Abstract
Keywords
Introduction
Tumor-related morbidity and mortality are increasing annually, endangering human life and health. For example, in 2020, the International Agency for Research on Cancer reported 19.3 million new cancer cases and nearly 10 million deaths worldwide, of which colorectal cancer (CRC), a digestive system cancer, accounted for approximately 9.4% of the deaths, ranking second. 1 Additionally, the agency estimates that by 2030, CRC will kill approximately 1.1 million people worldwide. Moreover, disease onset is insidious; the early stage is not easy to detect, and there are no typical symptoms. Therefore, clinical diagnoses are often made in the middle and late stages. Consequently, the five-year relative survival rate for patients with middle- and advanced-stage CRC is only 14% despite improved preventive screening methods since most cases are not detected until symptoms have developed.2,3
CRC poses a serious threat to public health and increases the social burden. General CRC risk factors include excessive drinking, obesity, gene mutations, and chronic intestinal inflammation. 4 The primary treatment methods are surgery, chemotherapy, radiotherapy, and immunotherapy, with surgery as the gold standard treatment. However, surgical treatment comes with high intraoperative risk. Furthermore, the postoperative recurrence rate is high, and the postoperative survival rate is low. Auxiliary treatments, such as radiotherapy, chemotherapy, targeted therapy, and immunotherapy, can improve the long-term prognosis but have adverse effects, including vomiting, appetite loss, leukopenia, thrombocytopenia, hand-foot syndrome, and neurotoxicity. Therefore, auxiliary treatments have not significantly improved the prognosis of patients with CRC.5,6 As a result, there is no ideal diagnostic or treatment plan for CRC, and new therapeutic methods are needed. Compared with chemical drugs, traditional Chinese medicines (TCMs) have multiple components and thus targets, affect several pathways, and have minimal adverse reactions 7 ; thus, they are frequently used as a cancer treatment.
TCMs have been used for thousands of years in China and form a unique and comprehensive theoretical framework. Guided by holistic concepts, TCM focuses on the body's ability to resist evil, which enhances the effectiveness and reduces the toxicity of the medicine. TCMs have been demonstrated to be safe, minimally toxic, well-tolerated, and have few adverse reactions, suggesting that TCM's role in cancer treatment cannot be underestimated.8,9 In other words, TCM treatments can complement modern medicine strategies, such as surgery, radiation, and chemotherapy. In TCM, there is no technical term for CRC, but the terms “mushroom,” “lump of the intestine,” and “caked disease,” among others, exist. These descriptions are similar to the clinical manifestations of CRC, such as abscesses and bloody mucous stool, changes in stool characteristics, stool-related difficulties, and anal swelling.
Compound Scutellariae Radix Granules (CSRG) contain the following TCMs: Fructus Jujubae (FJ) and licorice (LI) (oning the spleen and stomach, relieving pain), Paeoniae Radix Alba (PRA) (acidic taste, collects Yin and camp pain relief), and Scutellariae Radix (SR) (bitter cold, clear heat). The herbs are used together to clear heat and dysentery and relieve pain. Primarily, CSRG are used to treat intestinal ulcers and pus. However, Chinese medical master Liu Shangyi creatively suggested diagnosis and treatment ideas, such as “introducing ulcers into tumors” and “treating tumors from membranes;” this bold approach of selecting the department for diagnosis and treating tumors has achieved good clinical effects. Consequently, the academic thought of “choosing a rational diagnosis for the tumor,” “choosing a legal tumor,” and “choosing the drug therapy” was created. 10 Studies have shown that as the most commonly used of the herbs in China to treat cancer, SR and PRA have shown high anti-tumor efficacy and safety. Tumor cell apoptosis was induced, the cell cycle was arrested, proliferation was inhibited, invasion and metastasis were blocked, and the development of drug resistance was delayed, through different molecular mechanisms, It mainly involves the nuclear transcription factor NF-κB family, microRNAs, p53 signaling pathway and so on.11,12 JF has been consumed as a fruit in China for thousands of years, and is also the representatives of homology of medicine and food, It is included in many formulas used for the treatment of CRC. Studies have demonstrated that jujube triterpenes can be used to fight CRC through the PI3K/Akt/NF-κB signaling pathway. 13 LI is a well-known natural herb, and the main component of its extract is triterpenoids, the role in tumor prevention and therapy by participating in the regulation of tumor cell death, oxidative stress and inflammation. 14 CSRG contain SR:PRA:LI:FJ (3:3:2:4, w/w/w/w). The mixture was decocted twice with water: first with 12 times the amount of water and then with ten times the amount of water, with each decoction lasting for 1 hour. The filtrate was concentrated to a suitable amount (ρ = 1.03-1.08 g/cm−3, 60 °C), mixed well with dextrin and sucrose, and dried into granules.
The composition of CSRG is complex, and thus its anti-tumor mechanisms, particularly those related to CRC, have not been fully elucidated. Therefore, we performed a network pharmacological analysis to identify the active CSRG ingredients involved in CRC treatment. We conducted cytology experiments to identify potential therapeutic targets and explore potential signaling pathways to systematically understand the correlation between CSRG and CRC and provide new evidence for future research and clinical applications.
Materials and Methods
Schematic Diagram
Figure 1 shows a schematic representation of the study design and workflow. We used network pharmacology and molecular docking techniques to determine the prevention and treatment mechanisms and the material basis of CSRG in CRC.
