Abstract
Objectives
In this study, the efficacy of and mechanism of Qingrekasen Granules (QRKSG) is evaluated by metabolomics and transcriptomics using adriamycin (ADR)-induced nephrotic syndrome (NS) in rat model.
Methods
The model, benazepril, and QRKSG group received a single injection of 6.5 mg/kg ADR via the tail vein of the rats. The untreated group received an equal saline injection. The administration of drugs by gavage began after completing the modeling for one week. Benazepril was given at 0.9 mg/kg/d to the benazepril group and QRKSG was given at 1.62 g/kg/d to the QRKSG group. During gavage, 24 h urine was collected weekly. After four weeks of gavage, rats were anesthetized, and we collected the serum, feces, and kidney samples. The protective effect of QRKSG on NS was assessed by the detection of proteins in the urine at 24 h and serum biochemical indexes, as well as histopathological observation, TUNEL assay, and Western Blot of kidney samples. Moreover, gas chromatography-mass spectrometry (GC-MS) metabolomics sequencing of kidney metabolites helped investigate the significant differential metabolites produced by QRKSG that are implicated in ameliorating the pathological damage to the kidneys of NS rats. Subsequently, the significant targets of QRKSG that had an impact on the action of drugs were investigated by a transcriptome sequencing analysis. The correlation between both sets was also investigated. In addition, the effect of QRKSG on the gut microbiota of NS rats was investigated. Finally, the core targets were validated by molecular docking.
Results
The results indicated that QRKSG possess significant renoprotective effects in NS rats by reversing the abnormal urinary protein content and serum biochemical disorders, as well as improving renal pathological damage. Multiple genes and metabolites were shown to be back-regulated after QRKSG delivery, based on further multi-omics studies. The integrated metabolomics and transcriptomics analysis showed that QRKSG alleviated NS mainly by regulating amino acid metabolic pathways. Gut microbiota analysis demonstrated that QRKSG could alleviate NS by improving gut microbiota. The molecular docking results showed good binding ability of the QRKSG active ingredient to the core target, and among them, the ingredients with the best docking effect were mainly flavonoids and phenolic acid ingredients. The combined metabolomics and transcriptomics study findings were validated through a TUNEL assay and a Western Blot analysis.
Conclusion
We have concluded that ADR-induced NS in rats can be treated with QRKSG. The mechanisms used by QRKSG to reduce the symptoms of NS are through the multi-component, multi-target, and multi-pathway therapeutic modulation of inflammation, oxidative stress, energy homeostasis, and apoptosis. Additionally, QRKSG could alleviate NS by regulating gut microbiota.
Keywords
Introduction
A typical disorder of the urinary system is called nephrotic syndrome (NS), which causes edema, proteinuria, hypoproteinemia, and hyperlipidemia. Kidney failure caused by the ongoing progress of NS may even require a kidney transplant to reach a clinical cure. 1 However, the mechanism of NS is complicated and may be related to inflammation.2,3 The occurrence and development of NS are usually significantly correlated with oxidative stress and apoptosis.4,5 Apoptosis leads to the loss of resident renal cells, followed by the occurrence of kidney injury. 6 Moreover, NS is easily induced by the decrease in plasma albumin (ALB) in the blood due to the increased permeability of glomerular-filtered macromolecular proteins. 7 Phosphoinositide 3-kinase-protein kinase B (PI3K-Akt) and its downstream signaling pathway regulate various signaling pathways, such as those that regulate oxidative stress, inflammatory response, and apoptosis of renal podocytes caused by NS.8,9 The common medical treatment for NS is hormone therapy, but patients are prone to develop drug resistance and other side effects, eventually developing renal failure anyway. 10 Therefore, the pathogenesis of NS should be explored to find more effective treatment strategies. Modern pharmacology study shows that traditional medicine has protective effects on kidney and other tissues. It has been shown that hesperidin, as a citrus flavonoid, has protective effects against carbon monoxide-induced cardiac injury. 11 Hesperidin also exerts a protective effect against dust storm particulate matter-induced kidney injury in rats. 12 Importantly, traditional Chinese medicine effectively protects renal function and delays renal failure in NS patients. 13 It also reduces the recurrence rate of this disease. 14
Chicory (Cichorium intybus L.) is a perennial herb of the Asteraceae family widely distributed in Europe, Asia, and North Africa. It facilitates functions such as diuresis, anti-inflammation, digestion, strengthening the heart, and protecting the liver and kidney. 15 It also treats a variety of diseases, including malaria, liver disease, diabetes, and kidney disease.16-19 Based on the available data, cyclooxygenase 2 protein expression and prostaglandin E2 synthesis are inhibited by chicory extract, which therefore exerts anti-inflammatory effects, 20 and alleviates monosodium urate-induced ankle edema. 21 Some studies 22 revealed that chicory extract reduces serum creatinine and urea level and alleviates renal injury in NS patients. Qingrekasen Granules (QRKSG) represent a remedy obtained from chicory and approved by the State Food and Drug Administration based on Approval Number Z65020172 and included in the 2020 Edition of the People's Republic of China Pharmacopoeia.
Metabolomics consists of the analysis of metabolites extracted from biological fluids, cells, and tissues, which provides relevant information on disease pathogenesis and pharmacological effects. 23 Transcriptomics consists of the analysis of the genome of an organism to reveal the molecular mechanisms involved in a specific biological process. 24 Therefore, the aim of this study was to use a multi-omics strategy that integrates metabolomics and transcriptomics to explore the mechanism of action of QRKSG in the reduction of NS. The research focused on the evaluation of biological processes such as energy homeostasis regulation, amino acid metabolism, apoptosis, as well as differentially expressed genes (DEGs) and differential metabolites between the model and QRKSG group. In addition, it has been shown that kidney disease can cause changes in the gut microbiota, which can be involved in the body's material and energy metabolism. 25 Therefore, our study could be a reference for clinical usage and approval of QRKSG to treat NS.
Materials and Methods
Materials and Reagents
The QRKSG was purchased from Xinjiang Uygur Pharmaceutical Co., Ltd (NO.1807521, Xinjiang, China). Benazepril was purchased from Novartis (No. 2007, Beijing, China). Adriamycin (ADR) was purchased from Beijing Solarbio Science & Technology Co., Ltd (D8740, Beijing, China).
Ultra Performance Liquid Chromatography-Tandem Mass Spectrometry (UPLC-MS/MS) Analysis of QRKSG
The qualitative analysis of QRKSG used integrated UPLC (Nexera X2, Shimadzu, Japan) and MS/MS (4500 QTRAP, Applied Biosystems, MA, United States). The analytical conditions were: column from Agilent, 181.8 µm, 2.1 mm*100 mm; mobile phases from liquids A and B (Solution A: pure water containing 0.1% formic acid; Solution B: acetonitrile containing 0.1% formic acid); 0.35 mL/min flow rate, a 40 °C column, and a 4 µL injection volume.
