Abstract
Coronavirus disease (COVID-19), caused by the novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), is a highly infectious viral disease. Clinical observations have shown that Qing-Fei-Da-Yuan (QFDY) granules have good anti-COVID-19 effects, but the underlying molecular mechanisms are unclear. In this study, we explored the potential mechanism of QFDY with regard to its anti-COVID-19 effect. We first screened the active chemical constituents of QFDY based on the pharmacodynamic activity parameters, followed by screening with the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform. The Uniprot database was used for querying the corresponding genes of the target, and Cyoscape 3.6.1 software was used to construct the network of herb-compound-target. Protein interaction analysis, target gene function enrichment analysis, and signal pathway analysis were performed via STRING database, Database for Annotation, Visualization, and Integrated Discovery, and KEGG Pathway database. Molecular docking was used to predict the binding capacity of the core compound with COVID-19 hydrolase 3CL and angiotensin converting enzyme 2 (ACE2). The results showed that a network of herb-compound-target was successfully constructed, with key targets involving PTGS2, HSP90AA1, CAMKK2, NCOA2, and ESR1. Major metabolic pathways affected were those in cancer, procancer, nonsmall cell lung cancer, and apoptosis. The core compounds, such as quercetin, luteolin, and naringenin, showed a strong binding ability with COVID-19 3CL hydrolase; compounds such as anemasaponin C and medicocarpin showed a strong binding ability with ACE2. Thus, it is predicted that QFDY has the characteristics for multicomponent, multitarget, and multichannel overall control. The mechanism of action of QFDY in the treatment of COVID-19 may be associated with the regulation of genes co-expressed with ACE2, the regulation of inflammation and immune-related signaling pathways, and the influence of COVID-19 3CL hydrolase and ACE2 binding ability.
In late December 2019, acute pneumonia caused by a novel coronavirus (severe acute respiratory syndrome coronavirus 2, SARS-CoV-2) was observed in Wuhan, Hubei Province, China, and has been raging for over 2 months. As of April 26, 2020, there have been reported 83 338 confirmed cases with 4642 deaths in China, 1 254 122 confirmed cases with 120 143 deaths in Europe, 1 032 278 confirmed cases with 58 534 deaths in North America, and 131 130 confirmed cases with 6010 deaths in South America. The source of the novel coronavirus remains to be identified; however, with the spread of COVID-19 to 209 countries and regions, the global pandemic is becoming extremely serious and the global risk level has been raised to the highest. The route of transmission of COVID-19 is mainly through respiratory droplets and aerosols, but the virus can also be transmitted through contact. It has the characteristics of rapid and widespread transmission, strong infectivity, and general susceptibility to all populations. COVID-19 patients with a mild infection have fever, fatigue, dry cough, and other symptoms; most severe patients have dyspnea, respiratory distress syndrome, or septic shock, and other symptoms. In addition, some COVID-19 patients have headache, nausea, and vomiting and other symptoms of the meridian system. There is growing evidence indicating that coronavirus infections are not limited to the respiratory tract. Coronaviruses may also invade the central nervous system and cause nervous system diseases. At present, no specific medicine is available for the treatment of this viral infection. 1 -3 Severe acute respiratory syndrome coronavirus 2 is a newly discovered β-coronavirus, the infectivity of which is closely associated with angiotensin converting enzyme 2 (ACE2) in human cells. The 3-dimensional structure of SARS-CoV-2 has been determined by Zihe et al at Shanghai University of Science and Technology (high resolution [2.1 Å, 0.21 nm] crystal structure of the 3CL hydrolase of the virus is shown in Figure 1), who have designed a potentially active compound N3. In the course of the diagnosis and treatment of this disease, chemical drugs such as arbidol, remdesivir, and chloroquine have shown good effects, but there is still not enough evidence to prove whether a certain chemical drug can completely eradicate COVID-19. 4,5 Thus, there is an urgent need to develop a vaccine and new drugs for treating COVID-19, to treat pneumonia caused by COVID-19, and to effectively control the spread of this pandemic.

Protein structure of COVID-19 3CL hydrolase and angiotensin converting enzyme 2. Crystal structure of (a) COVID-19 3CL hydrolase (Mpro) (protein data bank: 6LU7) and (b) angiotensin converting enzyme 2 (protein data bank: 1R42).
