules and receptors and found that the binding capacities of quercetin to AR and PRKAC were −4.56 and −6.80 kcal/mol, respectively. The binding capacities of kaempferol to AR and PRKACA were −6.72 and −4.83 kcal/mol, respectively. The binding capacities of wogonin to AR, PRKACA, and GSK3B were −5.65, −6.74, and −6.12 kcal/mol, respectively. The binding capacities of isorhamnetin to AR and GSK3B were 0.81 and 0.73 kcal/mol, respectively (Table 3). Kaempferol and wogonin can bind to AR, PRKACA, and GSK3B strongly. These are the key target genes of COVID-19 and diabetes, and, thus, have an important role in managing diabetes and COVID-19. Quercetin, kaempferol, and wogonin are the key active ingredients of QFPDD; their pharmacological effects are enhanced when used together. The treatment strategy using QFPDD also follows the “multi-components, multi-targets, multi-approaches, and synergy” concept of TCM treatment.

Herb–ingredient–target network of the key targets of Qing-Fei-Pai-Du Decoction (QFPDD). (A) Herb–ingredient–co-target regulatory network of QFPDD. (B–E) CytoHubba analysis of the TOP 10 components in the network; (B) MCC-TOP10, (C) Degree-TOP10, (D) Closeness-TOP10, and (E) Betweenness-TOP10. (F) Co-target genes associated with the TOP 10 components of the QFPDD network.
Binding Free Energy of Key Compounds in QFPDD.

Abbreviations: QFPDD, Qing-Fei-Pai-Du Decoction; AR, Androgen receptor; PRKACA, PKA Catalytic Subunit C-alpha; GSK3B, Glycogen synthase kinase-3 beta; GLU, glutamic; PHE, phenylalanine; ASN, asparagine; GLN, glutamine; LYS, lysine; LEU, leucine; ASP, aspartate; ILE, isoleucine; ARG, arginase.
The inflammatory response in COVID-19 cases increases after infection, and the levels of various plasma pro-inflammatory cytokines increase dramatically. 28 During the early stages of infection, TNF-α, IL-1β, and IL-8 increase rapidly with the aggravation of the disease. This leads to an imbalance of pro-inflammatory and anti-inflammatory signals, which can trigger the release and activation of inflammatory cytokines, such as IL-6 and IL-8, due to an uncontrolled inflammatory response.29,30 Based on the results of the PPI enrichment analysis of the co-target genes of QFPDD, COVID-19, and diabetes, we found that AR, PRKACA, and GSK3B were important target nodes for their therapeutic effects. These genes have a good regulatory effect on the inflammatory response in diabetic patients suffering from the COVID-19 infection, and the effect might be caused primarily by the positive regulation of inflammatory reaction, apoptosis, and other biological processes. They are also closely related to Toll-like receptors and the pathways associated with TNF.
Conclusions
To summarize, in this study, we used NP and MD techniques to conduct a preliminary analysis of potentially effective ingredients and critical targets of QFPDD for treating diabetes in conjunction with COVID-19. We found that AR, PRKACA, and GSK3B, which are important regulated factors, can directly be associated with the main effective components (quercetin, kaempferol, and wogonin) present in QFPDD. This might be the main way by which QFPDD alleviates the problems of diabetes and COVID-19. Because the mechanism of action of many components in Chinese medicine is very complex, the mechanism of coronavirus pathogenesis and NP remains unknown. The MD technology itself is limited, and the experimental results obtained from the MD may has some deviation. In future study, the results should be confirmed by clinical and experimental studies.
Materials and Methods
Screening TCMs and Their Active Components in QFPDD
In this study, we retrieved Flos farfarae, Rhizoma Alismatis, Rhizoma Atractylodis macrocephalae, Poria, Polyporus umbellatus, Rhizoma Pinelliae, Pogostemon cablin, Radix Glycyrrhizae, Asteris Radix et Rhizoma, Fructus Aurantii Immaturus, Herba Asari, Rhizoma Zingiberis Recens, Rhizoma Belamcandae, Rhizoma Dioscoreae, Herba Ephedrae, Radix Scutellariae, Ramulus Cinmomi, Pericarpium Citri Reticulatae, Radix Bupleuri, gypsum, and apricot kernel, present in QFPDD from the TCMSP database (https://old.tcmsp-e.com/tcmsp.php) (Table 4). There was no information on the active ingredients of gypsum and apricot kernel, which were not included in the analysis. Patchouli was replaced by Pogostemon cablin, and Pinellia ternata was replaced by Rhizoma Pinelliae because their main ingredients are identical. The main components of each Chinese medicine were selected based on the thresholds of DL ≥0.18 and oral OB ≥30%.
Plants in Qing-Fei-Pai-Du Decoction (QFPDD).

