Abstract
Introduction
Knee osteoarthritis (KOA) is a universal problem worldwide and a common disease encountered in primary care settings. A survey showed that more than 200 million people worldwide suffer from osteoarthritis (OA), and the knee joint is one of the most frequent sites. 1 According to statistics, 6.5 of every 1000 people suffer from KOA. 2 In addition to pain, KOA also causes joint stiffness, restricted daily activities, joint swelling, joint instability, and lower limb muscle weakness, thereby reducing the patient's quality of life and causing serious social and economic burdens. 3 With the extension of the life cycle, the severity of the disease and the symptoms worsen. 3 The aging of the global population is becoming increasingly prominent and plans to maintain and promote the management of chronic diseases are becoming increasingly important. In the treatment of KOA, drugs are mainly related to the relief of pain symptoms, which is a main appeal for many patients with KOA.
Medicinal plants have been used in traditional herbal medicine for thousands of years as a promising alternative treatment, and the incidence of adverse events is usually equal to that of traditional medicines.
3
In Ayurvedic medical systems, natural medicines have been used to treat chronic diseases for many years. In traditional Chinese medicine, Frankincense and myrrh (FM) has been used in combination in the same prescription for thousands of years, and their combination exerts a better therapeutic effect on diseases.
4
FM generally has a stronger synergistic effect than a single drug.
5
Deng et al summarized the frequency of use of herbal medicines for the treatment of KOA, and the results showed that the frequency of use of FM was 43.2% and 40.6%.
6
The association rule analysis conducted by Gao et al showed that
Frankincense is a resin exuded from the bark of
The mechanism of action of FM in the treatment of KOA has been partially explored, but the complexity of the herbal medicine and the diversity of its targets require further elucidation of the mechanism underlying its therapeutic effect on KOA. Natural medicine treats diseases with multiple targets and multiple channels. Current basic research still focuses on the treatment of a disease with a single monomer or a natural drug compound. Thus, exploration of how complex pharmaceutical ingredients act in the complex human environment is difficult. In recent years, with the rise of network science, such as systems biology and network pharmacology, combination drugs have become a hot topic in modern drug research, and new trends in the development and use of modern drugs have been reported. 17 Based on this information, this article adopts the network pharmacology research method to reveal the mysteries underlying the synergistic effects of FM on KOA and provide further evidence for the use of FM as a treatment for diseases. By exploring the molecular mechanisms and targets of FM in the treatment of KOA, new ideas are expected to guide drug research and clinical applications.
Methods
Network Pharmacology Analysis
Compound Composition, Collection, and Screening
TCMSP
18
(https://tcmspw.com/tcmsp.php) and BATMAN-TCM
19
(http://bionet.ncpsb.org.cn/batman-tcm/) were used to collect herbal and drug target components. We searched the chemical composition information of FM in TCMSP using the following screening conditions to obtain active drug ingredients: oral bioavailability (OB) ≥30% and drug likelihood (DL) ≥0.18. According to the BATMAN-TCM operation steps, the score cutoff of “drug-target” was set to ≥20, with an adjusted
FM and KOA-Related Target Acquisition
In our previous study, we searched and screened genes related to KOA. 20 Using the UniProt database (https://www.uniprot.org/UniProt/), we converted the information and names of the KOA-related targets. Venn diagrams were constructed to show the logical connections between drugs and disease targets. Mapping was carried out between the target of the active ingredient of the drug and the target of KOA, and the overlapping targets were selected as the common targets of FM for KOA.
Construction of the Protein Interaction Network and Screening of Core Genes
We imported the obtained common targets into the STRING database, analyzed the protein interaction network (PPI), and drew the PPI network diagram. The following conditions were used: multiple proteins, Homo sapiens, and a confidence level of 0.7. The TSV file was imported into Cytoscape software, and the “CytoNCA” App was used to analyze the topological properties and obtain the degree centrality (DC), betweenness centrality (BC), closeness centrality (CC), eigenvector centrality (EC), network centrality (NC), and local average connectivity (LAC)-based method. The core target was selected based on the values of DC, BC, CC, EC, NC, and LAC. First, the median of the six indicators was calculated, and the median of the obtained results was calculated to obtain the core target. We independently screened pivotal genes using the CytoHubba App to verify the reliability of the results . CytoHubba generates the attribute ranking of nodes in the network. This research used the maximal clique centrality (MCC) topology analysis algorithm.
Construction of the Network of Active Drug Ingredients and Disease Targets
The active ingredients of herbs and the screening results of KOA targets were imported into Cytoscape software to construct a network of active ingredients of FM and KOA targets. A visual analysis was performed on the built network. In this network, each node represents a target or active ingredient, and each edge represents the relationship between biomolecules.
GO and KEGG Enrichment Analyses
A relevant analysis was carried out using R software to further explore the biological processes (BPs) of FM in the treatment of KOA and its regulatory signaling pathways. The bioconductor collection toolkit in R software was used to perform the GO functional enrichment analysis and KEGG pathway enrichment analysis of common genes with
Molecular Docking
The active ingredients of the drug were used as ligands, and the target protein was used as the receptor. DS software was used for molecular docking analysis. Ligand information was obtained from the Chemicalbook database (https://www.chemicalbook.com/) and TCMSP. The crystal structure of the receptor was downloaded from the RCSB PDB database. First, the receptor and ligand were preprocessed using DS software. Then, the binding site was defined and edited, and the Dock Ligands (LibDock) module was run. The following settings were used: high quality, BEST conformation method, parallel processing, and the default values for the other parameters. The docking results are displayed as LibDock scores. The higher the score, the more likely the active ingredient of the drug will bind to the target.