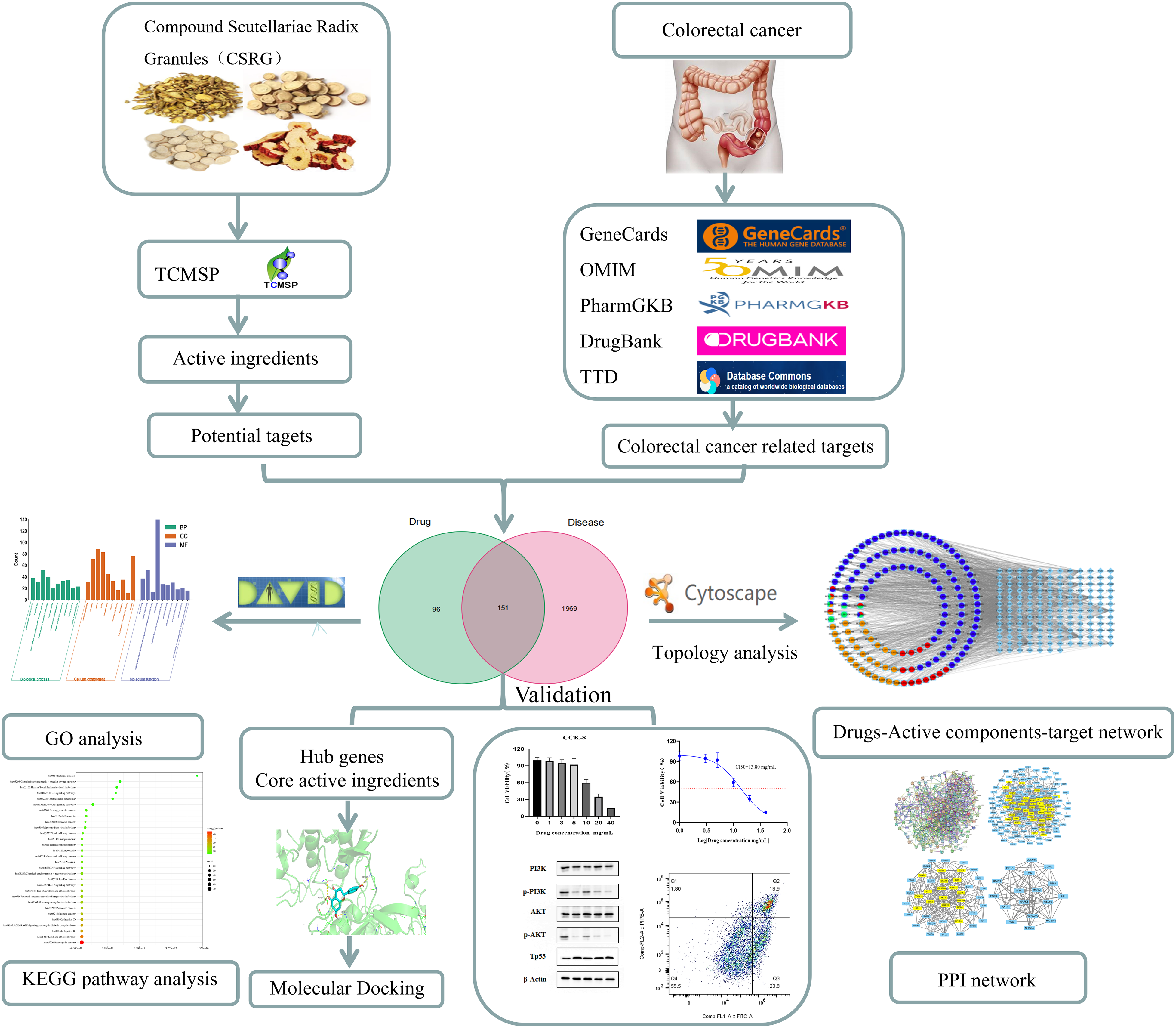
Work flowchart.
Screening of the Main Active Ingredients and CSRG Target
The Latin name of each herb in the CSRG was input into the TCMSP database. Oral bioavailability (OB) ≥ 30% and drug-like drug (DL) ≥ 0.18, important indexes for determining pharmacokinetic absorption, distribution, metabolism and excretion (ADME), were used as the criteria to screen potential active substances. Potential active ingredients and action targets of CSRG were obtained from the TCMSP database simultaneously, and information on the major chemical components was collected and screened from the PubChem database.
CRC Disease Target Collection
We used “colorectal cancer” as the key search term in the following databases: GeneCards (filter criteria: >10 score), OMIM, PharmGKB, Pharmacogenetics and Pharmacogenomics Knowledge Base, DrugBank, TTD. A target corresponding to CRC was searched and duplicate targets were deleted to obtain the target corresponding to the disease.
Identification of the Potential Target of CSRG in the Treatment of CRC
We used the online tool Venny2.1.0 (https://bioinfogp.cnb.csic.es/tools/venny/) to draw a Venn diagram to obtain targets of CSRG ingredients and CRC. The potential target of CSRG for CRC was obtained.
Construction of Protein–Protein Interaction (PPI) Network
A protein interaction network was constructed using the STRING database (https://cn.string-db.org/). In the Multiple proteins module, the common target of the active ingredient of CSRG and CRC was introduced, Homo sapiens was selected as the target species, and the minimum required interaction score was set to ≥0.9. Then, uncorrelated nodes in the network were hidden, the remaining parameters were set as default, and the string_interaction_short.tsv file was exported. The string_interaction_short.tsv file was imported into Cytoscape 3.8.0 for visualization of PPI results, and NetworkAnalyzer was used to analyze node degree values. The more the nodes are associated with each other, the greater the degree value is, and the greater the significance of the node in the network.
Construction of the Interaction Network Between Active Ingredients of CSRG and CRC Targets
Active ingredients and intersection target genes of CSRG were introduced into Cytoscape3.8.0 software to construct a visual interaction network of “herbs-active ingredient-target.”
