Abstract
Objective
SARS-CoV-2 coinfection with Streptococcus pyogenes and S pneumoniae increases disease severity, thus deserving more in-depth understanding.
Methods
The potential of garlic extract against the bacteria has been theoretically examined and extended to synergic predictions on SARS-CoV-2 Delta and Omicron variants.
Results
Gas chromatography–mass spectrometry characterized five main components in the extract: benzeneacetaldehyde (
Conclusion
The consistency of a variety of computational outputs justifies the dietary–supplemental potential of garlic essential oil based on its organosulfur ingredients.
Keywords
Introduction
The respiratory system has been commonly known to be vulnerable to bacterial and viral coinfections, often resulting in significantly higher mortality and morbidity. Because of its severe impacts globally, the current SARS-CoV-2 pandemic and associated coinfections have gained special attention from the scientific community. In particular, for the latter, an estimate indicated that 94.2% of COVID-19-confirmed patients (in Jiangsu Province) were coinfected with one or more (up to 24) respiratory pathogens within 1–4 days of onset of the first infection; 1 Streptococcus pneumoniae was the most common, followed by Klebsiella pneumoniae and Haemophilus influenzae. Pal et al reported in detail three cases with SARS-CoV-2 complicated by S pneumoniae coinfection, revealing that all three patients had respiratory failure and required intensive oxygen supplementation. 2 Many bacterial species that cause disease in humans and animals belong to the Streptococci group. Various diseases are typically caused by this group in humans, eg arthritis, neonatal sepsis, meningitis, and pneumonia; they are also known mainly to cause mastitis in animals. Therefore, accumulating knowledge about the SARS-CoV-2-related coinfection, in general, and the respiratory potential bacteria, in particular, is of importance.
Streptococcus pyogenes, group A streptococcal (GAS), is a major causal agent of acute bacterial pharyngitis, popularly known as strep throat. 3 This illness is known to affect ca 30% of children and over 10% of adults. Nearly 1000 million new cases of pharyngitis are diagnosed globally each year. As a result, S pyogenes infection is one of the leading healthcare costs. 4 LuxS is essential for various S pyogenes virulence features, especially its capacity to build biofilm. Lyon et al conducted research that involved the mutation of the luxS gene, which resulted in altered virulence activity, increasing expression of the sagA gene and decreasing SpeB proteolytic activity. 5 In another work on a mutant luxS gene, Kang et al revealed the importance of luxS in S pyogenes pathogenicity. 6 These findings altogether demonstrated the important role of luxS in S pyogenes activities. 6 The structure database of luxS protein of S pyogenes can be found and retrieved from UniProtKB with ID P0C0C7 (S-ribosylhomocysteinase from S suis; S-ribosylhomocysteine lyase). The biological assembly of protein P0C0C7 (UniProtKB: P0C0C7 (LUXS_STRPY)) of S pyogenes is shown in Figure 1a.

Crystal structures of (a) LuxS protein of S pyogenes (UniProtKB: P0C0C7 (LUXS_STRPY)), (b) LuxS protein of S pneumoniae (UniProtKB: Q8DQF8 (Q8DQF8_STRR6)), (d) spike protein of SARS-CoV-2 Delta variant (DOI:10.2210/pdb7V7N/pdb), spike protein of (e) SARS-CoV-2 Omicron variant (DOI:10.2210/pdb7T9J/pdb), and structural formula of (c) benzylpenicillin (penicillin G) (D1) and (f) structural formula of molnupiravir (D2).
Infection with S pneumoniae has caused millions of deaths worldwide over decades. In fact, pneumonia is still the primary cause of juvenile death in poor countries, where infections kill 1.6 million people worldwide each year, ca 0.7–1 million children under the age of five, predominantly recorded in Asia and Africa, according to World Health Organization (WHO). 7 Like other Gram-positive bacteria, S pneumoniae also releases autoinducers, ie Autoinducer-2 (AI-2) directly produced by the protein luxS. More particularly, AI-2 belongs to a family of signaling molecules activated in quorum sensing (QS) and is essential to build biofilm. The structure database of LuxS protein of S pneumoniae can be found and retrieved from UniProtKB: Q8DQF8 (Q8DQF8_STRR6) of S pneumoniae (strain ATCC BAA-255/R6). The biological assembly of protein Q8DQF8 (UniProtKB: Q8DQF8 (Q8DQF8_STRR6)) of S pneumoniae (strain ATCC BAA-255/R6) is shown in Figure 1b.
Benzylpenicillin (penicillin G), known as the “natural” penicillin, was used as a positive control. Penicillins possess a wide antibacterial spectrum. Billions of doses have been administered since the 1940s in different regions. 8 S pneumoniae and Lancefield Group A, β-hemolytic streptococcus (S pyogenes) are known to be very sensitive to penicillin G. The structural formula of penicillin G is given in Figure 1c.
The SARS-CoV-2 pandemic has become a worldwide concern to public health because of its high incidence of infection and fatality. Because of unknown effective specific medications or vaccinations, the mortality toll has been escalating at an alarming pace. To control the overrapid spread of life-threatening infectious outbreaks, the idea of drug repositioning and alternative therapy are important strategies. Among the strains encountered, Delta and Omicron variants have gained special attention from the scientific community. Delta variant B.1.167.2, which has 23 mutations (12 are spike-based mutations), is the most well-known and deadliest SARS-CoV-2 variant. 9 The key mutation of B.1.167.2 is L452R and P681R spike protein mutations. In detail, L452R improves binding affinity to Angiotensin-converting enzyme 2 (ACE2) receptor and evading CD8 T lymphocytes, while P681R cleaves the precursor to form two new spike proteins called S1 and S2, which in turn induce superior fusion on the virus-bound host cells. Omicron has 32 mutations, mainly at receptor binding domain (RBD), N-terminal domain (NTD), and receptor binding motif (RBM). These changes are thought to be able to neutralize antibodies produced by previous infections or vaccinations. In theory, if the inactivation of spike proteins is of efficient success, SARS-CoV-2 itself cannot bind to the host receptors, meaning that the virus is not able to interact with or enter human cells. The crystal structure of spike proteins of SARS-CoV-2 Delta and Omicron variants was recently determined and published in Protein Data Bank under the entries PDB-7V7N (http://doi.org/10.2210/pdb7V7N/pdb) and PDB-7V7N (http://doi.org/10.2210/pdb7T9J/pdb), respectively. Figure 1e–d shows their biological assemblies and that of molnupiravir (Figure 1f), an antiviral medication used to treat COVID-19 in those infected by SARS-CoV-2.
For centuries, garlic (Allium sativum L.), the most abundantly produced bulb after onion, has been serving nutritional and medicinal roles of importance. It is known that garlic essential oil contains a wide variety of special organosulfur compounds, which exhibit a variety of antioxidant, antibacterial, antifungal, antitumor, and antimicrobial properties. Among the main components of garlic essential oil, allicin is a typical reactive sulfur-containing organic compound that has received enormous attention.10,11 In our preceding work, garlic essential oil was analyzed for chemical characterization, revealing the presence of 17 organosulfur compounds, accounting for 99.4% of its contents. These compounds were predicted to exhibit strong interactions with the amino acids of the protein ACE2 and the main protease PDB6LU7 of SARS-CoV-2 (wild variant). 11 This suggests, to a certain degree, that the organosulfur family still retains its potential for the mutated variant of SARS-CoV-2 (eg Delta and Omicron), in particular, and be effective against bacterial microorganisms (eg S pyogenes and S pneumoniae), in general.
In this work, evidence for the antibiotic activity of garlic extract was collected by experimental attempts, while the biological and pharmacological potentiality of individual components was predicted by cross-platform computational iterations. The theoretical consistency obtained can justify the potential of garlic as a supplemental food for the fight against SARS-CoV-2 coinfections.
Results and Discussion
Experimental
Chemical Composition
The chemical composition of the garlic extract, determined by gas chromatography–mass spectrometry (GC-MS) analysis, is summarized in Table 1; five components were characterized, and their structural formulae are shown in Figure 2. In particular, diallyl trisulfide (DTS) (