In Vivo Experiments
We obtained 24 male Sprague Dawley (SD) rats from Liaoning Changsheng Biotechnology Co., Ltd (SCXK-2020-0001, Liaoning, China) at 6 weeks and weighed in the range of 180 and 200 g. All SD rats were housed in an SPF-grade animal room at the Hubei University of Chinese Medicine with a 50% humidity level, an average temperature of 25 °C, and a 12-h cycle of light and darkness. All the rats had unrestricted access to food and water. We followed the study methodology of the Hubei University of Chinese Medicine's Animal Ethics Code (HBUCMS202206002). Based on the random number table approach, the rats were split into 4 groups of 6 rats each: the control group, model group, benazepril group, and QRKSG group.
After one week of acclimatization, the NS modeling was induced by tail vein injection of 6.5 mg/kg ADR 26 in the model, benazepril, and QRKSG group, and the same volume of saline was injected into the tail vein of the control group. The administration of benazepril, QRKSG, and saline was at 1 week after ADR administration; the benazepril group received a clinically equivalent dose of 0.9 mg/kg/d, the QRKSG group received a clinically equivalent dose of 1.62 g/kg/d, 27 and the control and model group received an equal volume of saline by gavage. Gastric gavage was performed once daily for four weeks. The 24 h urine was collected once a week during the entire experiment. Urine was collected using a urine collection container, mineral oil was added to the container to avoid evaporation, the collected urine was centrifuged and the supernatant was obtained. Post-trial, the rats were put to sleep to obtain their blood, colon contents, and kidney samples. The obtained blood samples were centrifuged at 3000 rpm for 10 min after incubating at 4 °C for 30 min to separate the serum for examination. The intestinal contents were stored at −80 °C for subsequent gut microbiota analysis. For subsequent Western Blot or histological investigation, kidney samples were sliced and promptly frozen at −80 °C or fixed in 4% paraformaldehyde.
Serum Biochemistry Assay and 24-h Urine Protein Test
The serum concentrations of ALB, total triglycerides (TG), creatinine (Cr), cholesterol (CHO), total protein (TP), and urea nitrogen (BUN) were determined with a biochemical analyzer (Chemray240, Rayto, Guangdong, China). The urine collected at 24 h was centrifuged and the total protein concentration was determined using the BCA protein assay kit (Beyotime, P0012, Shanghai, China) and enzyme marker (Epoch, Bio Tek, VT, United States) according to the manufacturer's instructions.
Histopathological Analysis and TUNEL Assay
Hematoxylin and eosin (H&E) helped stain paraffin slices of kidney samples, and were observed under a light microscope (BH-2, Olympus, Japan) for analysis. The manufacturer's instructions (Servicebio, China) indicated that the TUNEL labeling could identify renal apoptosis, and the green fluorescence represented the apoptotic renal cells.
Metabolomics Analysis by GC-MS
Kidney Sample Preparation
After freezing at −80 °C, kidney tissue samples were defrosted at ambient temperature, weighed at 50 mg in a centrifuge tube, added 1 mL of methanol, and vortexed for 1 min. We obtained the metabolite-containing supernatant after centrifuging and separating 600 μL of kidney tissue homogenate for 10 min while keeping it in a cold bath for 15 min. Centrifugation helped concentrate the supernatant, which was dried through evaporation. Afterward, 40 μL of methoxypyridine mixture and 10 μL of D27-myristate were mixed and reacted for 90 min at 30 °C while shaking. To derivatize and end the derivatization reaction, 80 μL N-Methyltrimethylsilyl trifluoroacetamide + 1% Methyl Chlorosilane (MSTFA + 1% TMCS) (Sigma-Aldrich, MO, United States) and 20 μL hexane were added. After centrifugation, 1 μL of the supernatant was removed and kept in a micro-sample tube for analysis.
GC-MS Data Acquisition
The GC-MS was performed using a GC-MS (7,890B–5977B, Agilent Technologies, CA, United States) outfitted with a DB-5MS quartz capillary column (0.25 mm × 30 mm, 0.25 μm), as previously mentioned. 28 A 1 µL injection volume with 99.999% pure helium as the carrier gas was utilized for the analysis. The split ratio was 10:1, and the flow rate was 1.0 mL/min. The column temperature was 60 °C for one minute, then raised by 10 °C/min to 250 °C and kept there for 10 min. The injection temperature was 250 °C, while the electron bombardment ion source's temperature was 230 °C. The solvent delay was 5.9 min, and the scanning range of the mass spectrum was m/z 50.0–600.0.
Metabolites and Metabolic Pathway Analysis
Agilent Masshunter software (Qualitative Navigator B.08.00; Agilent Technologies, Inc.) helped collect the GC-MS raw data. Differential metabolites were chosen if P < 0.05 and variable importance in the projection (VIP) > 1 were present. The principal component analysis (PCA), route analysis, and partial least squares-discriminant analysis (PLS-DA) were performed using MetaboAnalyst 5.0. 29 Q2 represents the prediction rate of the model, and R2 represents the degree of fit of the model; the closer the Q2 and R2 to 1, the better the model construction.
RNA Extraction and Transcriptomic Data Analysis
Samples from 4 kidneys were randomly selected from the control, model, and QRKSG group. RNA was extracted, its purity (OD260/280 ratio) was assessed using NanoDrop (Wilmington, DE, United States), its concentration was accurately quantified by Qubit, and its integrity was accurately assessed using an Agilent 2100 Bioanalyzer. The extracted mRNA was reverse transcribed by enzymatic reaction to synthesize the first and second strand of complementary deoxyribonucleic acid (cDNA), the double-stranded cDNA was purified, and the junctions were connected after adding A and repairing at the end, and finally the cDNA library was enriched by PCR, and the original downstream data were filtered and compared to the reference genome of rats. The expression of each gene was calculated based on the comparison results to facilitate further screening and analysis of the samples for differential genes, enrichment, and clustering analysis. DEGs were selected using a value of P ≤ 0.05 and |Fold Change| ≥ 2 as thresholds. GO functional analysis and Gene Set Enrichment Analysis (GSEA) analysis of DEGs were performed using the Omicshare platform, and volcano map, heat map and KEGG enrichment analyses were performed using the bioinformatics website.
Integrative Analysis of Metabolomics and Transcriptomics
The differential metabolites and DEGs obtained from the selection were imported into MetaboAnalyst 5.0 to perform the integration analysis to identify potential relationships between genes and metabolites and to explore the mechanism of action of QRKSG in NS.
