Abstract
Introduction
SP can be defined as idiopathic chronic itching in individuals over the age of 60 1 or systemic itching in patients without primary skin lesions. 2 Pruritus generally originates on the skin, is an unpleasant sensation provoking a desire to scratch and may result in pain, skin damage, and infection. 3 Although scratching may generate pleasant sensations, it does not relieve symptoms. On the contrary, it tends to result in damaging cycles of itching and scratching. 4 Literature reports indicate an SP prevalence of 11% to 78%5–7 and, as individuals age, immune aging ensues and the incidence rate of pruritus increases. 8
Due to the unclear pathogenesis of SP, treatment usually involves anti-allergic drugs such as diphenhydramine or loratadine. Corticosteroids and calcineurin inhibitors can also be used to control itching but are accompanied by side effects such as drowsiness and weight gain. 9 Therefore, the development of innovative treatments for SP is imperative.
YGZ223 is a formula comprised of 7 traditional Chinese medicines including Paeoniae Radix Rubra, Angelicae Sinensis Radix, Cortex Erythrinae, Tribuli Fructus, Sophorae Flavescentis Radix, Dendrobii Caulis, and Zaocys. The formula originates from the clinical experience of the Hubei Provincial Hospital of Traditional Chinese Medicine and has proven therapeutic efficacy against skin pruritus among middle-aged and elderly individuals. This study utilizes network pharmacology methods to uncover the mechanism of YGZ223 in treating SP. UPLC-MS reveals the chemical constituents of YGZ223, molecular docking assesses the bonding ability of the primary constituents to the targets, and the effectiveness of YGZ223 is determined in a 4-aminopyridine-induced mouse itching model.
Materials and Methods
Materials
YGZ223 (batch number 20230417) was obtained from Hubei Provencial Hospital of Traditional Chinese Medicine. Xiaofeng Zhiyang granules (batch number 220107) were obtained from Nanjing Tongrentang Pharmaceutical Co., Ltd. Diphenhydramine hydrochloride tablets (batch numbers 14W10081 and 14W10041) were provided by Yichang Humanwell Pharmaceutical Co., Ltd. Sodium chloride injection (0.9%, batch number 220511522) was purchased from Wuhan Fuxing Biopharmaceutical Co., Ltd. 4-Aminopyridine (4-AP, batch number PWAWH-GU) was provided by Shanghai Tixi Aihua Industrial Development Co., Ltd.
A high-speed freezing centrifuge (Eppendorf AG, 22331 Hamburg), chromatographic system (ACQUITY UPLC M-Class, Waters, USA), and mass spectrometer (Waters Xevo G2-XS, Waters, USA) apparatus were applied.
Animals
SPF grade Kunming mice aged 3 to 4 weeks and weighing 21.67 to 25.70 g were purchased from Hunan Slake Jingda Experimental Animal Co., Ltd, Animal Ethics Number: IACUC (Approved)—YJY-2022-016.
Methods
Chemical Component Acquisition and Target Protein Screening of YGZ223
The chemical composition information of the 7 medicinal herbs of YGZ223 (Paeoniae Radix Rubra, Angelicae Sinensis Radix, Cortex Erythrinae, Tribuli Fructus, Sophorae Flavescentis Radix, Dendrobii Caulis, and Zaocys) was screened using the Herb database, 10 SymMap database, 11 the Chemical Professional Database of the Shanghai Institute of Organic Chemistry, and additional literature screening. PubChem 12 was used to identify compounds meeting the 3 SwissADME rules 13 and conforming with “yes” and “high” requirements for gastrointestinal absorption. Active YGZ223 constituents were retrieved from the SwissTargetPrediction database with a probability >0.1, and target duplication was eliminated. 14
Prediction of SP Targets and Construction of Protein–Protein Interaction Networks
Using the keyword “senile pruritus,” the obtained targets were integrated and de-duplicated with the GeneCards database, 15 DrugBank database, 16 and the TTD database. 17 On the Bioinformatics platform, jvenn 18 was used to create Venn diagrams illustrating the overlap between component targets and disease targets. To obtain preliminary protein–protein interaction networks (PPIs), protein interaction analysis was carried out on the online website STRING, 19 the TSV data file was imported from STRING into Cytoscape, 20 and the Centiscape2.2 plugin in Cytoscape and the network analysis option were applied to simplify analysis and plotting.
GO and KEGG Pathway Enrichment Analysis
Genes common to drugs and diseases were imported into the DAVID database 21 for enrichment analysis of Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG). The GO analysis identified the top 10 significant results with P < .01 while KEGG screened out the top 20 significant pathways. Column charts for the GO analysis and bubble charts for the KEGG analysis were created using the Bioinformatics platform.