Structural formulae of compounds (A1–A5) from the ethanol 70% extract of A sativum.
Identification of Major Volatile Bioactive Compounds in the Ethanol 70% Extract of Allium sativum.
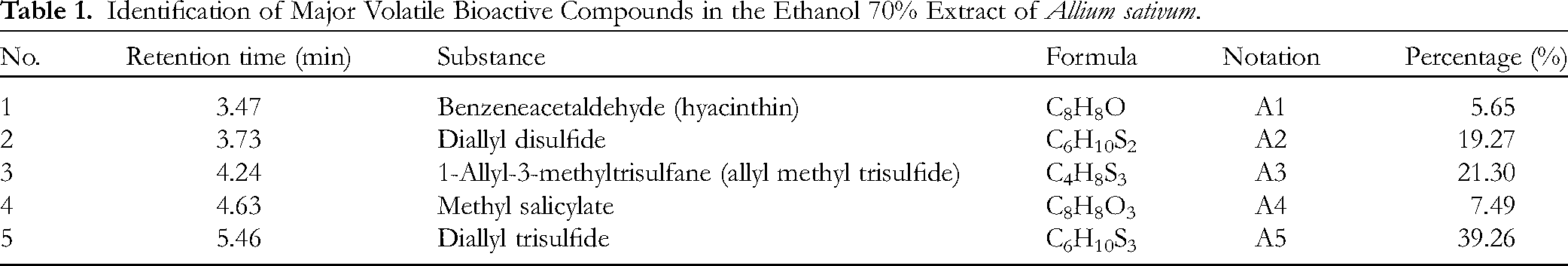
Garlic oil is known to contain ca 20 organosulfur compounds, eg diallyl sulfide (DAS), DDS, DTS, and diallyl tetrasulfide (DATTS), likely to correlate with the biological activities of the plant. Regarding the essential oils (EOs) obtained from fresh bulbs of garlic more particularly, significant antibacterial activity was in vitro proven quantitatively against S aureus (inhibition zone 14.8 mm), P aeruginosa (inhibition zone 21.1 mm), and E coli (inhibition zone 11.0 mm). 13 In this work, the main constituents determined were diallyl monosulfide, DDS, DTS, and DATTS.
From an experimental standpoint, it seems transparent that there is a solid correlation between garlic extract organosulfur compounds and biological activities of significance; yet, the related explanation for the factual observations has not been looked at in-depth from the standpoint of molecular electronic structures. Therefore, the chemical components determined in this work were also subjected to quantum-based analysis (ie density functional theory [DFT] calculation) in the attempt to provide a better understanding of their biological behavior from the view of the molecular constitution.
Antimicrobial Activity
Garlic extract inhibited the growth of S pneumoniae and S pyogenes, as shown in the agar diffusion method. Inhibition zones of 8 ± 2 mm were recorded, which is regarded as moderate. The positive control, penicillin G, produced an inhibition zone ≥ 30 ± 2 mm. The inhibition zones were observed for 18 h. The final result for S pneumoniae was two inhibitory circles, one showing complete inhibition and the other partial inhibition. This suggests that the bacteria were inhibited for the first few hours in the initial growth phase, but growth renewed after the garlic extract volatile components had vaporized at atmospheric pressure.
Therefore, disc volatilization assays were utilized in an attempt to give a more in-detail view of the antibacterial activity of these volatile components. In brief, garlic extract–impregnated filter paper discs were inoculated with microorganism-cultured agars in Petri dishes and sealed with parafilm to prevent leakage of vaporizable components into the atmosphere. In the absence of garlic extract (negative control), the growth of the two Streptococcus strains could be observed by the naked eye and through the beta-hemolysis phenomena, given by the uniform breakdown of red blood cells leaving clearer regions around the bacterial growth. In contrast, the presence of the garlic extract vapor phase resulted in no colonies of the two test strains on the plates after incubation. This suggests that the bacterial growth was inhibited even at the initial growth phase. In detail, most bacterial cells were killed after treatment with 0.0036 mg garlic extract per cm2 in the area unit or 3.6 mg per L in the volume unit. This value is calculated based on the growth area of bacteria in the 90 mm Petri disc, which is 58 cm2 in area and 0.058 L in volume. The results observed from the disc volatilization assays also indicate that the nonvolatile compounds did not play a significant role in the observed antibacterial activity.