Gut microbiota Analysis
Six randomly selected samples of rat intestinal contents from control, model, and QRKSG groups, respectively, were used to extract microbial genomic DNA for microbial community analysis. Amplification was performed using primers in the V3 and V4 regions of the 16S rRNA gene, followed by purification, quantification and sequencing. After obtaining the sequencing data, it was modified based on the official tutorial of QIIME2 2019.4. Finally, the obtained sequencing data were used for relevant bioinformatics analysis. 30
Molecular Docking
The processed target proteins and QRKSG active ingredient structures were imported inside the molecular docking tool Auto Dock, and a docking score table was obtained after the completion of the docking; compounds with a score < -5.0 were considered as having a good binding ability. Finally, the selected targets and active ingredients were visualized and analyzed using PyMOL.
Western Blot Analysis
The kidney samples were cut into fine pieces on ice, 1 mL lysis solution was added to 100 mg kidney and homogenized to completely lyse it. The proteins were separated by 12% SDS-PAGE electrophoresis and transferred to PVDF membranes. The membrane was blocked with 5% skim milk in TBST solution (Tris-buffered saline, 0.1% Tween 20) for 1 h at room temperature and then incubated overnight at 4 °C with the following primary rabbit antibodies: PI3 K (1:1000 dilution, all the antibodies were provided by Bioworld Technology Inc., MN, USA), p-PI3 K, AKT, p-AKT, Caspase 9, spleen tyrosine kinase (SYK), monoamine oxidase (MAOB), aldehyde dehydrogenase 3 family member A1 (ALDH3A1), glutaminase 2 (GLS2) (all diluted 1:1000). Next, the membranes were washed three times with TBST solution, rinsed thrice using TBST water before being infused with the secondary antibody with horseradish peroxidase (HRP)-labeled goat anti-rabbit IgG (1:20,000) for 30 min at 37 °C. Immunoreactive areas on the membrane were observed using the Enhanced Chemiluminescence (ECL) kit (WBKLS0100, Millipore, MA, United States) according to the manufacturer's instructions. Densitometric and immunoreactive analyses using a Tanon-5200 imaging system (Tanon Science and Technology Co. Ltd, Shanghai, China) and ImageJ software (NIH, MD, United States). The internal basis for qualitative analysis was β-action.
Statistical Analysis
The outcomes were presented as mean ± standard deviation with the GraphPad Prism 9.5 statistical software and used for data analyses. One-way ANOVA helped compare between groups. P < 0.05 indicated statistical significance.
Results
Determination of the Active Ingredient of QRKSG
The active components of QRKSG were analyzed by UPLC-MS/MS. A total of 422 compounds were identified from the metware database according to the secondary spectral information of the substance representation. They mainly included phenolic acids, flavonoids, terpenoids, phenylpropanoids, alkaloids, and tannins (Figure 1A).

Component analysis and 24 h urine protein test. (A) Chart of QRKSG active ingredient composition ratio; (B) Urine protein quantitation. Results are presented as mean ± standard deviation. n = 6, **P < 0.01 compared to the control group; ##P < 0.01 compared to the model group.
Urine Protein Quantitation and serum Biochemical Indexes
After developing the NS model, the protein level of rat urine steadily increased from week one to four (Figure 1B). The model group's urine levels of protein were greater than the absence group (P < 0.01), suggesting that the NS model was successfully established. No significant change in the 24 h urinary protein content in the first week of gavage in the QRKSG group and benazepril group compared with the model group, while the urinary protein content in the second, third, and fourth weeks of gavage was significantly lower in the QRKSG and benazepril group compared with the model group (P < 0.01), indicating that the two drugs reduced the urinary protein content in NS rats, and the effect of QRKSG was similar to that of benazepril. In addition, the levels of TP and ALB in the model group were lower than those in the control group, while the levels of Cr, BUN, TG, and CHO were higher than those in the control group. QRKSG inhibited the NS-induced decrease in TP and ALB levels and the NS-induced increase in Cr, BUN, TG, and CHO levels compared with the model group, with results similar to those in the benazepril group (Figure 2).
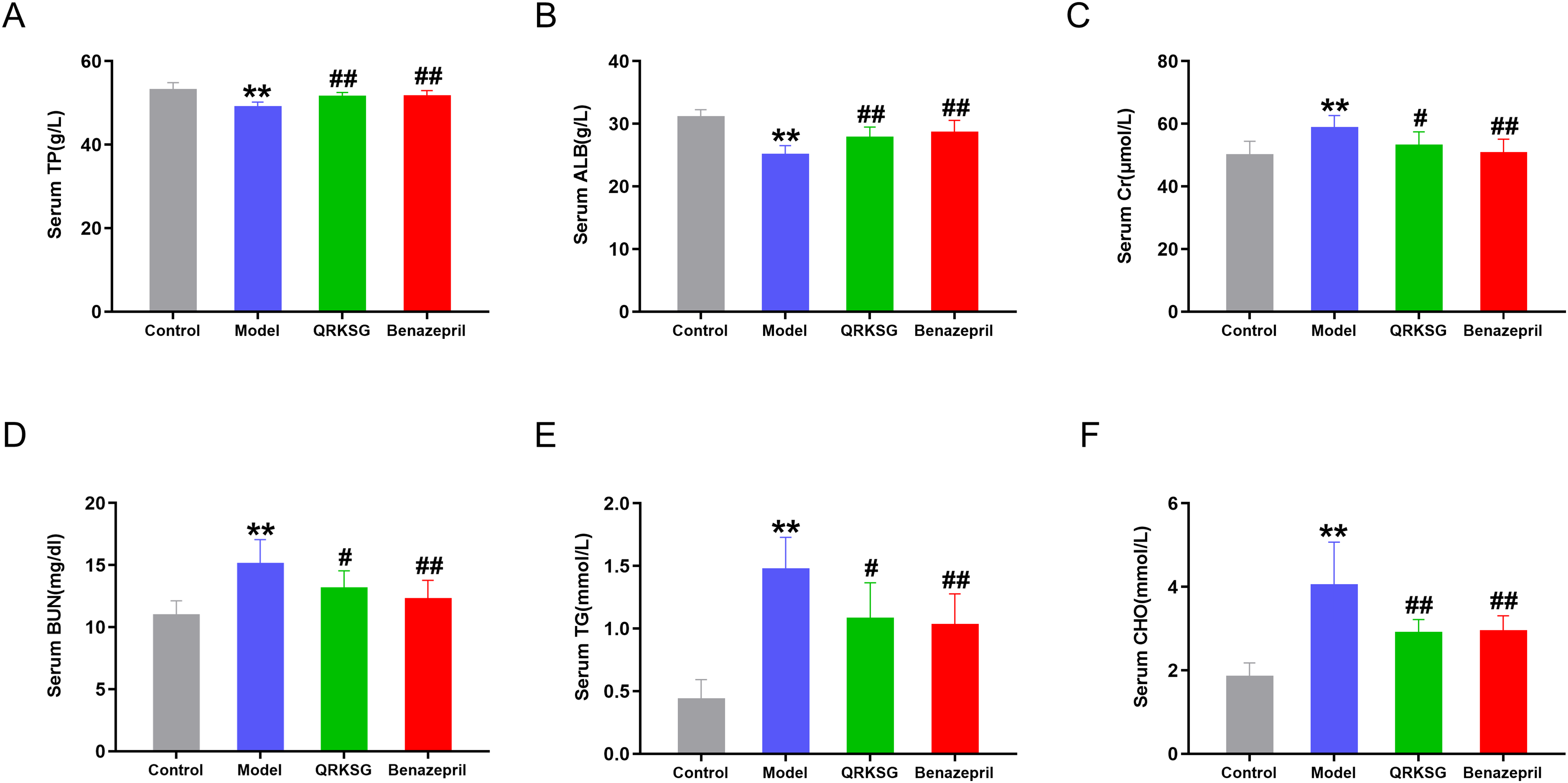
Changes in serum biochemical indexes. (A) Total protein (TP) level; (B) albumin (ALB) level; (C) creatinine (Cr) level; (D) urea nitrogen (BUN) level; (E) total triglycerides (TG) level; (F) cholesterol (CHO) level. n = 6, **P < 0.01 compared to the control group; #P < 0.05 and ##P < 0.01 compared to the model group.