Chemicals and Reagents
Ru Xiang (Frankincense,
Drug Preparation
According to the experimental design, 6 g of frankincense (Ru Xiang) and 6 g of myrrh (Mo Yao) were obtained from the Affiliated Hospital of Nanjing University of Chinese Medicine. These drugs were identified by Prof. Wang Peimin. The herbs were mixed, and the mixture was boiled in distilled water and filtered. The filtrate was evaporated with a rotary evaporator and then dried to a constant weight in a vacuum drying oven for further in vitro experiments. FM was dissolved in phosphate buffered saline (PBS) to a stock concentration of 40 mg/mL and stored at −80 °C. The working concentration of FM was prepared by diluting the stock solution in PBS, filtering through a 0.22-µm filter, and storing at 4 °C.
LC–MS for Component Identification
Ten milligrams of the lyophilized drug was weighed and dissolved in 0.5 ml of ultrapure water. Then, 80 µl of the solution was pipetted, mixed with 20 µl of pure acetonitrile, and placed at −20 °C overnight. After centrifugation (14 000 rpm*10 min), the supernatant was used for detection.
The detection system was ultrafast liquid chromatography (UFLC, Shimadzu, Kyoto, Japan) with Triple-ToFTM 5600 (Q-TOF, AB Sciex, Framingham, MA, USA). The ion source was electron spray ionization (ESI), and the control system was Analyst TF 1.6 software (AB Sciex, Singapore). The chromatographic column was a Waters Symmetry ShieldTM RP18 C18 (250 mm × 4.6 mm, 5 μm, Waters, USA), and the column temperature was 30 °C. The mobile phase was 0.2% formic acid water (A) and acetonitrile (B), with a flow rate of 1 ml/min. Elution gradient: 0-2 min, 2% B; 2-18 min, 30% B; 18-30 min, 60% B; 30-38 min, 100% B; 38-40 min, 100% B; 40-44 min, 2% B; 44-48 min, 2% B. The samples were analyzed in positive and negative ion modes, and the injection volume was 10 µL. Mass spectrometry parameters: data-dependent acquisition (IDA) in dynamic background subtraction mode (DBS), acquisition range of m/z 50-1500 Da, and an error of 50 mDa. The corresponding top 10 substances were fragmented, and MS2 information was collected. Ion source temperature: ± 550 °C; CE: ± 40 V; DP: ± 100 V; CUR: 40 V; GS1, GS2: 40 V; ISVF: 5500 V (pos)/4500 V (neg).
Peak picking and alignment were performed by MS-DIAL (ver. 4.60). The pubchem database was used to search the MS/MS spectra, remove the recognition results of |ppm|>20 and invalid comparison, and reconfirm the compounds with SN > 3 and a total score of 80 points or more.
Obtaining Primary Chondrocytes
This study was approved by the Animal Ethics Committee of Nanjing University of Chinese Medicine (202104A017) and complied with the National Institutes of Health Guide for the Care and Use of Laboratory Animals. Twenty C57BL/6 male mice (9 weeks) were obtained from the Experimental Animal Center of Nanjing University of Chinese Medicine (Nanjing, China) with license number SCXK (ZHE)-2019-0001. The C57BL/6 male mice were sacrificed with an overdose of sodium pentobarbital. The articular cartilage of the femoral condyle and tibial plateau was extracted under aseptic conditions. The tissue was cut into pieces (1 mm3) and digested with 0.2% collagen II solution for 4 h. Chondrocytes were seeded in a 10 cm2 Petri dish at a density of 2 × 105/mL and placed in an environment at 37 °C and 5% CO2. The cells used in the subsequent experiments were in the second generation and entered the logarithmic growth phase.
Chondrocyte Identification
Chondrocytes (second generation) were seeded into 6-well plates at 2 × 104 cells/well. When the cells were more than 50% confluent, they were rinsed twice with PBS and toluidine stain solution and dye were added for 5 min at room temperature. The same amount of distilled water was added and mixed for 15 min. Finally, the cells were washed twice with distilled water and observed under a microscope (DMi1, Leica, Germany).
CCK-8 Assay
The cells were seeded in a 96-well plate at a density of 6000 cells/well. Then, different concentrations (0, 25, 50, 100, 200 and 400 μg/mL) of FM were used for 24/48 h of treatment. Then, 10 μL of CCK-8 solution was added to each well and incubated at 37 °C for 2 h. The absorbance was detected at 450 nm with multimode plate reader (EnSpire, PerkinElmer, USA).
Experimental Design
Except for the blank group, all other groups were treated with 10 ng/mL IL-1β. After that, FM (100, 200, 400 μg/mL) was added to the drug groups and incubated at 37 °C in a humidified atmosphere containing 5% CO2 for 24 h. The protein was collected from the supernatant and stored at −80 °C.
Immunofluorescence
Chondrocytes were treated in 6-well plates (2 × 104), fixed with 4% (v/v) paraformaldehyde, permeabilized using 0.5% Triton X-100, blocked with 3% goat serum, incubated overnight at 4 °C with antibodies against p65 and collagen II, and incubated with anti-mouse or anti-rabbit Alexa Fluor 594 sary antibodies (Abcam, USA) for 30 min. The cells were counterstained with DAPI for 5 min and visualized using a fluorescence microscope (DMi8, Leica, Germany).