The efficient susceptibility of the bacteria to the extract is assessed by minimum inhibitory concentration (MIC) values. The results reveal that S pneumoniae was inhibited by the garlic extract at a concentration of 64 mg·mL−1; the corresponding figure for S pyogenes was 128 mg·mL−1. This also indicates the greater susceptibility of the former to the garlic extract, in comparison to that of the latter. However, the MIC value of garlic extract exhibited a large variation compared with that of penicillin G against Streptococcus spp., which was 0.03 μg·mL−1. This value was in line with NCCLS-recommended interpretive criteria for penicillin (ie ≤0.06 μg·mL−1, susceptible; 0.12–1 μg·mL−1, intermediate; and ≥2 μg·mL−1, resistant).
Sulfur-containing compounds found in garlic extract, eg allicin, DDS, and DTS, were reportedly able to interact with enzyme-based thiol groups, eg alcohol dehydrogenase or thioredoxin reductase, and degrade the bacterial membrane stability. 14 In general, garlic extract had already been shown to be active both in vitro and in vivo against S mutans, the principal etiological bacterium in dental caries, with MIC values ranging from 0.5 to 128 mg·mL−1. 15 Herein, the antibacterial activities against two other pathogenic Streptococcus, eg S pyogenes and S pneumoniae, are reported with MIC values ranging from 64 to 128 mg·mL−1.
The effectiveness of the garlic extract against the two Streptococcus spp. is experimentally supported; however, the knowledge of the biological behavior of the candidate–microorganism interactions is still inadequate from the theoretical view. Nevertheless, the evidence from the existing literature seems to direct the responsibility of significance toward sulfur-containing compounds. Therefore, another computational technique (ie molecular docking simulation) was utilized in an attempt to reach an appropriate explanation. Furthermore, due to its infectiousness and transmissibility, in this work, SARS-CoV-2 was not subjected to direct in vitro tests but to in silico predictions. The results can serve as supplemental understanding of the overall potential of garlic extract organosulfur compounds against the dangerous respiratory micropathogens.
Computational
Density Functional Theory–Based Chemical Properties
Results from DFT calculation can be utilized either to check the unusual bonding parameters (if any) in the structural formulae of the bioactive compounds (
The optimized structures are shown in Figure 3; in-detail parameters of the structures are presented in Table 2. Overall, the computation can reach a geometry convergence regarding all the formulae, implying the compounds’ in-nature stable existence. No unusual bonding parameters were detected either in lengths or angles. In particular, all the figures meet the characteristic values, eg 1.54 Å for C–C, 1.09 Å for C-H, 1.43 Å for C-O (oxatriquinane), 1.23 Å for C=O (carbonyl compounds), 1.82 Å for C-S, and 1.8–3.0 Å for S-S. The corresponding ground state electronic energies register in the range between −1429.226248 au and −384.931174 au, which are commonly agreed to be highly stable. All the structures are also considered polarized molecules, especially
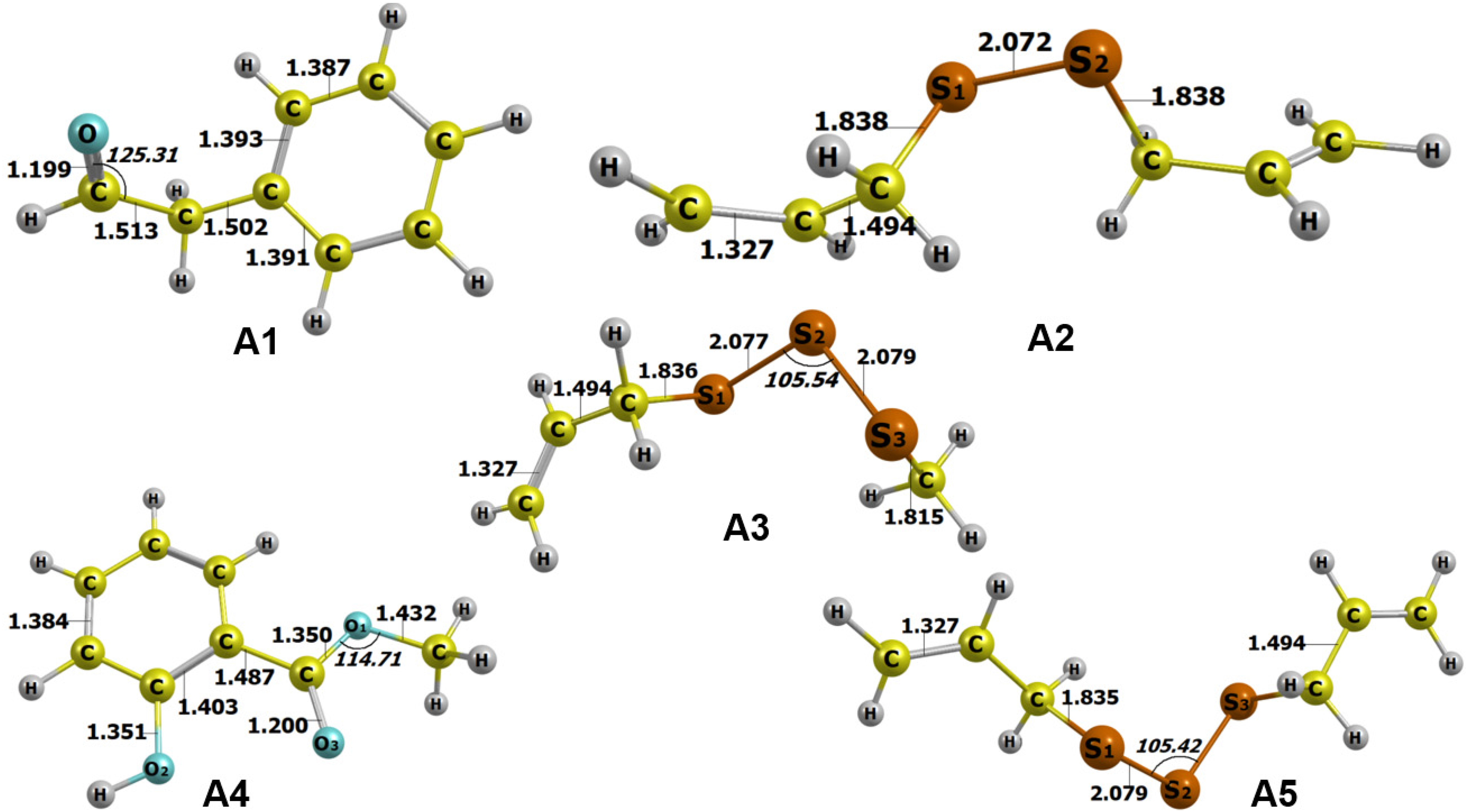
Optimized structures of A1–A5 calculated by DFT at level of theory M052X/6-311++g (d,p). Bond lengths are given in Å and angles in °.
Ground State Electronic Energy and Dipole Moment Values of A1–A5 Calculated by DFT at Level of Theory M052X/6-311++g(d,p).