Histopathological Analysis
Parts of the H&E staining (Figure 3) of kidneys showed that the glomerular and tubular structures were normal in the control group, and no evident kidney injury was present. The model group possessed abnormal tubular structures, tubular dilatation, and edema compared to those in the control group, demonstrating the viability of the NS model. Notably, the benazepril and QRKSG group showed less luminal dilatation and reduced edema compared with the model group, and were similar to the control group, indicating that QRKSG ameliorated kidney injury in ADR-induced NS rats.
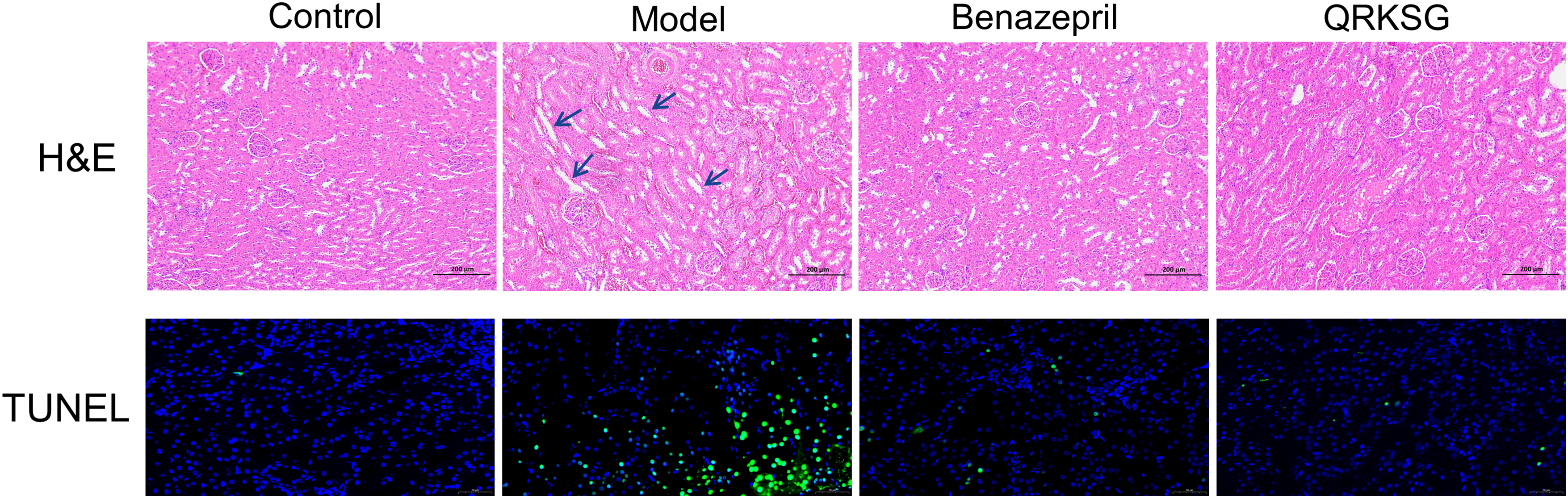
H&E and TUNEL staining in the kidneys of control, model, benazepril, and QRKSG rats. The green fluorescent cells were apoptosis-positive cells.
TUNEL Fluorescent Staining
The model group showed more dead cells than the control group. Compared to the model group, benazepril and QRKSG lowered apoptosis, suggesting that QRKSG had a mitigating impact on renal apoptosis in NS rats (Figure 3).
Metabolomics Analysis of Kidney Samples
Various metabolite levels differed significantly in each group, based on total ion flow plots of kidney samples from the control, model, benazepril, and QRKSG group (Figure 4A). The multivariate analysis showed a clear separation among the control, model, QRKSG, and benazepril group (Figure 4B, C), with R2 and Q2 values of 0.99963 and 0.84692 respectively, for PLS-DA cross-validation (Figure 4D), indicating that the NS model was well constructed and had a good predictive ability.