Western Blotting
Western blotting was used to detect the protein levels of p38, p-p38, JNK, p-JNK, ERK1/2, p-ERK1/2, P65, p-P65, and IκB. RIPA Lysis Buffer (RIPA) and Phenylmethylsulfonyl fluoride (PMSF) were used to extract the total protein from chondrocytes. After brief sonication, the cell lysates were centrifuged at 12 000 rpm at 4 °C for 20 min. The BCA Protein Assay Kit (Beyotime) was used to determine the protein concentration. Forty micrograms of protein was loaded on SDS-PAGE gels and transferred to polyvinylidene fluoride(PVDF) membranes (Bio-Rad, USA). After blocking with 5% skimmed milk powder for 2 h at room temperature, the membranes were washed with tris-buffered saline Tween-20 (TBST) 3 times for 10 min each. Then, the membranes were incubated with the targeted primary antibody overnight at 4 °C. After washing, the blots were incubated with horseradish peroxidase-conjugated goat antirabbit or anti-mouse IgG secondary antibodies for 2 h at room temperature. ImageQuant LAS 4000 mini (LAS 4000, Cytiva, USA) was used for signal detection. Image J software was used to analyze the gray value of each band.
Statistical Analysis
Data analysis was conducted using SPSS version 22.0 software (SPSS) and GraphPad Prism 8.0. The results are expressed as the mean ± standard deviation (SD). Statistical comparisons were performed using Student's
Results
Constructing a “Drug Component-Disease Target” Network
According to the screening conditions and after deleting duplicate data, 31 active ingredients were finally included, as shown in Table 1. The drug targets were mapped with the disease targets to obtain 94 common target genes, as shown in Figure 1A and Table 2.
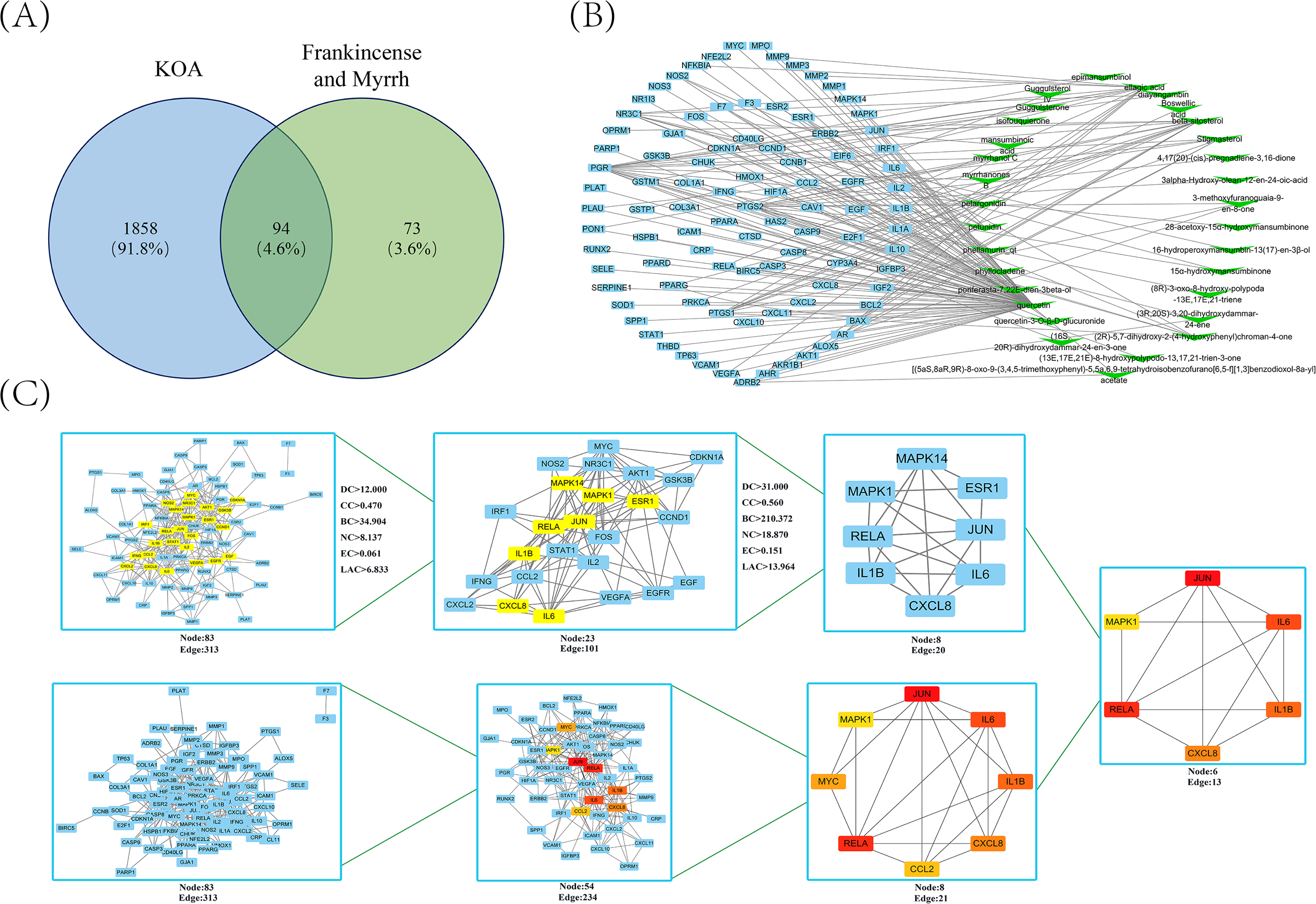
Drug-disease target network construction and core target screening. (A) Common target acquisition. (B) The blue quadrilateral represents the common gene, and the green inverted triangle represents the active ingredient of frankincense and myrrh. (C) Core gene screening in the PPI network.