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Enrichment Analyses
To further explore the potential role of CSRG target genes in the treatment of CRC and their role in the signaling pathway, we imported common target genes from DAVID (https://david.ncifcrf.gov/) for comments, visualization, and integration analysis. The species was restricted to Homo sapiens (screening criteria P-value: P < 0.05). Enrichment analyses of GO and KEGG pathways were performed. Pathways were screened based on the number of enrichment targets to obtain more important pathways. Using the bioinformatics cloud platform (http://www.bioinformatics.com.cn/), GO and visualization for the second-order classification histogram KEGG enrichment and visualization as a bubble chart analysis were performed.
Molecular Docking
AutoDock Vina software was used to verify the molecular docking between the core targets and key components. First, the 3D structures of key components were obtained from the PubChem database, crystal structures of core target proteins were downloaded from the PDB database, and operations such as dewatering and delignification were performed on key targets using PyMOL software. AutoDockTools was used for hydrogenation and charge processing and AutoDockVina was used for molecular docking. Visualization was performed using PyMOL, and the docking effect was evaluated based on binding energy.
In Vitro Cell Experiments
Materials
CSRG were produced by Anhui Golden Sun Biochemical Pharmaceutical Co. LTD (Fuyang, Anhui, China). The four herbs in the recipe were prepared by water extraction, so the granules could be used for the subsequent experiments. Cell Counting Kit-8 and AnnexinV-FITC/propidium iodide (PI) reagent were supplied by Bomei Biotechnology; BCA protein concentration determination kit and Hypersensitive ECL Chemiluminescence Kit were supplied by Biosharp; Protein Marker was supplied by Beyotime Biotechnology; p-Akt, Akt, p-PI3K, and PI3K primary antibodies, and β-Actin and Tp53 were purchased from CST (USA). All of the above five primary antibodies were rabbit antibodies. HRP-labeled Sheep Anti-Rabbit IgG and murine β-Actin antibody secondary antibodies were provided by Servicebio. IGF-I was purchased from Sigma-Aldrich (USA).
Experimental Cells
Human colon cancer HT-29 cells were purchased from ATCC Company and cultured in Dulbecco's Modified Eagle Medium (DMEM) with high glucose medium supplemented with 100 mg/L fetal bovine serum and 10 g/L penicillin and streptomycin double antibody. The cells were incubated at 37 °C in a 5% CO2 incubator.
Cell Viability Assay
HT-29 cells were cultured in an incubator at a constant temperature of 37 °C with 5% CO2 to logarithmic growth phase. Cell density was adjusted to 5 × 104 cells/mL, the cells were inoculated in 96-well plates. Then, 100 μL of cell suspension was added to each well and the plate was placed in a 37 °C 5% CO2 incubator. HT29 cells were digested and counted using a counting plate. The concentration of the cell suspension was 5 × 104 cells/mL and the cells were inoculated in 96-well plates after sufficient suspension. When the cell density reached 60% to 80%, the culture medium was aspirated and culture media containing different drugs, including CSRG and IGF-I, were added. 15 Six replicate wells in each group were set up after incubation for 48 h. Then 10 μL of CCK-8 solution was added (avoiding light) to each well, and the plate was placed in the incubator for 1 h at 37°C. Absorbance (A) was measured at 450 nm using an enzyme-labelling instrument as follows: Cell viability = (A value (drug addition) - A value (blank))/(A (0 drug addition) - A value (blank)) × 100%.
Cell Apoptosis Analysis
HT29 cells were inoculated in a 6-well plate at 2 × 105 cells per well and cultured in complete culture medium. When the cell density reached 60% to 80%, the culture medium in the wells was aspirated and 2 mL of DMEM culture solution was added to each well. The cells were divided into groups. 15 and placed at 37°C. The cells were cultured in an incubator with 5% CO2 for 24 h. The cells were collected in 10 mL EP tubes, centrifuged at 1500 r/min for 10 min and the supernatant was discarded. The cells were resuspended by adding 200 μl Annexin V conjugate per tube, 3 μl Annexin V-FITC per tube, and incubated on ice for 15 min. Then 5 μl of PI staining solution was added to each tube and the cells were incubated on ice for 10 min. Next, 5 μL PI staining solution was added to each tube and mixed, and then the cells were incubated on ice for 10 min. Finally, 200 μL Annexin V conjugate was added to each tube and mixed well. The cells were filtered through gauze into the flow tube, and the samples were detected by flow cytometry within 1 h.
Western Blotting Experiment
Human CRC cells HT-29 in the logarithmic growth phase were collected and inoculated in 6 cm dishes at a density of 3 × 105 cells per dish. After 24 h the cells were divided into five groups: the control group, the positive drug group (5-FU), the CSRG-L group, the CSRG-M group, and the CSRG-H group. The cells from each group were collected after 48 h and washed with PBS. Next, 30 μL of cell lysate was added, at 4 °C for 30 min, and centrifuged at 4 °C, 12,000 r/min for 15 min. The supernatant was extracted and the protein was quantified by the BCA method. Next, 10% SDS-PAGE electrophoresis was performed, the separated proteins were transferred to a membrane. The membrane was sealed with skimmed milk powder for 1.5 h and the primary antibodies Akt, p-Akt (1:500), PI3K, p-PI3K (1:1000) and β-actin (1:1000) were added and incubated overnight at 4 °C. The membrane was washed thrice with 1× TBST for 5 min each time. Secondary antibodies were added and incubated at room temperature for 2 h and the membrane was then washed thrice with 1× TBST for 10 min each time. Following membrane exposure in a dark room, the protein bands were analyzed by ImageJ software. Protein expression was calculated as follows: expression of target protein = gray value of target protein/gray value of internal reference protein.