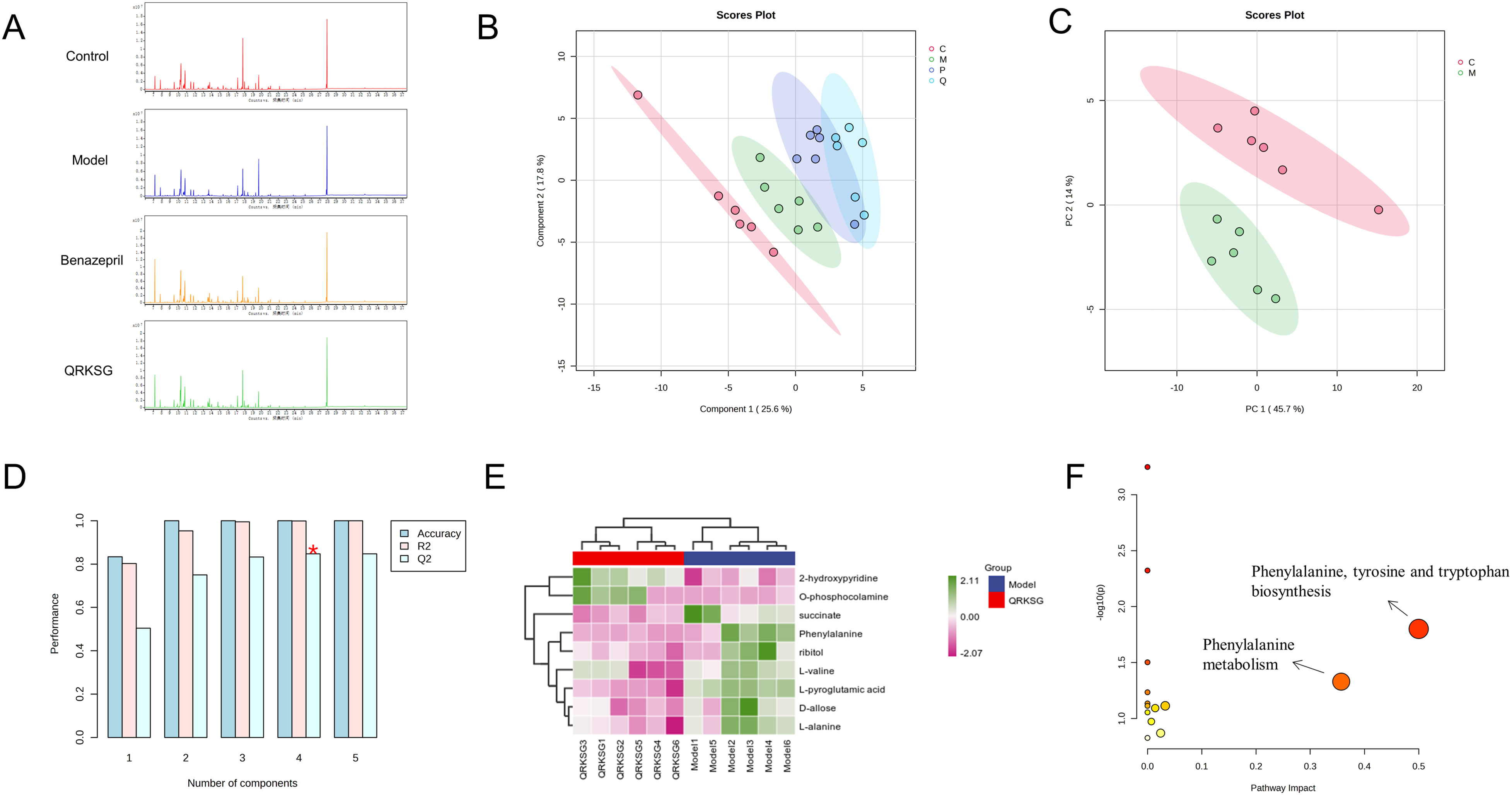
Metabolomics analysis. (A) Total ion flow chromatograms of kidney metabolites in the control, model, benazepril, and QRKSG group of rats; (B) PLS-DA score plot; (C) PCA analysis of the control and model group; (D) R2 and Q2 in PLS-DA; (E) heatmap visualization of 9 metabolites; (F) metabolic pathways analysis. n = 6.
The selection was performed using VIP > 1 and P < 0.05, and a total of 9 potential differential metabolites were selected in the model and QRKSG group. The expression of two metabolites was up-regulated, while that of seven metabolites was down-regulated in the QRKSG group compared with the model group (Table 1). The metabolite clustering heat map (Figure 4E) showed that the differential metabolites in the QRKSG group significantly changed compared to those in the model group. The above results suggested that QRKSG reversed NS to some extent. The pathway enrichment analysis of the nine selected differential metabolites (Figure 4F) revealed that two key pathways were involved by impact > 0.1, mainly phenylalanine, tyrosine and tryptophan biosynthesis, phenylalanine metabolism, suggesting that NS might be related to amino acid metabolism disorder.
Differentially Expressed Metabolites in the Model and QRKSG Group. n = 6.

Transcriptomics Analysis of Kidney Samples
A total of 687 genes were significantly expressed in the QRKSG group compared to the model group (|Fold Change| ≥ 2 and pval ≤ 0.05), which were divided into 288 up-regulated genes and 399 down-regulated genes, as shown in the volcano plot (Figure 5A). The expression of 301 DEGs was restored after QRKSG administration (Figure 5B). The heat map (Figure 5C) showed the significantly altered expression of some genes after modeling and QRKSG administration, indicating that the disorder of some genes may return to normal after QRKSG treatment. In addition, KEGG pathway analysis on DEGs revealed that they were mainly enriched in the PI3K-Akt signaling pathway and B cell receptor (BCR) signaling pathway (Figure 5D) and the GSEA analysis revealed that the PI3K-Akt signaling pathway and BCR signaling pathway were down-regulated after QRKSG administration (Figure 5F). GO enrichment analysis (Figure 5E) showed that the biological functions that DEGs were involved were mainly hemopoiesis, regulation of leukocyte-mediated immunity, immune system process, hematopoietic or lymphoid organ development, and immune system development.

Transcriptomics analysis. (A) Volcano plot of DEGs in the model and QRKSG group; (B) Venn diagram of DEGs among groups; (C) heatmap visualization of DEGs in the control, model, and QRKSG group; (D) KEGG pathway analysis of DEGs in the model and QRKSG group; (E) GO terms of the biological processes in which DEGs are involved; (F) GSEA analysis of the model group and QRKSG group. n = 4.
Integration Analysis of Metabolomics and Transcriptomics
The results in Figure 6A show that the pathways involved in DEGs, and differential metabolites mainly included linoleic acid metabolism, phenylalanine metabolism, ascorbate and alternate metabolism, pyruvate metabolism, alpha-Linolenic acid metabolism, alanine, aspartate and glutamate metabolism, butanoate metabolism, and phenylalanine, tyrosine and tryptophan biosynthesis. Like the results of metabolomic pathway analysis, amino acid metabolism was the main relevant metabolic pathway. Among them, the phenylalanine metabolism pathway and the alanine, aspartate and glutamate metabolism pathway involved both DEGs and differential metabolites. Further analysis revealed that L-phenylalanine, MAOB, and ALDH3A1 matched to the compounds and gene libraries related to the phenylalanine metabolism pathway (Figure 6B); L-alanine, succinate, and GLS2 were present among the compounds and gene libraries related to the alanine, aspartate and glutamate metabolism pathway (Figure 6C). The above results suggested that QRKSG might regulate metabolic pathways involved in NS by altering the expression of DEGs.

Integrated analysis of metabolomics and transcriptomics. (A) Joint pathway analysis based on transcriptomics and metabolomics; (B) phenylalanine metabolism pathway; (C) alanine, aspartate and glutamate metabolism pathway. Metabolites are represented by square hexagons and genes are represented by circles. Those marked by dark blue and dark red are respectively potential targets and metabolites regulated by QRKSG.