Action Target Collection and Network Construction
We obtained the targets of all Chinese medicines present in QFPDD from TCMSP. Then, the target names were converted using the UniProt database (https://www.uniprot.org/). The GeneCards (https://www.genecards.org/) database was used to retrieve disease targets with the keywords “COVID-19” and “diabetes,” and all these targets were analyzed using Venn diagrams (http://bioinformatics.psb.ugent.be/webtools/Venn/). The targets were intersected to obtain the co-targets for further analysis.
Analysis of the GO and KEGG Signaling Pathways of Common Target Proteins
The GO and KEGG analyses were performed using R software (3.6.3 version). The clusterProfiler package [version 3.14.3] was used for clustering and enrichment analyses and the org.hs.eg.db package [version 3.10.0] was used for ID translation.
Interaction Analysis of Common Target Proteins
The co-target genes of drugs and diseases were uploaded into the Metascape database (https://metascape.org/gp/index.html#/main/step1), and PPI and the key targets were analyzed. The protein interaction network map thus obtained was imported into the Cytoscape software for graphical analysis.
Co-Target-Active Ingredient-Drug Network Interaction Analysis
The drug and disease co-target genes were matched with the active ingredients. Importing the data into Cytoscape yielded a target-active ingredient-drug network diagram, and the key targets were analyzed using the CytoHubba plugin. The key target genes obtained from Metascape and CytoHubba were compared, and the key co-targets were identified. Next, we matched the co-targets with the Chinese medicine and imported the data into Cytoscape to obtain a key target-active ingredient-drug network diagram and analyzed the data using CytoHubba. By performing the analysis, we identified the final key target genes and active ingredients.
Docking of the Key Target Genes With the Main Active Ingredients
The crystal structure of the key target genes was retrieved from the protein data bank (https://www.rcsb.org/) database. PyMoL was then used to remove water molecules from the receptor. The pretreatment was performed by hydrogenation. The key active ingredients were retrieved using the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) for 2D file structures. Additionally, corrections were performed and the 3D structures were derived from the Chem3D software files. The key target gene was identified as a receptor, whereas the effective component was identified as a ligand. MD was performed using the AutoDock tool, followed by recording and analysis of the binding energy value. If the value was negative, the ligand and acceptor could bind freely.
Supplemental Material
sj-rar-1-npx-10.1177_1934578X221124769 - Supplemental material for Investigating the Potential Bioactive Components of Qing-Fei-Pai-Du Decoction Against COVID-19 in Diabetes/Diabetic Patients Based on Network Pharmacology and Molecular Docking
Supplemental material, sj-rar-1-npx-10.1177_1934578X221124769 for Investigating the Potential Bioactive Components of Qing-Fei-Pai-Du Decoction Against COVID-19 in Diabetes/Diabetic Patients Based on Network Pharmacology and Molecular Docking by Yang Liu and Huilian Huang in Natural Product Communications
Footnotes
Author Contributions
Yang Liu and Huilian Huang designed the study, Yang Liu wrote the manuscript, Huilian Huang collected and analyzed the data, Yang Liu and Huilian Huang reviewed and edited the manuscript. All authors have read and agreed to the published version of the manuscript.
Authors’ Note
All relevant data are within the manuscript and its Supporting Information files.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Huzhou Municipal Science and Technology Bureau, Natural Science Foundation of Zhejiang Province (grant number 2018GZ35, GF20H150001, LY17H160004).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