Constructing “Herb-Ingredient-Target” and “Components-Targets-Pathways” Networks
Both “herb-ingredient-target” and “constituent-target-pathway” networks were created using Cytoscape software, and analyzed with the Network Analyzer plug-in in Cytoscape, with emphasis on the node degree values and arranging the graphs in order of degrees of freedom.
Preparation of YGZ223 Sample
YGZ223 extract (0.10 g) was accurately measured and transferred into a 10 mL volumetric flask. In a suitable amount of pure water, ultrasound was applied at 53 kHz and 350 W for 30 min. Once cooled down to room temperature, pure water was added to the flask up to the mark. After shaking, a 1 mL sample was placed in a 1.5 mL centrifuge tube. Centrifugation for 10 min at room temperature at 12000 rpm· min−1 was followed by microporous membrane filtration (0.22 μM) to obtain the filtrate.
Chromatography and Mass Spectrometry Conditions
Using a Waters ACQUITY BEH C18 column (1.7 μm, 100 × 2.1 mm) and 0.1% formic acid in water as mobile phase A and acetonitrile as mobile phase B, the following elution procedure was applied: 0 to 8 min, 5% B; 8 to 9 min, 5% to 9% B; 9 to 14 min, 9% B; 14 to 30 min, 9% to 19% B; 30 to 45 min, 19% to 42% B; 45 to 50 min, 42% to 95% B; 50 to 60 min, 95% B; applying flow rate 0.3 mL/min, injection volume 2 μL, detection wavelength 205 nm, and column temperature 30 °C.
Mass spectrometry conditions comprised an ESI ion source, collecting data in both positive and negative ion modes. The MSE mode collects primary and secondary mass spectrometry ions, with a scanning range of m/z 50 to 1500 Da, at electric spray voltage 3000 V, cone hole voltage 20 V, ESI ion source temperature 100 °C, curtain gas 50 L/h, and N2 desolvation gas at a flow rate 1000 L/h.
Data Collection, Statistics, and Chemical Composition Identification
UPLC-MS/MS data was collected and analyzed using a Waters MassLynx V4.1 and MSE DataViewer 1.2 software. The literature on the chemical constituents and mass spectrometry fragmentation patterns of the 7 medicinal herbs of YGZ223 was reviewed, summarizing the classes of compounds in YGZ223 with reference to the MassBank and PDMB databases, and according to reverse phase chromatography retention times.
Molecular Docking
The 3D structure SDF files of potentially active constituents were downloaded from the PubChem database and were converted into PDB files using OPENBabel software. The complete sequence and high-resolution 3D crystal structures of target proteins were downloaded from RCSB 22 and AlphaFold protein database. 23 PyMOL software was used to remove solvent ligands, and hydrogenate the proteins to act as receptors. Small molecule compounds were introduced as ligands to obtain the optimal conformation through conformational binding energy and interaction force analysis. Finally, PyMOL software was used to visualize the molecular docking results.
Molecular Dynamics Simulation of Formononetin-AKT1 complex
Small molecules and proteins were described using GAFF small molecule force field and AMBER-14SB protein force field, respectively. Using the pdb2gmx subroutine to add hydrogen atoms to the system, a truncated cubic TIP3P solvent box was added at a distance of 10 Å in the system, 24 and Na+/Cl− was added to the system for charge balance. Finally, the topology and parameter files for simulation were output.
Molecular dynamics simulation was conducted using Gromacs 2020.6 softwareff, 25 with a simulation duration of 80 ns. Before simulation, the system was optimized for energy by using the mdrun command and the steepest descent method for energy minimization (canonical ensemble), with a starting step size of 0.01 nm and a maximum force tolerance of 1000 kJ/mol • nm. After optimization of the system energy, a 100 ps NVT (isothermal-isochoric) lineage simulation was performed on the system at a fixed volume and constant heating rate, causing the system temperature to slowly rise from 0 K to 310.15 K, further unifying the distribution of solvent molecules in the solvent box. Subsequently, a 100 ps NpT (isothermal-isobaric) ensemble simulation was conducted using a Berendsen constant pressure device to reach pressure equilibrium on the solvent composite system, raising the system pressure to 1 bar. In the MD simulation process, all hydrogen bonds involved are constrained using the LINCS algorithm, with an integration step of 2 fs. The electrostatic interaction was calculated using the Particle mesh Ewald method, with a cutoff value of 1.2 nm. The non-bond interaction truncation value was set to 10 Å and was updated every 10 steps. Periodic elimination was performed on the simulated trajectory and subsequent analysis on RMSD, RMSF, Rg, hydrogen bonding, MM/GBSA, etc. The binding free energy of small molecules and proteins was calculated using the MM/GBSA method.26–28 This article uses the GB model developed by Nguyen et al for calculation (igb = 2). 29
Pharmacodynamic Experiment of YGZ223 on 4-Aminopyridine Induced Skin Itching Model in Mice
Forty SPF grade KM mice, half male and half female, were randomly divided into five groups including the model control group (Model, given physiological saline), the Xiaofeng Zhiyang granule group (XFZY, 10.8 g/kg), the diphenhydramine hydrochloride group (DF, 0.0226 g/kg), the YGZ223 group (19.6 g/kg), and the blank control group (Control, given physiological saline). The YGZ223 group and 2 positive control groups were administered with the corresponding drugs according to body weight every morning. The Model control group and blank Control group were given physiological saline with a dosage volume of 0.2 mL/10 g, and were administered daily for 7 days. After the last 60 min of administration, itching test was conducted: except for the blank control group, mice in other groups were subcutaneously injected with 0.02% 4-AP solution on the back of the tail, 0.1mL per mouse. The blank control group was given an equal amount of physiological saline, 0.1 mL per mouse. Mouse licks itself until it stops counting once (lasting less than 5 s). Longer lasting licking (exceeding 5 s) is recorded as 2 licks during the statistical analysis. The number of licks per mouse is recorded within a 20-min period.