The efficacy and safety of traditional Chinese medicine (TCM) have been tested in the 5000 years of history of China. Since January 27, 2020, China’s National Health Commission and the State Administration of Traditional Chinese Medicine have included TCM in the treatment of coronavirus-induced pneumonia and have achieved remarkable results. Qing-Fei-Da-Yuan (QFDY) is a TCM formula developed by experts at the Hubei Provincial Hospital of Traditional Chinese Medicine under the emergency response mechanism. This formula has shown a significant effect against this new viral pneumonia during clinical observation. Currently, TCM laboratories are conducting related basic research, clinical observation, and molecular docking, and other experimental procedures for rapid screening of monomers and compound prescriptions of TCMs. 6,7 However, given the current urgency, the identification and mechanism of effective ingredients of TCM compound prescriptions are being explored according to the clinical feedback in the treatment of COVID-19-induced pneumonia, and rapid optimization and secondary development of TCM formulations may be an effective way to deal with this clinical emergency. Traditional Chinese medicine combined with chemical drugs may be an effective means to treat COVID-19 and in turn control the pandemic.
Qing-Fei-Da-Yuan is made from the addition and subtraction of Chaihu Xianxiong Decoction and Dayuanyin, and contains 13 Chinese herbal medicines, including Radix Bupleuri (Chaihu), Radix Scutellariae (Huangqin), Rhizoma Pinelliae (Banxia), Radix Codonopsis (Dangshen), Fructus Trichosanthis (Gualou), Semen Arecae (Binglang), Fructus Tsaoko (Caoguo), Cortex Magnoliae Officinalis (Houpo), Rhizoma Anemarrhenae (Zhimu), Radix Paeoniae Alba (Baishao), Radix Glycyrrhizae (Gancao), Pericarpium Citri Reticulatae (Chenpi), and Rhizoma Polygoni Cuspidati (Huzhang). Qing-Fei-Da-Yuan, also known as “Pneumonia No. 1,” is one of the four TCM prescriptions officially recommended by the Hubei Provincial Epidemic Prevention and Control Command; it has the effect of detoxification and expelling dampness, and has good curative effects on fever, cold, heat, cough, chest tightness, and fatigue. As of February 22, 2020, 517 COVID-19 patients treated at Hubei Provincial Hospital of Traditional Chinese Medicine have been cured and 158 patients have been discharged. Most of these patients were treated with QFDY, with a total efficacy of over 90%. In a multicenter clinical study led by Professor Ba Yuanming, it was found that the therapeutic effect of “Pneumonia No. 1” on the new COVID-19-induced pneumonia is specifically manifested in the following 3 aspects: (1) obvious effect in relieving symptoms: improvement in the main symptoms including fever, cough, and fatigue, in addition to other symptoms such as chills, stuffy nose, runny nose, sneezing, itching, sore throat, chest tightness, anorexia, vomiting, nausea, bloating, and diarrhea; (2) significant changes in clinical observations: the red or crimson color on the patient’s tongue gradually fades; white greasy moss, thick greasy moss, and yellow greasy moss become thinner with an improvement rate of over 65%; (3) core indicators: the average fever duration is 3 days, the average time taken for nucleic acids to test negative is 8 days, and 92% of patients show improved lung imaging indicators and relieved lung inflammation. These results show that “Pneumonia No. 1” combined with conventional Western medicine treatment has a good therapeutic effect on COVID-19-induced pneumonia, can quickly stabilize the condition, improve the clinical symptoms of patients, and relieve lung inflammation; further, it is safe and reliable for use. Thus, “Pneumonia No. 1” can be recommended as an important reference prescription for treating new COVID-19-induced pneumonia. 8
Network pharmacology was introduced by Hopkins in 2008 using a visualization network software and a variety of algorithms combined with integrated analysis of existing related databases. This is a big data integration method based on a large number of database resources and statistical algorithms used to observe the synergistic effect of multiple components, targets, and mechanisms of diseases, which break the previous single component/single target/disease concept; it also follows the TCM theory of multiple components/multiple targets/more complicated disease. Network pharmacology is a novel field based on the development of an “active component-target-disease” interaction network multidisciplinary fusion theory, and provides a novel way for modern medical interpretation of the responsible system of TCM. 9 -11 Molecular docking is a drug design method used for directly analyzing the characteristics of receptors and determining the interactions between receptors and drug molecules. It is a theoretical simulation method that mainly studies the interaction between molecules (such as ligands and receptors) and predicts their binding mode and affinity. In recent years, subdocking has become an important technology in computer-aided drug research. 12 In this study, through network pharmacological analysis of QFDY, we report an effective TCM granule against COVID-19; we further predict its core active compounds and speculate the biological pathway. The molecular docking technique was used to predict the activity of the key compounds in the granules against closely related targets of COVID-19, and the signal pathway analysis was performed to predict the mechanism of QFDY in the treatment of COVID-19.