Figure 4 presents the highest occupied molecular orbital (HOMO) and lowest unoccupied molecular orbital (LUMO) of the optimized structures; Table 3 gives the in-detail quantum parameters. Except for

HOMO and LUMO of A1–A5 calculated by DFT at level of theory M052X/def2-TZVPP.
Quantum Chemical Parameters of Compounds A1–A5 Calculated by NBO at the Level of Theory M052X/def2-TZVPP.
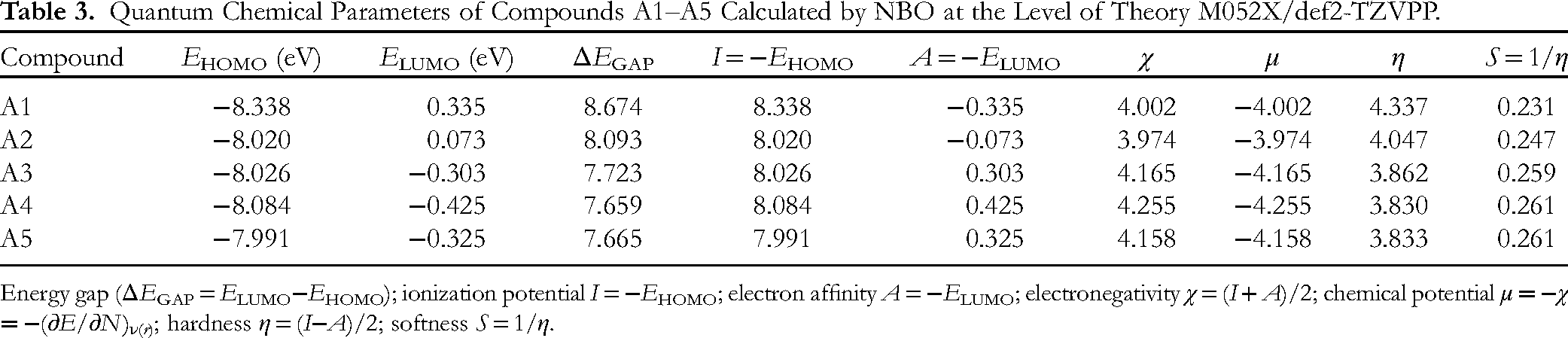
Energy gap (ΔEGAP = ELUMO−EHOMO); ionization potential I = −EHOMO; electron affinity A = −ELUMO; electronegativity χ = (I + A)/2; chemical potential μ = −χ = −(∂E/∂N)ν( r ); hardness η = (I−A)/2; softness S = 1/η.
Molecular electronic potential (MEP) maps of the optimized structures given in Figure 5 can provide another view of their chemical activities based on the distribution of potential energy; by convention, reddish shades represent the negative regions (related to electrophilic reactivity), while bluish shades represent positive regions (related to nucleophilic reactivity). Overall, the structures (

Molecular electrostatic potential (MEP) formed by mapping of total density over the electrostatic potential of A1–A5 calculated by DFT at level of theory M052X/def2-TZVPP.
The Fukui function can predict the reaction sites favored by nucleophilic (f+), electrophilic (f−), and radical (f0) attacks. In principle, reactivity of atoms is measured by the corresponding Fukui indices, which are summarized in Table 4. Similar to those retrieved from MEP analysis, organosulfur compounds (
Mulliken Atomic Charges and Fukui Indices of A1–A5.

This means that the reference is more useful for chemical reactions (based on electronic reconfiguration) than for intermolecular interaction (based on dipole–dipole forces); meanwhile, the latter is commonly accepted as the principal driving force for ligand–protein inhibition.
Docking-Based Inhibitory Properties
Retrieval from molecular docking simulation can be utilized either to verify the inhibitory effectiveness observed by experimental bioassays or to study the ligand–protein inhibitory behavior. In principle, the total docking score (DS) values and the number of hydrogen bonds are the most important parameters as the former represents Gibbs free energy of the inhibitory complexes, while the latter gives information on strong intermolecular bonds; besides, root-mean-square deviation (RMSD) (ie the average distance between backbone atoms; bioconformational rigidity or ligand–protein fitting) and van de Waals interactions (ie hydrophobic forces) may be considered to a certain degree. In each ligand–protein duo, only the most stable intermolecular structure is selected for more in-depth discussion since it most likely corresponds to the dominant product in reality. In brief, the computation can provide a certain insight into ligand–protein interaction potential from the standpoint of classical mechanics.
LuxS Proteins (Q8DQF8 and P0C0C7)
Figure 6 presents the most susceptible sites (sites 1–4) of the targeted proteins (Q8DQF8 and P0C0C7) to the ligands (

Quaternary structures of protein Q8DQF8 and protein P0C0C7 with approachable sites by A1–A5 and D1: site 1 (gray), site 2 (yellow), site 3 (blue), and site 4 (orange).
Prescreening Results on Inhibitability of A1–A5 and Controlled Drug (Penicillin G) (D1) toward the sites of proteins Q8DQF8 and P0C0C7.

P: protein; L: ligand; E: DS value (kcal·mol−1); N: number of hydrophilic interactions.
The selected data are summarized in Table 6 (ligand–Q8DQF8) and Table 7 (ligand–P0C0C7). According to the theoretical interpretation as given, the most effective ligand–protein inhibitory structures regarding S pneumoniae are in the order:
Molecular Docking Simulation Results for Ligand–Q8DQF8 Inhibitory Complexes.