Result
Obtaining Chemical Components and Targets
Through database and literature searches, 253 active constituents were identified in YGZ223. Use of SwissTargetPrediction to screen targets with a probability >0.1 and remove duplicates, 941 targets corresponding to the active constituents in YGZ223 were obtained. The GeneCards database, DrugBank database, and TTD database identified 1077 molecular targets for SP after merging and de-duplication. The common targets of the 2 are 258, which are potential targets for YGZ223 to act on SP, as shown in Figure 1A.
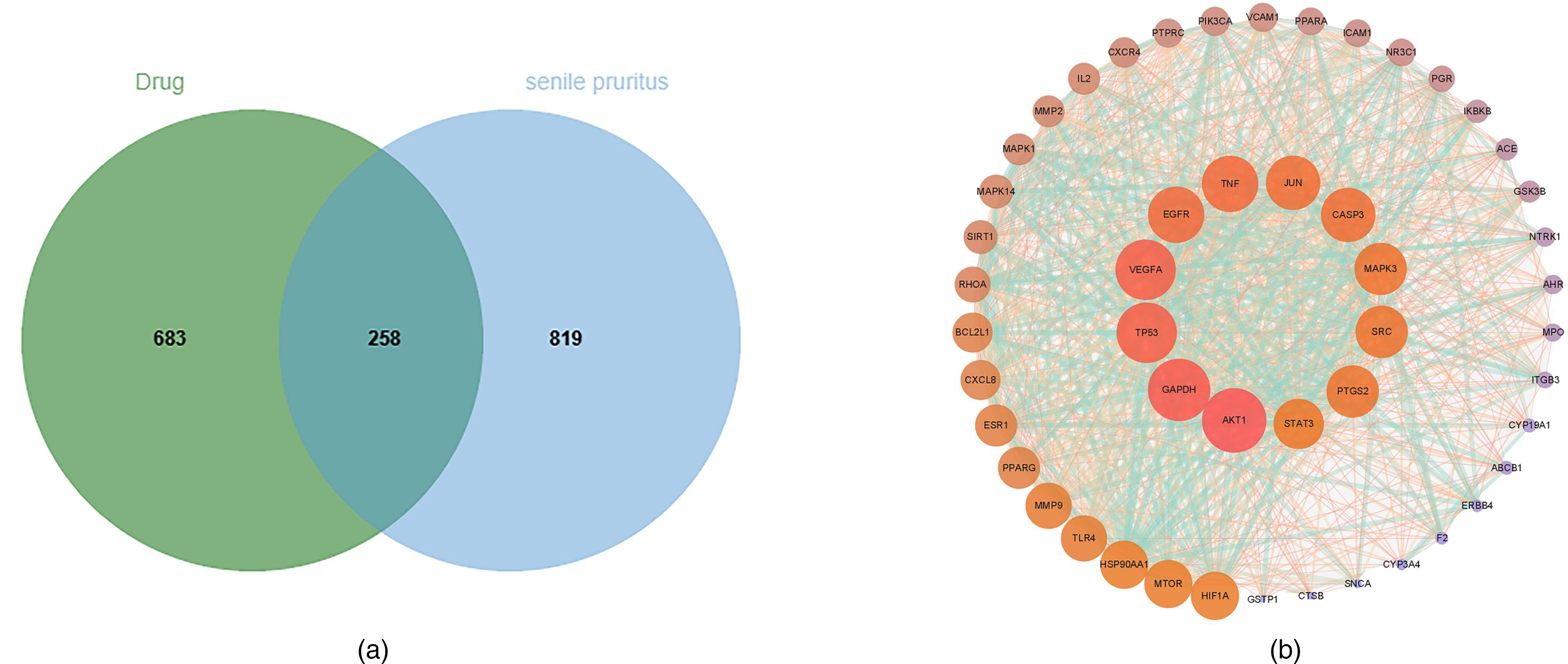
(A) YGZ223 and Venn map of SP targets. (B) Intersection target PPI network (red indicates a high score, while the larger the circle, the higher the score).