Materials and Methods
Collection of Active Ingredients
Figure 2 shows a workflow of a network pharmacological study of QFDY in the treatment of COVID-19. Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, http://tcmspw.com/index.php, Version 2.3) is a systematic pharmacology platform that can identify the relationship between Chinese herbal medicines, targets, and diseases, and is used for new drug discovery. 13 The platform also provides pharmacokinetic properties of natural compounds such as oral bioavailability (OB) and drug-likeness (DL). The active ingredients of QFDY were obtained from the TCMSP.

Workflow of the network pharmacological study of Qing-Fei-Da-Yuan in the treatment of COVID-19.
Screening of Active Compounds and Target Proteins
Oral bioavailability is an important pharmacokinetic parameter in drug absorption, distribution, metabolism, and excretion (ADME). It indicates the speed and degree at which the active components or active groups of the oral drug are absorbed into the systemic circulation. Further, DL refers to the similarity of a compound with a known drug, and the potential of this class of compounds to become drugs. Higher OB and DL values usually indicate better biological activity of the drug. 14,15 Therefore, we screened each of the retrieved compounds based on their OB and DL properties, with OB ≥ 30% and DL ≥ 0.18. Potential active ingredients were obtained after screening. At the same time, the target information related to these compounds was downloaded from the TCMSP database. The target related to viral pneumonia was obtained by searching “Viral pneumonia” as the keyword in the Genecard database (https://www.Genecards.org/). An intersection with the predicted target of the obtained compound was used to obtain the potential target of QFDY to treat viral pneumonia.
Network Construction
Gene names were standardized through the Uniprot (https://www.uniprot.org/) database. The herb-compound-target network was constructed using Cytoscape3.6.1 software (https://cytoscape.org/download.html). Topological analysis of the constructed pharmacological network was performed to obtain the betweenness centrality, closeness centrality, and degree of each node.
Construction of Protein-Protein Interaction Network
The STRING database (https://string-db.org/) collects, scores, and integrates all publicly available protein-protein interaction (PPI) information, including direct (physical) and indirect (functional) interactions. The potential action targets of QFDY in the treatment of viral pneumonia were recorded on the STRING website (https://string-db.org/cgi/input.pl), and the PPI network of the potential action targets was obtained. Simultaneously, the genes interacting with ACE2 were found through the STRING database.
Gene Ontology and Pathway Enrichment
The Database for Annotation, Visualization, and Integrated Discovery (DAVID, https://david.ncifcrf.gov/) provides systematic, comprehensive biofunctional annotations for a large number of genes or proteins, enabling the identification of the most significantly enriched biological annotations. We introduced disease-related genes into the DAVID, and set the parameters as follows: “Select Identifier” was set to “OFFICIAL GENE SYMBOL,” “List Type” was set to “Gene List,” and the species was defined as “Homo sapiens” in the “Background” and “List.” Gene ontology function and KEGG pathway were run on the gene targets. The most important biological process (BP), cellular component (CC), molecular function (MF), and KEGG pathway results were screened and stored, with a threshold of P < .05. An online drawing website OmishareTools (http://www.omicshare.com/tools/index.php/) was used to visualize the results.
Molecular Docking
The core components selected from the active ingredients of QFDY were molecularly docked with COVID-19 coronavirus 3CL hydrolase and ACE2 (protein data bank [PDB]: 1r42, crystal structure shown in Figure 1). Subsequently, the structures of related proteins were downloaded from the RSCB PDB database (https://www.rcsb.org/) and compound structures were downloaded from the Pubchem database (https://pubchem.ncbi.nlm.nih.gov/). DiscoveryStudio software was used for solvent molecules and ligand removal, hydrogenation, electron addition, and other operations. Molecular docking was then performed (Discovery Studio 2016; BIOVIA; San Diego, United States). A LiDock Score greater than 100 indicates potential activity.