DS: Docking score energy (kcal·mol−1); RMSD: root-mean-square deviation (Å); L: ligand; P: protein; T: type; D: distance (Å); E: energy (kcal·mol−1).
Molecular Docking Simulation Results for Ligand–P0C0C7 Inhibitory Complexes.

DS: docking score energy (kcal·mol−1); RMSD: root-mean-square deviation (Å); L: ligand; P: protein; T: type; D: distance (Å); E: energy (kcal·mol−1).
The corresponding visual projections (3D in-pose morphology and 2D interaction map) are presented in Figure 7 (ligand-Q8DQF8) and Figure 8 (ligand-P0C0C7). Regarding

Visual presentation and in-pose interaction maps with ligand-Q8DQF8 (A1–A5 and D1).

Visual presentation and in-pose interaction maps with ligand-P0C0C7 (A1–A5 and D1).
Spike Proteins (7V7N and 7T9J)
Figure 9 presents the most susceptible sites (sites 1–4) of the targeted proteins (7V7N and 7T9J) to the ligands (

Quaternary structures of protein 7V7N and protein 7T9J with approachable sites by A1–A5 and D2: site 1 (gray), site 2 (yellow), site 3 (blue), and site 4 (orange).
Prescreening Results on Inhibitability of A1–A5 and Controlled Drug Molnupiravir (D2) Toward the Sites of Proteins 7V7N and 7T9J.

P: protein; L: ligand; E: DS value (kcal·mol−1); N: number of hydrophilic interactions.
The selected data are summarized in Table 9 (ligand–7V7N) and Table 10 (ligand–7T9J). Regarding the former, the difference in inhibitory effectiveness is rather marginal, varying between −13 and −15 kcal·mol−1. In detail, the order is
Molecular Docking Simulation Results for Ligand–7V7N Inhibitory Complexes.

DS: docking score energy (kcal·mol−1); RMSD: root-mean-square deviation (Å); L: ligand; P: protein; T: type; D: distance (Å); E: energy (kcal·mol−1).
Molecular Docking Simulation Results for Ligand–7T9J Inhibitory Complexes.

DS: docking score energy (kcal·mol−1); RMSD: root-mean-square deviation (Å); L: ligand; P: protein; T: type; D: distance (Å); E: energy (kcal·mol−1).
The corresponding visual projections (3D in-pose morphology and 2D interaction map) are presented in Figure 10 (ligand-7V7N) and Figure 11 (ligand-7T9J). Similarly, the consistency between the results retrieved from the quantum-based calculation (DFT) and that of Newton's mechanics (docking) is explicitly noticeable. The aromatic ring of