PPI Construction and Core Target Screening
The 258 common targets were uploaded to the STRING database for retrieval of the TSV files which were loaded to Cytoscape, adjusting the visualization parameters Degree >37.39, Closeness >0.002, and Betweenness >263.66, ultimately obtaining 50 nodes and 825 edges (Figure 1B). The top 20 selected key molecular targets are shown in Table 1.
Information on the top 20 key Molecular Targets.

Analysis of YGZ223 Effective Constituents Action Target Network Diagram
The YGZ223 active constituent target network constructed using Cytoscape3.9.1 comprises 429 nodes and 2057 edges. The circle in the graph represents the active constituents, different colors represent different traditional Chinese medicines, and the blue diamond represents 258 common drug disease targets (Figure 2). Cytoscape was used to calculate the degree values of each node, representing the number of correlations between the constituent and the molecular target. The higher the degree value, the greater the likelihood of the constituent's interaction with the disease. Through network analysis, the 10 constituents with the highest degree values were selected. The specific constituents, names, and rankings are shown in Table 2.

Herb-ingredient-target network. Circles represent herbs and ingredients, while diamonds represent targets. A, B, and C are classification numbers for common constituents of medicinal herbs.
Top 10 Active Constituents in Degree Values.

GO and KEGG Enrichment Analysis of Potential Targets
The GO function obtained after filtering with the DAVID database is shown in Figure 3A. In terms of biological processes, these targets mainly involve response to xenobiotic stimulus, negative regulation of apoptotic processes, response to hypoxia, and inflammatory response. In terms of cell composition, plasma membrane, cell surface, integral component of plasma membrane, and receptor complex are represented. From a functional molecular perspective, these molecular targets mainly participate in functions such as protein serine/threonine/tyrosine kinase activity, protein tyrosine kinase activity, and identical protein binding.
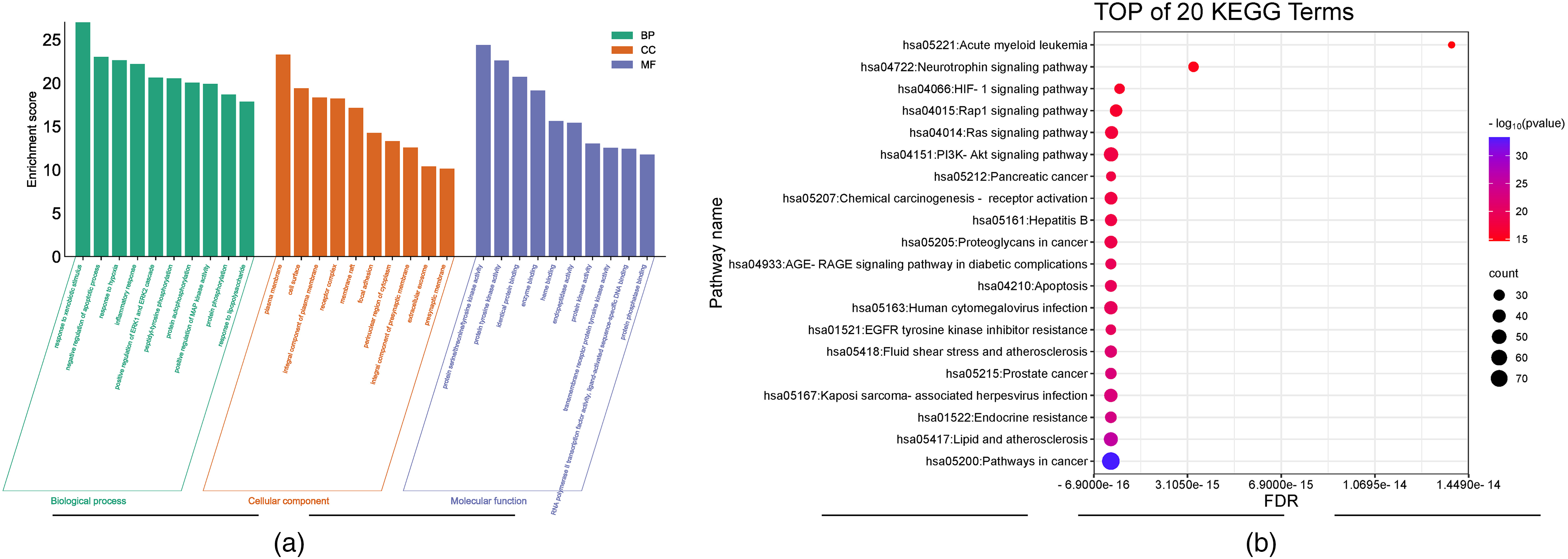
(A) Bar charts of GO enrichment analysis in the top 10 rankings. (B) Bubble chart analysis of the top 20 pathways of intersection target KEGG.