Gut microbiota Analysis
The microbiota of rat gut contents was analyzed for species annotation (Figure 7A, B). At the portal level, the major components of the gut microbiota were Firmicutes, Bacteroidetes, Proteobacteria, and Tenericutes. Firmicutes and Proteobacteria decreased and Bacteroidetes and Tenericutes increased in the model group compared to the control group, while QRKSG treatment showed the opposite trend. At the genus level, Lactobacillus, Prevotella, Oscillospira, Ruminococcus, Clostridium, Allobaculum, and Desulfovibrio showed large variations in relative abundance among the three groups. The sparse curves (Figure 7D) and box plots (Figure 7E) of the Alpha diversity index showed that the sequencing data of each group of samples were feasible and suitable for further study. Beta diversity analysis, principal coordinate analysis (PCoA) showed a clear separation between control, model, and QRKSG group (Figure 7C). The linear discriminant analysis effect size (LEfSe) analysis was used to identify differentially abundant microbiota in the three groups (Figure 7F), where the circles represent different levels of microbial communities, and the different diameters of the circles represent the different relative abundances of these microbial communities. The higher the LDA score, the more significant the differences in the taxa. Taking the LDA score > 3.0 for screening, a total of 22 microbiota were identified at the genus level, mainly including Actinomyces, Adlercreutzia, Paraprevotella, Bacteroides, Prevotella, Alistipes, Lactobacillus, Streptococcus, Turicibacter, Blautia, Coprococcus, Pseudobutyrivibrio, Roseburia, Ruminococcus, Phascolarctobacterium, Allobaculum, Sutterella, Delftia, Nitrosomonas, Desulfovibrio, Flexispira, Treponema. Functional prediction analysis of microorganisms showed that major changes occurred in the pathways of amino acid metabolism, carbohydrate metabolism and energy metabolism (Figure 8).

Results of gut microbiota analysis in rats. (A) Relative abundance of gut microbiota at the gate level in the control, model, and QRKSG group; (B) relative abundance of gut microbiota at the genus level in three groups; (C) principal coordinate analysis (PCoA) among control, model, and QRKSG group; (D) sparse curves of Alpha diversity index; (E) box plots of Alpha diversity index; (F) identification of significantly different microbiota in control, model, and QRKSG group by linear discriminant analysis effect size (LEfSe) analysis. n = 6.

Predictive analysis of the function of gut microorganisms.
Molecular Docking
Molecular docking was performed on the selected genes with some of the active ingredients of QRKSG to confirm the key genes. The results in Figure 9 showed that the docking of MAOB, ALDH3A1, PI3 K, and SYK with the active ingredients was the best, and the docking of phenolic acid components and flavonoid components with each target was the best. The four targets with the best overall docking effect and the components with the best docking effect were selected to visualize the graph.

Molecular docking results. (A) Heat map of docking fractions of targets obtained from the integration analysis and QRKSG active ingredients; (B) MAOB docking with 1,3-O-Dicaffeoylquinic acid; (C) ALDH3A1 docking with Quercetin-3-O-robinobioside; (D) PI3 K docking with Isochlorogenic acid B; (E) SYK docking with Isochlorogenic acid B.
Expression of Proteins Involved in the Pathways Obtained from DEG Enrichment
The major signaling pathways obtained from DEG enrichment were further investigated, as well as the effect of QRKSG on metabolic pathways obtained from metabolomics and transcriptomics integration analysis. The results in Figure 10 showed that NS induced a significant upregulation of MAOB, ALDH3A1, GLS2, SYK, p-PI3 K, p-AKT, and caspase 9 compared with the control group (P < 0.01). The expression of MAOB, ALDH3A1, GLS2, SYK, p-PI3 K, p-AKT, and caspase 9 was significantly decreased after QRKSG administration compared with the model group (P < 0.01). The above results suggested that QRKSG might regulate the related pathways by down-regulating the expression of these proteins to treat NS.

MAOB, ALDH3A1, GLS2, SYK, p-PI3 K/PI3 K, p-AKT/AKT, and caspase 9 protein expression in rat kidney by Western Blot. n = 3, **P < 0.01 compared to the control group; ##P < 0.01 compared to the model group.
Discussion
QRKSG effectively improved renal function indexes and reduced renal injury in an ADR-induced NS rat model. In addition, QRKSG treatment resulted in a significant reduction of apoptosis in the kidney. Thus, QRKSG has a protective effect on NS. By integrating metabolomics and transcriptomics, it was found that QRKSG can be used to treat NS by modulating phenylalanine metabolism, alanine, aspartate and glutamate metabolism, and the PI3K-Akt signaling pathway (Figure 11). The results of the intestinal flora analysis showed that QRKSG also affected the abundance of some gut microbiota.
Current studies suggest that the main mechanism regulating NS is the increased permeability of glomeruli to proteins, resulting many proteins entering the urine causing proteinuria, which leads to many symptoms including hydrops, renal inflammation, coagulation, and immune disorders in patients. 31 At present, corticosteroids, cyclophosphamide, chloramphenicol, cyclosporine, and angiotensin-converting enzyme inhibitors are commonly used to treat NS. However, drug resistance and other side effects of hormone drugs have not been effectively solved.1,32 There are many studies showing that traditional medicines can exert a protective effect on renal tissues by reducing oxidative stress, anti-apoptosis, and anti-inflammation. For example, gallic acid may protect the kidney from cisplatin-induced nephrotoxicity through antioxidant, anti-inflammatory, and anti-apoptotic properties. 33 Naringin protects the kidney by reducing pro-inflammatory factors, inhibiting apoptosis, and limiting oxidative stress. 34 Vanillic acid protects the kidney from methotrexate-induced nephrotoxicity through anti-apoptotic, antioxidant and anti-inflammatory properties. 35 Therefore, this study aimed to find treatment options from natural medicines. Various components contained in chicory have a protective effect on the kidney and attenuate the apoptosis of kidney cells.36,37 QRKSG is a drug obtained from chicory processing, but its mechanism of action on NS is not clear. Therefore, there is an urgent need to explore the mechanism of action of QRKSG in the treatment of NS and to provide a theoretical basis for the clinical use of QRKSG in the treatment of NS. Among multi-omics studies, metabolomics occupies a unique position and is receiving increasing attention in comprehensive analyses. 38 The transcriptomic analysis accelerates target discovery by measuring gene expression and contributes to the understanding of human diseases. 39 In addition, loads of evidence in multi-omics studies provide more reliable results and reduce the risk of false positive results. 40 Therefore, in this study, a multi-omics strategy combining metabolomics and transcriptomics was used to investigate the mechanism of action of QRKSG on NS. Kidney disease causes changes in the intestinal microbiota, and polysaccharides rich in chicory help to improve the intestinal microbiota. Hence, this study used intestinal flora analysis to explore the mechanism of action of QRKSG for NS.