Statistical Analysis
Statistical analysis was conducted using GraphPad Prism 8.0 (ANOVA). The results are presented as the mean ± SD (standard deviation). Differences among the groups were analyzed. A P-value <0.05 was considered statistically significant.
Results
Active CSRG Constituents and Targets
In total, 170 active compounds were obtained from the Traditional Chinese Medicine Systems database, including 29 from FJ, 92 from LI, 13 from PRA, and 36 from SR. After removing the repeats, 247 targets were obtained. The primary core active substances were naringenin, quercetin, licochalcone A, acacetin, wogonin, baicalein, kaempferol, and isorhamnetin (Table 1).
Core Active Compounds of CSRG.

Potential Targets of CSRG in CRC
In total, 1838 targets were obtained from the GeneCards database, 102 from the Therapeutic Target Database, 410 from the Online Mendelian Inheritance in Man database, 46 from the DrugBank database, and 285 from the Pharmacogenomics Knowledge Base database.
Overall, 2681 disease targets were identified (Figure 2A); 2120 remained after removing the duplicates. Targets of the potentially active components of CSRG crossed with genes related to CRC were the potential targets for CSRG resistance; we identified 151 cross-targets (Figure 2B), constituting the potential targets of CSRG for CRC treatment.

Venn diagrams of (A) the disease targets in colorectal cancer in various databases and (B) compound Scutellariae Radix Granules (ie, drug) targets with disease (ie, colorectal cancer) targets. Abbreviations: OMIM, Online Mendelian Inheritance in Man; PharmGKB, Pharmacogenomics Knowledge Base; TTD, Therapeutic Target Database.
Protein–Protein Interaction (PPI) Network Analysis and Core Target Identification
The 151 intersecting genes were uploaded to the Search Tool for the Retrieval of Interacting Genes/Proteins (ie, STRING) database; with a minimum required interaction score of 0.9, 773 edges and 151 nodes were identified (Figure 3A). Cytoscape 3.8.0 software was used to establish the protein interaction network of common targets, analyze the degree value of each node, and determine the core target based on its ranking(Figure 3B). 16 The top five target genes (Table 2) were screened based on their degree values; these genes played crucial roles in the PPI network. Thus, the active components in CSRG regulate the key CRC targets, which was verified by molecular docking.

Protein–protein interaction (PPI) network topology analysis. (A) Target PPIs. (B) First 17 major target genes are obtained after a step-by-step analysis of the topological network. The more the nodes are connected, the greater the degree value. Abbreviations: BC, Betweenness centrality; CC, Closeness centrality; DC, Degree centrality; EC, Eigenvector centrality; LAC, Local average connectivity-based method; NC, Network centrality.
Top Five Target Genes Related to Colorectal Cancer.

Abbreviations: MAPK3, mitogen-activated protein kinase 3, MAPK1, mitogen-activated protein kinase; TP53, tumor protein 53; STAT3, signal transducer and activator of transcription 3; RELA, RELA proto-oncogene, NF-kB subunit; BC, betweenness centrality; CC, closeness centrality; DC: degree centrality; EC, eigenvector centrality; LAC: local average connectivity-based method; NC: network centrality.
Herb-Active Ingredient–CRC Target Interaction Network of CSRG
Cytoscape_v3.8.0 software was used to construct the herb-active ingredient-target interaction network of CSRG based on the 151 target genes and 170 active components. The network consisted of 285 nodes and 1550 edges (Figure 4). The greater the number of connections between the components and target proteins, the higher the degree value, indicating a closer connection between the nodes.

Compound Scutellariae Radix Granules (herb)–active ingredient–potential colorectal cancer target network. Round orange, green, blue, and red circles represent active compounds derived from licorice (LI), Fructus Jujubae (FJ), Scutellariae Radix (SR), and Paeoniae Radix Alba (PRA), respectively. Blue diamonds indicate the target genes for the interaction between Compound Scutellariae Radix Granules (CSRG) and colorectal cancer (CRC).
Enrichment Analysis
To further elucidate the CSRG mechanisms in CRC, we performed GO and KEGG enrichment analyses on 161 intersecting genes. The numbers of GO annotated terms associated with biological processes, cellular components, and molecular function were 805, 78, and 170, respectively (P < 0.05). The top ten entries were visualized and analyzed separately (Figure 5A). The target genes were involved in the positive regulation of gene expression, response to drugs, positive regulation of transcription from RNA polymerase II promoter, positive regulation of transcription, positive regulation of apoptotic processes, negative regulation of apoptotic processes, apoptotic processes, response to hypoxia, nucleoplasm, nucleus, cytosol, extracellular space, chromatin, transcription factor complex, RNA polymerase II transcription factor complex, cytoplasm, RNA polymerase II transcription factor activity, protein binding, transcription factor activity, transcription factor binding, and RNA polymerase II sequence-specific DNA binding transcription factor binding, among others.