Results
Compounds Collection
The compounds in Chaihu, Huangqin, Banxia, Dangshen, Gualou, Binglang, Caoguo, Houpo, Zhimu, Baishao, Gancao, Chenpi, and Huzhang were searched through the TCMSP database. With OB ≥ 30% and DL ≥ 0.18, 223 active compounds were selected, of which 17 were from Chaihu, 36 from Huangqin, 13 from Banxia, 21 from Dangshen, 11 from Gualou, and 8 from Binglang, 8 from Caoguo, 2 from Houpo, 15 from Zhimu, 13 from Baishao, 92 from Gancao, 5 from Chenpi, and 10 from Huzhang. There were 17 ingredients contained in more than 2 herbs. The basic information of some compounds in QFDY is shown in Table 1. All the collected compound information is shown in Supplemental Table S1.
Basic Information of Some Components in Qing-Fei-Da-Yuan.

DL, drug-likeness; MW, molecular weight; OB, oral bioavailability.
Attachments—Composition of QFDY per day: Radix Bupleuri (Chaihu): 20 g; Radix Scutellariae (Huangqin): 10 g; Rhizoma Pinelliae (Banxia): 10 g; Radix Codonopsis (Dangshen): 15 g; Fructus Trichosanthis (Gualou): 10 g; Semen Arecae (Binglang): 10 g; Fructus Tsaoko (Caoguo): 15 g; Cortex Magnoliae Officinalis (Houpo): 15 g; Rhizoma Anemarrhenae (Zhimu): 10 g; Radix Paeoniae Alba (Baishao): 10 g; Radix Glycyrrhizae (Gancao): 10 g; Pericarpium Citri Reticulatae (Chenpi): 10 g; Rhizoma Polygoni Cuspidati (Huzhang): 10 g.
Topology Analysis Network
The constructed network included 496 nodes (13 herb nodes, 195 compound nodes, and 288 target nodes); 28 of the 223 compounds were not included in the network construction (Figure 3(a)). There are 2913 edges between compounds and targets, and each compound interacted with 14.94 targets on an average; further, each target interacted with 10.11 compounds on an average; thus, each compound in QFDY interacted with multiple targets. At the same time, we observed a phenomenon where different compounds interacted with the same target, reflecting the mechanism of interaction between multicomponents and multitargets in TCM. With regard to compounds, 30.5% compounds were observed to have ≥20 targets and 20% have ≥30 targets. By analyzing the betweenness centrality, closeness centrality, and degree between the compound and the target, we found that the top 10 compounds were MOL000098: quercetin, MOL000422: kaempferol, MOL000449: stigmasterol, MOL000358: β-sitosterol, MOL000006: luteolin, MOL003896: 7-methoxy-2-methyl isoflavone, MOL004328: naringenin, MOL002714: baicalein, MOL000354: isorhamnetin, and MOL000173: wogonin, and these can generate 616, 252, 155, 115, 114, 86, 74, 74, 74, and 45 edges, respectively (Table 2). The top 10 targets were found to be PTGS2, HSP90AA1, PTGS1, CAMKK2, NCOA2, AR, ESR1, PPARG, NOS2, and PRSS1, which can interact with 168, 130, 114, 109, 109, 105, 10, 95, 94, and 92 compounds, respectively (Table 3).

Network constructed by Cytoscape. (a) Herbs-compound-target network; (b) herbs-compound-viral pneumonia-related target network; and (c) Wayne for compound predicted target and viral pneumonia target illustration. Purple represents herbs, green represents compounds, and light blue represents target genes.
Top 10 Compounds Information in the Herbs-Compound-Target Network.

Top 10 Targets Information in the Herbs-Compound-Target Network.

In addition, we screened the top 100 genetic targets related to viral pneumonia from the Genecard database and intersected them with the possible targets of QFDY to obtain 24 common targets (Figure 3(c)). We used Cytoscape3.6.1 software to construct a herb-compound-common target network (Figure 3(b)). From the network, 15 compounds related to 24 targets, but these 15 compounds were obtained from only 10 herbs, of which the contributions of Gancao, Huangqin, Huzhang, and Baishao were the most.
Protein-Protein Interaction Network
The obtained potential targets were analyzed using the STRING website, and the PPI network was obtained (Figure 4(a)). We also constructed a PPI network for the top 10 genes in the herb-compound-common target network (Figure 4(b)). The network diagram shows the close interactions between genes. We further found interactions among AGT, AGTR1, AGTR2, DPP4, MET1A, MET1B, MME, PRCP, XPNPEP2, REN, and ACE2 (Figure 4(c)), whereas the target of isorhamnetin, areapillin, quercetin, kaempferol, panicolin, rivularin, acacetin, baicalein, wogonin, luteolin, and 70 other compounds was DPP4. It is predicted that the mechanism of action of QFDY in the treatment of COVID-19 may be related to the regulation of genes co-expressed with ACE2.