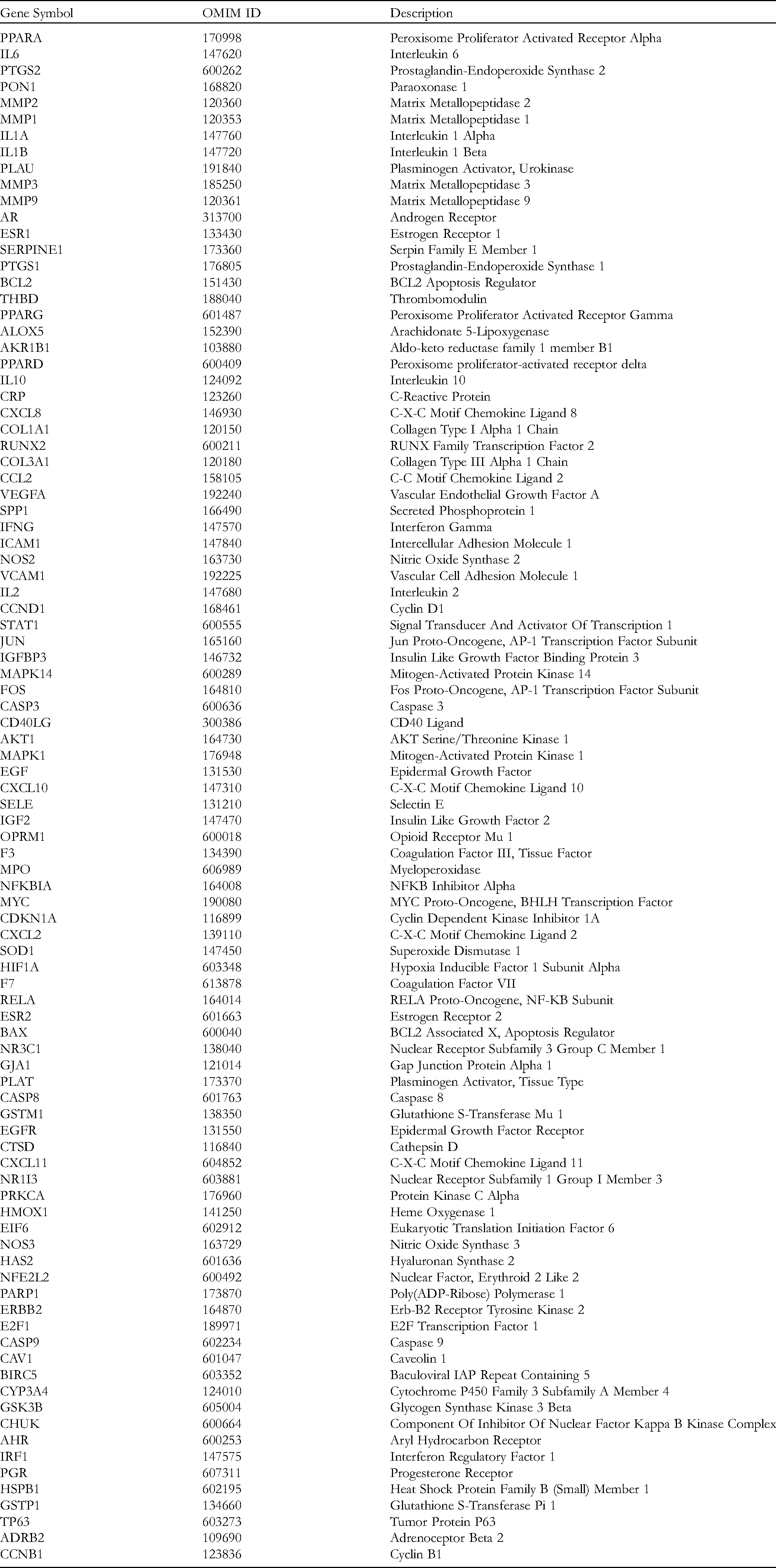
Active Ingredients Parameters of Frankincense and Myrrh.

The Common Gene of Frankincense and Myrrh in the Treatment of KOA.
Cytoscape software was used to construct a network of active ingredients and common gene relationships. After importing the data, a network of 125 points and 174 connections was obtained. The blue nodes represent a common gene, and the green nodes represent the active ingredient, as shown in Figure 1B. Taking the degree as the main reference basis for the topological analysis, the top 5 active ingredients were quercetin, beta-sitosterol, ellagic acid, pelargonidin, and petunidin, and the parameters used for the topological analysis of the remaining active ingredients are shown in Supplementary Table 1. The topological analysis results are shown in Table 3. These compounds with higher degrees may be the key compounds in FM that treat KOA.
Topology Parameters in the PPI Network.

Analysis of the Target Protein Interaction Network
The STRING database platform was used to predict protein interactions and draw protein relationship network diagrams. Ninety-four common genes were imported into the STRING database to obtain FM PPI networks acting on KOA, with a total of 83 nodes and 313 interactive connections. Cytoscape software was used for visualization. The topological parameters of the network nodes were calculated using the CytoNCA App, and the DC, CC, BC, EC, NC and LAC values were obtained. The first screening parameter settings were DC≥6.000, EC≥0.065, LAC≥2.250, BC≥39.849, CC≥0.220, NC≥3.000, a total of 23 nodes, and 101 interaction connections. The second screening parameter settings were DC≥14.000, EC≥0.165, LAC≥4.824, BC≥229.115, CC≥0.246, NC≥6.589, a total of 8 nodes, and 20 interaction connections. In addition, we searched for the core target again using the CytoHubba App to predict the most relevant core target. First, the target network node values (node scores) were calculated, the top 8 CytoHubba nodes were selected according to the MCC algorithm, the first-stage nodes were assessed, and the shortest path and the expanded subnetwork were displayed. The screening process is shown in Figure 1C.