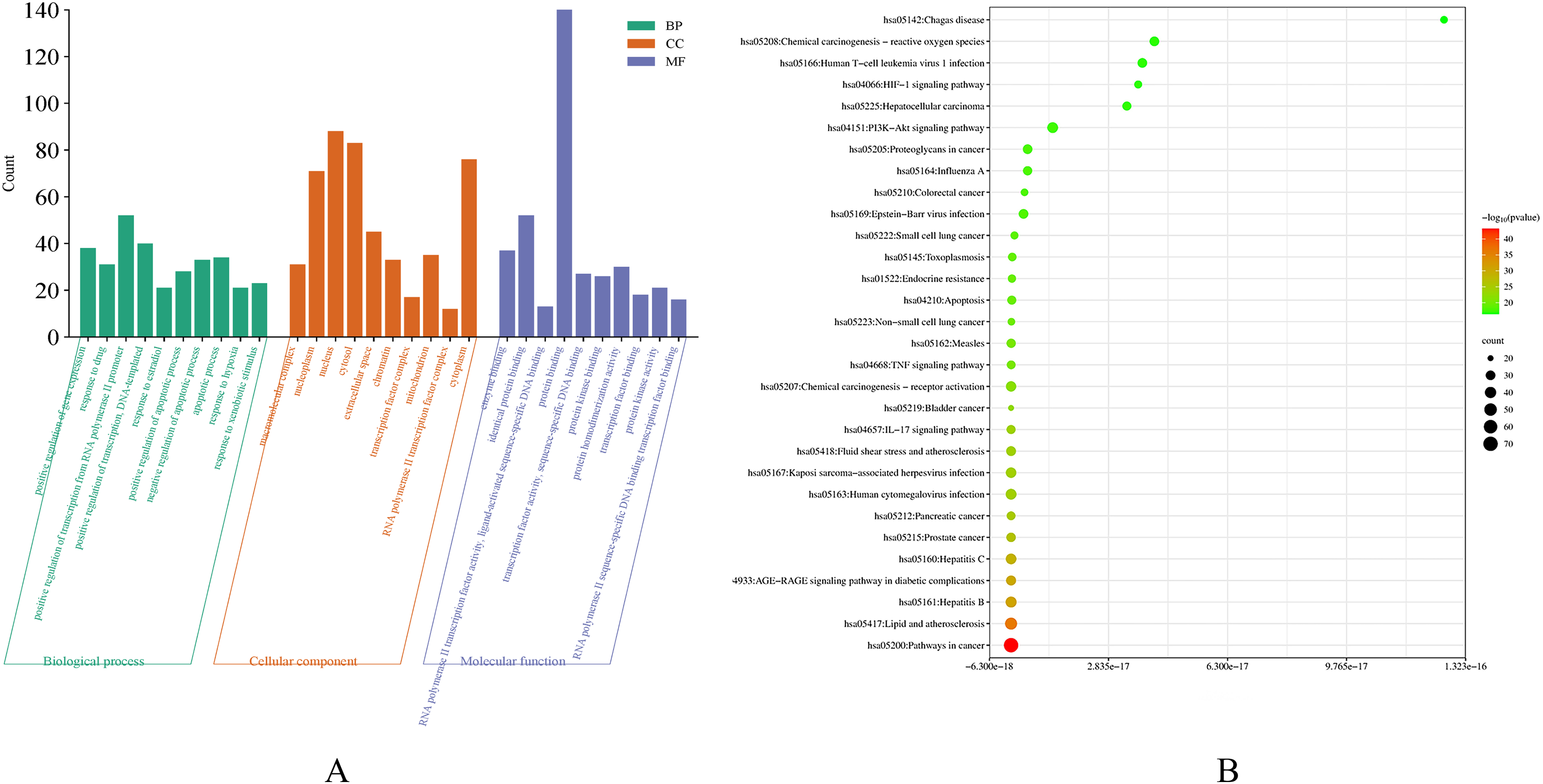
Visualizations of the key (A) Gene Ontology (GO) and (B) Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis results based on P-values. (A) The top 10 biological process (BP), cellular component (CC), and molecular function (MF) components. The bar graph's height indicates enrichment level of the target. (B) The top 30 KEGG signaling pathways. As the P-values increase, the colors of the circles change from green to red, and their sizes change from small to large, corresponding to the number of genes moving from small to large.
Figure 5B illustrates the top 30 of 178 important pathways linked to CRC in the KEGG analysis. Several were strongly associated with CRC and could be the primary targets of CSRG for CRC treatment, including the phosphoinositide-3-kinase/protein kinase B (PI3K/Akt), tumor necrosis factor (TNF), interleukin (IL)-17, and hypoxia-inducible factor 1 (HIF-1) signaling pathways. Figure 6 presents the associated target genes of the PI3K/Akt signaling pathway in detail.

Key targets of the phosphoinositide-3-kinase/protein kinase B (ie, PI3K/Akt) signaling cascade. Red represents Compound Scutellariae Radix Granules (CSRG) targets.
Molecular Docking
The top five target genes in the PPI core network were sorted and screened using degree values for molecular docking with core components. The more stable the binding conformation, the lower the binding energy required. Binding energy values of ≤0 kcal/mol indicated no binding activity, values of ≤–5.0 kcal/mol indicated excellent binding activity, and values of ≤–7.0 kcal/mol indicated strong binding activity. 17 Molecular docking was performed on the top five target genes and their corresponding compounds; Table 3 presents the results. PyMOL 2.4.0 was used to draw a three-dimensional docking diagram of the results. 18
Molecular Docking Results Displaying the Binding Energy Between the Main Core Targets and Correlated Compounds.

Effects of CSRG on HT-29 Cells
To elucidate the effect of CSRG on cancer HT-29 cells, we chose the CCK-8 assay to detect the cell viability of HT29 cells after treatment with different concentrations of CSRG and a PI3K/AKT agonist. The results showed that 0 to 40 mg/mL CSRG inhibited the proliferation of HT-29 cells in a dose-dependent manner. Compared with that in the 20 mg/mL CSRG group, combined treatment with 100 ng/mL IGF-I and 20 mg/mL CSRG significantly increased HT-29 cells proliferation, and the cell survival rates of the two groups at the same time point were significantly different. The IC50 values of HT-29 cells were 13.80 mg/mL at 48 h after CSRG treatment (Figure 7A). Three concentrations (5, 10, and 20 mg/mL) of CSRG and combined treatments (100 ng/mL IGF-I and 20 mg/mL CSRG-H) were selected for subsequent experiments. Annexin V-FITC/PI double staining and flow cytometric art analysis showed that all concentrations of CSRG above 5 mg/mL induced apoptosis to some extent in a concentration-dependent manner. Compared with that in the 20 mg/mL CSRG group, combined treatment (100 ng/mL IGF-I, 20 mg/mL CSRG) significantly reduced the rate of apoptosis in HT-29 cells, and the cell apoptosis rates of the two groups at the same time point were significantly different (Figure 7C).