Protein-protein interaction network. (a) Interaction of potential targets of Qing-Fei-Da-Yuan for the treatment of viral pneumonia; (b) interaction of top 10 targets in the herbs-compounds-targets network; and (c) genes that interact with angiotensin converting enzyme 2.
Module Analysis
Gene ontology functional enrichment analysis in DAVID yielded 516 GO entries (P < .05, count > 10), including 402 BP entries, 62 cell composition (CC) entries, and 52 MF entries, accounting for 77.9%, 12.1%, and 10.0%, respectively (Figure 5).
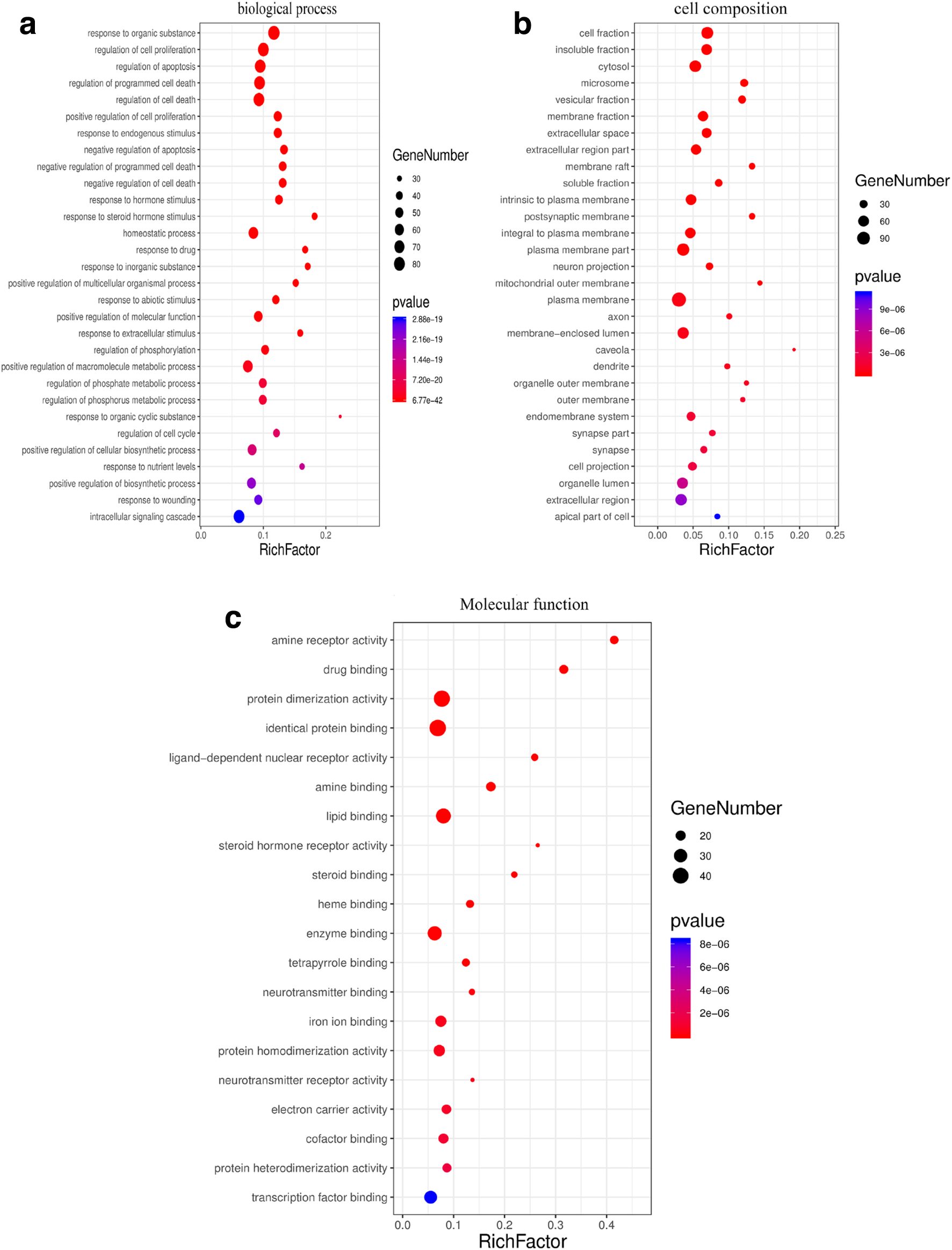
The enrichment analysis in biological processes, cellular components and molecular functions of the identified target proteins by Database for Annotation, Visualization, and Integrated Discovery. The top 30 significantly enriched terms in (a) biological process, (b) cellular component (CC), and (c) molecular function.