The first 20 pathways obtained from KEGG enrichment analysis by importing common targets into the DAVID database are plotted as bubble plots in Figure 3B. The target is significantly enriched in multiple pathways such as cancer, lipid and atmospheric clarity, endocrine resistance, PI3K-AKT signaling pathway, fluid shear stress, and atmospheric clarity.
Constituent-Target-Pathway Network Analysis
Based on the 258 common targets and 20 pathways screened through KEGG enrichment, a constituent-target-pathway diagram was constructed using Cytoscape software (Figure 4). The targets were sorted and plotted according to the degree value, and core targets such as GSK3B, PIK3CB, SRC, CSF1R, and FLT1 were selected from the 20 pathways. These may be important targets for YGZ223 to exert therapeutic effects through the first 20 pathways.

Constituent-target-pathway network. Circles represent constituents, squares represent targets.
UPLC-MS/MS Results
Using MassLynx V4.1 software for peak extraction from raw mass spectrometry data, the mass deviation of the feature peak, element matching, molecular formula prediction, and isotope distribution matching is set within 10 ppm. After extracting the secondary mass spectrometry, the retention time and secondary mass spectrometry information of each compound were analyzed using MassLynx V4.1 software. The chromatographic behavior was compared with relevant references, the MassBank database, and reference materials.
Chemical Composition Analysis of YGZ223
According to the UPLC-MS results, 91 chemical constituents were ultimately identified. Most were detected in positive ion mode, some in negative ion mode, and some in both, such as paeoniflorin and paeoniflorin. The identified compounds include flavonoids, alkaloids, terpenoids, and others classes, among which up to 30 alkaloid compounds are mainly derived from Sophorae flavescentis Radix and Cortex erythrie. Among the top 10 compounds screened by network pharmacology, phaseolin, formononetin, isoxanthohumol, erysodienone, and (-)-estetol were identified. Of these, formononetin and erysodienone had the highest ionic strengths, as shown in Table 3 and Figure 5.

YGZ223 UPLC-MS/MS total ion flow diagram: (A) positive ion mode (upper: sample; lower: blank control); and (B) negative ion mode (upper: sample; lower: blank control).
UPLC-MS/MS Identification and Analysis of YGZ223 Related Constituents.

Fragmentation and Structural Analysis of Main Constituents
Compounds
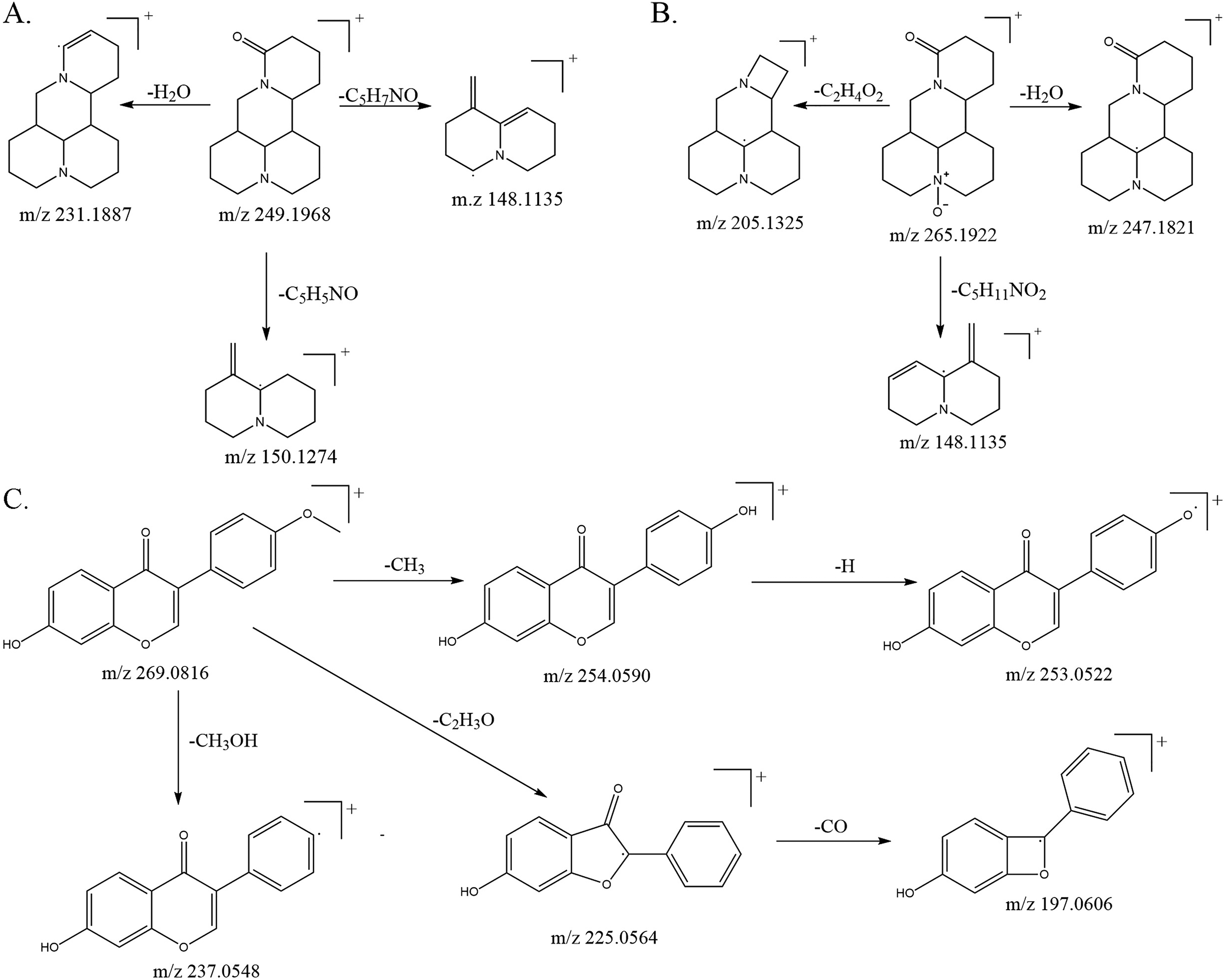
Pyrolysis mechanisms in positive ion mode for: (A) matrine; (B) oxymatrine; and (C) formononetin.
Compound
Compound

Fragmentation mechanism of albiflorin in positive ion mode.
Compound

Fragmentation mechanism of hecogenin in positive ion mode.
Molecular Docking Result