GO Enrichment Analysis
According to the results of the GO enrichment analysis, 2213 BPs were enriched, and were involved in the negative regulation of apoptosis signaling pathways, regulation of reactive oxygen species metabolism, response to tumor necrosis factor, mechanical stimulation, cellular responses to biological stimulation, and oxidative stress. One hundred thirty-five items were enriched in the molecular function (MF) category, among which DNA binding transcription activation activity, cytokine receptor binding, cytokine, receptor ligand, nuclear receptor activity, and RNA polymerase II transcription factor binding were the top enriched terms. Based on the results of the cell fraction (CC) enrichment analysis, 54 items were enriched, which were mainly related to cell membrane parts such as membrane rafts, membrane microdomains, membrane regions, and caveolae. The 20 most notable items are presented in a bubble chart and a GO enrichment network diagram in Figure 2A.

Go and KEGG enrichment analyses. (A) GO biological function enrichment analysis. The size of the
Analysis of Enriched KEGG Pathways
The KEGG pathway enrichment analysis results reveal 143 enriched pathways. Among the top 20 signaling pathways, fluid shear stress and atherosclerosis and the TNF, IL-17, HIF-1, and toll-like receptor signaling pathways are closely related to the mechanism of FM treatment in KOA. In addition, apoptosis, the NF-κB signaling pathway, the PI3K-Akt signaling pathway, cellular senescence, the MAPK signaling pathway, the p53 signaling pathway, the mTOR signaling pathway, arachidonic acid metabolism, and the Hippo signaling pathway were included in the enrichment results, as shown in Figure 2B. The TNF signaling pathway is shown in Figure 2C.
Molecular Docking Analysis
By screening the core targets, 6 important targets were obtained, including JUN, IL6, IL1B, CXCL8, RELA, and MAPK1. Using Discovery Studio software, these six intersecting targets and the active ingredients obtained from the screen were subjected to molecular docking analysis. The software evaluates the binding ability of the ligand and the receptor through the binding between the targets and generates a LibDock score after calculation and fitting. The larger the value, the higher the binding activity, and the easier it is for the drug to bind to the receptor. Topological analysis showed that quercetin ranked first in the prediction. We show the docking process of quercetin and 6 core targets in Figure 3A and B. Other docking results are shown in Supplementary Table 2.
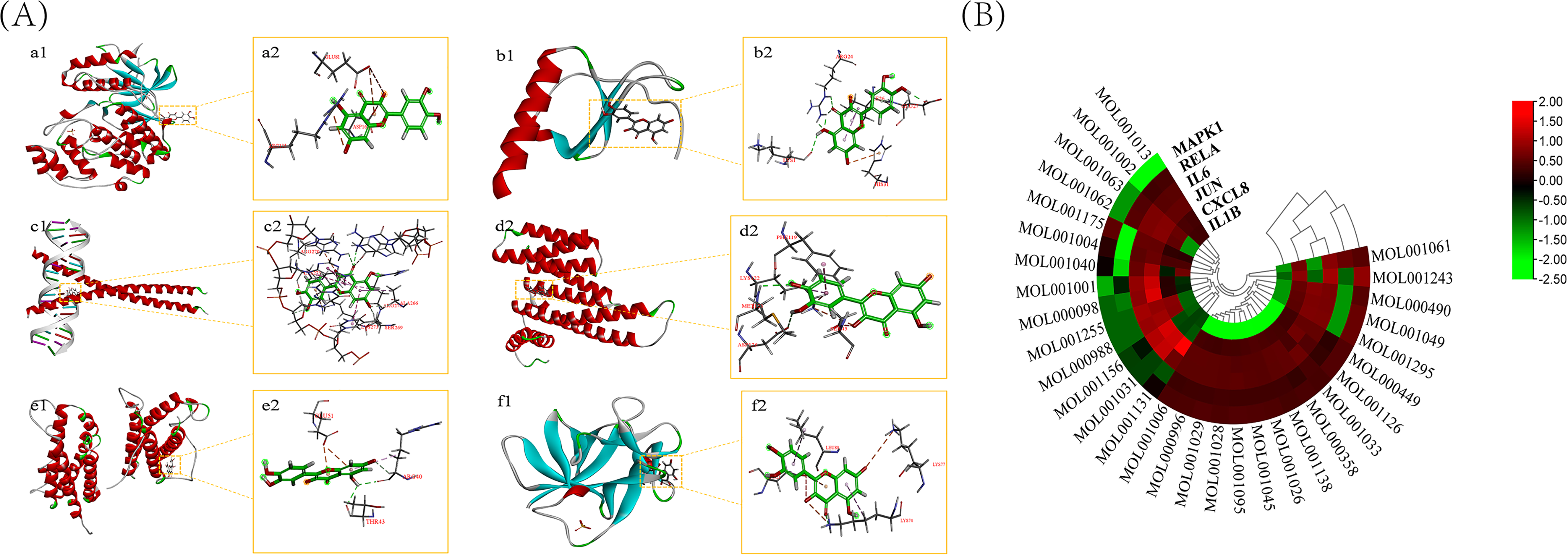
Molecular docking of compounds with core targets. (A) a1 and a2: Docking process of quercetin with MAPK1; b1 and b2: Docking process of quercetin with CXCL8; c1 and c2: Docking process of quercetin with JUN; d1 and d2: Docking process of quercetin with RELA; e1 and e2: Docking process of quercetin with IL6; f1 and f2: Docking process of quercetin with IL1B. (B) Heatmap of the LibDock Score for molecular docking. The redder the color, the higher the LibDock Score, and the greener the color, the lower the LibDock Score.