The KEGG pathway enrichment and screening yielded 47 signaling pathways (P < .05, count > 10), which are involved in various other signaling pathways, such as those in cancer, prostate cancer, nonsmall cell lung cancer, apoptosis, and small cell lung cancer. Among them, pathways in cancer involved 65 genes including E2F1, E2F2, PPARD, PTGS2, MMP9, PPARG, FASLG, PTEN, MMP2, MMP1, TGFB1, and AKT1. The prostate cancer pathway involved 32 genes including ERBB2, NFKBIA, PTEN, AKT1, CASP9, and BCL2. Nonsmall cell lung cancer involved 21 genes including PRKCA, EGFR, PIK3CG, RXRB, RXRA, ERBB2, and TP53. Further, the apoptosis pathway involved 20 genes including TNF, RELA, TP53, NFKBIA, and FASLG (Figure 6). We predicted that some compounds in QFDY could act on E2F1, E2F2, PTEN, AKT1, TNF, RELA, TP53, and other genes to regulate nonsmall cell lung cancer, apoptosis, small cell lung cancer, and other signaling pathways, to achieve an anti-COVID-19 effect. At the same time, QFDY may regulate multiple signaling pathways related to inflammation and immunity, and achieve the purpose of restoring immune homeostasis and helping the body return to normal.

Bubble chart of the results of KEGG pathway enrichment analysis of the identified target proteins by Database for Annotation, Visualization, and Integrated Discovery.
Molecular Docking Display
It is a general belief that the lower the energy of ligand and receptor binding, the higher the LiDock Score, the greater the possibility of interaction, and the stronger the potential activity of the compound. In this study, molecular docking was performed on core compounds in QFDY; remdesivir, chloroquine, and arbidol, which have shown positive effects for treating COVID-19; and N3 designed by Professor Rao’s group. The results showed that the LiDock Score of the core active compounds in QFDY with COVID-19 3CL hydrolase was over 100. Although the potential activity of most compounds was not as good as that of remdesivir and N3, it was almost the same as that of chloroquine and arbidol. In addition, through molecular docking, we found that compounds such as medicarcarpin and anemarsaponin C can stably bind to ACE2 (with activities much higher than those of chloroquine and arbidol). The details are shown in Table 4 and Figures 7 and 8.
Results of Molecular Docking.

ACE2, angiotensin converting enzyme 2; MW, molecular weight.

Results of molecular docking of each compound with COVID-19 3CL hydrolase. (a) Arbidol-6LU7, (b) chloroquine-6LU7, (c) N3-6LU7, (d) remdesivir-6LU7, (e) quercetin-6LU7, (f) luteolin-6LU7, and (g) naringenin-6LU7.

Results of molecular docking of each compound with angiotensin converting enzyme 2. (a) Chloroquine-1R42, (b) N3-1R42, (c) remdesivir-1R42, (d) anemarsaponin C-1R42, and (e) medicocarpin-1R42.
Discussion
Network pharmacology combines network biology with polypharmacology, based on the poor efficacy of highly selective single-target drugs. Through network pharmacology, we can directly identify drugs and disease targets from a vast amount of data and understand the mechanisms and pathways between them, making it an effective method. Nowadays, the application spectrum of network pharmacology is expanding and it includes exploring the basic pharmacological effects of drugs on diseases and their mechanisms, analyzing TCM theory, and studying TCM application. 16,17 However, network pharmacology is still in the early stages of development and has many problems. First, appropriate computing software has not been developed for network pharmacology. Further, systematic screening, integration, and processing of data is another problem. The data on various drugs, genes, proteins, etc., are not comprehensive. The databases on Chinese herbal medicines, proprietary Chinese medicines, and TCM are incomplete, and their accuracy and integrity have not been verified. Network pharmacology uses computer network screening to achieve target selection, and many results do not have enough experimental data support. The known compound targets do not completely correspond to the complex mechanism of TCM formulae. 18,19 The prediction of the potential side effects of drugs is an important part of the drug development process. Most drugs come with some side effects and, apart from lack of efficacy, these are the leading cause of attrition in clinical trials of new drugs. Thus, it is clear that although the current methods already show great promise, they are still constantly developing. 20 Continuously developing network pharmacology technology surely has huge potential.