Visual presentation and in-pose interaction maps with ligand-7V7N (A1–A5 and D2).
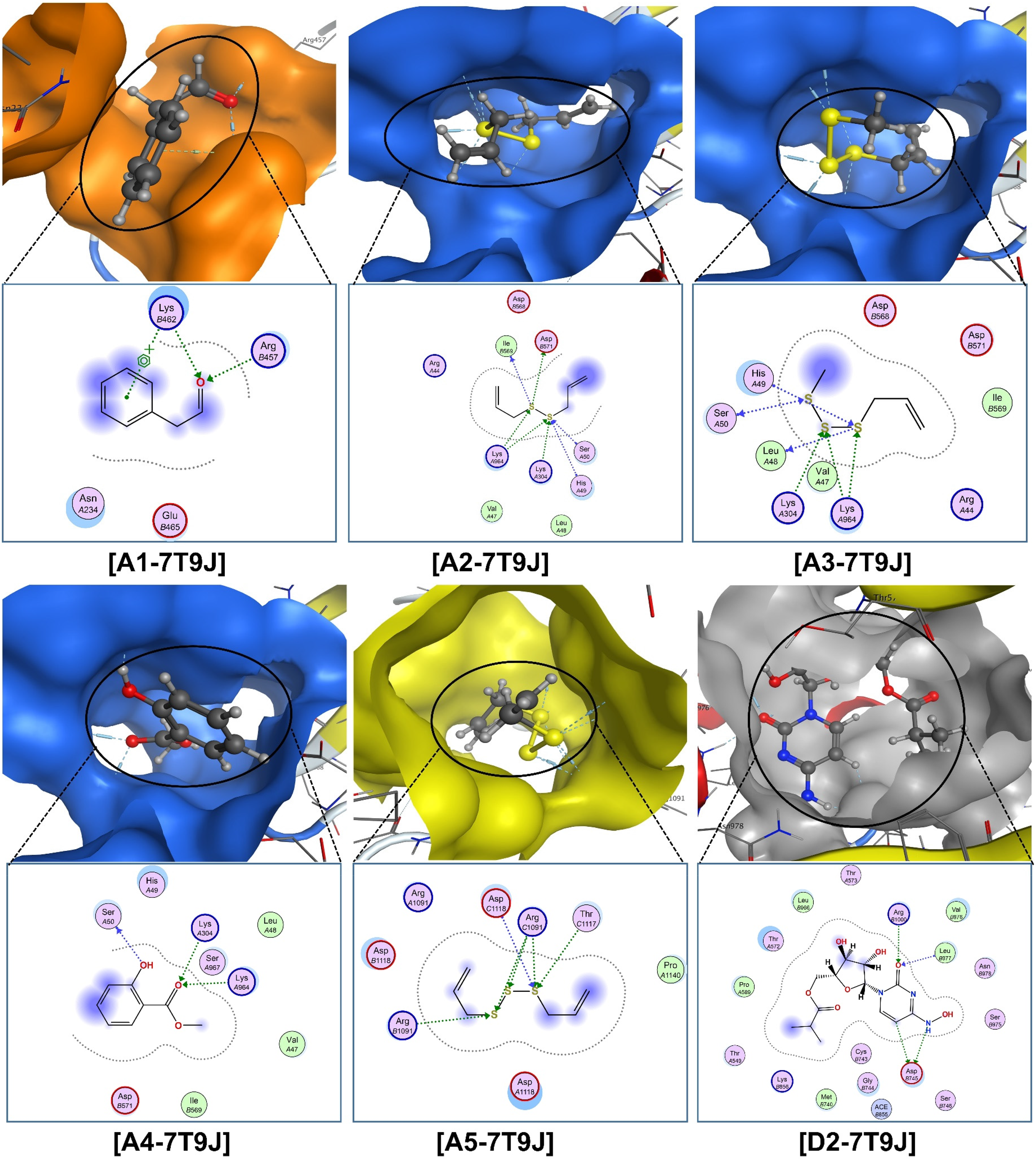
Visual presentation and in-pose interaction maps with ligand-7T9J (A1–A5 and D2).
It is also noteworthy that quantum computations are purely based on the wavelike behavior of molecular electrons, while those of classical mechanics are established on the assumption that each molecular atom is a rigid particle and assigned with arbitrary coefficients representing its individual physical properties. Therefore, the consistency seen should be of great interest; yet, more observations collected from this approach are still needed in order to reach a solid conclusion.
QSARIS-Based Physicochemical Properties
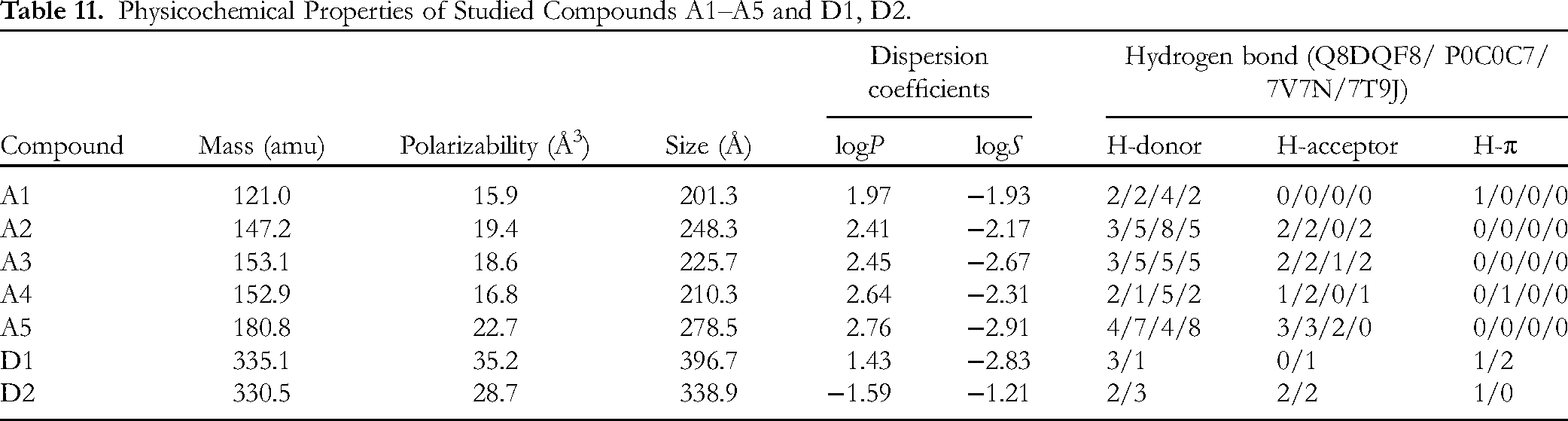
Table 11 summarizes the QSARIS-based physicochemical properties of the investigated compounds (
Physicochemical Properties of Studied Compounds A1–A5 and D1, D2.
ADMET-Based Pharmacokinetics and Pharmacology
Table 12 summarizes the absorption, distribution, metabolism, excretion, and toxicity (ADMET)-based output for the candidates (
ADMET-Based Pharmacokinetics and Pharmacology of the Studied Compounds A1–A5 and D1, D2.
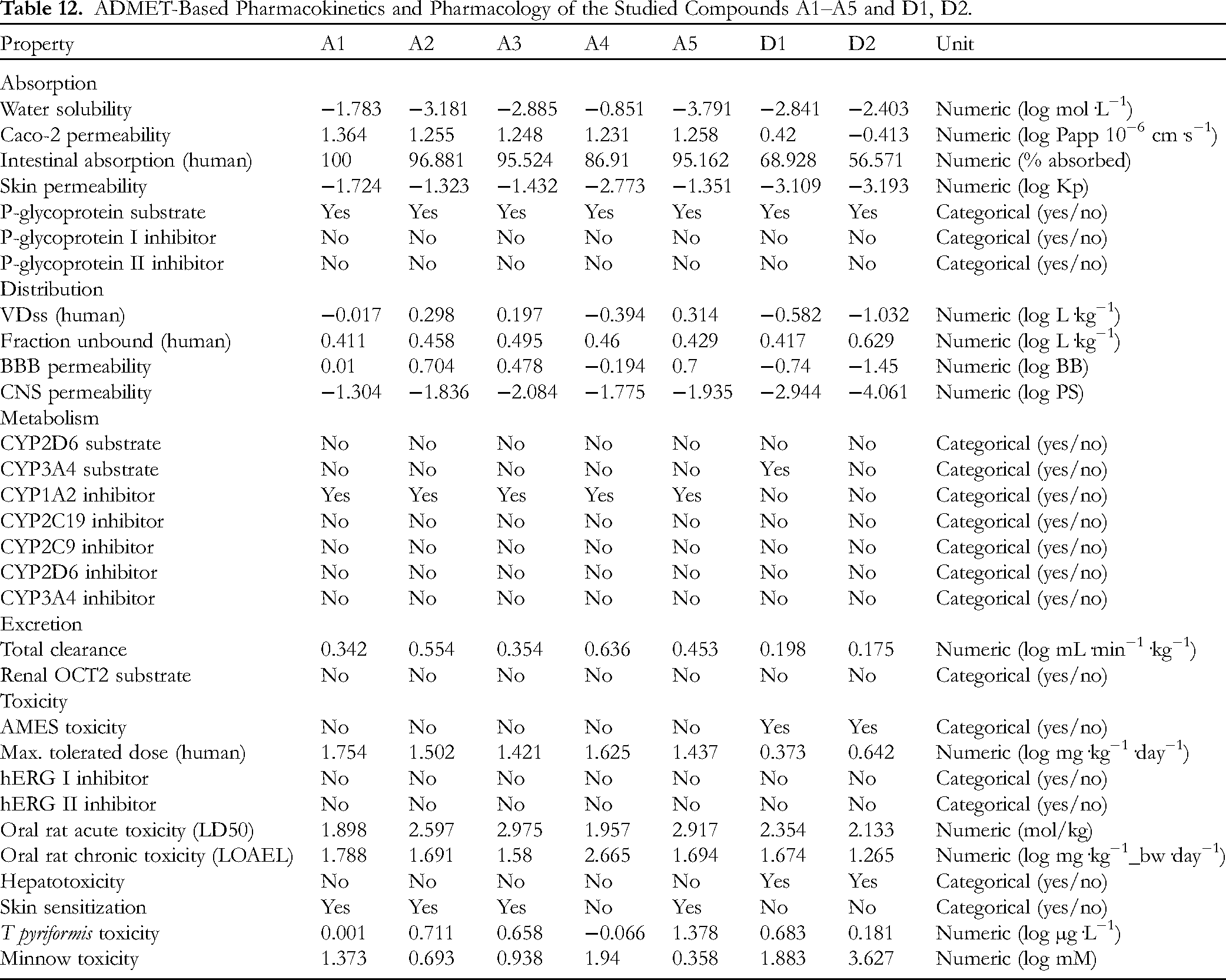
First, a compound is considered to have poor intestinal absorption if less than 30%. This means that all the candidates possess significant effectiveness of intestine absorbance regarding any compound by over 85%; especially, the model predicts complete absorption (ie 100%) regarding
Conclusions
This study, for the first time, includes both experimental and computational approaches to examine the potentiality of garlic extract against S pyogenes and S pneumoniae, extended to prediction against SARS-CoV-2 Delta and Omicron Spike proteins. GC-MS characterized five bioactive compounds: benzeneacetaldehyde (
Material and Methods
Materials
Plant Materials
Fresh peeled garlic was milled, then incubated at 4 °C for 10 min; afterward, ethanol (70%) was used for a further 15-min soaking, with a ratio of material: solvent of 1:10 g/mL. The mixture was then extracted with the assistance of ultrasonic radiation (frequency 37 kHz; temperature < 30 °C; duration 30 min). The obtained extract was homogenized in a flask (ca 1–2 min), then diluted with the original solvent to appropriate material/solvent ratios. The supernatant was collected after centrifugation (speed 3500 rpm; duration 10 min; temperature 4 °C) and subjected to a secondary extraction with similar experimental settings. These alcoholic extracts were collected and low-temperature evaporated (below 30 °C; atmospheric pressure) to obtain the final extract. The volatile part isolated from the total extract was further subjected to compositional analysis (GC-MS) and antibacterial tests.
Bioassay Materials
Microorganisms S pyogenes (ATCC 19615) and S pneumoniae (ATCC 49619) were supplied by the Microbiology and Parasitology Department, Faculty of Pharmacy, Nguyen Tat Thanh University.
Chemicals and reagents include ethanol (OPC Pharmaceutical Company); n-hexane (VN-Chemsol, Co., Ltd); dimethyl sulfoxide (DMSO), Mueller–Hinton broth (MHB), and Mueller–Hinton agar (MHA) (Merck Co., Ltd, Germany); and penicillin G (Nam Khoa Biotek Co., Ltd). Equipment includes a test tube, micropipette, Eppendorf, Petri dish, 96-well microplate, water bath, and biosafety cabinet class II. All other chemical reagents and solvents, purchased from Sigma–Aldrich (USA), were of analytical grade and used without further purification.
Experimental Methods
Chemical Characterization
The garlic extract was analyzed by GC-MS, with an Agilent DB-5MS column (30 m × 0.25 mm × 0.25 µm). Helium was used as the carrier gas. In a typical procedure, the sample (1.0 µL) was injected with a split ratio of 20:1 and at a temperature of 250 °C. The GC temperature configuration was (i) initiation at 80 °C and (ii) linear increase to 300 °C (rate 20 °C min−1). The MS scanning was set from 29 to 650 amu for mass analysis. The compounds in the garlic extract were identified by reference to the NIST-17 database.
Antimicrobial assay
Microorganisms