To determine the viability of chemical constituent binding to target proteins, AutoDock Vina software was used to screen the top 5 proteins from the network pharmacology, and to dock the top 2 ion strength compounds detected by UPLC-MS. Of the proteins, PI3 K is a complex, so the 3 subunits PIK3CA, PIK3CB, and PIK3CD were docked separately. The binding energies of the 5 proteins with the 2 compounds are all less than −5.0 Kcal· mol−1, indicating good binding ability (Table 4). PyMOL software was then used to visualize the docking results, some of which are shown in Figure 9. Formononetin has the lowest binding energy with AKT1, indicating the strongest binding force. The receptor ligand interaction site is located at amino acid residues ILE-290 and THR-211, which are hydrogen bonded. The region where these 2 residues are located may be the key region of action.

Molecular docking diagrams of formononetin and erysodienone with related targets (binding energy below −7 Kcal· mol−1): (A) formononetin and EGFR; (B) formononetin and AKT1; (C) formononetin and GSK3β; (D) formononetin and mTOR; (E) erysodienone and AKT1; and (F) erysodienone and GSK3β.
Docking Binding Energies of Formononetin and Erysodienone With Various Targets.

Molecular Dynamics Simulation Results
The RMSD results show that the overall fluctuations between 0-20 and 50-80 ns are relatively small and stable (Figure 10A). The fluctuations between 20 and 50 ns are caused by conformational adjustments of small molecules in the pocket, which did not escape the protein during this period, remaining bound. The overall RMSF of the protein is at a relatively low level (Figure 10B). The N- and C-termini of the protein are less restricted by other peptide segments and have greater flexibility, resulting in significant fluctuations at both ends in the figure. The RMSF values around Res100-150 are relatively high since this peptide segment belongs to a random curled secondary structure with greater flexibility caused by bending under the influence of the surrounding environment. However, it is distant from the small molecule binding region, so it will not have a significant impact on the docking. Radius of gyration (Rg) characterizes protein structure compactness and changes in the degree of peptide chain looseness during the simulation process. The Rg value fluctuated to some extent during the first 10 ns of simulation, showing an overall downward trend. After 10 ns, the Rg value decreased to around 2.5 nm, remaining vibrationally stable throughout the following 70 ns, with reduced fluctuations. Hence, the protein can be considered to remain stable during this period (Figure 10C). The HBond results showed that during the 0 to80 ns period, a certain number of hydrogen bonds were maintained between small molecules and proteins, and there was no absence of hydrogen bonds or detachment from protein binding cavities, indicating relatively stable binding (Figure 10D).