Molecular docking is the application of mathematical, biological, and computer-based models to predict the binding affinity of small molecules toward a particular receptor. In pharmacology, molecular docking can predict the affinity of drugs within the binding site of the target of interest. Further, with the help of docking study, thousands of molecules can be evaluated for potential efficacy and safety at a small cost in a short time span. 21 Although docking technology is of great help in pharmacology, certain limitations remain. It is well known that individual software has its own shortcomings. Further, software cannot provide the same information with the same level of reliability as a measurement. In silico models might not perform well if a predicted chemical is beyond the chemical space within which the software is developed. First, the accuracy of docking may be affected by the use of a protein structure. In most cases, protein structures available in the PDB are complexed with an internal ligand. To use these proteins, researchers normally delete the ligand from the protein and dock the molecule. However, this process can affect the accuracy of docking. Second, the environment of the binding site may affect the accuracy of results. For example, a drug molecule has to bind with an intracellular receptor to show its response. In this case, even docking results show high binding under an in silico environment but, when these molecules are tested in labs, they show false results. Third, a different pH of the protein may affect the outcomes by altering the binding of the ligand with receptors. In the process of drug discovery and pharmacological research, discovery of active compounds has always been challenging. 22 With the help of docking technology, an increasing number of compounds will be screened at a faster pace, which will greatly save costs. We hope to accelerate the development cycle of future forecasting applications, to aid an increasing number of researchers to appropriately and reasonably use these technologies.
The novel coronavirus disease, named “COVID-19” by the World Health Organization on February 11, 2020, belongs to the category of epidemic diseases in TCM. In Chinese history, Chinese medicine has combated a variety of diseases (including plague, malaria, and cholera), showing extremely positive effects. Traditional Chinese medicine also proved to be effective against SARS in 2003. 23 Two months ago, an outbreak of COVID-19 occurred. 24 However, no specific drugs have yet been developed for the treatment of COVID-19-induced pneumonia, and the existing chemical drugs can only alleviate some symptoms. Recent studies have reported that remdesivir is effective, but the sample sizes were too small; accordingly, the drug is still at the stage of research and development and cannot be used clinically in the near future. 25 It is encouraging that after careful discussion by the Chinese government, the intervention of Chinese medicine in the prevention and treatment of this epidemic has achieved good results. There are sufficient recent arguments in the literature to explain the feasibility of using TCM in the prevention of epidemic diseases. 26 Traditional Chinese medicine not only can prevent epidemic diseases but also has a good therapeutic effect on diagnosed patients. In the process of interventional treatment of COVID-19 with TCM, the use of TCM prescriptions for several groups of patients has shown significant curative effects. Qing-Fei-Da-Yuan is a TCM granule preparation based on clinical observation.
We used the network pharmacology method and molecular docking technique to screen the core compounds in QFDY and predicted its possible action mechanism in COVID-19 treatment. The results showed that the top 10 compounds with the highest comprehensive score in QFDY were flavonoids. Compared with the clinically recommended chemicals, the core compounds in this granule have a good binding strength with COVID-19 3CL hydrolase; in particular, quercetin, luteolin, naringenin, and some other compounds show very similar binding abilities to those of arbidol and chloroquine. According to reports, both SARS-CoV-2 and SARS-CoV virus infection pathways cause the virus to invade the body through its expressed S-protein and angiotensin-converting enzyme in the human body. 27,28 Therefore, we conducted molecular docking with ACE2 on the core compounds in QFDY, and the results showed that compounds such as anemarsaponin C and medicocarpin were strongly bound to ACE2, especially anemarsaponin C. It is predicted that some core compounds in QFDY can directly act on SARS-CoV-2, making it either lose or reduce its virus activity; in addition, some compounds can act on ACE2, which makes it difficult for the virus to bind with ACE2, thus reducing the infection rate of the virus within the human body.