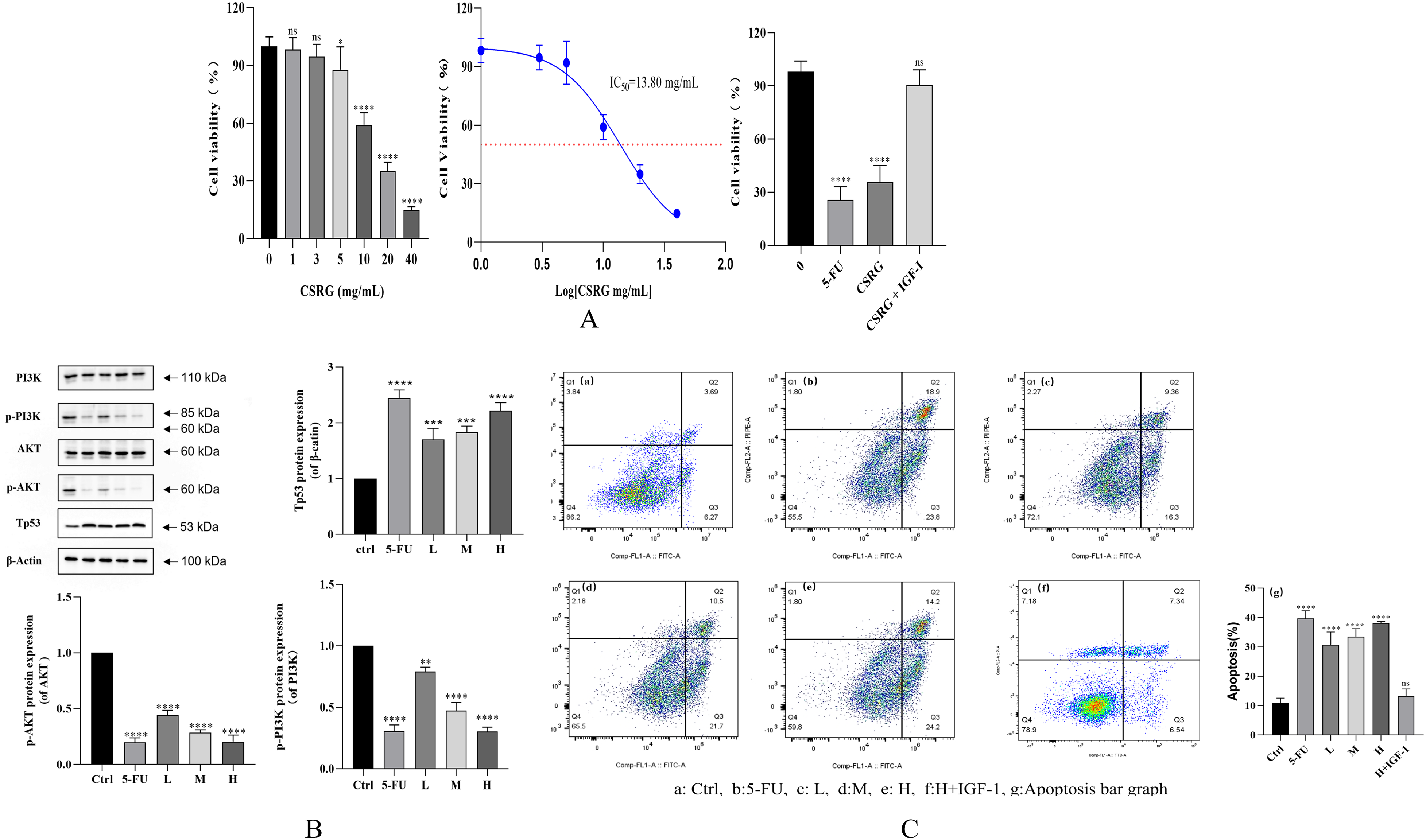
(A) Effect of CSRG and IGF-I on HT29 cell survival (n = 6). (B) Effects of CSRG on core protein expression in HT-29 cells (n = 3). (C) Effects of CSRG and IGF-I on HT29 cell apoptosis (n = 3).
The expressions of PIK, p-PI3K, Akt, p-Akt, and Tp53 proteins were detected by Western blotting. The results showed that CSRG inhibited the phosphorylation of PIK and Akt and up-regulated the expression of Tp53 protein (Figure 7B). The above results suggest that the PI3K/Akt signaling pathway may be an important pathway involved in CSRG-induced apoptosis.
Discussion
Many doctors use CSRG to treat moist and hot intestinal carbuncles, as well as dysentery in China. The symptoms of dysentery are similar to those of enteritis in Western medicine; they are also risk factors for CRC, given the processes of inflammatory cancer transformation. Specifically, long-term inflammatory stress causes genetic damage to the intestinal mucosa and excessive self-tissue repair, manifesting as hyperplasia of the colon epithelial tissue, eventually leading to bowel cancer. 19 Animal studies have demonstrated that components in CSRG promote normal colorectal stem cell proliferation, inhibits CRC tumor cell proliferation, and accelerates intestinal mucosal repair.20,21 Additionally, clinical trials have reported significantly superior results in those treated with combined chemotherapy and CSRG than in those receiving the control treatment; CSRG decreased the toxic side effects of the chemotherapy drugs, improving quality of life, which highlights its effectiveness as an adjuvant drug for treating tumors.22,23 However, the material basis and mechanisms of CSRG as a CRC treatment remain unclear.