FM Component Separation and Identification
Through component annotation of the data, a total of 72 possible compounds were identified, of which 28 compounds were only recognized in the positive ion mode, 36 metabolites were only recognized in the negative ion mode, and were 8 common components, as shown in Figure 4A to C. Supplementary Table 3 provides details and some representative MS/MS spectra. The 72 kinds of ingredients included 13 kinds of flavonoids, 11 kinds of amino acids, 9 kinds of terpenes, 8 kinds of phenylpropanoids, 4 kinds of phenols, 4 kinds of fatty acids, and 11 kinds of other ingredients, as shown in Figure 4D. The positive and negative ion identification results are shown in Figure 4E and F.

FM ingredients identified by LC–MS. (A) Twenty-eight ingredients were identified in positive ion mode. (B) Thirty-six ingredients were identified in negative ion mode. (C) Shared ingredients of the positive and negative modes. (D) Major categories of identified ingredients. (E) Confirmation of the mass spectrogram obtained in positive ion mode. (F) Confirmation of mass spectrograms obtained in negative ion mode.
Toluidine Blue Staining to Identify Chondrocytes
Second-generation chondrocytes with good growth were selected. Adherent blue chondrocytes were scattered and distributed like paving stones. After staining, the cell cytoplasm was light blue, and the nucleus was dark blue and spherical, as shown in Figure 5A and B. Since collagen II is mainly produced by chondrocytes, collagen II immunofluorescence was used to identify the chondrocytes. Under the inverted phase contrast microscope, the cytoplasm of chondrocytes was blue (Figure 5C and D).

Identification of mouse articular chondrocytes and cytotoxicity assay of FM on chondrocytes. (A) Toluidine blue staining of chondrocytes (magnification, up: × 100) (magnification, down: × 200). (B) Identification of chondrocytes by collagen II immunofluorescence (magnification: × 100). (C-D) 24/48 h detection of chondrocyte viability by CCK-8 analysis.
Cytotoxicity Assay of FM Using Chondrocytes
The CCK-8 method was used to detect the cytotoxicity of FM to chondrocytes. The results show (Figure 5E and F) that FM did not inhibit the viability of chondrocytes after treatment at 25, 50, 100, 200, and 400 μg/mL for 24/48 h, which indicates that FM has no cytotoxicity to chondrocytes at the selected dose and exposure time.
FM Inhibits IL-1β-Induced Chondrocyte Damage in Mice by Inhibiting MAPK Signaling Pathways
Combined with the pathway map and core target prediction, we chose the NF-κB and MAPK signaling pathways for verification. The NF-κB and MAPK pathways are involved in the expression of inflammatory mediators induced by IL-1β. First, we investigated whether the inhibitory effect of FM on its expression is mediated by the MAPK signaling cascade. As shown in Figure 6A to D, compared with the untreated group, p38, JNK, and ERK were significantly phosphorylated under IL-1β stimulation. FM inhibited the activation of p38, JNK, and ERK. Inhibition of the activation of the MAPK pathway may be one of the mechanisms by which FM protects mouse chondrocytes.
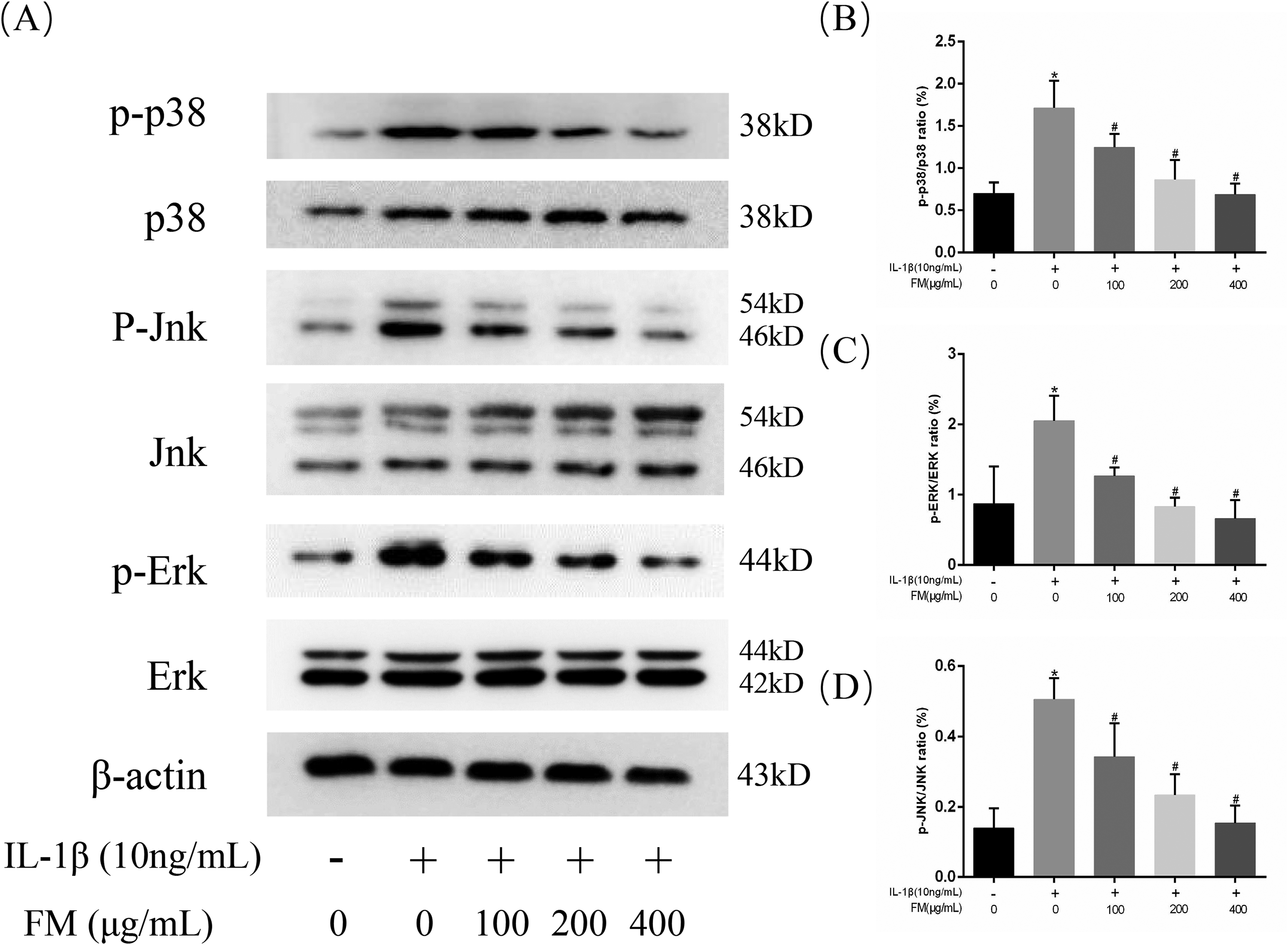
Effect of FM on IL-1β-induced MAPK activation in chondrocytes. Chondrocytes were pretreated with FM at different concentrations for 4 h and then incubated with or without IL-1β (10 ng/mL) for 24 h. Western blot analysis (A) and relevant quantitative analysis (B-D) of p38, p-p38, Jnk, p-Jnk, Erk, and p-Erk. *