The pathways most relevant to the lungs obtained by KEGG analysis are nonsmall cell lung cancer, small cell lung cancer, and pathways in cancer, all of which involve E2F1, E2F2, RXRB, RXRA, TP53, CDK4, AKT1, and CCND1 genes, and these genes are mostly targets of quercetin, luteolin, and some other compounds. However, whether the core compounds of QFDY regulate nonsmall cell lung cancer and small cell lung cancer pathways by acting on these gene targets to play the anti-COVID-19 role remains to be studied further. Nevertheless, we found that among the targets of the QFDY core compounds, there are at least 24 related to common viral pneumonia (most of these targets include the action targets of quercetin, luteolin, kaempferol, and baicalein). 29,30 These genes contain AKT1 and TP53, which predict that QFDY can regulate nonsmall cell lung cancer, small cell lung cancer, and other signaling pathways by acting on AKT1 and TP53 genes, subsequently playing a role in fighting viral pneumonia.
Traditional Chinese medicine plays an important role in COVID-19 treatment. However, because of too many ingredients in TCM and the low content of some effective medicinal substances, the short-term efficacy of TCM is limited, which also limits its global acceptance. With the current research on TCM, it is difficult to determine a manner in which TCM can be adapted for local conditions, find TCM prescriptions for symptomatic treatment, and to discover the specific active substances to form a reasonable combination of multicomponents. At present, the best solution for this stubborn epidemic is early protection, early diagnosis, early isolation, and early treatment. Rational use of TCM and COVID-19 treatment with a combination of TCM and Western medicine may be the most effective measure, and may also become an effective method to control the epidemic quickly.
In this study, we evaluated the chemical constituents, action targets, and the binding ability of the core compounds of QFDY to COVID-19 3CL hydrolase and ACE2 in QFDY via network pharmacology and molecular docking. From the research results, it can be seen that QFDY can exert its curative effect through the synergistic effect of multicomponents, multitargets, and multipathways in the treatment of diseases. Because of the limitations of network pharmacology and molecular docking, experimental studies can be performed with regard to material basis, pharmacodynamics, metabonomics, and pathway verification at later stages to provide the theoretical and experimental basis for the use of QFDY in the treatment of COVID-19. In addition, future studies can be conducted with regard to drug development and the use of QFDY as a reference for dealing with invasions of other stubborn viruses.
Conclusion
Qing-Fei-Da-Yuan can regulate multiple signaling pathways and play a role in treating COVID-19; further, it can be very effective in the rehabilitation of patients with COVID-19, so that a higher number of individuals who are aware of and have used QFDY to treat COVID-19-induced pneumonia may benefit other patients. In summary, we screened the active ingredients of QFDY and explored its complex pharmacological mechanisms for the rehabilitation therapy of COVID-19 through network pharmacology, referencing the development of new drugs and elucidation of the mechanisms of QFDY against COVID-19.
Supplemental Material
Table S1 - Supplemental material for Network Pharmacology Integrated Molecular Docking Reveals the Anti-COVID-19 Mechanism of Qing-Fei-Da-Yuan Granules
Supplemental material, Table S1, for Network Pharmacology Integrated Molecular Docking Reveals the Anti-COVID-19 Mechanism of Qing-Fei-Da-Yuan Granules by Zongchao Hong, Xueyun Duan, Songtao Wu, Yang Yanfang and Hezhen Wu in Natural Product Communications
Supplemental Material
Table S2 - Supplemental material for Network Pharmacology Integrated Molecular Docking Reveals the Anti-COVID-19 Mechanism of Qing-Fei-Da-Yuan Granules
Supplemental material, Table S2, for Network Pharmacology Integrated Molecular Docking Reveals the Anti-COVID-19 Mechanism of Qing-Fei-Da-Yuan Granules by Zongchao Hong, Xueyun Duan, Songtao Wu, Yang Yanfang and Hezhen Wu in Natural Product Communications
Supplemental Material
Table S3 - Supplemental material for Network Pharmacology Integrated Molecular Docking Reveals the Anti-COVID-19 Mechanism of Qing-Fei-Da-Yuan Granules
Supplemental material, Table S3, for Network Pharmacology Integrated Molecular Docking Reveals the Anti-COVID-19 Mechanism of Qing-Fei-Da-Yuan Granules by Zongchao Hong, Xueyun Duan, Songtao Wu, Yang Yanfang and Hezhen Wu in Natural Product Communications
Footnotes
Availability of Data and Materials
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (No. 31570343).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