This study used a network pharmacology analysis to explore for the first time the material basis, molecular targets, and possible mechanisms of CSRG as a CRC treatment; we also predicted the underlying pharmacological mechanisms through molecular docking and cell experiments. First, we identified eight key active components in CSRG based on their high degree values in the complex regulatory network: naringenin, quercetin, licochalcone A, acacetin, wogonin, baicalein, kaempferol, and isorhamnetin. These components have antioxidant, anti-inflammatory, and anti-tumor effects and thus are crucial for the anti-CRC effects of CSRG. We also found that these active components potentially acted on 151 CRC-related targets and predicted that they could treat CRC via multiple targets and pathways. Finally, GO and KEGG analyses and cell experiments showed that CSRG may influence CRC development through the PI3K–Akt signaling pathways and apoptosis.
In this study, quercetin had the highest degree value. This component belongs to a class of flavanol compounds with anti-inflammatory, anti-tumor, and antioxidant effects, making it the most important and extensively studied. For example, it elicits anti-cancer effects by inhibiting transforming growth factor-alpha (ie, TGF-α) and cycloocesterase-2 (COX-2) expression in CRC by increasing caspase-3 (Casp-3) activity and inducing apoptosis. 24 Furthermore, quercetin inhibits the nuclear factor kappa light chain enhancer of activated B cells (NF-kB) signaling pathway activity, down-regulates B-cell lymphoma 2 (Bcl-2) protein expression, promotes Bax protein expression, and inhibits the inflammatory response but promotes apoptosis in Caco-2 and SW-620 CRC cells. In addition, it promotes the phosphorylation of AMP-activated protein kinase (ie, AMPK) and p53 and induces apoptosis in HT-29 rectal cancer cells. 25 Naringenin inhibits the proliferation of HCT116 and SW480 human CRC cells by down-regulating cyclin D1. 26 Furthermore, it promotes DNA breakage and cell contraction, causing severe damage to tumor cells and increasing the number of apoptotic cells. 27 Naringenin also promotes S and G2/M phase cell cycle arrest and induces time- and dose-dependent apoptosis in SW1116 and SW837. 28 Licochalcone A inhibits the expression of thioredoxin reductase 1 (ie, TrxR1) in HCT-116 cells, leading to increased reactive oxygen species (ROS) levels and apoptosis. 29 Furthermore, it inhibits the release of the angiogenic factors IL-6 and IL-8 and the vascular endothelial growth factor receptor 2 (ie, VEGFR-2) signaling pathway to inhibit neovascularization. 30 Acacetin induces mitochondrial ROS-mediated apoptosis in SW480 and HCT-116 colon cancer cells in a caspase-independent manner by inducing apoptosis-inducing factor (ie, AIF). 31 Additionally, it inhibits the PI3K/Akt /inhibitor of nuclear factor-κB (IκB) kinase (ie, IKK) and MAPK signaling pathways of RAW264.7 macrophages, subsequently inhibiting NF-κB activity. Nitric oxide synthase (ie, iNOS) and COX-2 expression are also down-regulated to prevent inflammation-related tumors. 32
Other ingredients also influence the treatment of CRC. In addition to those mentioned above, Wogonin induces autophagy in CRC-related cells by inhibiting the PI3K/Akt/mammalian target of rapamycin (mTOR) signaling pathway, eliciting an anti-cancer effect. It also causes SW48 cells to stagnate in the G2/M phase, inhibiting the growth of intestinal cancer cells. 33 Wogonin also inhibits β-catenin-mediated transcription by interfering with T cell factor/lymphoid enhancer factor family (ie, TCF/Lef) transcriptional activity and inhibiting the kinase activity of CDK8, which is a suspected oncogene involved in CRC development. 34 Baicalein effectively inhibits the proliferation, migration, and invasion abilities of CRC cells dose- and time-dependently. 35 Furthermore, it regulates Bcl-2 and Bax protein expression, promoting mitochondrial dysfunction, Casp-3 activation, poly (ADP-ribose) polymerase (ie, PARP) cleavage, and tumor cell apoptosis. 36 Additionally, toll-like receptor 4 (TLR4) binds to and inhibits the TLR4/HIF-1α/vascular epidermal growth factor (VEGF) signaling pathway, reducing cancer metastasis and thus playing an anti-colon cancer role. 37 Kaempferol decreases ROS production and blocks the activation of TNF inflammatory pathways.38,39 Furthermore, it inhibits thymidylate synthase and reduces p-Akt activation to inhibit proliferation and induce apoptosis in colon cancer cells, which acts synergistically with 5-Fluorouracil to reduce toxicity and delay drug resistance. 40 Additionally, in HT29 cells, kaempferol increased mitochondrial membrane permeability and the cytochrome C concentration, which activated cell surface apoptosis receptors and the mitochondrial apoptosis pathway, inducing apoptosis. 41 Finally, isorhamnetin inhibits the proliferation of the human CRC cell lines HT-29, HCT116, and SW480, induces cell cycle arrest in the G2/M phase, and inhibits the PI3K–Akt–mTOR pathway, inhibiting cell proliferation. 42 Furthermore, heat shock protein (HSP) 70 family protein expression, including HSPa1a, HSPa1b, and HSPa8, is inhibited by the upregulation of apoptosis-related genes (eg, apoptotic protease activating factor 1 [ie, Apaf1], Casp-3, and caspase-9) in colon cancer cells, preventing tumor cell growth. 43 All these important active ingredients are derived from CSRG, which is, thus, a synergistic treatment for CRC.