Transcriptomic analysis showed that QRKSG could alleviate NS by regulating the BCR signaling pathway and the PI3K-Akt signaling pathway. The PI3K-Akt signaling pathway has a mechanism ahead of it known as the BCR transmission pathway. SYK plays a critical role in the adaptive immune receptor signaling; phosphorylated tyrosine residues trigger the activation of SYK and its direct binding to PI3 K, thereby activating downstream signaling components and various cellular responses. 41 SYK is crucial in inflammatory response onset because it transmits the signal of immune cell receptors to intracellular signaling pathways that control cellular responses to extracellular antigens and antigen-immunoglobulin complexes. Inhibiting SYK expression hence reduces the signs and effects of both acute and chronic inflammation. 42 In addition, results from clinical studies suggest that SYK is involved in the pathogenesis of multiple kidney diseases. 43 The tyrosine kinase receptor in order G protein-coupled receptors/cytokine receptor, and Ras protein-related GDP enzyme receptor function together to activate the neuro-signaling enzyme PI3 K. 44 AKT is a PI3 K downstream serine/threonine kinase that is activated to control various proteins vital to cell survival, proliferation, migration, digestion, and angiogenesis. 45 PI3K-Akt signaling system in mammalian cells regulates metabolism, cell migration, expansion, and reproduction, 46 and is involved in the treatment of NS.9,47 Caspases are intracellular proteases working as initiators and effectors of apoptosis, and AKT is activated to induce caspase 9 expression. 48 Caspase 9 is a member of the cysteine protease family of cysteine proteases and is involved in apoptosis and cytokine processing. 49 The inhibition of caspase 9 expression reduces apoptosis in the kidney. 50 Therefore, the inhibition of PI3 K and AKT expression might in turn reduce the expression of caspase 9, thus protecting kidney cells. Our findings indicated that QRKSG affected the expression of the genes SYK, PI3 K, and AKT and that these genes were associated with the PI3K-Akt and BCR signaling pathways. The expression of SYK, p-PI3 K, p-AKT, and caspase 9 increased with the establishment of the NS model, suggesting that the inflammatory pathway and apoptotic pathway were activated. Thus, renal cells gradually underwent apoptosis, leading to kidney injury. QRKSG administration resulted in a decreased expression of SYK, p-PI3 K, p-AKT, and caspase 9, suggesting that the BCR signaling pathway and PI3K-Akt signaling pathway were inhibited and renal injury was attenuated. Our hypothesis was that QRKSG might significantly reduce the release of caspase 9 and attenuate the apoptosis of renal cells by inhibiting the BCR signaling pathway and PI3K-Akt signaling pathway in NS rats, thus alleviating NS, which was also confirmed by the TUNEL staining results.
Integrated analysis of metabolomics and transcriptomics revealed that QRKSG could improve NS by regulating alanine, aspartate and glutamate metabolic pathways. Renal disease is closely associated with an abnormal amino acid metabolism. 51 Succinate causes tubular dilatation as well as tubular epithelial cell apoptosis and induces significant renal injury. 52 Reactive oxygen species (ROS) can damage cells, primarily generated by mitochondria and NADPH oxidases, and are involved in various processes. 53 Succinate induces the accumulation of ROS in the kidney, which excessive production causes oxidative stress and consequent tissue damage.54,55 The accumulation of ROS also activates pathways associated with cell proliferation and survival, including MAPK and PI3K-Akt pathways. 56 Therefore, a close relationship exists between oxidative stress and inflammation. Studies have shown that NS is closely related to oxidative stress,57,58 and we surmise that succinate-induced oxidative stress may exacerbate renal inflammation of NS. GLS2 is localized in the inner mitochondrial membrane and catalyzes the γ-amino hydrolysis of glutamine to form glutamate and ammonia. 59 GLS2 is the key enzyme in the alanine, aspartate and glutamate metabolic pathways, and it mediates the conversion of L-glutamine to L-glutamate, eventually forming succinate after a series of reactions. GLS2 expression was down-regulated and succinate was reduced after QRKSG administration, suggesting that QRKSG might affect the synthesis of succinate by affecting GLS2 expression, ultimately regulating this metabolic pathway, reducing oxidative stress, alleviating inflammation, and thus protecting the kidney from injury.
Integrated analyses showed that QRKSG also improved NS by affecting the phenylalanine metabolic pathway. MAO is an enzyme that catalyzes the degradation of different monoamines by oxidative deamination. Two isoforms exist, MAOA and MAOB, both involved in the regulation of cell growth and repair, as well as in proliferative kidney diseases. 60 The inhibition of MAO expression reduces vascular oxidative stress in patients with end-stage renal disease. 61 Phenylalanine is metabolized in the human kidney and converted to tyrosine. 62 The tyrosine plays an important role in mediating energy metabolism. 63 QRKSG treatment decreased the expression of MAOB and ALDH3A1 and phenylalanine was reduced in the phenylalanine metabolic pathway, probably because the genes with reduced expression and reduced phenylalanine might lead to reduced tyrosine production. In contrast, the increased tyrosine content in the model group might be due to the disturbance of energy metabolism caused by NS, since the body needs to produce more tyrosine to meet the energy demand. The decreased tyrosine after the administration of QRKSG suggested that it improved energy metabolism, leading to its gradual stabilization. Targeting podocyte energy metabolism has been proposed as an intervention in the treatment of NS. 64 Therefore, QRKSG might regulate tyrosine synthesis and improve energy metabolism by restoring the normal expression of MAOB and ALDH3A1 in the phenylalanine metabolic pathway, thereby protecting renal tissues. In addition, most of the targets obtained by the selection had good binding ability with the active ingredients of QRKSG, with the best binding with phenolic acid and flavonoid components. Thus, our speculation was that the active ingredients of QRKSG might play a therapeutic role in NS.
QRKSG was found to alleviate NS by improving the gut microbiota through gut flora analysis. The gut microbiota is closely linked to human health, and a healthy gut microbiota consists mainly of Firmicutes and Bacteroidetes. 65 When the intestinal microbiota is dysbiotic, the Firmicutes-Bacteroidetes ratio is altered, and a decrease in the Firmicutes-Bacteroidetes ratio can be observed in some chronic autoimmune diseases. 66 In our study, we found that NS rats had a reduced Firmicutes-Bacteroidetes ratio compared to the control group, and QRKSG administration reversed the change in the Firmicutes-Bacteroidetes ratio. At the genus level, Lactobacillus acts as a probiotic that promotes a stable gut microbiota. 67 Our results showed that Lactobacillus in the intestinal tract of NS rats was significantly less than that of the control group, and QRKSG administration increased the number of Lactobacillus, which facilitated the homeostasis of the intestinal microbiota. Recent studies have shown that increased abundance of Prevotella is associated with a number of inflammatory diseases and metabolic disorders, and that it may be a clinically important virulence factor that can participate in human disease by promoting chronic inflammation. 68 Our results showed that the abundance of Prevotella in the intestines of NS rats was significantly higher than that of the control group, and the abundance of Prevotella was reduced after QRKSG administration, and we hypothesized that QRKSG might reduce the inflammatory response by decreasing the abundance of Prevotella to achieve the alleviation of NS. As a member of the Firmicutes, Ruminococcus produces the short-chain fatty acid butyrate, which, in addition to supplying energy to the cells of the colon, reduces inflammation and helps to maintain the health of the intestinal tract.69-71 Our study showed that the abundance of Ruminococcus in the intestines of NS rats was reduced compared to the control group and elevated after QRKSG administration. Therefore, we concluded that the occurrence of NS might be closely related to gut microbiota dysbiosis, and that QRKSG might ameliorate NS by maintaining the homeostasis of the gut microbiota.