Molecular dynamics simulation of various analysis indicators: (A) RMSD; (B) RMSF; (C) Rg; and (D) Hbond.
After periodic elimination of the simulated trajectory, the conformations of the formononetin-AKT1 complex at 0 and 80 ns were extracted and compared for overlap. In Figure 11A, the green part represents the initial structure at 0 ns, while the blue part represents the simulated final structure at 80 ns. Comparison of the 2 shows that the flexible side chains of the protein undergo distorted conformational changes after simulation. At the same time, there are also changes in the conformation and position of the small molecules. Based on the analysis of RMSD and Rg, it can be concluded that the 80 ns composite is relatively stable. Hence, the interaction force was analyzed between small molecules and the proteins at 80 ns. Formononetin formed salt bridge interactions with residues Asp292, Thr291, and Met281, and hydrogen bond and thiol residue Π-bond interactions. The binding free energy (MM/GBSA) analysis of the trajectory of the formononetin-AKT1 complex during the 50 to 80 ns period showed an overall binding energy of −20.65 kcal/mol, indicating a strong tendency for small molecules to bind to the proteins with a stable binding. Based on MM/GBSA, the key amino acids that interact with the small molecules during the simulation process were analyzed. Among these, Lys158, Gly159, Val164, Lys179, Lys276, Glu278, Asn279, Asp292, Thr312, and Tyr315 are key amino acids that mainly interact with the small molecules during the stable 50 to 80 ns period (Figure 11B).

Structure diagram of (A) 0 ns (green) and 80 ns (blue) composite; and (B) energy of each residue at 80 ns.
YGZ223 Pharmacodynamic Test Results
No abnormalities were observed in any mice during the experiment, the weight of each group of animals showed a stable growth trend, and no abnormal weight phenomena were observed. During monitoring of the pruritus response, the number of licks in the Model control group after injection of 4-AP significantly increased (P < .01) compared with the blank Control group. Compared with the Model group, the DF group and YGZ223 group significantly reduced the number of 4-AP-induced licks (P < .05), while the other groups of mice showed no statistical difference (P > .05). Compared to the Model control, YGZ223 and DF groups showed a significant decrease in scratching frequency (P < .05), while there was no significant difference between the XFZY group and the Model control group. The results are shown in Table 5 and Figure 12. The efficacy of YGZ223 is somewhat superior to that of DF, yet the two are not significantly different.

Number of tickles induced by 4-AP in each group of mice: * P < .05, ## P < .01.
Number of Tickles Results in Mouse Tickles Test (n = 8; unit: times; X ± SD).

Significantly different from the blank control (P <.01).
Significantly different from the model comparison (P <.05).
Discussion
A total of 91 compounds were identified in YGZ223 using UPLC-MS/MS. The majority of these were alkaloids, such as matrine found in Sophorae Flavescentis Radix, erythridine in Cortex erythrie, and alkaloids lotusine, and harmine. While flavonoids, terpenoids, and phenylpropanoids were also present, they were relatively uncommon. The specific constituents of Radix Paeoniae Rubra, namely paeoniflorin and albiflorin, showed a high level of ion strength. Matrine, sophocarpine, and oxymatrine possess various pharmacological activities such as antibacterial, antiviral, anti-tumor, and protective effects on internal organs and blood vessels. 42 Erysodine is a competitive antagonist of neuronal nicotinic acetylcholine receptors, 43 and it exhibits a DPPH free radical scavenging effect. 44 Furthermore, flavonoids exhibit anti-inflammatory, liver protective, antioxidant, and antibacterial effects associated with skin pruritus. 45
Data mining revealed that, among the top 10 active compounds, formononetin and erysodienone had high ionic strength in the UPLC-MS assay, indicating that their content is not low and that they may be key constituents for exerting drug efficacy. Among the intersection targets obtained through screening, 2 potential targets for formononetin appeared in the top 20 key targets, namely EGFR and ESR1. The key targets that erysodienone may act on are EGFR, FRC, PTGS2, and mTOR. Further analysis and screening was performed on the top 20 pathways obtained from the KEGG analysis, finding that EGFR and mTOR were involved in the multiple pathways: pancreatic cancer, endocrine resistance, PI3K-AKT signaling pathway et al. Therefore, it is speculated that EGFR and mTOR are key targets for drug efficacy. Among the 10 GO functions, protein autophysiography, identity protein binding, protein kinase activity and negative regulation of apoptotic process are functions shared by the 2 compounds. The endocrine resistance pathway is the pathway of highest score in the KEGG analysis, and the PI3K-AKT signaling pathway also plays a role in endocrine resistance. EGFR acts on PI3 K and further affects the PI3K-AKT pathway, thereby affecting the downstream protein mTOR. The AKT1 with the highest score in the PPI analysis happens to be on this pathway, so EGFR-PI3K-AKT-mTOR may be a key pathway for the action of formononetin and erysodienone.