In this study, we identified 151 cross-targets with CRC and then further screened the top five targets of CSRG for CRC treatment, specifically MAPK3, MAPK1, TP53, STAT3, and RELA. These target genes are related to cancer and gene expression, cell proliferation, apoptosis, the cell cycle, and metabolism. In molecular docking experiments, TP53 and corresponding core components showed high affinity, indicating that TP53 may be one of the key targets of CSRG in the treatment of CRC.
TP53 is a tumor suppressor gene and a key regulator of programmed cell death, the cell cycle, DNA repair, and genomic stability. 44 Mutations in this gene activate immune pathways and cause inflammation, which is crucial in CRC. 45 TP53 tumor suppressors can be activated by various cellular stresses, including oncogene expression and DNA damage, and implement appropriate responses to combat cancer. 46
The GO enrichment analysis showed that CSRG-related genes were mainly associated with transcription, gene expression regulation, apoptosis regulation, and other biological processes. Furthermore, the KEGG analysis indicated that the PI3K–Akt, TNF, IL-17, and HIF-1 signaling pathways were closely related to CRC and involved in tumor cell survival, proliferation, invasion, metabolism, hypoxia, and inflammatory responses.
The pathway with the greatest number of enriched genes was the PI3K–Akt signaling pathway, with 37 targets enriched in this pathway, including MAPK1, TP53, MCL1, MAPK3, and HSP90AA1. Phosphatidylinositol 3′-kinase (PI3K)-Akt is a signaling pathway that regulates the basic functions of cells, such as transcriptional translation and growth and proliferation, and can be activated by toxic damage or cell stimulation. PI3K catalyzes the formation of phosphatidylinositol 34,5-triphosphate (PIP3) on the cell membrane, and PIP3 acts as a second messenger to activate Akt. Finally, Akt regulates the key processes of cell growth and development by phosphorylating proteins involved in cell synthesis, metabolism, the cell cycle, and apoptosis. 47 48This pathway is involved in regulating various cell cycle processes, promoting the occurrence and development of tumors, regulating multiple downstream targets, phosphorylating various proteins, and causing abnormal pathway activation for CRC cell survival and proliferation. This pathway promotes matrix metalloproteinase 2 and 9 (ie, MMP-2 and MMP-9) and urokinase plasminogen activator (ie, uPA) expression, thereby promoting tumor cell invasion and metastasis. 49
Finally, the PI3K–Akt signaling pathway and TP53 were used as targets to observe the anti-cancer effects of CSRG in vitro based on KEGG pathway enrichment analysis and molecular docking results. HT29 cells, CSRG and PI3K/AKT agonist IGF-I were used for in vitro experiments with verification. The results showed that CSRG inhibited the proliferation of HT-29 cells, induced apoptosis in a dose-dependent manner, inhibited the phosphorylation of PI3K and Akt, and up-regulated the expression of TP53 protein. Meanwhile, the results showed that the PI3K/AKT agonist IGF-I could inhibit cell proliferation and apoptosis mediated by CSRG. This result agrees with that reported in the literature. 50 Overall, the active ingredients in CSRG elicit an anti-CRC effect by simultaneously regulating various signaling pathways (eg, cell proliferation, apoptosis) through multiple targets. Furthermore, cell experiments verified that CSRG components have some anti-cancer effects in CRC treatment, confirming the accuracy of the network pharmacology prediction results.
This study relied on various existing databases, which may not be updated regularly, and TCM components are complex. Therefore, the preliminary exploration of the role of CSRG in CRC treatment has some limitations. The present study focuses on the apoptosis-inducing role of CSRG in HT29 cells and does not examine other roles of CSRG in CRC, such as gene expression regulation and metastasis. However, these findings provide important preliminary information on the role of CSRG in CRC therapy, suggesting that CSRG may be a potentially effective drug for the treatment of CRC. Further in vitro and in vivo experiments are required to clarify the molecular targets and signaling pathways of CSRG in CRC. Nonetheless, this study provides a theoretical basis for further research on the pharmacodynamic components of CSRG in the treatment of CRC and for optimizing experimental design.
Conclusions
Using network pharmacology, we determined that the main active ingredients in CSRG were naringenin, quercetin, licochalcone A, acacetin, wogonin, baicalein, kaempferol, and isorhamnetin. These components act on MAPK3, MAPK1, TP53, STAT3, RELA, and other potential targets to regulate various biological processes and molecular functions. These components help regulate the PI3K–Akt and TP53 signaling pathways to treat CRC. This study systematically analyzed the active components, target genes, and CSRG signaling pathways of CSRG in the treatment of CRC, providing a theoretical basis for further research on CSRG.
Footnotes
Acknowledgments
Authors Contributions
Hui Zhang and Aiping Zheng conceptualized and designed the study. Material preparation, data collection, and analysis were performed by Shijie Zhong and Min Wu. The first draft of the manuscript was written by Yadong Xing, and all authors reviewed the original draft of the manuscript. All authors have read and agreed to the published version of the manuscript.
Data Availability Statement
Relevant data are contained in the paper and Supplemental materials.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Not applicable.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Natural Science Foundation of China (81573357); National New Drug Key Technology Research and Development Program (2018ZX09721003⁃007/2018ZX09J18107); National Key Research and development Program (2023YFC2706100; 2023YFC2605000); Key scientific research projects of Anhui Provincial Department (2023AH052017).
Informed Consent Statement
Not applicable. No human subjects were used in this article.
Statement of Human and Animal Rights
Not applicable. No animal subjects were used in this article.