However, there are some limitations in this study, the main components found by molecular docking that QRKSG exerts therapeutic effects are flavonoid and phenolic acid components, which can be followed up by testing the components of the animal's blood intake in order to further validate the active components.
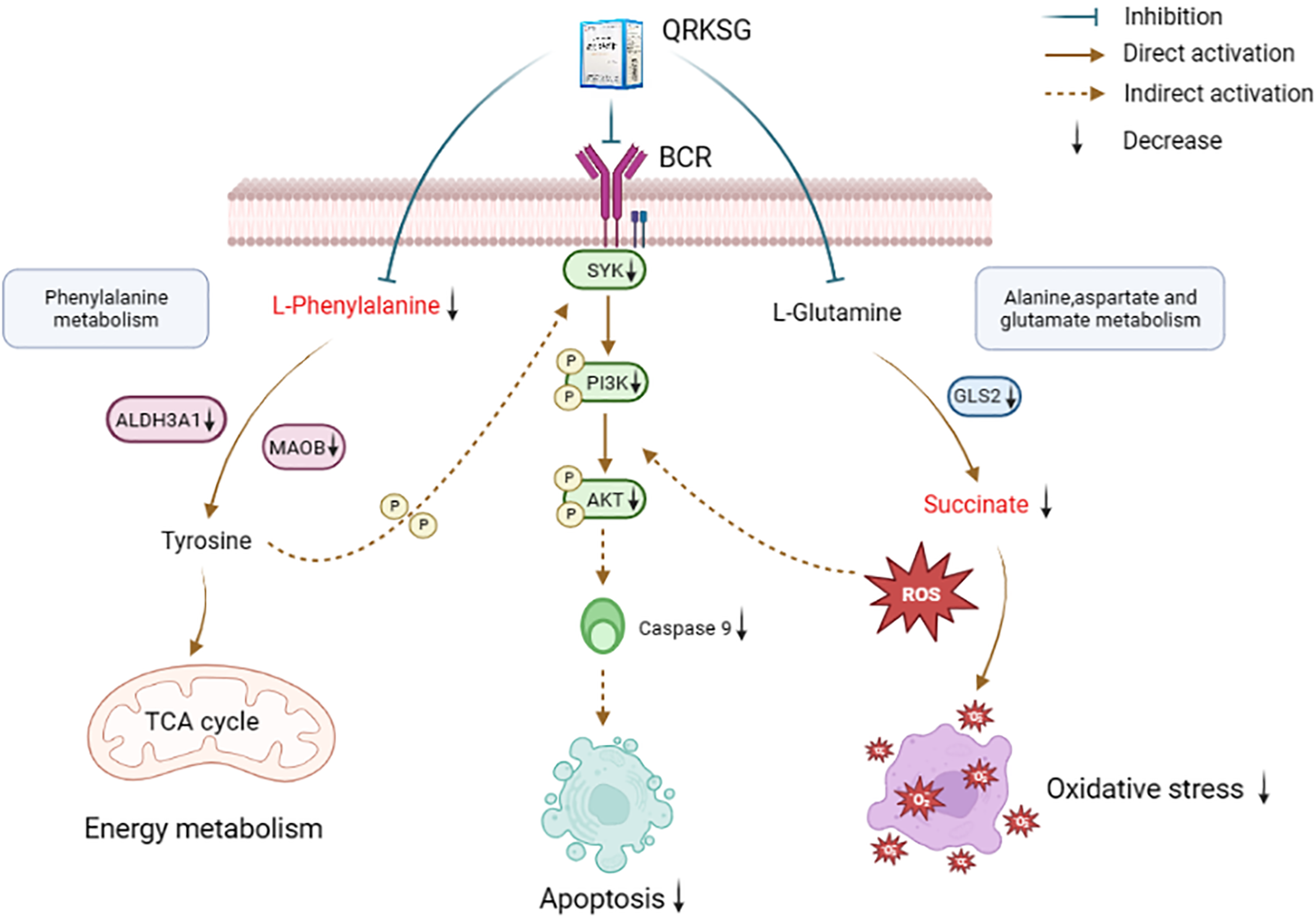
Mechanism of action of QRKSG on NS.
Conclusions
The present study indicated that QRKSG was effective in rats with ADR-induced NS. A comprehensive examination using metabolomics and transcriptomics revealed that QRKSG was mainly involved in the control of the energy balance, oxidative stress, inflammation, and apoptosis, as well as being linked to the control of amino acid metabolism, all factors reducing NS. The therapeutic effects of QRKSG were mediated by a variety of components, pathways, and targets. In addition, QRKSG might also achieve relief of NS by improving the gut microbiota. These results implied that a multi-omics approach might be an effective approach to evaluate the complex mechanisms used by herbal remedies to treat diseases.
Footnotes
Abbreviations
Acknowledgments
This study was financially supported by the Science and Technology Innovation Team Project of Hubei Provincial Department of Education for Young and Middle-aged Scientists (Grant No. T201608), and the Funds for Key Program for Traditional Chinese Medicine of Hubei University of Chinese Medicine (No. 2022ZZXZ004).
Author Contributions
Lintao Han and Jingjing Li designed the research project; Shanshan Wang, Hailing Huang, Zhuang Huang, and Yan Ye performed the experiments; Shanshan Wang, Wangqiang Dai, Jiajing Liu, Pengyu Chen, Bailu Duan, Qi Jiang, and Yuxin Wen collected and analyzed the data; Shanshan Wang, Wangqiang Dai, and Zhuang Huang wrote the manuscript. All the authors contributed to the preparation of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This study was approved by the Hubei Provincial Laboratory Animal Management Committee.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the the Science and Technology Innovation Team Project of Hubei Provincial Department of Education for Young and Middle-aged Scientists, the Funds for Key Program for Traditional Chinese Medicine of Hubei University of Chinese Medicine, (grant number T201608, 2022ZZXZ004).
Statement of Human and Animal Rights
All experimental procedures involving animals were conducted by the Code of Animal Ethics of Hubei University of Traditional Chinese Medicine (HBUCMS202206002) and approved by the Hubei Provincial Laboratory Animal Management Committee.
Statement of Informed Consent
There were no human subjects in this study and informed consent did not apply.