Effects of FM on IL-1β-Induced NF-κB Signal Activation in Chondrocytes
Since p65 is an indicator of NF-κB activation after IL-1β stimulation, the protein level of p65 was measured. As shown in Figure 7A to C, we found that p-p65 increased after IL-1β stimulation, while FM inhibited its phosphorylation. In addition, we found that IL-1β stimulated the degradation of IkBα in mouse chondrocytes. The immunofluorescence results show that p65 protein was mainly located in the cytoplasm of mouse chondrocytes not treated with IL-1β; after IL-1β treatment, p65 protein translocated into the nucleus (Figure 7D). FM treatment reversed IL-1β-induced p65 expression and prevented the translocation of p65 from the cytoplasm to the nucleus, as shown in Figure 7D and E. These results indicate that FM has a protective effect on mouse OA chondrocytes by inhibiting the activation of the NF-κB pathway.

Effect of FM on IL-1β-induced NF-κB signal activation in chondrocytes. Chondrocytes were pretreated with FM at different concentrations for 4 h and then incubated with or without IL-1β (10 ng/mL) for 24 h. Western blot analysis (A) and relevant quantitative analysis
(B & C) of p65, p-p65, and IκB in chondrocytes. *
Discussion
According to some previous studies,
Quercetin is a plant-derived polyphenol with a wide range of biological effects and anti-inflammatory, antiviral and anticancer activities. 24 In particular, quercetin exerts a beneficial effect on inflammation and immune function.25,26 Studies have shown that quercetin inhibits the inflammation and apoptosis of chondrocytes, regulates the polarization of synovial macrophages to M2 macrophages, weakens the oxidative stress response, inhibits the degradation of chondrocyte extracellular matrix, and protects articular cartilage. 27 β-Sitosterol is a phytosterol and tetracyclic triterpenoid. 28 β-Sitosterol shows good pharmacological antioxidant, anti-inflammatory, immunoregulatory, antitumor, and other effects. According to basic research, β-sitosterol treatment reduces LPS-induced increases in the levels of inflammatory mediators, including IL-6, iNOS, TNF-α, and COX-2. 29 Stigmasterol is also a phytosterol with various pharmacological activities, such as antitumor, blood cholesterol-lowering, and anti-OA properties. Stigmasterol reduces the degradation of cartilage caused by OA by inhibiting the degradation of inflammatory mediators and the matrix.30,31 Ellagic acid is a natural antioxidant that inhibits the IL-1β-induced expression of iNOS, COX-2, NO, TNF-α, PGE2, and IL-6 and downregulates the expression of MMP-13 and ADAMTS-5.29,32 Pelargonidin has antioxidant, anti-inflammatory, and antidiabetic activities.33,34 In addition, PEL has the potential to improve Alzheimer's disease and atherosclerosis.35,36 Mansumbinoic acid is an extract of myrrh with strong anti-inflammatory activity. Studies have shown that the molar titer of mansumbinoic acid is on the same order of magnitude as those of indomethacin and prednisolone. 37
According to the analysis of the PPI network topology, 6 core targets were identified: JUN, IL-6, IL-1β, CXCL8, RELA, and MAPK1. These may be potential targets on which FM acts to treat OA. According to the enrichment analysis of the KEGG pathway and the results of previous studies, among the selected pathways for FM treatment of KOA, the NF-κB and MAPK signaling pathways have a strong correlation.