The antibacterial activity of garlic extract on S pyogenes (ATCC 19615) and S pneumoniae (ATCC 49619) was evaluated by well diffusion assay and broth dilution methods. The procedures followed the standards issued by the Clinical and Laboratory Standards Institute, with modifications (CLSI supplement M100, 2019). 24 The bacteria were cultured in MHB (Merck, Germany) medium (temperature 37 °C; duration 5–8 h) until the 0.5 McFarland turbidity (approximately 5.108 CFU·mL−1). This provided the standardized bacterial suspension containing a concentration of 5.108 CFU·mL−1. 24
Well Diffusion Test
The standardized bacterial suspension was diluted to 1:10 in PBS before (100 µL used) being swabbed evenly over the surface of blood agar; the medium was allowed to dry (ca 5 min). Circular holes were punched in the agar for the diffusion of the antibacterial samples. Garlic extract (volume 70 µL; diluted 1:5 w/v in DMSO 10%) was injected into a hole for each experiment. Penicillin G (20 U) and DMSO (10% w/v) were used as the positive and negative controls, respectively. Afterward, the dishes were incubated (duration 24 h; temperature 37 °C) before the inhibition zone diameter (IZD) was measured. 25 The procedure was in triplicate in order for the mean IZD to be determined26,27 and interpreted as 0 mm for no sensitivity or resistance, 1–6 mm for low sensitivity, 7–14 mm for moderate sensitivity, and >14 for high sensitivity.
Vapor Phase Test
The in vitro vapor phase test was based on the method described by Kloucek et al with some adjustments. 28 The test system was carried out in three-section Petri dishes (90 mm diameter), consisting of a blood agar medium. The standardized bacterial suspension (100 µL) was swabbed evenly over the surface of the blood agar with two sections of the plate and allowed to dry (duration ca 5 min); the two different strains were spread into two different sections; the third section was left uninoculated as the negative control. Garlic extract (210 μL) was distributed on the surface of a sterile filter paper disc (thickness 0.18 mm; diameter 84 mm); the paper disc was placed on the separating wall of the plate. This means the distance between the disc and the inoculated agar surface was ca 2 mm. The plates were hermetically closed with parafilm adhesive tape to avoid evaporation and incubated (temperature 37 °C; duration 16–18 h). The visible growth of the test bacteria (if observed) was captured. All tests were carried out in a class II microbiological safety cabinet and in triplicate.
Agar Dilution Assay
Agar dilution involves the incorporation of different concentrations of the extract into an agar medium followed by a standardized microbial inoculum onto the surface of the agar plate. 29 In this study, the method was utilized for the evaluation of garlic extract MIC, ie the lowest concentration of the drug that inhibited the organism's growth. The assay followed a twofold dilution of the garlic extracts, ie 1, 2, 4, 8, 16, 32, 64, 128, and 256 mg·mL−1, in molten blood agar media; the standardized bacterial suspension was under 50-fold dilution (concentration ca 107 CFU·mL−1) before 2 mL was applied onto the agar surface (approximately 104 CFU per spot). One agar plate was used as a control seeded without garlic extract. After incubation (temperature 37 °C; duration 16–18 h), the plates were observed for visual comparison of microbial growth. Regarding this experimental scheme, MICs were determined as the lowest concentration of garlic extract that stopped the growth of bacteria.
Computational Methods
Quantum Chemical Calculation
Molecular quantum properties of the studied compounds (
Molecular Docking Simulation
A typical procedure for molecular docking simulation (by MOE 2015.10
35
) follows four steps,36–38 providing predictions on ligand–protein complex structures.
Predocking preparation: Sources for simulation input: crystal structures of LuxS proteins of S pneumoniae (UniProtKB: Q8DQF8 (Q8DQF8_STRR6)) and of S pyogenes (UniProtKB: P0C0C7 (LUXS_STRPY)) were referenced from UniProt under the respective entry ID; those of spike proteins of SARS-CoV-2 Delta (PDB-7V7N) and Omicron (PDB-7T9J) variants were downloaded from Protein Data Bank; chemical formulae of ligands were from experimental findings by GC-MS in this work; structural formulas of benzylpenicillin (penicillin G) ( Docking investigation: Configuration for docking simulation: poses retaining for intermolecular interaction probing = 10; maximum solutions per iteration = 1000; maximum solutions per fragmentation = 200; output format for ligand–protein inhibitory structures in *.sdf. Redocking iteration: The reliability of the docking protocol was validated by redocking iterations and recalculating RMSD. To implement, the ligand was removed from its inhibitory complexes and redocked to the protein structures. To validate, the accuracy of the docking protocol is justified if RMSD values (of docked and redocked conformations) are all under 2 Å. Postdocking analysis: In principle, the inhibitory effectiveness of a ligand–protein inhibitory system was primarily indicated by DS energy, the pseudo-Gibbs free energy. The value is contributed by attractive forces (hydrophilic binding and hydrophobic interaction) and geometrical complementarity (RMSD, ie the average between backbone atoms).