EGFR is a tyrosine kinase involved in cell proliferation, mitosis, and carcinogenesis. 46 EGF binds to EGFR and activates it through a series of protein interactions, thereby affecting downstream signaling cascades such as PI3K-AKT. 47 EGFR is currently the therapeutic target of first-line anti-tumor drugs, which often have significant side effects such as pruritus, rash, and hypertension. Significant thinning of the epidermal stratum corneum has been observed in patients treated with EGFR inhibitors. 48 EGFR phosphorylation and MAPK activation were inhibited, resulting in a strong inflammatory response, 49 further inhibiting downstream signals such as the PI3K-AKT pathway and STATs. TGF-α/EGFR signaling defects are associated with skin barrier damage. 50 Studies have shown that the skin of AKT1/AKT2 knockout mice is semi-transparent, similar to the skin condition of elderly people, and that mTOR activity in AKT1/AKT2 cells is inhibited. 51 A decrease in AKT1 activity has been observed in skin with specific dermatitis, 52 which can cause itching. GFR1 knockout mice also have the same skin damage condition, 53 and IGFR1 can bind to EGFR, activate the downstream PI3K-AKT pathway, and then activate mTOR to maintain the skin condition. Overall, the proteins EGFR, AKT1 and mTOR may play a key role in alleviating senile pruritus. Analysis of the constituent-target-pathway diagram identified GSK3B and PIK3B as the 2 target genes with highest scores. GKS3B encodes the protein GSK3β and PIL3CB encodes the production of p110β, one of the catalytic subunits of PI3 K involved in the regulation of the PI3K-AKT-mTOR pathway. Activation of the PI3K-AKT-mTOR pathway can phosphorylate GSK3β and inhibit its activity, 54 and studies have shown that GSK3B inhibition plays a critical role in skin wound healing. 55 Analysis of the effects of these targets provides further evidence that EGFR, PI3 K, GSK3β, AKT1 and mTOR may be the targets for YGZ223 to exert therapeutic effects in SP.
The molecular docking results showed that formononetin had low binding energy with various proteins, the smallest with AKT1, indicating the strongest binding ability with AKT1 and relatively weak binding ability with other targets. Compared with formononetin, erysodienone has significant binding force with various targets, and a better binding with mTOR. According to the docking results, formononetin is a key compound and AKT1 may be a key target.
In-vivo pharmacological tests showed that YGZ223 has a good anti-pruritic effect in the 4-AP-induced mouse pruritus model, and that the effect is superior to the positive control drug.
This study is a preliminary exploratory study, and experimental verification has not yet been conducted on the screened compounds and corresponding targets. Subsequent experimental verification is required. In-vivo experiments have confirmed the effectiveness of YGZ223, but the 4-AP-induced itching model has limitations and is different from SP, which may be related to the complex pathogenesis of SP. Consequently, in the future, we will consider utilizing in-vitro experiments or other more appropriate animal models for the validation of drug efficacy and mechanisms.
Conclusion
This study is based on network pharmacology and explores the possible mechanism of YGZ223 in the treatment of SP. Compounds with the top 10 scores and targets and pathways with the top 20 scores were screened. Formononetin and erysodienone are potentially key compounds, EGFR, PI3 K, AKT1, GSK3β and mTOR may be key targets, and EGFR-PI3K-mTOR may be a key pathway. Molecular docking shows that both compounds have some binding ability to the target, while formononetin and AKT1 may be more critical. In parallel, pharmacological experiments were conducted, indicating that YGZ223 can alleviate 4-AP-induced itching in mice. The above research provides the basis for further development of YGZ223.
Footnotes
Acknowledgments
We are grateful to Professor Liu Bo for his support of this research, and to the other authors for their assistance at various stages of this study.
Author Contributions
Bo Liu and Sa Ling conceived and designed the whole experiment. Xiankui Shu, Ling Ye, and Shiqi Han participated in and completed all experiments. Xiankui Shu, Ling Ye, and Shiqi Han wrote the manuscript. Lu Liang, Hancheng Song, Shuang Xue, and Yilang Zhao proposed modification suggestions. Bo Liu and Sa Ling reviewed and edited this manuscript. All the authors read and approved final version of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Ethics Approval
This study has been approved and recognized by the Experimental Animal Management and Use Committee (IACUC) of Hubei Tianqin Biotechnology Research Institute Co., Ltd. Animal Ethics Number: IACUC (Approved) - YJY-2022-016.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