MAPK is an activated serine/threonine protein kinase that is mainly involved in the regulation of cell growth, differentiation, apoptosis, and the inflammatory response. 38 Jun is a member of the mitogen-activated protein kinase family that plays a role in cell proliferation, apoptosis, survival, stress, and apoptosis by inducing transcription or the activation of transcription factors for downstream target genes. 39 The downregulation of Jun reduces the expression of inflammatory factors and inhibits endothelial cell apoptosis. 40 In OA, IL-6 released from joint tissues binds to IL-6R, which activates the immune system, and monocytes are recruited to the inflamed joint area. 41 IL-1β participates in a series of cellular processes, such as cell proliferation, differentiation, and apoptosis. 38 IL-1β plays an important role in the pathogenesis of OA and is currently used to induce OA models in vitro. NF-κB, which is triggered by proinflammatory cytokines and ECM degradation products, plays an important role in the pathogenesis of OA.42,43 Activated NF-κB is an important part of OA events because NF-κB regulates the expression of a variety of cytokines, chemokines, and matrix-degrading enzymes. 44 Under physiological conditions, p65 exists in the cytoplasm. When stimulated by IL-1β, p65 is released, phosphorylation increases, and it rapidly transfers to the nucleus and promotes the expression of a large number of inflammatory factors. Our results show that FM inhibits p65 phosphorylation and nuclear translocation, indicating that the NF-κB pathway has an inhibitory effect in chondrocytes. The MAPK pathway mainly includes ERK1/2, Jnk, and p38, which can be activated by phosphorylation of upstream signals. According to the western blotting results, FM can reduce the phosphorylation of Erk, p38, and JNK induced by IL-1β, thereby inhibiting MAPK signaling. Therefore, the potential mechanism by which FM prevents cartilage degeneration is related to the inhibition of the NF-κB and MAPK pathways.
Conclusions
In summary, this study preliminarily explored the potential mechanism of FM as a treatment for KOA through the combined actions of multiple targets and pathways. Our results show that FM can inhibit chondrocyte inflammatory factors in vitro by inhibiting the NF-κB and MAPK signaling pathways. Furthermore, this study provides a theoretical basis and research data for the clinical application of FM in the treatment of KOA and reveals the material basis and potential mechanism underlying the efficacy of FM in the KOA treatment.
Supplemental Material
sj-xls-1-npx-10.1177_1934578X221116984 - Supplemental material for Comprehensive Network Pharmacological Analysis and In Vitro Verification Reveal the Potential Active Ingredients and Potential Mechanisms of Frankincense and Myrrh in Knee Osteoarthritis
Supplemental material, sj-xls-1-npx-10.1177_1934578X221116984 for Comprehensive Network Pharmacological Analysis and In Vitro Verification Reveal the Potential Active Ingredients and Potential Mechanisms of Frankincense and Myrrh in Knee Osteoarthritis by Xiaoqing Shi, Lishi Jie, Peng Wu, Jun Mao, Peimin Wang, Zixiu Liu and Songjiang Yin in Natural Product Communications
Supplemental Material
sj-xls-2-npx-10.1177_1934578X221116984 - Supplemental material for Comprehensive Network Pharmacological Analysis and In Vitro Verification Reveal the Potential Active Ingredients and Potential Mechanisms of Frankincense and Myrrh in Knee Osteoarthritis
Supplemental material, sj-xls-2-npx-10.1177_1934578X221116984 for Comprehensive Network Pharmacological Analysis and In Vitro Verification Reveal the Potential Active Ingredients and Potential Mechanisms of Frankincense and Myrrh in Knee Osteoarthritis by Xiaoqing Shi, Lishi Jie, Peng Wu, Jun Mao, Peimin Wang, Zixiu Liu and Songjiang Yin in Natural Product Communications
Supplemental Material
sj-docx-3-npx-10.1177_1934578X221116984 - Supplemental material for Comprehensive Network Pharmacological Analysis and In Vitro Verification Reveal the Potential Active Ingredients and Potential Mechanisms of Frankincense and Myrrh in Knee Osteoarthritis
Supplemental material, sj-docx-3-npx-10.1177_1934578X221116984 for Comprehensive Network Pharmacological Analysis and In Vitro Verification Reveal the Potential Active Ingredients and Potential Mechanisms of Frankincense and Myrrh in Knee Osteoarthritis by Xiaoqing Shi, Lishi Jie, Peng Wu, Jun Mao, Peimin Wang, Zixiu Liu and Songjiang Yin in Natural Product Communications
Footnotes
Acknowledgements
We gratefully acknowledge the contributions of Juan Zhao (Nanjing University of Chinese Medicine) to the data analysis.
Author Contributions
SXQ and JLS contributed equally to this paper and designed the study. Experimental work and data collection were conducted by SXQ and LZX. SXQ and JLS analyzed and interpreted the data. SXQ and WP drafted the manuscript. WPM and YSJ provided critical comments and revised the manuscript. LZX, YSJ and ZNS helped perform the analysis and with constructive discussions. All authors read and approved the final version of the manuscript.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present study was supported by the Natural Science Foundation of Jiangsu Province (SBK2020041404), the National Natural Science Foundation of China (82074460), the Universities Natural Science Research Project of Jiangsu Province (20KJB360003), and the Research and Practice Innovation Program for Postgraduates in Jiangsu Province (KYCX21_1684). The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Data Availability Statement
The original contributions presented in the study are included in the article, and further inquiries can be directed to the corresponding authors.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