QSARIS-Based Analysis
Druglikeness properties of phytochemicals were under in silico analysis by the QSARIS system using the Gasteiger–Marsili method; 39 as output, the main parameters were molecular mass (Da), polarizability (Å3) and volume or size (Å), and dispersion coefficients (logP: partition coefficient, and logS: aqueous solubility). Afterwards, they were subjected for assessment of orally pharmacological compatibility based on Lipinski's rule of five. 40 Accordingly, a good membrane-permeable candidate should satisfy the following criteria: (1) Molecular mass < 500 Da, (2) no more than five groups for hydrogen bonds, (3) no more than 10 groups receiving hydrogen bonds, and (4) the value of logP is less than +5 (logP < 5).41,42
ADMET-Based Analysis
ADMET properties were obtained from a web-based regressive model developed and maintained by the Molecular Modeling Group, Swiss Institute of Bioinformatics, ie SwissADME (http://www.swissadme.ch/; May 17, 2023). The theoretical interpretations of output pharmacokinetic parameters were described by Pires et al. 43
Supplemental Material
sj-docx-1-npx-10.1177_1934578X231194440 - Supplemental material for Potentiality of Organosulfur Compounds Against SARS-CoV-2-Coinfected Bacteria Streptococcus pyogenes and S pneumoniae: A Cross-Platform Analysis from Computational Chemistry
Supplemental material, sj-docx-1-npx-10.1177_1934578X231194440 for Potentiality of Organosulfur Compounds Against SARS-CoV-2-Coinfected Bacteria Streptococcus pyogenes and S pneumoniae: A Cross-Platform Analysis from Computational Chemistry by Nguyen Minh Thai, Thanh Q. Bui, Phan Tu Quy, Nguyen Thi Thanh Hai, Dao-Cuong To, Duong Tuan Quang, Nguyen Quang Co, Nguyen Thanh Triet, Nguyen Thi Ai Thuan, Nguyen Thanh To Nhi, Vo Mong Tham and Nguyen Thi Ai Nhung in Natural Product Communications
Footnotes
Authors’ Contribution
All authors contributed to the study conception and design. All authors read and approved the final manuscript. Nguyen Minh Thai worked on methodology (experimental), formal analysis (experimental), and writing—original draft (experimental). Thanh Q. Bui worked on conceptualization (computational), methodology (computational), and writing—review and editing. Phan Tu Quy worked on investigation (computational) and software (computational). Nguyen Thi Thanh Hai worked on investigation (extraction) and visualization. Dao-Cuong To worked on conceptualization (experimental), formal analysis (experimental), and writing—original draft (experimental). Duong Tuan Quang worked on conceptualization (experimental), formal analysis (experimental), writing—review and editing (experimental), and supervision (experimental). Nguyen Quang Co worked on data curation (computational) and writing—original draft (computational). Nguyen Thanh Triet worked on resources (protein structures) and writing—original draft (computational). Nguyen Thi Ai Thuan worked on investigation (factionalization) and visualization. Nguyen Thanh To Nhi worked on investigation (bioassay) and visualization. Vo Mong Tham worked on investigation (bioassay) and visualization. Nguyen Thi Ai Nhung worked on conceptualization (computational), investigation (computational), data curation (computational), writing—review and editing (experimental), and supervision (computational).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical approval is not applicable to this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Vietnam National Foundation for Science and Technology Development (NAFOSTED) (Grant number 02/2020/ĐX) and Hue University (Grant number DHH2022-01-198; Grant number NCM.DHH.2020.04).
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article, and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
